Genotyping Analysis of Cryptococcus deuterogattii and Correlation with Virulence Factors and Antifungal Susceptibility by the Clinical and Laboratory Standards Institute and the European Committee on Antifungal Susceptibility Testing Methods
Abstract
:1. Introduction
2. Materials and Methods
2.1. Isolates and Main Clinical Information
2.2. Determination of Mating Types
2.3. Phylogenetic Analysis
2.4. Minimum Spanning Tree, Multidimensional Scaling Plots, and Split Decomposition Analysis
2.5. Nucleotide Diversity
2.6. Genetic Differentiation Based on Sequences of Populations
2.7. Virulence Factors and Antifungal Susceptibility
2.8. Statistical Analyses
3. Results
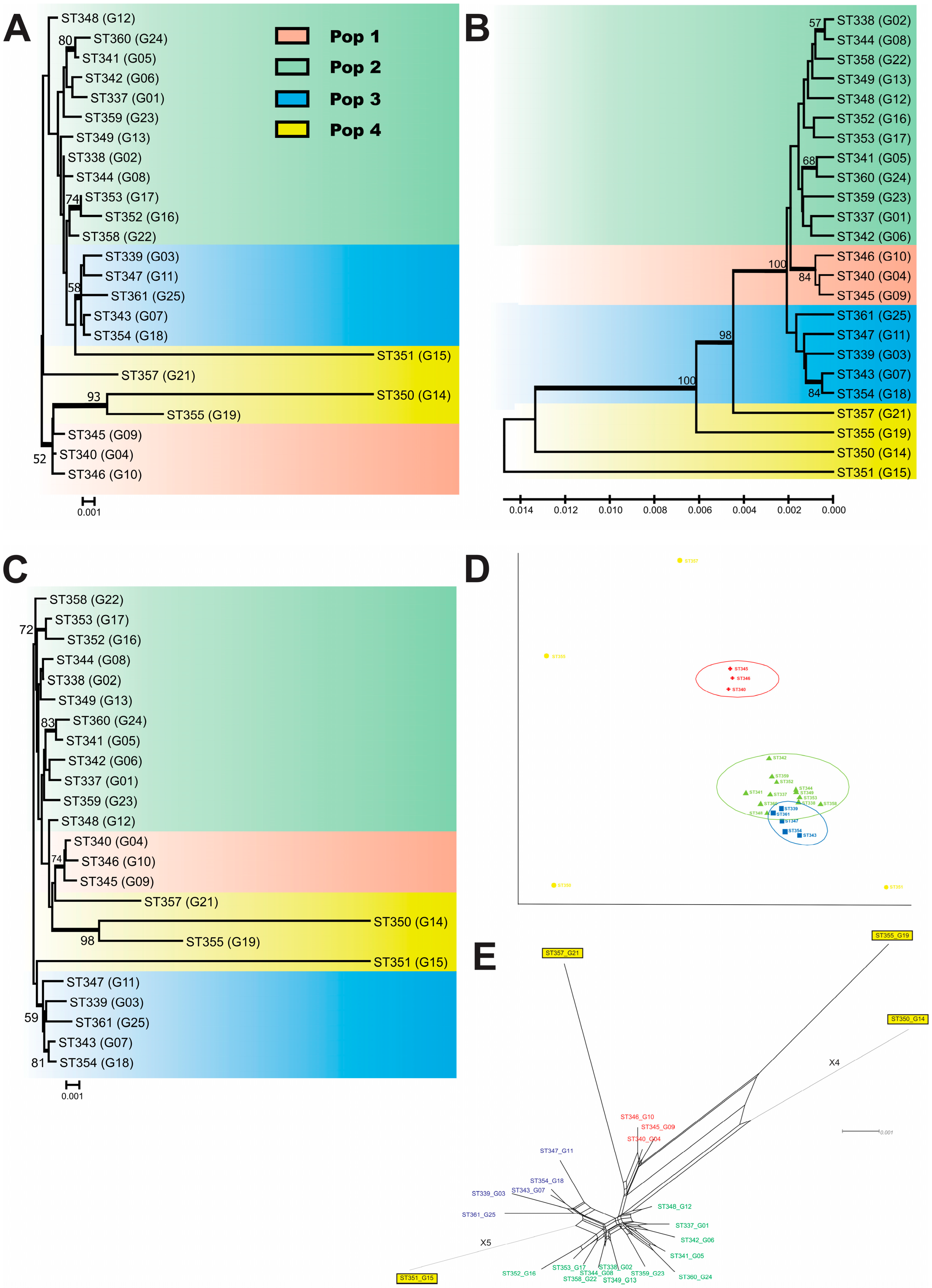
3.1. MLST, Population Structure, and Genetic Variability
3.2. Virulence Factors
3.3. Antifungal Susceptibility
3.4. The Minimum Spanning Tree
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kwon-Chung, K.J.; Fraser, J.A.; Doering, T.L.; Wang, Z.; Janbon, G.; Idnurm, A.; Bahn, Y.S. Cryptococcus neoformans and Cryptococcus gattii, the etiologic agents of cryptococcosis. Cold Spring Harb. Perspect. Med. 2014, 4, a019760. [Google Scholar] [CrossRef]
- Chen, S.C.; Meyer, W.; Sorrell, T.C. Cryptococcus gattii infections. Clin. Microbiol. Rev. 2014, 27, 980–1024. [Google Scholar] [CrossRef]
- Ngamskulrungroj, P.; Gilgado, F.; Faganello, J.; Litvintseva, A.P.; Leal, A.L.; Tsui, K.M.; Mitchell, T.G.; Vainstein, M.H.; Meyer, W. Genetic diversity of the Cryptococcus species complex suggests that Cryptococcus gattii deserves to have varieties. PLoS ONE 2009, 4, e5862. [Google Scholar] [CrossRef]
- Meyer, W. Cryptococcus gattii in the Age of Whole-Genome Sequencing. mBio 2015, 6, e01761-15. [Google Scholar] [CrossRef]
- Hagen, F.; Khayhan, K.; Theelen, B.; Kolecka, A.; Polacheck, I.; Sionov, E.; Falk, R.; Parnmen, S.; Lumbsch, H.T.; Boekhout, T. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. FG B 2015, 78, 16–48. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J.; Bennett, J.E.; Wickes, B.L.; Meyer, W.; Cuomo, C.A.; Wollenburg, K.R.; Bicanic, T.A.; Castaneda, E.; Chang, Y.C.; Chen, J.; et al. The Case for Adopting the “Species Complex” Nomenclature for the Etiologic Agents of Cryptococcosis. mSphere 2017, 2, e00357-16. [Google Scholar] [CrossRef] [PubMed]
- Hagen, F.; Lumbsch, H.T.; Arsic Arsenijevic, V.; Badali, H.; Bertout, S.; Billmyre, R.B.; Bragulat, M.R.; Cabanes, F.J.; Carbia, M.; Chakrabarti, A.; et al. Importance of Resolving Fungal Nomenclature: The Case of Multiple Pathogenic Species in the Cryptococcus Genus. mSphere 2017, 2, e00238-17. [Google Scholar] [CrossRef] [PubMed]
- Farrer, R.A.; Chang, M.; Davis, M.J.; van Dorp, L.; Yang, D.H.; Shea, T.; Sewell, T.R.; Meyer, W.; Balloux, F.; Edwards, H.M.; et al. A New Lineage of Cryptococcus gattii (VGV) Discovered in the Central Zambezian Miombo Woodlands. mBio 2019, 10, e02306-19. [Google Scholar] [CrossRef]
- Beardsley, J.; Dao, A.; Keighley, C.; Garnham, K.; Halliday, C.; Chen, S.C.; Sorrell, T.C. What’s New in Cryptococcus gattii: From Bench to Bedside and Beyond. J. Fungi 2022, 9, 41. [Google Scholar] [CrossRef]
- Monroy-Nieto, J.; Bowers, J.R.; Montfort, P.; Adame, G.; Taverna, C.G.; Yaglom, H.; Sykes, J.E.; Brady, S.; Mochon, A.B.; Meyer, W.; et al. Phylogenomic Placement of American Southwest-Associated Clinical and Veterinary Isolates Expands Evidence for Distinct Cryptococcus gattii VGVI. Microorganisms 2022, 10, 1681. [Google Scholar] [CrossRef]
