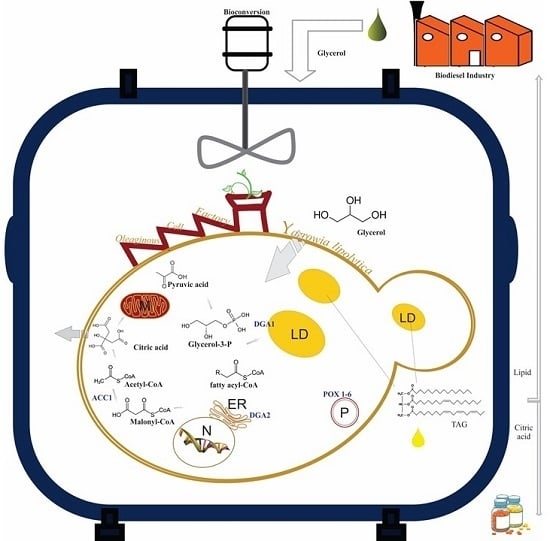
Engineering Yarrowia lipolytica for Enhanced Production of Lipid and Citric Acid
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Culture Condition
2.2. Batch Fermentation
2.3. Genetic Techniques
2.4. Analytical Methods
2.4.1. Dry Biomass
2.4.2. Glycerol and Citric Acid Concentrations
2.4.3. Qualitative and Quantitative Analysis of Lipids
3. Results
3.1. Comparative Time-Course Study
3.1.1. Glycerol Consumption
3.1.2. Biomass Production
3.1.3. Citric Acid Production
3.1.4. Lipid Production
3.2. Fermentation Study
3.3. Comparative Study of LEU2 Expression and Leucine Addition
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Karamerou, E.E.; Theodoropoulos, C.; Webb, C. A biorefinery approach to microbial oil production from glycerol by rhodotorula glutinis. Biomass Bioenergy 2016, 89, 113–122. [Google Scholar] [CrossRef]
- Kolouchová, I.; Maťátková, O.; Sigler, K.; Masák, J.; Řezanka, T. Lipid accumulation by oleaginous and non-oleaginous yeast strains in nitrogen and phosphate limitation. Folia Microbiol. 2016, 61, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Silverman, A.M.; Qiao, K.; Xu, P.; Stephanopoulos, G. Functional overexpression and characterization of lipogenesis-related genes in the oleaginous yeast Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2016, 100, 3781–3798. [Google Scholar] [CrossRef] [PubMed]
- Bellou, S.; Triantaphyllidou, I.-E.; Mizerakis, P.; Aggelis, G. High lipid accumulation in Yarrowia lipolytica cultivated under double limitation of nitrogen and magnesium. J. Biotechnol. 2016, 234, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Seip, J.; Jackson, R.; He, H.; Zhu, Q.; Hong, S.-P. Snf1 is a regulator of lipid accumulation in Yarrowia lipolytica. Appl. Environ. Microbiol. 2013, 79, 7360–7370. [Google Scholar] [CrossRef] [PubMed]
- Probst, K.V.; Schulte, L.R.; Durrett, T.P.; Rezac, M.E.; Vadlani, P.V. Oleaginous yeast: A value-added platform for renewable oils. Crit. Rev. Biotechnol. 2015, 36, 942–955. [Google Scholar] [CrossRef] [PubMed]
- Koutinas, A.A.; Vlysidis, A.; Pleissner, D.; Kopsahelis, N.; Garcia, I.L.; Kookos, I.K.; Papanikolaou, S.; Kwan, T.H.; Lin, C.S.K. Valorization of industrial waste and by-product streams via fermentation for the production of chemicals and biopolymers. Chem. Soc. Rev. 2014, 43, 2587–2627. [Google Scholar] [CrossRef] [PubMed]
- Madzak, C. Yarrowia lipolytica: Recent achievements in heterologous protein expression and pathway engineering. Appl. Microbiol. Biotechnol. 2015, 99, 4559–4577. [Google Scholar] [CrossRef] [PubMed]
- Abghari, A.; Chen, S. Yarrowia lipolytica as oleaginous cell factory platform for the production of fatty acid based biofuel and bioproducts. Front. Bioenergy Res. 2014. [Google Scholar] [CrossRef]
- Liu, H.-H.; Ji, X.-J.; Huang, H. Biotechnological applications of Yarrowia lipolytica: Past, present and future. Biotechnol. Adv. 2015, 33, 1522–1546. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Qiao, K.; Ahn, W.S.; Stephanopoulos, G. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc. Natl. Acad. Sci. USA 2016, 113, 10848–10853. [Google Scholar] [CrossRef] [PubMed]
- Harzevili, F.D. Biotechnological Applications of the Yeast Yarrowia lipolytica; Springer: New York, NY, USA, 2014. [Google Scholar]
