Unanticipated Large-Scale Deletion in Fusarium graminearum Genome Using CRISPR/Cas9 and Its Impact on Growth and Virulence
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification of Initial F. graminearum Genes for Targeted Deletion and Disruption Strategy
2.2. sgRNA Design for Target Genes
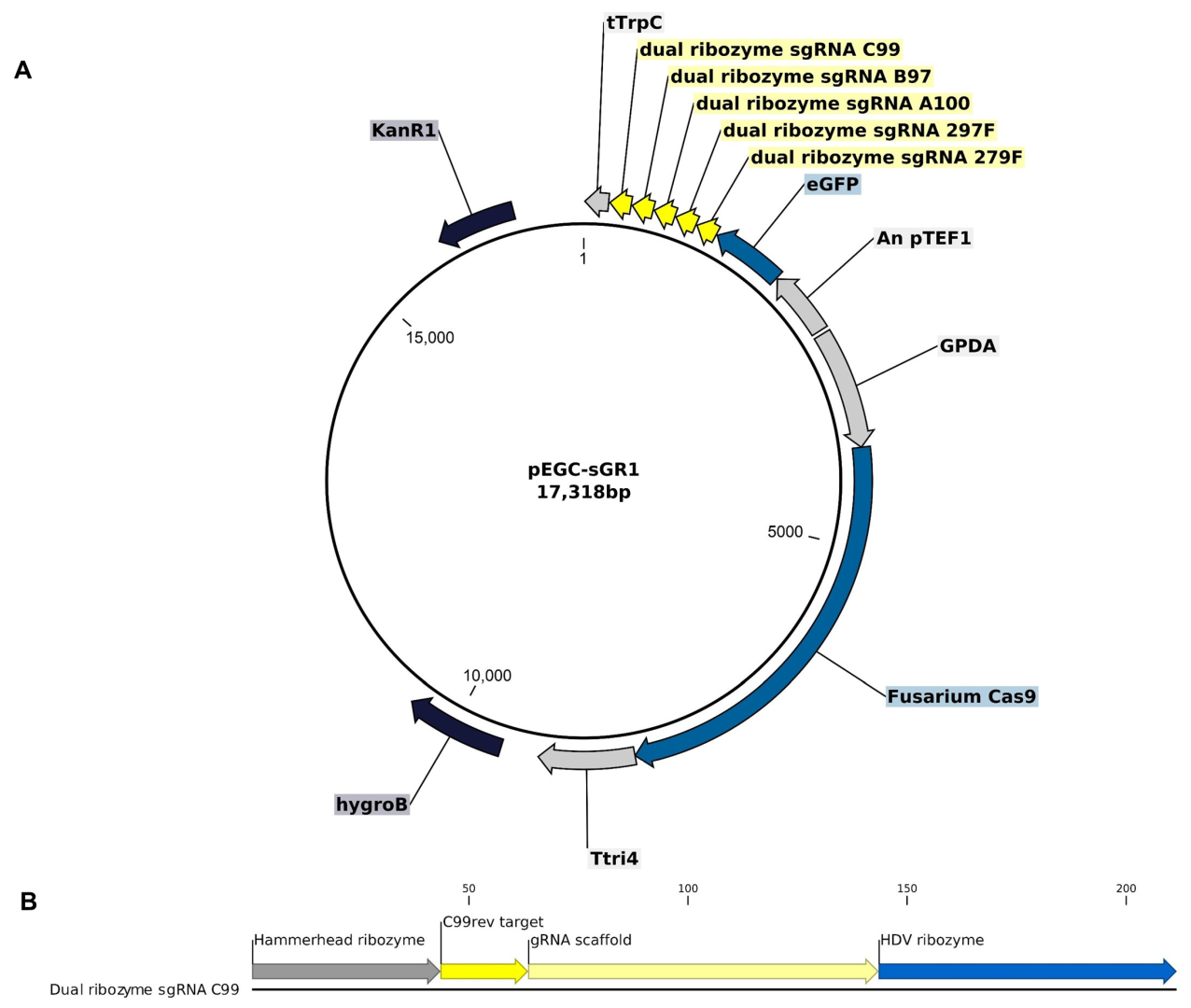
2.3. Vector Construct for CRISPR Editing
- GPDA_exp_cassette: Containing the gpdA promoter from Aspergillus nidulans, a 20 bp spacer with the AscI cut site, and the F. graminearum Tri4 terminator, synthesized by GeneArt (Life Technologies Inc., Toronto, ON, Canada).
