Survey of Lichenized Fungi DNA Barcodes on King George Island (Antarctica): An Aid to Species Discovery
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Site and Handling of Samples
2.2. DNA Extraction from Lichen Thalli
2.3. PCR Amplification of Fungal Barcode Marker
2.4. Sequencing and Sequence Analysis
2.5. Sequence-Based Grouping and Identification
3. Results and Discussion
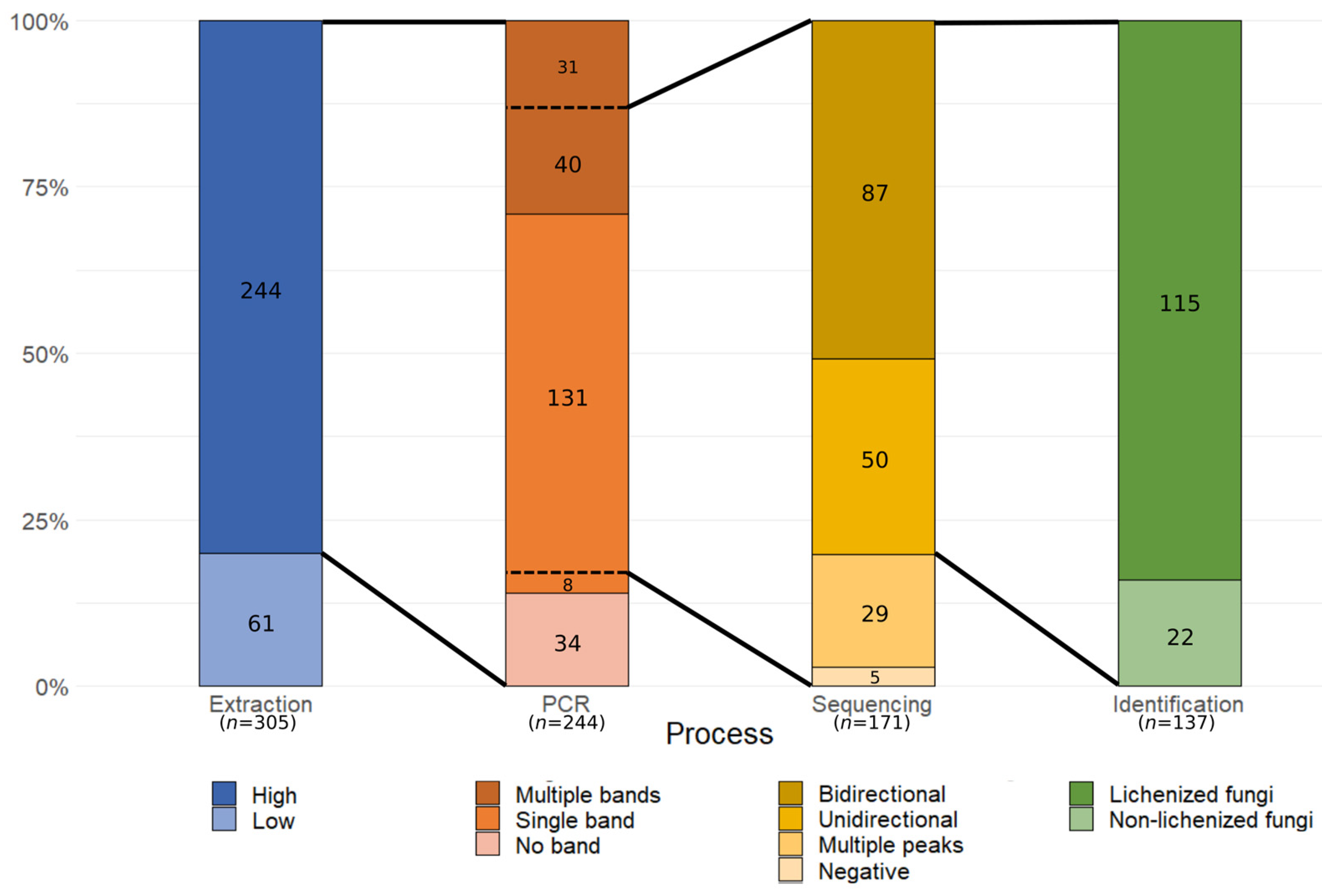
3.1. SAMPLE Heterogeneity and Molecular Processing Performance
3.2. Lichenized Fungi Identification
3.2.1. DNA-Based Identification at the Species Level
3.2.2. DNA-Based Identification at the Genus Level
3.2.3. Unidentified Lichenized Fungi Groups
3.3. Value of Molecular Surveys in Understudied Regions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Honegger, R. The Symbiotic Phenotype of Lichen-Forming Ascomycetes. In Fungal Associations; Hock, B., Ed.; The Mycota; Springer: Berlin, Heidelberg, 2001; pp. 165–188. ISBN 978-3-662-07334-6. [Google Scholar]
- Lücking, R.; Hodkinson, B.P.; Leavitt, S.D. Corrections and Amendments to the 2016 Classification of Lichenized Fungi in the Ascomycota and Basidiomycota. Bryologist 2017, 120, 58–69. [Google Scholar] [CrossRef]
- Lutzoni, F.; Kauff, F.; Cox, C.J.; McLaughlin, D.; Celio, G.; Dentinger, B.; Padamsee, M.; Hibbett, D.; James, T.Y.; Baloch, E.; et al. Assembling the Fungal Tree of Life: Progress, Classification, and Evolution of Subcellular Traits. Am. J. Bot. 2004, 91, 1446–1480. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, R.A. Adaptation of Lichens to Extreme Conditions. In Plant Adaptation Strategies in Changing Environment; Shukla, V., Kumar, S., Kumar, N., Eds.; Springer: Singapore, 2017; pp. 1–27. ISBN 978-981-10-6744-0. [Google Scholar]
- Zedda, L.; Rambold, G. The Diversity of Lichenised Fungi: Ecosystem Functions and Ecosystem Services. In Recent Advances in Lichenology: Modern Methods and Approaches in Lichen Systematics and Culture Techniques; Springer: New Delhi, India, 2015; Volume 2, pp. 121–145. ISBN 978-81-322-2234-7. [Google Scholar]
- de Carvalho, C.R.; Santiago, I.F.; da Costa Coelho, L.; Câmara, P.E.A.S.; Silva, M.C.; Stech, M.; Rosa, C.A.; Rosa, L.H. Fungi Associated with Plants and Lichens of Antarctica. In Fungi of Antarctica: Diversity, Ecology and Biotechnological Applications; Rosa, L.H., Ed.; Springer International Publishing: Cham, Switzerland, 2019; pp. 165–199. ISBN 978-3-030-18367-7. [Google Scholar]
- Arnold, A.E.; Miadlikowska, J.; Higgins, K.L.; Sarvate, S.D.; Gugger, P.; Way, A.; Hofstetter, V.; Kauff, F.; Lutzoni, F. A Phylogenetic Estimation of Trophic Transition Networks for Ascomycetous Fungi: Are Lichens Cradles of Symbiotrophic Fungal Diversification? Syst. Biol. 2009, 58, 283–297. [Google Scholar] [CrossRef] [PubMed]
- Lücking, R.; Aime, M.C.; Robbertse, B.; Miller, A.N.; Ariyawansa, H.A.; Aoki, T.; Cardinali, G.; Crous, P.W.; Druzhinina, I.S.; Geiser, D.M.; et al. Unambiguous Identification of Fungi: Where Do We Stand and How Accurate and Precise Is Fungal DNA Barcoding? IMA Fungus 2020, 11, 14. [Google Scholar] [CrossRef]