- Papanikolaou, S.; Muniglia, L.; Chevalot, I.; Aggelis, G.; Marc, I. Yarrowia lipolytica as a potential producer of citric acid from raw glycerol. J. Appl. Microbiol. 2002, 92, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Moeller, L.; Strehlitz, B.; Aurich, A.; Zehnsdorf, A.; Bley, T. Optimization of citric acid production from glucose by Yarrowia lipolytica. Eng. Life Sci. 2007, 7, 504–511. [Google Scholar] [CrossRef]
- Çelik, G.; Bahriye Uçar, F.; Akpınar, O.; Çorbacı, C. Production of citric and isocitric acid by Yarrowia lipolytica strains grown on different carbon sources. Turkish J. Biochem. 2014, 39, 285–290. [Google Scholar]
- Morgunov, I.G.; Kamzolova, S.V.; Lunina, J.N. The citric acid production from raw glycerol by Yarrowia lipolytica yeast and its regulation. Appl. Microbiol. Biotechnol. 2013, 97, 7387–7397. [Google Scholar] [CrossRef] [PubMed]
- Pfleger, B.F.; Gossing, M.; Nielsen, J. Metabolic engineering strategies for microbial synthesis of oleochemicals. Metab. Eng. 2015, 29, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Amaral, P.F.F.; Ferreira, T.F.; Fontes, G.C.; Coelho, M.A.Z. Glycerol valorization: New biotechnological routes. Food Bioprod. Proc. 2009, 87, 179–186. [Google Scholar] [CrossRef]
- André, A.; Chatzifragkou, A.; Diamantopoulou, P.; Sarris, D.; Philippoussis, A.; Galiotou-Panayotou, M.; Komaitis, M.; Papanikolaou, S. Biotechnological conversions of bio-diesel-derived crude glycerol by Yarrowia lipolytica strains. Eng. Life Sci. 2009, 9, 468–478. [Google Scholar] [CrossRef]
- Rakicka, M.; Lazar, Z.; Dulermo, T.; Fickers, P.; Nicaud, J.M. Lipid production by the oleaginous yeast Yarrowia lipolytica using industrial by-products under different culture conditions. Biotechnol. Biofuels 2015, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kuttiraja, M.; Douha, A.; Valéro, J.R.; Tyagi, R.D. Elucidating the effect of glycerol concentration and C/N ratio on lipid production using Yarrowia lipolytica SKY7. Appl. Biochem. Biotechnol. 2016, 180, 1586–1600. [Google Scholar] [CrossRef] [PubMed]
- Dobrowolski, A.; Mituła, P.; Rymowicz, W.; Mirończuk, A.M. Efficient conversion of crude glycerol from various industrial wastes into single cell oil by yeast Yarrowia lipolytica. Bioresour. Technol. 2016, 207, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Rywińska, A.; Juszczyk, P.; Wojtatowicz, M.; Robak, M.; Lazar, Z.; Tomaszewska, L.; Rymowicz, W. Glycerol as a promising substrate for Yarrowia lipolytica biotechnological applications. Biomass Bioenergy 2013, 48, 148–166. [Google Scholar] [CrossRef]
- Sara, M.; Brar, S.K.; Blais, J.F. Lipid production by Yarrowia lipolytica grown on biodiesel-derived crude glycerol: Optimization of growth parameters and their effects on the fermentation efficiency. RSC Adv. 2016, 6, 90547–90558. [Google Scholar] [CrossRef]
- Ledesma-Amaro, R.; Nicaud, J.-M. Metabolic engineering for expanding the substrate range of Yarrowia lipolytica. Trends Biotechnol. 2016, 34, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Poli, J.S.; da Silva, M.A.N.; Siqueira, E.P.; Pasa, V.M.; Rosa, C.A.; Valente, P. Microbial lipid produced by Yarrowia lipolytica QU21 using industrial waste: A potential feedstock for biodiesel production. Bioresour. Technol. 