- Illnait-Zaragozi, M.T.; Martinez-Machin, G.F.; Fernandez-Andreu, C.M.; Perurena-Lancha, M.R.; Hagen, F.; Meis, J.F. Cryptococcus and cryptococcosis in Cuba. A minireview. Mycoses 2014, 57, 707–717. [Google Scholar] [CrossRef] [PubMed]
- Meyer, W.; Aanensen, D.M.; Boekhout, T.; Cogliati, M.; Diaz, M.R.; Esposto, M.C.; Fisher, M.; Gilgado, F.; Hagen, F.; Kaocharoen, S.; et al. Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med. Mycol. 2009, 47, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Saijo, T.; Chen, J.; Chen, S.C.; Rosen, L.B.; Yi, J.; Sorrell, T.C.; Bennett, J.E.; Holland, S.M.; Browne, S.K.; Kwon-Chung, K.J. Anti-granulocyte-macrophage colony-stimulating factor autoantibodies are a risk factor for central nervous system infection by Cryptococcus gattii in otherwise immunocompetent patients. mBio 2014, 5, e00912-14. [Google Scholar] [CrossRef] [PubMed]
- Byrnes, E.J., 3rd; Bildfell, R.J.; Frank, S.A.; Mitchell, T.G.; Marr, K.A.; Heitman, J. Molecular evidence that the range of the Vancouver Island outbreak of Cryptococcus gattii infection has expanded into the Pacific Northwest in the United States. J. Infect. Dis. 2009, 199, 1081–1086. [Google Scholar] [CrossRef]
- Byrnes, E.J., 3rd; Li, W.; Lewit, Y.; Ma, H.; Voelz, K.; Ren, P.; Carter, D.A.; Chaturvedi, V.; Bildfell, R.J.; May, R.C.; et al. Emergence and pathogenicity of highly virulent Cryptococcus gattii genotypes in the northwest United States. PLoS Pathog. 2010, 6, e1000850. [Google Scholar] [CrossRef]
- Engelthaler, D.M.; Hicks, N.D.; Gillece, J.D.; Roe, C.C.; Schupp, J.M.; Driebe, E.M.; Gilgado, F.; Carriconde, F.; Trilles, L.; Firacative, C.; et al. Cryptococcus gattii in North American Pacific Northwest: Whole-population genome analysis provides insights into species evolution and dispersal. mBio 2014, 5, e01464-01414. [Google Scholar] [CrossRef]
- Kidd, S.E.; Hagen, F.; Tscharke, R.L.; Huynh, M.; Bartlett, K.H.; Fyfe, M.; Macdougall, L.; Boekhout, T.; Kwon-Chung, K.J.; Meyer, W. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc. Natl. Acad. Sci. USA 2004, 101, 17258–17263. [Google Scholar] [CrossRef]
- Bielska, E.; May, R.C. What makes Cryptococcus gattii a pathogen? FEMS Yeast Res. 2016, 16, fov106. [Google Scholar] [CrossRef]
- Kronstad, J.W.; Attarian, R.; Cadieux, B.; Choi, J.; D’Souza, C.A.; Griffiths, E.J.; Geddes, J.M.; Hu, G.; Jung, W.H.; Kretschmer, M.; et al. Expanding fungal pathogenesis: Cryptococcus breaks out of the opportunistic box. Nat. Rev. Microbiol. 2011, 9, 193–203. [Google Scholar] [CrossRef]
- Steenbergen, J.N.; Casadevall, A. The origin and maintenance of virulence for the human pathogenic fungus Cryptococcus neoformans. Microbes Infect. 2003, 5, 667–675. [Google Scholar] [CrossRef]
- Barcellos, V.A.; Martins, L.M.S.; Fontes, A.C.L.; Reuwsaat, J.C.V.; Squizani, E.D.; de Sousa Araujo, G.R.; Frases, S.; Staats, C.C.; Schrank, A.; Kmetzsch, L.; et al. Genotypic and Phenotypic Diversity of Cryptococcus gattii VGII Clinical Isolates and Its Impact on Virulence. Front. Microbiol. 2018, 9, 132. [Google Scholar] [CrossRef] [PubMed]
- O’Meara, T.R.; Alspaugh, J.A. The Cryptococcus neoformans capsule: A sword and a shield. Clin. Microbiol. Rev. 2012, 25, 387–408. [Google Scholar] [CrossRef] [PubMed]
- Hagen, F.; Illnait-Zaragozi, M.T.; Bartlett, K.H.; Swinne, D.; Geertsen, E.; Klaassen, C.H.; Boekhout, T.; Meis, J.F. In vitro antifungal susceptibilities and amplified fragment length polymorphism genotyping of a worldwide collection of 350 clinical, veterinary, and environmental Cryptococcus gattii isolates. Antimicrob. Agents Chemother. 2010, 54, 5139–5145. [Google Scholar] [CrossRef] [PubMed]