- Leavitt, S.D.; Moreau, C.S.; Thorsten Lumbsch, H. The Dynamic Discipline of Species Delimitation: Progress Toward Effectively Recognizing Species Boundaries in Natural Populations. In Recent Advances in Lichenology: Modern Methods and Approaches in Lichen Systematics and Culture Techniques; Upreti, D.K., Divakar, P.K., Shukla, V., Bajpai, R., Eds.; Springer India: New Delhi, India, 2015; Volume 2, pp. 11–44. ISBN 978-81-322-2235-4. [Google Scholar]
- Lücking, R.; Leavitt, S.D.; Hawksworth, D.L. Species in Lichen-Forming Fungi: Balancing between Conceptual and Practical Considerations, and between Phenotype and Phylogenomics. Fungal Divers. 2021, 109, 99–154. [Google Scholar] [CrossRef]
- Jiang, S.-H.; Lücking, R.; Liu, H.-J.; Wei, X.-L.; Xavier-Leite, A.B.; Portilla, C.V.; Ren, Q.; Wei, J.-C. Twelve New Species Reveal Cryptic Diversification in Foliicolous Lichens of Strigula s.Lat. (Strigulales, Ascomycota). J. Fungi 2022, 8, 2. [Google Scholar] [CrossRef]
- Crespo, A.; Pérez-Ortega, S. Cryptic Species and Species Pairs in Lichens: A Discussion on the Relationship between Molecular Phylogenies and Morphological Characters. An. Jardín Botánico Madr. 2009, 66, 71–81. [Google Scholar] [CrossRef]
- Lücking, R.; Dal-Forno, M.; Sikaroodi, M.; Gillevet, P.M.; Bungartz, F.; Moncada, B.; Yánez-Ayabaca, A.; Chaves, J.L.; Coca, L.F.; Lawrey, J.D. A Single Macrolichen Constitutes Hundreds of Unrecognized Species. Proc. Natl. Acad. Sci. USA 2014, 111, 11091–11096. [Google Scholar] [CrossRef]
- Lücking, R.; Forno, M.D.; Moncada, B.; Coca, L.F.; Vargas-Mendoza, L.Y.; Aptroot, A.; Arias, L.J.; Besal, B.; Bungartz, F.; Cabrera-Amaya, D.M.; et al. Turbo-Taxonomy to Assemble a Megadiverse Lichen Genus: Seventy New Species of Cora (Basidiomycota: Agaricales: Hygrophoraceae), Honouring David Leslie Hawksworth’s Seventieth Birthday. Fungal Divers. 2017, 84, 139–207. [Google Scholar] [CrossRef]
- Kelly, L.J.; Hollingsworth, P.M.; Coppins, B.J.; Ellis, C.J.; Harrold, P.; Tosh, J.; Yahr, R. DNA Barcoding of Lichenized Fungi Demonstrates High Identification Success in a Floristic Context. New Phytol. 2011, 191, 288–300. [Google Scholar] [CrossRef]
- Divakar, P.K.; Leavitt, S.D.; Molina, M.C.; Del-Prado, R.; Lumbsch, H.T.; Crespo, A. A DNA Barcoding Approach for Identification of Hidden Diversity in Parmeliaceae (Ascomycota): Parmelia Sensu Stricto as a Case Study. Bot. J. Linn. Soc. 2016, 180, 21–29. [Google Scholar] [CrossRef]
- Bradshaw, M.; Grewe, F.; Thomas, A.; Harrison, C.H.; Lindgren, H.; Muggia, L.; St. Clair, L.L.; Lumbsch, H.T.; Leavitt, S.D. Characterizing the Ribosomal Tandem Repeat and Its Utility as a DNA Barcode in Lichen-Forming Fungi. BMC Evol. Biol. 2020, 20, 2. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Consortium, F.B. Nuclear Ribosomal Internal Transcribed Spacer (ITS) Region as a Universal DNA Barcode Marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef]