- Cas9-2018ABP2XP: A F. graminearum codon-optimized gene encoding the native Cas9 protein, also synthesized by GeneArt synthetic gene synthesis (Life Technologies Inc., Canada).
- RNA_exp_cassette: Comprising five sgRNAs embedded between hepatitis delta virus (HDV) and hammerhead ribozymes, followed by an enhanced green fluorescent protein (GFP)-encoding marker gene. The expression of the sgRNA is driven by the A. nidulans translation elongation factor promoter (pTef1) and followed by the A. nidulans trpC terminator. This cassette was synthesized by General Biosystems (Duram, NC, USA).
2.4. Protoplast Transformation
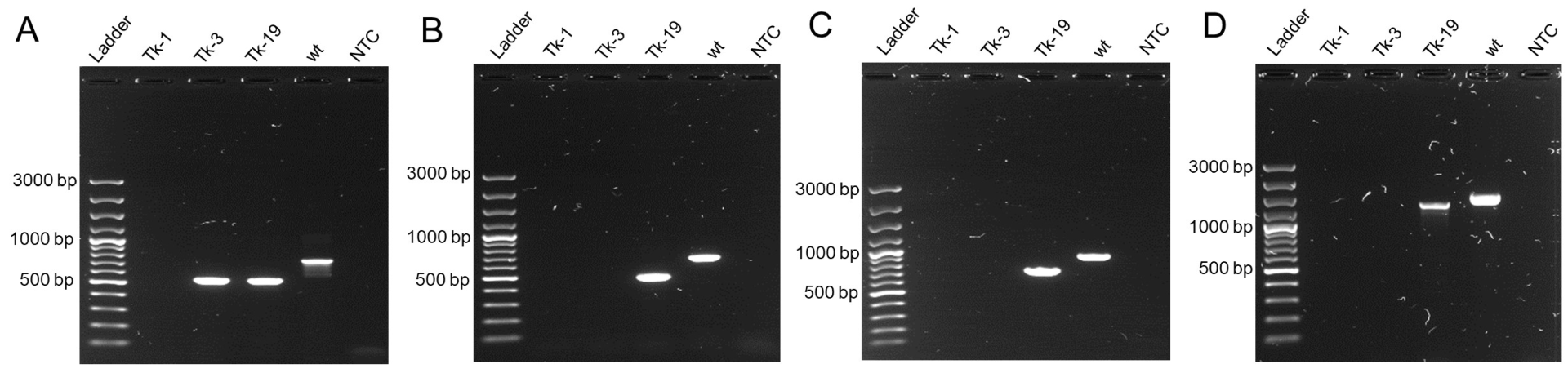
2.5. Genome Sequencing of F. graminearum Isolates to Investigate Gene Disruption
2.6. Growth Rate Assessment of F. graminearum Isolates
2.7. Effect of Gene Disruption on F. graminearum Virulence of Wheat
- Root disease assay: Wheat seeds were planted in Pro-Mix soil-filled 50 mL tubes and inoculated with a suspension of mycelium. The mycelium was cultivated in PDB liquid media for a week at 25 °C, mildly shaken at 50 RPM, centrifuged at 3200 CFU for 5 min, and the resultant mycelial pellet was resuspended in RO water. After disrupting the mycelium in a blender, the suspension was normalized to 200,000 CFU mL−1. Each seed was treated with 500 µL of the disrupted isolate, and seedling germination, emergence, and visual disease were documented at 3, 7, and 14 days after inoculation DAI.
- Clip dipping assay: Clip dipping protocol was modified from Shin et al. [24]. Wheat plants were grown in 50 mL tubes in Pro-Mix soil with four seeds per pot until they reached the Feekes 1.3 growth stage with three expanded leaves. Leaves were trimmed with scissors and dipped into a suspension of disrupted mycelium, produced as described above. Measurements of disease progression were taken at 3 and 7 DAI.