2014, 161, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Gatter, M.; Förster, A.; Bär, K.; Winter, M.; Otto, C.; Petzsch, P.; Ježková, M.; Bahr, K.; Pfeiffer, M.; Matthäus, F. A newly identified fatty alcohol oxidase gene is mainly responsible for the oxidation of long-chain ω-hydroxy fatty acids in Yarrowia lipolytica. FEMS Yeast Res. 2014, 14, 858–872. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed.; Coldspring-Harbour Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Gajdoš, P.; Nicaud, J.M.; Čertík, M. Glycerol conversion into a single cell oil by engineered Yarrowia lipolytica. Eng. Life Sci. 2016. [Google Scholar] [CrossRef]
- Fickers, P.; Le Dall, M.T.; Gaillardin, C.; Thonart, P.; Nicaud, J.M. New disruption cassettes for rapid gene disruption and marker rescue in the yeast Yarrowia lipolytica. J. Microbiol. Methods 2003, 55, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Lõoke, M.; Kristjuhan, K.; Kristjuhan, A. Extraction of genomic DNA from yeasts for PCR-based applications. Biotechniques 2011, 50, 325. [Google Scholar] [CrossRef] [PubMed]
- O’fallon, J.; Busboom, J.; Nelson, M.; Gaskins, C. A direct method for fatty acid methyl ester synthesis: Application to wet meat tissues, oils, and feedstuffs. J. Anim. Sci. 2007, 85, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Coelho, M.A.Z.; Amaral, P.F.F.; Belo, I. Yarrowia lipolytica: An industrial workhorse. Curr. Res. Technol. Educ. Top. Appl. Microbiol. Microb. Biotechnol. 2010, 2, 930–940. [Google Scholar]
- Kamzolova, S.; Anastassiadis, S.; Fatyhkova, A.; Golovchenko, N.; Morgunov, I. Strain and process development for citric acid production from glycerol-containing waste of biodiesel manufacture. Appl. Microbiol. Biotechnol. 2010, 1020–1028. [Google Scholar]
- Tchakouteu, S.; Kalantzi, O.; Gardeli, C.; Koutinas, A.; Aggelis, G.; Papanikolaou, S. Lipid production by yeasts growing on biodiesel-derived crude glycerol: Strain selection and impact of substrate concentration on the fermentation efficiency. J. Appl. Microbiol. 2015, 118, 911–927. [Google Scholar] [CrossRef] [PubMed]
- Silverman, A.M. Metabolic Engineering Strategies for Increasing Lipid Production in Oleaginous Yeast; Massachusetts Institute of Technology: Cambridge, MA, USA, 2015. [Google Scholar]
- Papanikolaou, S.; Beopoulos, A.; Koletti, A.; Thevenieau, F.; Koutinas, A.A.; Nicaud, J.-M.; Aggelis, G. Importance of the methyl-citrate cycle on glycerol metabolism in the yeast Yarrowia lipolytica. J. Biotechnol. 2013, 168, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Wojtatowicz, M.; Rymowicz, W.; Kautola, H. Comparison of different strains of the yeast Yarrowia lipolytica for citric acid production from glucose hydrol. Appl. Biochem. Biotechnol. 1991, 31, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Fontanille, P.; Kumar, V.; Christophe, G.; Nouaille, R.; Larroche, C. Bioconversion of volatile fatty acids into lipids by the oleaginous yeast Yarrowia lipolytica. Bioresour. Technol. 2012, 114, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Louhasakul, Y.; Cheirsilp, B. Industrial waste utilization for low-cost production of raw material oil through microbial fermentation. Appl. Biochem. Biotechnol. 2013, 169, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Makri, A.; Fakas, S.; Aggelis, G. Metabolic activities of biotechnological interest in Yarrowia lipolytica grown on glycerol in repeated batch cultures. Bioresour. Technol. 2010, 101, 2351–2358. [Google Scholar] [CrossRef] [PubMed]