- Lücking, R.; Nadel, M.R.A.; Araujo, E.; Gerlach, A. Two Decades of DNA Barcoding in the Genus Usnea (Parmeliaceae): How Useful and Reliable Is the ITS? Plant Fungal Syst. 2020, 65, 303–357. [Google Scholar] [CrossRef]
- Zhao, X.; Fernández-Brime, S.; Wedin, M.; Locke, M.; Leavitt, S.D.; Lumbsch, H.T. Using Multi-Locus Sequence Data for Addressing Species Boundaries in Commonly Accepted Lichen-Forming Fungal Species. Org. Divers Evol. 2017, 17, 351–363. [Google Scholar] [CrossRef]
- Orock, E.A.; Leavitt, S.D.; Fonge, B.A.; Clair, L.L.S.; Lumbsch, H.T. DNA-Based Identification of Lichen-Forming Fungi: Can Publicly Available Sequence Databases Aid in Lichen Diversity Inventories of Mount Cameroon (West Africa)? Lichenologist 2012, 44, 833–839. [Google Scholar] [CrossRef]
- Halıcı, M.G.; Bartak, M.; Güllü, M. Identification of Some Lichenised Fungi from James Ross Island (Antarctic Peninsula) Using NrITS Markers. N. Z. J. Bot. 2018, 56, 276–290. [Google Scholar] [CrossRef]
- Lücking, R.; Truong, B.V.; Huong, D.T.T.; Le, N.H.; Nguyen, Q.D.; Nguyen, V.D.; Raab-Straube, E.V.; Bollendorff, S.; Govers, K.; Vincenzo, V.D. Caveats of Fungal Barcoding: A Case Study in Trametes s.Lat. (Basidiomycota: Polyporales) in Vietnam Reveals Multiple Issues with Mislabelled Reference Sequences and Calls for Third-Party Annotations. Willdenowia 2020, 50, 383–403. [Google Scholar] [CrossRef]
- Hofstetter, V.; Buyck, B.; Eyssartier, G.; Schnee, S.; Gindro, K. The Unbearable Lightness of Sequenced-Based Identification. Fungal Divers. 2019, 96, 243–284. [Google Scholar] [CrossRef]
- Paul, F.; Otte, J.; Schmitt, I.; Dal Grande, F. Comparing Sanger Sequencing and High-Throughput Metabarcoding for Inferring Photobiont Diversity in Lichens. Sci. Rep. 2018, 8, 8624. [Google Scholar] [CrossRef]
- Faluaburu, M.S.; Nakai, R.; Imura, S.; Naganuma, T. Phylotypic Characterization of Mycobionts and Photobionts of Rock Tripe Lichen in East Antarctica. Microorganisms 2019, 7, 203. [Google Scholar] [CrossRef]
- Park, C.H.; Kim, K.M.; Elvebakk, A.; Kim, O.-S.; Jeong, G.; Hong, S.G. Algal and Fungal Diversity in Antarctic Lichens. J. Eukaryot. Microbiol. 2015, 62, 196–205. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Naganuma, T.; Nakai, R.; Imura, S.; Tsujimoto, M.; Convey, P. Microbiomic Analysis of Bacteria Associated with Rock Tripe Lichens in Continental and Maritime Antarctic Regions. J. Fungi 2022, 8, 817. [Google Scholar] [CrossRef]
- Fernández-Mendoza, F.; Fleischhacker, A.; Kopun, T.; Grube, M.; Muggia, L. ITS1 Metabarcoding Highlights Low Specificity of Lichen Mycobiomes at a Local Scale. Mol. Ecol. 2017, 26, 4811–4830. [Google Scholar] [CrossRef] [PubMed]
- Dal Forno, M.; Lawrey, J.D.; Moncada, B.; Bungartz, F.; Grube, M.; Schuettpelz, E.; Lücking, R. DNA Barcoding of Fresh and Historical Collections of Lichen-Forming Basidiomycetes in the Genera Cora and Corella (Agaricales: Hygrophoraceae): A Success Story? Diversity 2022, 14, 284. [Google Scholar] [CrossRef]