- Abegg, M.A.; Cella, F.L.; Faganello, J.; Valente, P.; Schrank, A.; Vainstein, M.H. Cryptococcus neoformans and Cryptococcus gattii isolated from the excreta of psittaciformes in a southern Brazilian zoological garden. Mycopathologia 2006, 161, 83–91. [Google Scholar] [CrossRef]
- Espinel-Ingroff, A.; Kidd, S.E. Current trends in the prevalence of Cryptococcus gattii in the United States and Canada. Infect. Drug Resist. 2015, 8, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, N.; DeBess, E.E.; Wohrle, R.; Sun, B.; Nett, R.J.; Ahlquist, A.M.; Chiller, T.; Lockhart, S.R. Correlation of genotype and in vitro susceptibilities of Cryptococcus gattii strains from the Pacific Northwest of the United States. J. Clin. Microbiol. 2010, 48, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Gomes, F.; Arantes, T.D.; Fernandes, J.A.L.; Ferreira, L.C.; Romero, H.; Bosco, S.M.G.; Oliveira, M.T.B.; Del Negro, G.M.B.; Theodoro, R.C. Polymorphism in Mitochondrial Group I Introns among Cryptococcus neoformans and Cryptococcus gattii Genotypes and Its Association with Drug Susceptibility. Front. Microbiol. 2018, 9, 86. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Zang, X.; Xiao, M.; Wang, L.; Wu, H.; Ma, X.; Wu, N.; Deng, H.; Zhou, M.; Pan, L.; et al. Significance of differential expression profiles of ABC transporters in azole susceptibility between Cryptococcus gattii VGI and VGII strains. Med. Mycol. 2022, 60, myac035. [Google Scholar] [CrossRef]
- CLSI (Ed.) Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts; Approved Standard, 3rd ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2008; p. 40. [Google Scholar]
- CLSI (Ed.) Epidemiological Cutoff Values for Antifungal Susceptibility Testing, CLSI Supplement M59 Document, 2nd ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018. [Google Scholar]
- Rodriguez-Tudela, J.L.; Arendrup, M.C.; Barchiesi, F.; Bille, J.; Chryssanthou, E.; Cuenca-Estrella, M.; Dannaoui, E.; Denning, D.W.; Donnelly, J.P.; Dromer, F.; et al. EUCAST definitive document EDef 7.1: Method for the determination of broth dilution MICs of antifungal agents for fermentative yeasts. Clin. Microbiol. Infect. 2008, 14, 398–405. [Google Scholar] [CrossRef]
- Chen, Y.C.; Kuo, S.F.; Lin, S.Y.; Lin, Y.S.; Lee, C.H. Epidemiological and Clinical Characteristics, Antifungal Susceptibility, and MLST-Based Genetic Analysis of Cryptococcus Isolates in Southern Taiwan in 2013–2020. J. Fungi 2022, 8, 287. [Google Scholar] [CrossRef]
- Bellet, V.; Roger, F.; Krasteva, D.; Gouveia, T.; Drakulovski, P.; Pottier, C.; Bertout, S. Multilocus sequence typing of strains from the Cryptococcus gattii species complex from different continents. Mycoses 2022, 65, 88–96. [Google Scholar] [CrossRef]
- Ponzio, V.; Chen, Y.; Rodrigues, A.M.; Tenor, J.L.; Toffaletti, D.L.; Medina-Pestana, J.O.; Colombo, A.L.; Perfect, J.R. Genotypic diversity and clinical outcome of cryptococcosis in renal transplant recipients in Brazil. Emerg. Microbes Infect. 2019, 8, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Nishikawa, M.M.; Almeida-Paes, R.; Brito-Santos, F.; Nascimento, C.R.; Fialho, M.M.; Trilles, L.; Morales, B.P.; da Silva, S.A.; Santos, W.; Santos, L.O.; et al. Comparative antifungal susceptibility analyses of Cryptococcus neoformans VNI and Cryptococcus gattii VGII from the Brazilian Amazon Region by the Etest, Vitek 2, and the Clinical and Laboratory Standards Institute broth microdilution methods. Med. Mycol. 2019, 57, 864–873. [Google Scholar] [CrossRef] [PubMed]