- Modified wheat point inoculation assay: This assay was modified from Feng and Tang [25]. Wheat plants were grown in a 1:1 mixture of sterilized field soil and Pro-Mix in 7.5-L pots until they reached anthesis, with 200 mL per pot of 1/4 Hoagland’s solution supplied weekly. The fifth anther from the base of the seed head was inoculated with a 2 mm plug of hyphae taken from the edge of a five-day-old culture grown on PDA at 25 °C. Plugs were attached with parafilm, and the entire plant was bagged to the base of the pot for 48 h to allow for disease development. Bags were removed, and disease was rated at 7 and 14 DAI by counting the number of infected spikelets showing bleached symptoms. A selection of diseased wheat tissues were surface sterilized with 70% ethanol and plated on PDA to confirm F. graminearum colonization.
2.8. Characterization of Mutant Isolates and Functional Analysis of Deleted Genes
3. Results and Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chongo, G.; Gossen, B.; Kutcher, H.; Gilbert, J.; Turkington, T.; Fernandez, M.; McLaren, D. Reaction of seedling roots of 14 crop species to Fusarium graminearum from wheat heads. Can. J. Plant Pathol. 2001, 23, 132–137. [Google Scholar] [CrossRef]
- Goswami, R.S.; Kistler, H.C. Heading for disaster: Fusarium graminearum on cereal crops. Mol. Plant Pathol. 2004, 5, 515–525. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Doudna, J.A.; Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096. [Google Scholar] [CrossRef]
- Atkins, A.; Chung, C.-H.; Allen, A.G.; Dampier, W.; Gurrola, T.E.; Sariyer, I.K.; Nonnemacher, M.R.; Wigdahl, B. Off-target analysis in gene editing and applications for clinical translation of CRISPR/Cas9 in HIV-1 therapy. Front. Genome Ed. 2021, 3, 673022. [Google Scholar] [CrossRef] [PubMed]
- Song, R.; Zhai, Q.; Sun, L.; Huang, E.; Zhang, Y.; Zhu, Y.; Guo, Q.; Tian, Y.; Zhao, B.; Lu, H. CRISPR/Cas9 genome editing technology in filamentous fungi: Progress and perspective. Appl. Microbiol. Biotechnol. 2019, 103, 6919–6932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, C.; Ma, X.; Gao, F.; Guo, Y. Off-target effects in CRISPR/Cas9 gene editing. Front. Bioeng. Biotechnol. 2023, 11, 1143157. [Google Scholar] [CrossRef] [PubMed]
- Ouedraogo, J.-P.; Tsang, A. CRISPR_Cas systems for fungal research. Fungal Biol. Rev. 2020, 34, 189–201. [Google Scholar] [CrossRef]
- Liu, R.; Chen, L.; Jiang, Y.; Zhou, Z.; Zou, G. Efficient genome editing in filamentous fungus Trichoderma reesei using the CRISPR/Cas9 system. Cell Discov. 2015, 1, 15007. [Google Scholar] [CrossRef] [Green Version]
- Ullah, M.; Xia, L.; Xie, S.; Sun, S. CRISPR/Cas9-based genome engineering: A new breakthrough in the genetic manipulation of filamentous fungi. Biotechnol. Appl. Biochem. 2020, 67, 835–851. [Google Scholar] [CrossRef]
- Zou, G.; Xiao, M.; Chai, S.; Zhu, Z.; Wang, Y.; Zhou, Z. Efficient genome editing in filamentous fungi via an improved CRISPR-Cas9 ribonucleoprotein method facilitated by chemical reagents. Microb. Biotechnol. 2021, 14, 2343–2355. [Google Scholar] [CrossRef]