- Wright, B.; St. Clair, L.L.; Leavitt, S.D. Is Targeted Community DNA Metabarcoding Suitable for Biodiversity Inventories of Lichen-Forming Fungi? Ecol. Indic. 2019, 98, 812–820. [Google Scholar] [CrossRef]
- Henrie, J.R.; Thomson, B.M.; Yungfleisch, A.A.; Kerr, M.; Leavitt, S.D. Characterizing Crustose Lichen Communities—DNA Metabarcoding Reveals More than Meets the Eye. Diversity 2022, 14, 766. [Google Scholar] [CrossRef]
- Mark, K.; Cornejo, C.; Keller, C.; Flück, D.; Scheidegger, C. Barcoding Lichen-Forming Fungi Using 454 Pyrosequencing Is Challenged by Artifactual and Biological Sequence Variation. Genome 2016, 59, 685–704. [Google Scholar] [CrossRef]
- Kim, J.H.; Ahn, I.Y.; Hong, S.G.; Andreev, M.; Lim, K.M.; Oh, M.J.; Koh, Y.J.; Hur, J.S. Lichen Flora around the Korean Antarctic Scientific Station, King George Island, Antarctic. J. Microbiol. 2006, 44, 480–491. [Google Scholar]
- Pérez-Ortega, S.; Ortiz-Álvarez, R.; Green, T.G.A.; de los Ríos, A. Lichen Myco- and Photobiont Diversity and Their Relationships at the Edge of Life (McMurdo Dry Valleys, Antarctica). FEMS Microbiol. Ecol. 2012, 82, 429–448. [Google Scholar] [CrossRef]
- Zúñiga, C.; Leiva, D.; Ramírez-Fernández, L.; Carú, M.; Yahr, R.; Orlando, J. Phylogenetic Diversity of Peltigera; Cyanolichens and Their Photobionts in Southern Chile and Antarctica. Microbes Environ. 2015, 30, 172–179. [Google Scholar] [CrossRef] [PubMed]
- Wagner, M.; Bathke, A.C.; Cary, S.C.; Green, T.G.A.; Junker, R.R.; Trutschnig, W.; Ruprecht, U. Myco- and Photobiont Associations in Crustose Lichens in the McMurdo Dry Valleys (Antarctica) Reveal High Differentiation along an Elevational Gradient. Polar Biol 2020, 43, 1967–1983. [Google Scholar] [CrossRef]
- Cubero, O.F.; Crespo, A. Isolation of Nucleic Acids from Lichens. In Protocols in Lichenology: Culturing, Biochemistry, Ecophysiology and Use in Biomonitoring; Kranner, I.C., Beckett, R.P., Varma, A.K., Eds.; Springer Lab Manuals; Springer: Berlin/Heidelberg, Germany, 2002; pp. 381–391. ISBN 978-3-642-56359-1. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. PCR Protoc. A Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Gardes, M.; Bruns, T.D. ITS Primers with Enhanced Specificity for Basidiomycetes—Application to the Identification of Mycorrhizae and Rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef]
- Corporation, G.C. Sequencher® Version 5.4.6 DNA Sequence Analysis Software; Sequencher: Ann Arbor, MI, USA, 2011. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML Version 8: A Tool for Phylogenetic Analysis and Post-Analysis of Large Phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Castresana, J. Selection of Conserved Blocks from Multiple Alignments for Their Use in Phylogenetic Analysis. Mol. Biol. Evol. 2000, 17, 540–552. [Google Scholar] [CrossRef]