- Silva, D.C.; Martins, M.A.; Szeszs, M.W.; Bonfietti, L.X.; Matos, D.; Melhem, M.S. Susceptibility to antifungal agents and genotypes of Brazilian clinical and environmental Cryptococcus gattii strains. Diagn. Microbiol. Infect. Dis. 2012, 72, 332–339. [Google Scholar] [CrossRef]
- Brito-Santos, F.; Reis, R.S.; Coelho, R.A.; Almeida-Paes, R.; Pereira, S.A.; Trilles, L.; Meyer, W.; Wanke, B.; Lazera, M.D.S.; Gremiao, I.D.F. Cryptococcosis due to Cryptococcus gattii VGII in southeast Brazil: The One Health approach revealing a possible role for domestic cats. Med. Mycol. Case Rep. 2019, 24, 61–64. [Google Scholar] [CrossRef] [PubMed]
- Grizante Bariao, P.H.; Tonani, L.; Cocio, T.A.; Martinez, R.; Nascimento, E.; von Zeska Kress, M.R. Molecular typing, in vitro susceptibility and virulence of Cryptococcus neoformans/Cryptococcus gattii species complex clinical isolates from south-eastern Brazil. Mycoses 2020, 63, 1341–1351. [Google Scholar] [CrossRef]
- Vilas-Boas, A.M.; Andrade-Silva, L.E.; Ferreira-Paim, K.; Mora, D.J.; Ferreira, T.B.; Santos, D.A.; Borges, A.S.; Melhem, M.S.C.; Silva-Vergara, M.L. High genetic variability of clinical and environmental Cryptococcus gattii isolates from Brazil. Med. Mycol. 2020, 58, 1126–1137. [Google Scholar] [CrossRef]
- Meyer, W.; Castaneda, A.; Jackson, S.; Huynh, M.; Castaneda, E. Molecular typing of IberoAmerican Cryptococcus neoformans isolates. Emerg. Infect. Dis. 2003, 9, 189–195. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Francisco, A.P.; Bugalho, M.; Ramirez, M.; Carrico, J.A. Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC Bioinform. 2009, 10, 152. [Google Scholar] [CrossRef] [PubMed]
- Francisco, A.P.; Vaz, C.; Monteiro, P.T.; Melo-Cristino, J.; Ramirez, M.; Carrico, J.A. PHYLOViZ: Phylogenetic inference and data visualization for sequence based typing methods. BMC Bioinform. 2012, 13, 87. [Google Scholar] [CrossRef] [PubMed]
- Tomasini, N.; Lauthier, J.J.; Llewellyn, M.S.; Diosque, P. MLSTest: Novel software for multi-locus sequence data analysis in eukaryotic organisms. Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2013, 20, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 2006, 23, 254–267. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Zaragoza, O.; Casadevall, A. Experimental modulation of capsule size in Cryptococcus neoformans. Biol. Proced. Online 2004, 6, 10–15. [Google Scholar] [CrossRef]
- Aoki, S.; Ito-Kuwa, S.; Nakamura, K.; Kato, J.; Ninomiya, K.; Vidotto, V. Extracellular proteolytic activity of Cryptococcus neoformans. Mycopathologia 1994, 128, 143–150. [Google Scholar] [CrossRef]
- Kanemitsu, K.; Nishino, T.; Kunishima, H.; Okamura, N.; Takemura, H.; Yamamoto, H.; Kaku, M. Quantitative determination of gelatinase activity among enterococci. J. Microbiol. Methods 2001, 47, 11–16. [Google Scholar] [CrossRef]
- Luo, G.; Samaranayake, L.P.; Yau, J.Y. Candida species exhibit differential in vitro hemolytic activities. J. Clin. Microbiol. 2001, 39, 2971–2974. [Google Scholar] [CrossRef]
- Price, M.F.; Wilkinson, I.D.; Gentry, L.O. Plate method for detection of phospholipase activity in Candida albicans. Sabouraudia 1982, 20, 7–14. [Google Scholar] [CrossRef]
- Samaranayake, L.P.; Raeside, J.M.; MacFarlane, T.W. Factors affecting the phospholipase activity of Candida species in vitro. Sabouraudia 1984, 22, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.; Escandon, P.; Castaneda, E. In vitro determination of virulence factors activity associated with several Cryptococcus neoformans clinical isolates. Rev. Iberoam. Micol. 2008, 25, 145–149. [Google Scholar] [CrossRef] [PubMed]