- Tomaszewska, L.; Rakicka, M.; Rymowicz, W.; Rywińska, A. A comparative study on glycerol metabolism to erythritol and citric acid in Yarrowia lipolytica yeast cells. FEMS Yeast Res. 2014, 14, 966–976. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, F.; Colen, G.; Takahashi, J. Yarrowia lipolytica and its multiple applications in the biotechnological industry. Sci. World J. 2014, 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Dulermo, T.; Nicaud, J.M. Involvement of the G3P shuttle and β-oxidation pathway in the control of TAG synthesis and lipid accumulation in Yarrowia lipolytica. Metab. Eng. 2011, 13, 482–491. [Google Scholar] [CrossRef] [PubMed]
- Beopoulos, A.; Mrozova, Z.; Thevenieau, F.; Le Dall, M.T.; Hapala, I.; Papanikolaou, S.; Chardot, T.; Nicaud, J.M. Control of lipid accumulation in the yeast Yarrowia lipolytica. Appl. Environ. Microbiol. 2008, 74, 7779–7789. [Google Scholar] [CrossRef] [PubMed]
- Fakas, S. Lipid biosynthesis in yeasts: A comparison of the lipid biosynthetic pathway between the model non-oleaginous yeast saccharomyces cerevisiae and the model oleaginous yeast Yarrowia lipolytica. Eng. Life Sci. 2016. [Google Scholar] [CrossRef]
- Tang, X.; Lee, J.; Chen, W.N. Engineering the fatty acid metabolic pathway in saccharomyces cerevisiae for advanced biofuel production. Metab. Eng. Commun. 2015, 2, 58–66. [Google Scholar] [CrossRef]
- Kohlwein, S.D. Triacylglycerol homeostasis: Insights from yeast. J. Biol. Chem. 2010, 285, 15663–15667. [Google Scholar] [CrossRef] [PubMed]
- Beopoulos, A.; Cescut, J.; Haddouche, R.; Uribelarrea, J.L.; Molina-Jouve, C.; Nicaud, J.M. Yarrowia lipolytica as a model for bio-oil production. Prog. Lipid Res. 2009, 48, 375–387. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Damude, H.G.; Yadav, N.S. Three diacylglycerol acyltransferases contribute to oil biosynthesis and normal growth in Yarrowia lipolytica. Yeast 2012, 29, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, J.; Tsakraklides, V.; Kamineni, A.; Greenhagen, E.H.; Consiglio, A.L.; MacEwen, K.; Crabtree, D.V.; Afshar, J.; Nugent, R.L.; Hamilton, M.A. Engineering of a high lipid producing Yarrowia lipolytica strain. Biotechnol. Biofuels 2016, 9, 1. [Google Scholar] [CrossRef] [PubMed]
- Blazeck, J.; Hill, A.; Liu, L.; Knight, R.; Miller, J.; Pan, A.; Otoupal, P.; Alper, H.S. Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Ledesma-Amaro, R.; Nicaud, J.-M. Yarrowia lipolytica as a biotechnological chassis to produce usual and unusual fatty acids. Prog. Lipid Res. 2016, 61, 40–50. [Google Scholar] [CrossRef] [PubMed]
- Tai, M.; Stephanopoulos, G. Engineering the push and pull of lipid biosynthesis in oleaginous yeast Yarrowia lipolytica for biofuel production. Metab. Eng. 2013, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Beopoulos, A.; Haddouche, R.; Kabran, P.; Dulermo, T.; Chardot, T.; Nicaud, J.-M. Identification and characterization of DGA2, an acyltransferase of the DGAT1 acyl-CoA: Diacylglycerol acyltransferase family in the oleaginous yeast Yarrowia lipolytica. New insights into the storage lipid metabolism of oleaginous yeasts. Appl. Microbiol. Biotechnol. 2012, 93, 1523–1537. [Google Scholar] [CrossRef] [PubMed]