- Nilsson, R.H.; Larsson, K.-H.; Taylor, A.F.S.; Bengtsson-Palme, J.; Jeppesen, T.S.; Schigel, D.; Kennedy, P.; Picard, K.; Glöckner, F.O.; Tedersoo, L.; et al. The UNITE Database for Molecular Identification of Fungi: Handling Dark Taxa and Parallel Taxonomic Classifications. Nucleic Acids Res. 2019, 47, D259–D264. [Google Scholar] [CrossRef]
- Gueidan, C.; Elix, J.A.; McCarthy, P.M.; Roux, C.; Mallen-Cooper, M.; Kantvilas, G. PacBio Amplicon Sequencing for Metabarcoding of Mixed DNA Samples from Lichen Herbarium Specimens. MycoKeys 2019, 53, 73–91. [Google Scholar] [CrossRef]
- Kistenich, S.; Halvorsen, R.; Schrøder-Nielsen, A.; Thorbek, L.; Timdal, E.; Bendiksby, M. DNA Sequencing Historical Lichen Specimens. Front. Ecol. Evol. 2019, 7, 5. [Google Scholar] [CrossRef]
- Lücking, R.; Lawrey, J.D.; Gillevet, P.M.; Sikaroodi, M.; Dal-Forno, M.; Berger, S.A. Multiple ITS Haplotypes in the Genome of the Lichenized Basidiomycete Cora inversa (Hygrophoraceae): Fact or Artifact? J. Mol. Evol. 2014, 78, 148–162. [Google Scholar] [CrossRef] [PubMed]
- Paloi, S.; Luangsa-ard, J.J.; Mhuantong, W.; Stadler, M.; Kobmoo, N. Intragenomic Variation in Nuclear Ribosomal Markers and Its Implication in Species Delimitation, Identification and Barcoding in Fungi. Fungal Biol. Rev. 2022, 42, 1–33. [Google Scholar] [CrossRef]
- Stocker-Worgotter, E. Experimental Studies of the Lichen Symbiosis: DNA-Analyses, Differentiation and Secondary Chemistry of Selected Mycobionts, Artificial Resynthesis of Two- and Tripartite Symbioses. Symbiosis 2001, 30, 207–227. [Google Scholar]
- Rosa, L.H.; Zani, C.L.; Cantrell, C.L.; Duke, S.O.; Van Dijck, P.; Desideri, A.; Rosa, C.A. Fungi in Antarctica: Diversity, Ecology, Effects of Climate Change, and Bioprospection for Bioactive Compounds. In Fungi of Antarctica: Diversity, Ecology and Biotechnological Applications; Rosa, L.H., Ed.; Springer International Publishing: Cham, Switzerland, 2019; pp. 1–17. ISBN 978-3-030-18367-7. [Google Scholar]
- Lagostina, E.; Dal Grande, F.; Andreev, M.; Printzen, C. The Use of Microsatellite Markers for Species Delimitation in Antarctic Usnea Subgenus Neuropogon. Mycologia 2018, 110, 1047–1057. [Google Scholar] [CrossRef]
- Schmitt, I.; Lumbsch, H.T. Molecular Phylogeny of the Pertusariaceae Supports Secondary Chemistry as an Important Systematic Character Set in Lichen-Forming Ascomycetes. Mol. Phylogenetics Evol. 2004, 33, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, I.; Yamamoto, Y.; Lumbsch, H.T. Phylogeny of Pertusariales (Ascomycotina): Resurrection of Ochrolechiaceae and New Circumscription of Megasporaceae. J. Hattori Bot. Lab. 2006, 100, 753–764. [Google Scholar] [CrossRef]
- Park, J.S.; Oh, S.-O.; Woo, J.-J.; Liu, D.; Park, S.-Y.; Hur, J.-S. First Report of the Lichen Ochrolechia akagiensis (Ochrolechiaceae, Ascomycota) in Korea. Korean J. Mycol. 2019, 47, 95–104. [Google Scholar] [CrossRef]
- Kukwa, M. The Lichen Genus Ochrolechia in Poland III with a Key and Notes on Some Taxa. Herzog 2009, 22, 43–66. [Google Scholar]