- Pedroso, R.S.; Lavrador, M.A.; Ferreira, J.C.; Candido, R.C.; Maffei, C.M. Cryptococcus neoformans var. grubii—Pathogenicity of environmental isolates correlated to virulence factors, susceptibility to fluconazole and molecular profile. Mem. Inst. Oswaldo Cruz 2010, 105, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Eisenman, H.C.; Mues, M.; Weber, S.E.; Frases, S.; Chaskes, S.; Gerfen, G.; Casadevall, A. Cryptococcus neoformans laccase catalyses melanin synthesis from both D- and L-DOPA. Microbiology 2007, 153, 3954–3962. [Google Scholar] [CrossRef] [PubMed]
- Mednick, A.J.; Nosanchuk, J.D.; Casadevall, A. Melanization of Cryptococcus neoformans affects lung inflammatory responses during cryptococcal infection. Infect. Immun. 2005, 73, 2012–2019. [Google Scholar] [CrossRef] [PubMed]
- Torres-Rodriguez, J.M.; Alvarado-Ramirez, E.; Gutierrez-Gallego, R. Urease activity in Cryptococcus neoformans and Cryptococcus gattii. Rev. Iberoam. Micol. 2008, 25, 27–31. [Google Scholar] [CrossRef]
- Costa, A.K.; Sidrim, J.J.; Cordeiro, R.A.; Brilhante, R.S.; Monteiro, A.J.; Rocha, M.F. Urban pigeons (Columba livia) as a potential source of pathogenic yeasts: A focus on antifungal susceptibility of Cryptococcus strains in Northeast Brazil. Mycopathologia 2010, 169, 207–213. [Google Scholar] [CrossRef]
- Springer, D.J.; Phadke, S.; Billmyre, B.; Heitman, J. Cryptococcus gattii, no longer an accidental pathogen? Curr. Fungal Infect. Rep. 2012, 6, 245–256. [Google Scholar] [CrossRef]
- Ngamskulrungroj, P.; Chang, Y.; Sionov, E.; Kwon-Chung, K.J. The primary target organ of Cryptococcus gattii is different from that of Cryptococcus neoformans in a murine model. mBio 2012, 3, e00103-12. [Google Scholar] [CrossRef]
- Ngamskulrungroj, P.; Price, J.; Sorrell, T.; Perfect, J.R.; Meyer, W. Cryptococcus gattii virulence composite: Candidate genes revealed by microarray analysis of high and less virulent Vancouver island outbreak strains. PLoS ONE 2011, 6, e16076. [Google Scholar] [CrossRef]
- Wiesner, D.L.; Moskalenko, O.; Corcoran, J.M.; McDonald, T.; Rolfes, M.A.; Meya, D.B.; Kajumbula, H.; Kambugu, A.; Bohjanen, P.R.; Knight, J.F.; et al. Cryptococcal genotype influences immunologic response and human clinical outcome after meningitis. mBio 2012, 3, e00196-12. [Google Scholar] [CrossRef] [PubMed]
- Andrade-Silva, L.E.; Ferreira-Paim, K.; Ferreira, T.B.; Vilas-Boas, A.; Mora, D.J.; Manzato, V.M.; Fonseca, F.M.; Buosi, K.; Andrade-Silva, J.; Prudente, B.D.S.; et al. Genotypic analysis of clinical and environmental Cryptococcus neoformans isolates from Brazil reveals the presence of VNB isolates and a correlation with biological factors. PLoS ONE 2018, 13, e0193237. [Google Scholar] [CrossRef] [PubMed]
- Khayhan, K.; Hagen, F.; Pan, W.; Simwami, S.; Fisher, M.C.; Wahyuningsih, R.; Chakrabarti, A.; Chowdhary, A.; Ikeda, R.; Taj-Aldeen, S.J.; et al. Geographically structured populations of Cryptococcus neoformans Variety grubii in Asia correlate with HIV status and show a clonal population structure. PLoS ONE 2013, 8, e72222. [Google Scholar] [CrossRef]
- Miglia, K.J.; Govender, N.P.; Rossouw, J.; Meiring, S.; Mitchell, T.G. Analyses of pediatric isolates of Cryptococcus neoformans from South Africa. J. Clin. Microbiol. 2011, 49, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Firacative, C.; Duan, S.; Meyer, W. Galleria mellonella model identifies highly virulent strains among all major molecular types of Cryptococcus gattii. PLoS ONE 2014, 9, e105076. [Google Scholar] [CrossRef] [PubMed]