- Gajdoš, P.; Nicaud, J.-M.; Rossignol, T.; Čertík, M. Single cell oil production on molasses by Yarrowia lipolytica strains overexpressing DGA2 in multicopy. Appl. Microbiol. Biotechnol. 2015, 99, 8065–8074. [Google Scholar] [CrossRef] [PubMed]
- Gajdoš, P.; Ledesma-Amaro, R.; Nicaud, J.-M.; Čertík, M.; Rossignol, T. Overexpression of diacylglycerol acyltransferase in Yarrowia lipolytica affects lipid body size, number and distribution. FEMS Yeast Res. 2016, 16, fow062. [Google Scholar] [CrossRef] [PubMed]
- Xue, Z.; Sharpe, P.L.; Hong, S.-P.; Yadav, N.S.; Xie, D.; Short, D.R.; Damude, H.G.; Rupert, R.A.; Seip, J.E.; Wang, J. Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat. Biotechnol. 2013, 31, 734–740. [Google Scholar] [CrossRef] [PubMed]
- Petrie, J.R.; Vanhercke, T.; Shrestha, P.; Liu, O.; Singh, S.P. Methods of Producing Lipids. U.S. Patent 9,127,288, September 2015. [Google Scholar]
- Athenstaedt, K.; Daum, G. The life cycle of neutral lipids: Synthesis, storage and degradation. Cell. Mol. Life Sci. CMLS 2006, 63, 1355–1369. [Google Scholar] [CrossRef] [PubMed]
- Dulermo, T.; Tréton, B.; Beopoulos, A.; Gnankon, A.P.K.; Haddouche, R.; Nicaud, J.-M. Characterization of the two intracellular lipases of Y. lipolytica encoded by TGL3 and TGL4 genes: New insights into the role of intracellular lipases and lipid body organisation. Biochim. Biophys. Acta 2013, 1831, 1486–1495. [Google Scholar] [CrossRef] [PubMed]
- Papanikolaou, S.; Chatzifragkou, A.; Fakas, S.; Galiotou-Panayotou, M.; Komaitis, M.; Nicaud, J.M.; Aggelis, G. Biosynthesis of lipids and organic acids by Yarrowia lipolytica strains cultivated on glucose. Eur. J. Lipid Sci. Technol. 2009, 111, 1221–1232. [Google Scholar] [CrossRef]
- Shi, S.; Chen, Y.; Siewers, V.; Nielsen, J. Improving production of malonyl coenzyme a-derived metabolites by abolishing snf1-dependent regulation of acc1. MBio 2014, 5, e01130-14. [Google Scholar] [CrossRef] [PubMed]
- Sabra, W.; Bommareddy, R.R.; Maheshwari, G.; Papanikolaou, S.; Zeng, A.-P. Substrates and oxygen dependent citric acid production by Yarrowia lipolytica: Insights through transcriptome and fluxome analyses. Microb. Cell Factories 2017, 16, 78. [Google Scholar] [CrossRef] [PubMed]
- Desfougères, T.; Haddouche, R.; Fudalej, F.; Neuvéglise, C.; Nicaud, J.-M. Soa genes encode proteins controlling lipase expression in response to triacylglycerol utilization in the yeast Yarrowia lipolytica. FEMS Yeast Res. 2009, 10, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Dulermo, R.; Gamboa-Meléndez, H.; Ledesma-Amaro, R.; Thévenieau, F.; Nicaud, J.-M. Unraveling fatty acid transport and activation mechanisms in Yarrowia lipolytica. Biochim. Biophys. Acta 2015, 1851, 1202–1217. [Google Scholar] [CrossRef] [PubMed]
- Kamisaka, Y.; Tomita, N.; Kimura, K.; Kainou, K.; Uemura, H. DGA1 (diacylglycerol acyltransferase gene) overexpression and leucine biosynthesis significantly increase lipid accumulation in the Δsnf2 disruptant of saccharomyces cerevisiae. Biochem. J. 2007, 408, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Lian, J.; Zhao, H. Metabolic engineering of saccharomyces cerevisiae to improve 1-hexadecanol production. Metab. Eng. 2015, 27, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Kerkhoven, E.J.; Pomraning, K.R.; Baker, S.E.; Nielsen, J. Regulation of amino-acid metabolism controls flux to lipid accumulation in Yarrowia lipolytica. Syst. Biol. Appl. 2016, 2, 16005. [Google Scholar] [CrossRef] [Green Version]