- Nødvig, C.S.; Nielsen, J.B.; Kogle, M.E.; Mortensen, U.H. A CRISPR-Cas9 system for genetic engineering of filamentous fungi. PLoS ONE 2015, 10, e0133085. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gardiner, D.M.; Kazan, K. Selection is required for efficient Cas9-mediated genome editing in Fusarium graminearum. Fungal Biol. 2018, 122, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Kosicki, M.; Tomberg, K.; Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 2018, 36, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Owens, D.D.; Caulder, A.; Frontera, V.; Harman, J.R.; Allan, A.J.; Bucakci, A.; Greder, L.; Codner, G.F.; Hublitz, P.; McHugh, P.J.; et al. Microhomologies are prevalent at Cas9-induced larger deletions. Nucleic Acids Res. 2019, 47, 7402–7417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, J.; Rowe, D.; Subedi, P.; Zhang, W.; Suelter, T.; Valent, B.; Cook, D.E. CRISPR-Cas12a induced DNA double-strand breaks are repaired by multiple pathways with different mutation profiles in Magnaporthe oryzae. Nat. Commun. 2022, 13, 7168. [Google Scholar] [CrossRef]
- Park, S.H.; Cao, M.; Pan, Y.; Davis, T.H.; Saxena, L.; Deshmukh, H.; Fu, Y.; Treangen, T.; Sheehan, V.A.; Bao, G. Comprehensive analysis and accurate quantification of unintended large gene modifications induced by CRISPR-Cas9 gene editing. Sci. Adv. 2022, 8, eabo7676. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Cook, D.E. The contribution of DNA repair pathways to genome editing and evolution in filamentous pathogens. FEMS Microbiol. Rev. 2022, 46, fuac035. [Google Scholar] [CrossRef] [PubMed]
- Cuomo, C.A.; Güldener, U.; Xu, J.-R.; Trail, F.; Turgeon, B.G.; Di Pietro, A.; Walton, J.D.; Ma, L.-J.; Baker, S.E.; Rep, M.; et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 2007, 317, 1400–1402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, C.; Waalwijk, C.; de Wit, P.J.; Tang, D.; van der Lee, T. RNA-Seq analysis reveals new gene models and alternative splicing in the fungal pathogen Fusarium graminearum. BMC Genom. 2013, 14, 21. [Google Scholar] [CrossRef] [Green Version]
- Concordet, J.-P.; Haeussler, M. CRISPOR: Intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 2018, 46, W242–W245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desmond, O.J.; Manners, J.M.; Stephens, A.E.; Maclean, D.J.; Schenk, P.M.; Gardiner, D.M.; Munn, A.L.; Kazan, K. The Fusarium mycotoxin deoxynivalenol elicits hydrogen peroxide production, programmed cell death and defence responses in wheat. Mol. Plant Pathol. 2008, 9, 435–445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guillemette, T.; Ram, A.F.; Carvalho, N.D.; Joubert, A.; Simoneau, P.; Archer, D.B. Methods for investigating the UPR in filamentous fungi. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2011; Volume 490, pp. 1–29. [Google Scholar]
- Shin, S.; Kim, K.-H.; Kang, C.-S.; Cho, K.-M.; Park, C.S.; Okagaki, R.; Park, J.-C. A simple method for the assessment of Fusarium head blight resistance in Korean wheat seedlings inoculated with Fusarium graminearum. Plant Pathol. J. 2014, 30, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, C.; Liu, H.; Tang, Z. Fusarium graminearum inoculation on wheat head. Bio. Protoc. 2018, 8, e2964. [Google Scholar] [CrossRef]
- Divon, H.H.; Fluhr, R. Nutrition acquisition strategies during fungal infection of plants. FEMS Microbiol. Lett. 2007, 266, 65–74. [Google Scholar] [CrossRef] [Green Version]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stergiopoulos, I.; de Wit, P.J. Fungal effector proteins. Annu. Rev. Phytopathol. 2009, 47, 233–263. [Google Scholar] [CrossRef] [Green Version]