- Alspaugh, J.A. Virulence mechanisms and Cryptococcus neoformans pathogenesis. Fungal Genet. Biol. FG B 2015, 78, 55–58. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Rhodes, J.C. Encapsulation and melanin formation as indicators of virulence in Cryptococcus neoformans. Infect. Immun. 1986, 51, 218–223. [Google Scholar] [CrossRef]
- Fu, M.S.; Coelho, C.; De Leon-Rodriguez, C.M.; Rossi, D.C.P.; Camacho, E.; Jung, E.H.; Kulkarni, M.; Casadevall, A. Cryptococcus neoformans urease affects the outcome of intracellular pathogenesis by modulating phagolysosomal pH. PLoS Pathog. 2018, 14, e1007144. [Google Scholar] [CrossRef]
- Olszewski, M.A.; Noverr, M.C.; Chen, G.H.; Toews, G.B.; Cox, G.M.; Perfect, J.R.; Huffnagle, G.B. Urease expression by Cryptococcus neoformans promotes microvascular sequestration, thereby enhancing central nervous system invasion. Am. J. Pathol. 2004, 164, 1761–1771. [Google Scholar] [CrossRef]
- Shi, M.; Colarusso, P.; Mody, C.H. Real-time in vivo imaging of fungal migration to the central nervous system. Cell Microbiol. 2012, 14, 1819–1827. [Google Scholar] [CrossRef]
- Baker, R.P.; Casadevall, A. Reciprocal modulation of ammonia and melanin production has implications for cryptococcal virulence. Nat. Commun. 2023, 14, 849. [Google Scholar] [CrossRef] [PubMed]
- Alanio, A.; Desnos-Ollivier, M.; Garcia-Hermoso, D.; Bretagne, S. Investigating Clinical Issues by Genotyping of Medically Important Fungi: Why and How? Clin. Microbiol. Rev. 2017, 30, 671–707. [Google Scholar] [CrossRef] [PubMed]
- Espinel-Ingroff, A.; Aller, A.I.; Canton, E.; Castanon-Olivares, L.R.; Chowdhary, A.; Cordoba, S.; Cuenca-Estrella, M.; Fothergill, A.; Fuller, J.; Govender, N.; et al. Cryptococcus neoformans-Cryptococcus gattii species complex: An international study of wild-type susceptibility endpoint distributions and epidemiological cutoff values for fluconazole, itraconazole, posaconazole, and voriconazole. Antimicrob. Agents Chemother. 2012, 56, 5898–5906. [Google Scholar] [CrossRef] [PubMed]
- Andrade-Silva, L.; Ferreira-Paim, K.; Mora, D.J.; Da Silva, P.R.; Andrade, A.A.; Araujo, N.E.; Pedrosa, A.L.; Silva-Vergara, M.L. Susceptibility profile of clinical and environmental isolates of Cryptococcus neoformans and Cryptococcus gattii in Uberaba, Minas Gerais, Brazil. Med. Mycol. 2013, 51, 635–640. [Google Scholar] [CrossRef]
- Fang, L.F.; Zhang, P.P.; Wang, J.; Yang, Q.; Qu, T.T. Clinical and microbiological characteristics of cryptococcosis at an university hospital in China from 2013 to 2017. Braz. J. Infect. Dis. Off. Publ. Braz. Soc. Infect. Dis. 2020, 24, 7–12. [Google Scholar] [CrossRef]
- Wu, S.Y.; Kang, M.; Liu, Y.; Chen, Z.X.; Xiao, Y.L.; He, C.; Ma, Y. Molecular epidemiology and antifungal susceptibilities of Cryptococcus species isolates from HIV and non-HIV patients in Southwest China. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2020, 40, 287–295. [Google Scholar] [CrossRef]
- Chen, Y.C.; Chang, T.Y.; Liu, J.W.; Chen, F.J.; Chien, C.C.; Lee, C.H.; Lu, C.H. Increasing trend of fluconazole-non-susceptible Cryptococcus neoformans in patients with invasive cryptococcosis: A 12-year longitudinal study. BMC Infect. Dis. 2015, 15, 277. [Google Scholar] [CrossRef]
- Firacative, C.; Escandon, P. Antifungal susceptibility of clinical Cryptococcus gattii isolates from Colombia varies among molecular types. Med. Mycol. 2021, 59, 1122–1125. [Google Scholar] [CrossRef]
- Trilles, L.; Meyer, W.; Wanke, B.; Guarro, J.; Lazera, M. Correlation of antifungal susceptibility and molecular type within the Cryptococcus neoformans/C. gattii species complex. Med. Mycol. 2011, 50, 328–332. [Google Scholar] [CrossRef]