- Galloway, D.J.; Lewis-Smith, R.I.; Quilhot, W. A New Species of Placopsis (Agyriaceae: Ascomycota) from Antarctica. Lichenologist 2005, 37, 321–327. [Google Scholar] [CrossRef]
- Beck, A.; Bechteler, J.; Casanova-Katny, A.; Dzhilyanova, I. The Pioneer Lichen Placopsis in Maritime Antarctica: Genetic Diversity of Their Mycobionts and Green Algal Symbionts, and Their Correlation with Deglaciation Time. Symbiosis 2019, 79, 1–24. [Google Scholar] [CrossRef]
- Spielmann, A.; Pereira, A. Lichens on the Maritime Antarctica (A Small Field Guide for Some Common Species). Glalia 2012, 4, 01–28. [Google Scholar]
- Davydov, E.A.; Peršoh, D.; Rambold, G. Umbilicariaceae (Lichenized Ascomycota)—Trait Evolution and a New Generic Concept. Taxon 2017, 66, 1282–1303. [Google Scholar] [CrossRef]
- Ott, S.; Brinkmann, M.; Wirtz, N.; Lumbsch, H.T. Mitochondrial and Nuclear Ribosomal DNA Data Do Not Support the Separation of the Antarctic Lichens Umbilicaria kappenii and Umbilicaria antarctica as Distinct Species. Lichenologist 2004, 36, 227–234. [Google Scholar] [CrossRef]
- McCune, B.; Meglio, J.D.; Curtis, M.J. An Unusual Ascospore Shape and a New Species, Umbilicaria nodulospora (Umbilicariaceae), from California and Oregon. Bryologist 2014, 117, 170–178. [Google Scholar] [CrossRef]
- Krzewicka, B.; García, M.A.; Johansen, S.D.; Sancho, L.G.; Martín, M.P. Morphological and Nuclear Ribosomal DNA Data Support Distinguishing Two New Species of Umbilicaria (Umbilicariaceae, Ascomycota) from Europe. Lichenologist 2009, 41, 631–648. [Google Scholar] [CrossRef]
- Øvstedal, D.O.; Smith, R.I.L. Lichens of Antarctica and South Georgia: A Guide to Their Identification and Ecology; Cambridge University Press: Cambridge, UK, 2001; ISBN 978-0-521-66241-3. [Google Scholar]
- Roca Valiente, B. Phylogenetic Study in the “Rhizocarpon geographicum” Group (Lichens, Rhizocarpaceae, Ascomycota). Contrasting Analysis of Morphological Characters and Biogeographic Patterns. Ph.D. Thesis, Universidad Complutense de Madrid, Madrid, Spain, 2013. [Google Scholar]
- Roca-Valiente, B.; Hawksworth, D.L.; Pérez-Ortega, S.; Sancho, L.G.; Crespo, A. Type Studies in the Rhizocarpon geographicum Group (Rhizocarpaceae, Lichenized Ascomycota). Lichenologist 2016, 48, 97–110. [Google Scholar] [CrossRef]
- Zhao, X.; Leavitt, S.D.; Zhao, Z.T.; Zhang, L.L.; Arup, U.; Grube, M.; Pérez-Ortega, S.; Printzen, C.; Śliwa, L.; Kraichak, E.; et al. Towards a Revised Generic Classification of Lecanoroid Lichens (Lecanoraceae, Ascomycota) Based on Molecular, Morphological and Chemical Evidence. Fungal Divers. 2016, 78, 293–304. [Google Scholar] [CrossRef]
- Ruprecht, U.; Lumbsch, H.T.; Brunauer, G.; Green, T.G.A.; Türk, R. Insights into the Diversity of Lecanoraceae (Lecanorales, Ascomycota) in Continental Antarctica (Ross Sea Region). Nova Hedwig. 2012, 94, 287–306. [Google Scholar] [CrossRef]
- Högnabba, F. Molecular Phylogeny of the Genus Stereocaulon (Stereocaulaceae, Lichenized Ascomycetes). Mycol. Res. 2006, 110, 1080–1092. [Google Scholar] [CrossRef]