- Latijnhouwers, M.; de Wit, P.J.; Govers, F. Oomycetes and fungi: Similar weaponry to attack plants. Trends Microbiol. 2003, 11, 462–469. [Google Scholar] [CrossRef]
- Paoletti, M.; Clave, C. The fungus-specific HET domain mediates programmed cell death in Podospora anserina. Eukaryot. Cell 2007, 6, 2001–2008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, F.-L.; Feng, T. Diterpenes specially produced by fungi: Structures, biological activities, and biosynthesis (2010–2020). J. Fungi 2022, 8, 244. [Google Scholar] [CrossRef]
- Gladieux, P.; Guérin, F.; Giraud, T.; Caffier, V.; Lemaire, C.; Parisi, L.; Didelot, F.; Le Cam, B. Emergence of novel fungal pathogens by ecological speciation: Importance of the reduced viability of immigrants. Mol. Ecol. 2011, 20, 4521–4532. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| ID | Sequence | Target 1 | Target 2 |
|---|---|---|---|
| A100 | ATAGACTCGATGGTCCACAT | FGSG 3455 | -- |
| B97 | ATGCTCTCGATGGTCCACTG | FGSC 8238 | -- |
| C99 | TGGCGTCGCGGATGGTCCAT | FGSC 4583 | -- |
| 279F | GGCGACTACACCGTCACCTC | FGSC 8238 | -- |
| 297F | TCTGGCTGGAGCGGCCAGTT | FGSG 3455 | FGSC 8238 |
| Target | ID | Primer Seq | Region Chromosome 2 | wt Product Size (bp) |
|---|---|---|---|---|
| 03445 | 4A-17-F | TACGCCTGGACCCTATCAAC | 5,177,854–5,177,873 | 740 |
| 4A-756-R | CCTGGTACTCAGAGCCCATC | 5,178,574–5,178,593 | ||
| 08238 | 4B-9-F | CAACTCAATCACTCGCTTCAA | 782,571–782,591 | 665 |
| 4B-673-R | CTCATTATGTATTGCCGCACA | 782,851–782,870 | ||
| 04583 | 4C-23-F | TCGTCTCTTTTCATCCTCATCA | 8,472,131–8,472,152 | 940 |
| 4C-962-R | TCCATCACTCTTTTGGGTGAG | 8,473,050–8,473,070 | ||
| 04583 | 4C-FL-F | CTGCGACATGCAGATGTACC | 8,471,771–8,471,790 | 1498 |
| 4C-FL-R | GGACACTGGGAGCAACTCTC | 8,473,249–8,473,268 |
| Isolate | Raw Reads | Reads after Trimming | Reads Mapped | Median Coverage |
|---|---|---|---|---|
| wt | 19,013,264 | 19,011,008 | 97.01% | 72× |
| Tk-1 | 14,778,038 | 14,776,430 | 96.83% | 55× |
| Tk-3 | 18,104,336 | 18,102,091 | 96.75% | 61× |
| Tk-19 | 16,428,990 | 16,427,116 | 96.27% | 72× |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Foster, A.J.; Johnstone, E.; Saunders, A.; Colic, E.; Lassel, N.; Holmes, J. Unanticipated Large-Scale Deletion in Fusarium graminearum Genome Using CRISPR/Cas9 and Its Impact on Growth and Virulence. J. Fungi 2023, 9, 673. https://doi.org/10.3390/jof9060673
Foster AJ, Johnstone E, Saunders A, Colic E, Lassel N, Holmes J. Unanticipated Large-Scale Deletion in Fusarium graminearum Genome Using CRISPR/Cas9 and Its Impact on Growth and Virulence. Journal of Fungi. 2023; 9(6):673. https://doi.org/10.3390/jof9060673
Chicago/Turabian StyleFoster, Adam John, Emily Johnstone, Abbey Saunders, Eva Colic, Nicole Lassel, and Janesse Holmes. 2023. "Unanticipated Large-Scale Deletion in Fusarium graminearum Genome Using CRISPR/Cas9 and Its Impact on Growth and Virulence" Journal of Fungi 9, no. 6: 673. https://doi.org/10.3390/jof9060673
APA StyleFoster, A. J., Johnstone, E., Saunders, A., Colic, E., Lassel, N., & Holmes, J. (2023). Unanticipated Large-Scale Deletion in Fusarium graminearum Genome Using CRISPR/Cas9 and Its Impact on Growth and Virulence. Journal of Fungi, 9(6), 673. https://doi.org/10.3390/jof9060673