- Guinea, J.; Hagen, F.; Pelaez, T.; Boekhout, T.; Tahoune, H.; Torres-Narbona, M.; Bouza, E. Antifungal susceptibility, serotyping, and genotyping of clinical Cryptococcus neoformans isolates collected during 18 years in a single institution in Madrid, Spain. Med. Mycol. 2010, 48, 942–948. [Google Scholar] [CrossRef]
- Cheong, J.W.; McCormack, J. Fluconazole resistance in cryptococcal disease: Emerging or intrinsic? Med. Mycol. 2013, 51, 261–269. [Google Scholar] [CrossRef] [PubMed]
- Trilles, L.; Lazera Mdos, S.; Wanke, B.; Oliveira, R.V.; Barbosa, G.G.; Nishikawa, M.M.; Morales, B.P.; Meyer, W. Regional pattern of the molecular types of Cryptococcus neoformans and Cryptococcus gattii in Brazil. Mem. Inst. Oswaldo Cruz 2008, 103, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Desnos-Ollivier, M.; Patel, S.; Raoux-Barbot, D.; Heitman, J.; Dromer, F. Cryptococcosis Serotypes Impact Outcome and Provide Evidence of Cryptococcus neoformans Speciation. mBio 2015, 6, e00311-15. [Google Scholar] [CrossRef] [PubMed]
- Dou, H.T.; Xu, Y.C.; Wang, H.Z.; Li, T.S. Molecular epidemiology of Cryptococcus neoformans and Cryptococcus gattii in China between 2007 and 2013 using multilocus sequence typing and the DiversiLab system. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2015, 34, 753–762. [Google Scholar] [CrossRef] [PubMed]
- Kassi, F.K.; Drakulovski, P.; Bellet, V.; Roger, F.; Chabrol, A.; Krasteva, D.; Doumbia, A.; Landman, R.; Kakou, A.; Reynes, J.; et al. Cryptococcus genetic diversity and mixed infections in Ivorian HIV patients: A follow up study. PLoS Negl. Trop. Dis. 2019, 13, e0007812. [Google Scholar] [CrossRef]
- Leao, C.A.; Ferreira-Paim, K.; Andrade-Silva, L.; Mora, D.J.; da Silva, P.R.; Machado, A.S.; Das Neves, P.F.; Pena, G.S.; de Almeida Teixeira, L.S.; Silva-Vergara, M.L. Primary cutaneous cryptococcosis caused by Cryptococcus gattii in an immunocompetent host. Med. Mycol. 2010, 49, 352–355. [Google Scholar] [CrossRef]
- Desnos-Ollivier, M.; Patel, S.; Spaulding, A.R.; Charlier, C.; Garcia-Hermoso, D.; Nielsen, K.; Dromer, F. Mixed infections and In Vivo evolution in the human fungal pathogen Cryptococcus neoformans. mBio 2010, 1, e00091-10. [Google Scholar] [CrossRef]
- Arsic Arsenijevic, V.; Pekmezovic, M.G.; Meis, J.F.; Hagen, F. Molecular epidemiology and antifungal susceptibility of Serbian Cryptococcus neoformans isolates. Mycoses 2014, 57, 380–387. [Google Scholar] [CrossRef]
- Bertout, S.; Drakulovski, P.; Kouanfack, C.; Krasteva, D.; Ngouana, T.; Dunyach-Remy, C.; Dongtsa, J.; Aghokeng, A.; Delaporte, E.; Koulla-Shiro, S.; et al. Genotyping and antifungal susceptibility testing of Cryptococcus neoformans isolates from Cameroonian HIV-positive adult patients. Clin. Microbiol. Infect. 2013, 19, 763–769. [Google Scholar] [CrossRef]



| Source (n) | Length | S | Π | K | h | Hd | D | FD | FF | FS | Theta-w | Rm | PHI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total (24) | 4180 | 282 | 0.00843 | 34.663 | 24 | 1.000 | −2.218 * | −3.5908 * | −3.708 * | −6.599 | 75.516 | 13 | <0.0001 * |
| Pop1 (3) | 4156 | 9 | 0.00144 | 6.000 | 3 | 1.000 | # | # | # | # | 6.000 | 0 | # |
| Pop2 (12) | 4165 | 38 | 0.00269 | 11.182 | 12 | 1.000 | −0.5062 | −0.2678 | −0.3766 | −4.522 | 12.583 | 8 | 0.07459 |
| Pop3 (5) | 4165 | 27 | 0.00274 | 11.400 | 5 | 1.000 | −0.8977 | −0.8977 | −0.9661 | −0.167 | 12.960 | 1 | 1.0 |
| Pop4 (4) | 4178 | 253 | 0.03192 | 131.33 | 4 | 1.000 | −0.6624 | −0.6125 | −0.6697 | 3.072 | 138.000 | 5 | 0.0004 * |
| BH (6) | 4180 | 235 | 0.02070 | 85.400 | 6 | 1.000 | −1.1961 | −1.2008 | −1.3194 | 1.673 | 102.920 | 6 | 0.0007 * |