- Högnabba, F.; Pino-Bodas, R.; Nordin, A.; Myllys, L.; Stenroos, S. Phylogenetic Position of the Crustose Stereocaulon Species. Lichenologist 2014, 46, 103–114. [Google Scholar] [CrossRef]
- Park, J.S.; Park, C.-H.; Park, S.-Y.; Oh, S.-O.; Jayalal, U.; Hur, J.-S. Revision of the Lichen Genus Stereocaulon (Stereocaulaceae, Ascomycota) in South Korea. Mycobiology 2018, 46, 101–113. [Google Scholar] [CrossRef]
- Redon Figueroa, J.; Instituto Antartico Chileno, S. Liquenes Antarticos; Instituto Antartico Chileno: Santiago, Chile, 1985. [Google Scholar]
- Sipman, H. Artificial Key to Marbach’s Tropical Buellia s.l. with Short Diagnoses. 2003. Available online: https://archive.bgbm.org/BGBM/STAFF/Wiss/Sipman/keys/Trobuellia.htm (accessed on 1 January 2023).
- Kalb, K.; Giralt, M. Orcularia, a Segregate from the Lichen Genera Buellia and Rinodina (Lecanoromycetes, Caliciaceae). Phytotaxa 2011, 38, 53. [Google Scholar] [CrossRef]
- Lücking, R.; Hodkinson, B.P.; Leavitt, S.D. The 2016 Classification of Lichenized Fungi in the Ascomycota and Basidiomycota—Approaching One Thousand Genera. Bryologist 2017, 119, 361–416. [Google Scholar] [CrossRef]
- Paknia, O.; Rajaei Sh, H.; Koch, A. Lack of Well-Maintained Natural History Collections and Taxonomists in Megadiverse Developing Countries Hampers Global Biodiversity Exploration. Org. Divers Evol. 2015, 15, 619–629. [Google Scholar] [CrossRef]
- Engel, M.S.; Ceríaco, L.M.P.; Daniel, G.M.; Dellapé, P.M.; Löbl, I.; Marinov, M.; Reis, R.E.; Young, M.T.; Dubois, A.; Agarwal, I.; et al. The Taxonomic Impediment: A Shortage of Taxonomists, Not the Lack of Technical Approaches. Zool. J. Linn. Soc. 2021, 193, 381–387. [Google Scholar] [CrossRef]






| Site Name | Transect Number | Sample Size | Coordinates | ||
|---|---|---|---|---|---|
| Latitude | Longitude | Elevation (m) | |||
| Scientific Campaign ANTAR XXVI–2019 | |||||
| Machu Picchu Base | 1 | 3 | 62°5′32″ S | 58°28′9″ W | 1 |
| 2 | 9 | 62°5′32″ S | 58°28′15″ W | 2 | |
| 3 | 32 | 62°5′43″ S | 58°29′19″ W | 201 | |
| 4 | 6 | 62°6′12″ S | 58°27′46″ W | 25 | |
| 5 | 48 | 62°5′38″ S | 58°28′25″ W | 41 | |
| 6 | 24 | 62°5′40″ S | 58°28′26″ W | 44 | |
| Scientific Campaign ANTAR XXVII–2020 | |||||
| Henryk Arctowski Base | 7 | 27 | 62°9′48″ S | 58°28′5″ W | 46 |
| 8 | 30 | 62°9′48″ S | 58°28′2″ W | 41 | |
| Machu Picchu Base | 9 | 25 | 62°5′36″ S | 58°28′12″ W | 21 |
| 10 | 30 | 62°5′36″ S | 58°28′28″ W | 38 | |
| Comandante Ferraz Base | 11 | 46 | 62°4′51″ S | 58°25′20″ W | 48 |
| 12 | 25 | 62°5′7″ S | 58°25′7″ W | 53 | |
| Sample Group | BLAST Top Hits | Identity Percentage Range | Number of Samples | Identification |
|---|---|---|---|---|
| I | Usnea aurantiacoatra | 98.36%−100% | 11 | Usnea aurantiacoatra |
| Usnea antarctica | 98.38%–100% | 27 | Usnea antarctica | |
| II | Ochrolechia frigida | 95.12%–100% | 13 | Ochrolechia frigida |
| Ochrolechia tartarea | 95.12%–100% | |||
| III | Placopsis antarctica | 98.2%–100% | 24 | Placopsis antarctica |
| Placopsis parellina | 99.44%–100% | |||