| SP (5) | 4163 | 58 | 0.00612 | 25.400 | 5 | 1.000 | −0.6626 | −06626 | −0.7184 | 0.796 | 27.840 | 7 | 0.8632 |
| TM (8) | 4164 | 36 | 0.00317 | 13.179 | 8 | 1.000 | −0.2710 | −0.1162 | −0.1700 | −1.628 | 13.844 | 6 | 0.0142 * |
| Skin (3) | 4164 | 12 | 0.00193 | 8.000 | 3 | 1.000 | # | # | # | # | 8.000 | 0 | @ |
| BAL (3) | 4163 | 16 | 0.00257 | 10.667 | 3 | 1.000 | # | # | # | # | 10.667 | 0 | @ |
| CSF (6) | 4164 | 33 | 0.00343 | 14.267 | 6 | 1.000 | −0.0816 | −01647 | −0.1608 | −0.440 | 14.453 | 5 | 0.0362 * |
| Pop1 | Pop4 | Pop3 | Pop2 | |
|---|---|---|---|---|
| Pop1 | 0.00000 | |||
| Pop4 | 0.02372 | 0.00000 | ||
| Pop3 | 0.59959 * | 0.12670 * | 0.00000 | |
| Pop2 | 0.49339 * | 0.28040 * | 0.24950 * | 0.00000 |
| SOD1 Locus | GPD1 Locus | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AT | Position | AT | Position | ||||||||||||||||
| 19 | 20 | 21 | 22 | 23 | 24 | 25 | 97 | 387 | 396 | 430 | 435 | 527 | 535 | 550 | 637 | 707 | 157 | ||
| AT14 (G8) | A | C | C | G | C | T | A | T | T | G | T | C | C | A | A | C | T | AT21 (G8) | G |
| AT58 (G9) | . | . | . | . | . | . | . | C | C | T | . | T | A | G | G | G | C | AT6 (G9) | . |
| Antifungal | Method | MIC 50 | MIC 90 | Range | WT (%) | Isolates SDD or Intermediate (%) | Isolates Resistants (%) | MIC Geometric Mean |
|---|---|---|---|---|---|---|---|---|
| Amphotericim B | CLSI | 0.12 | 0.25 | 0.03–0.5 | 100 | NA | 0 | 0.188 |
| EUCAST | 0.12 | 0.25 | 0.03–1 | NA | NA | NA | 0.14 | |
| Ketoconazole | CLSI | 0.06 | 0.03 | 0.03–0.12 | NA | NA | 0 | 0.04 |
| EUCAST | 0.03 | 0.06 | 0.03–1 | NA | NA | NA | 0.03 | |
| Itraconazole | CLSI | 0.06 | 0.12 | 0.03–0.12 | 100 | NA | 0 | 0.06 |
| EUCAST | 0.03 | 0.06 | 0.03–0.06 | NA | NA | NA | 0.03 | |
| Fluconazole | CLSI | 2 | 16 | 0.12–16 | 100 | 4 (16.7) | 0 | 2.78 |
| EUCAST | 2 | 8 | 0.12–32 | NA | 11 (45.8) | 4 (16.7) | 1.64 | |
| Voriconazole | CLSI | 0.12 | 0.25 | 0.03–0.25 | 100 | NA | 0 | 0.08 |
| EUCAST | 0.06 | 0.12 | 0.03–0.12 | NA | NA | NA | 0.06 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Andrade-Silva, L.E.; Vilas-Boas, A.; Ferreira-Paim, K.; Andrade-Silva, J.; Santos, D.d.A.; Ferreira, T.B.; Borges, A.S.; Mora, D.J.; Melhem, M.d.S.C.; Silva-Vergara, M.L. Genotyping Analysis of Cryptococcus deuterogattii and Correlation with Virulence Factors and Antifungal Susceptibility by the Clinical and Laboratory Standards Institute and the European Committee on Antifungal Susceptibility Testing Methods. J. Fungi 2023, 9, 889. https://doi.org/10.3390/jof9090889
Andrade-Silva LE, Vilas-Boas A, Ferreira-Paim K, Andrade-Silva J, Santos DdA, Ferreira TB, Borges AS, Mora DJ, Melhem MdSC, Silva-Vergara ML. Genotyping Analysis of Cryptococcus deuterogattii and Correlation with Virulence Factors and Antifungal Susceptibility by the Clinical and Laboratory Standards Institute and the European Committee on Antifungal Susceptibility Testing Methods. Journal of Fungi. 2023; 9(9):889. https://doi.org/10.3390/jof9090889
Chicago/Turabian StyleAndrade-Silva, Leonardo Euripedes, Anderson Vilas-Boas, Kennio Ferreira-Paim, Juliana Andrade-Silva, Daniel de Assis Santos, Thatiana Bragine Ferreira, Aercio Sebastião Borges, Delio Jose Mora, Marcia de Souza Carvalho Melhem, and Mario Léon Silva-Vergara. 2023. "Genotyping Analysis of Cryptococcus deuterogattii and Correlation with Virulence Factors and Antifungal Susceptibility by the Clinical and Laboratory Standards Institute and the European Committee on Antifungal Susceptibility Testing Methods" Journal of Fungi 9, no. 9: 889. https://doi.org/10.3390/jof9090889