| IV-a | Caloplaca regalis | 99.62% | 1 | Polycauliona regalis |
| Gondwania regalis | 99.03%–99.4% | |||
| IV-b | Gondwania sp. | 97.23% | 1 | Caloplaca sp. |
| Caloplaca sublobulata | 96.25% | |||
| Gondwania sejongensis | 96.01% | |||
| V-a | Umbilicaria krascheninnikovii | 99.79%–100% | 2 | Umbilicaria decussata |
| Umbilicaria decussata | 95.59%–98.79% | |||
| V-b | Umbilicaria aprina | 95.21%–100% | 2 | Umbilicaria sp. |
| Umbilicaria africana | 95.56%–99.8% | |||
| VI-a | Rhizocarpon geographicum | 95.02%–100% | 3 | Rhizocarpon aff. geographicum |
| Rhizocarpon nidificum | 99.75%–100% | |||
| VII-a | Psoroma hypnorum | 95.88%–100% | 2 | Psoroma hypnorum |
| VIII-a | Lecanora polytropa | 99.67% | 1 | Lecanora polytropa |
| VIII-b | Lecanora polytropa | 95%–98.32% | 2 | Lecanora sp. s.l. |
| Lecanora cf. polytropa | 95.41%–98.71% | |||
| Rhizoplaca aspidophora | 100% | |||
| Lecanora fuscobrunnea | 97.35%–99.1% | |||
| IX | Lecidella carpathica | 97.34%–99.73% | 2 | Lecidella carpathica |
| X | Candelariella flava | 96.79%–99.45% | 2 | Candelariella flava |
| XI | Steinera intricata | 95.61%–100% | 7 | Steinera intricata |
| XII | Lepraria elobata | 95.71%–97.46% | 1 | Lepraria sp. |
| Lepraria caesioalba | 95.77%–97.45% | |||
| Lepraria granulata | 97.35% | |||
| Lepraria neglecta | 95.09%–97.34% | |||
| XIII-a | Stereocaulon alpinum | 95.11%–100% | 2 | Stereocaulon sp. 1 |
| Stereocaulon grande | 99.32% | |||
| Stereocaulon saxatile | 96.10%–99.13% | |||
| XIII-b | Stereocaulon glabrum | 97.08%–100% | 2 | Stereocaulon sp. 2 |
| Stereocaulon sp. | 95.01%–100% | |||
| XIV | Myriospora signyensis | 95.33%–99.8% | 1 | Myriospora sp. |
| Lecanoromycetes sp. | 99.37%–99.55% |
| Sample Group | Number of Samples | Sample Codes | Morphological Determination | GenBank Accession Codes |
|---|---|---|---|---|
| VI-b | 1 | 51A6 | Rhizocarpon sp. | OP730856 |
| VII-b | 1 | 72A2_1 | Psoroma hypnorum | OP730855 |
| XV | 2 | 110A, 111A | Lecidea sp. s. l. | OP730849, OP730850 |
| XVI | 1 | 24C | Buellia sp. s. l. | OP730848 |
| XVII | 4 | 94A1, 103A, 107A, 108A | Austrolecia sp. | OP730851, OP730852, OP730853, OP730854 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
La Torre, R.D.; Ramos, D.; Mejía, M.D.; Neyra, E.; Loarte, E.; Orjeda, G. Survey of Lichenized Fungi DNA Barcodes on King George Island (Antarctica): An Aid to Species Discovery. J. Fungi 2023, 9, 552. https://doi.org/10.3390/jof9050552
La Torre RD, Ramos D, Mejía MD, Neyra E, Loarte E, Orjeda G. Survey of Lichenized Fungi DNA Barcodes on King George Island (Antarctica): An Aid to Species Discovery. Journal of Fungi. 2023; 9(5):552. https://doi.org/10.3390/jof9050552
Chicago/Turabian StyleLa Torre, Renato Daniel, Daniel Ramos, Mayra Doris Mejía, Edgar Neyra, Edwin Loarte, and Gisella Orjeda. 2023. "Survey of Lichenized Fungi DNA Barcodes on King George Island (Antarctica): An Aid to Species Discovery" Journal of Fungi 9, no. 5: 552. https://doi.org/10.3390/jof9050552