- Nambou, K.; Zhao, C.; Wei, L.; Chen, J.; Imanaka, T.; Hua, Q. Designing of a “cheap to run” fermentation platform for an enhanced production of single cell oil from Yarrowia lipolytica DSM3286 as a potential feedstock for biodiesel. Bioresour. Technol. 2014, 173, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.C.; Mi, L.; Pontrelli, S.; Luo, S. Fuelling the future: Microbial engineering for the production of sustainable biofuels. Nat. Rev. Microbiol. 2016, 14, 288–304. [Google Scholar] [CrossRef] [PubMed]
- Hofbauer, H.F.; Schopf, F.H.; Schleifer, H.; Knittelfelder, O.L.; Pieber, B.; Rechberger, G.N.; Wolinski, H.; Gaspar, M.L.; Kappe, C.O.; Stadlmann, J. Regulation of gene expression through a transcriptional repressor that senses acyl-chain length in membrane phospholipids. Dev. Cell 2014, 29, 729–739. [Google Scholar] [CrossRef] [PubMed]
- Qiao, K.; Abidi, S.H.I.; Liu, H.; Zhang, H.; Chakraborty, S.; Watson, N.; Ajikumar, P.K.; Stephanopoulos, G. Engineering lipid overproduction in the oleaginous yeast Yarrowia lipolytica. Metab. Eng. 2015, 29, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Pan, A.; Spofford, C.; Zhou, N.; Alper, H.S. An evolutionary metabolic engineering approach for enhancing lipogenesis in Yarrowia lipolytica. Metab. Eng. 2015, 29, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Egermeier, M.; Russmayer, H.; Sauer, M.; Marx, H. Metabolic flexibility of Yarrowia lipolytica growing on glycerol. Front. Microbiol. 2017, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Papanikolaou, S.; Aggelis, G. Selective uptake of fatty acids by the yeast Yarrowia lipolytica. Eur. J. Lipid Sci. Technol. 2003, 105, 651–655. [Google Scholar] [CrossRef]
- Petschnigg, J.; Wolinski, H.; Kolb, D.; Zellnig, G.; Kurat, C.F.; Natter, K.; Kohlwein, S.D. Good fat, essential cellular requirements for triacylglycerol synthesis to maintain membrane homeostasis in yeast. J. Biol. Chem. 2009, 284, 30981–30993. [Google Scholar] [CrossRef] [PubMed]
- Crown, S.B.; Marze, N.; Antoniewicz, M.R. Catabolism of branched chain amino acids contributes significantly to synthesis of odd-chain and even-chain fatty acids in 3T3-L1 adipocytes. PLoS ONE 2015, 10, e0145850. [Google Scholar] [CrossRef] [PubMed]
- Pomraning, K.R.; Kim, Y.-M.; Nicora, C.D.; Chu, R.K.; Bredeweg, E.L.; Purvine, S.O.; Hu, D.; Metz, T.O.; Baker, S.E. Multi-omics analysis reveals regulators of the response to nitrogen limitation in Yarrowia lipolytica. BMC Genomics 2016, 17, 138. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wu, C.; Wu, Q.; Dai, J.; Song, Y. Metabolic flux analysis of lipid biosynthesis in the yeast Yarrowia lipolytica using 13C-Labled glucose and gas chromatography-mass spectrometry. PLoS ONE 2016, 11, e0159187. [Google Scholar] [CrossRef] [PubMed]









| Y. Lipolytica Strain Names | Strain Genotypes | Gene Configurations | Reference |
|---|---|---|---|
| H222 (H) | MatA mating type | [27] | |
| H222ΔP leu+ ura− (HP-U) | MATA ura3-302::SUC2 ΔPOX1–6 | [27] | |
| H222ΔP leu+ ura+ (HP) | HP-U, ΔPOX3::URA3 | loxR-URA3-loxP flanked by POX3 homologous up/down stream sequences | This study |
| H222ΔP ΔL + DGA1 DGA2 leu− ura+ (HPDD) | HP, ΔLEU2 + DGA1 + DGA2::URA3 | loxR-URA3-loxP flanked by LEU2 homologous upstream and pFBA-DGA1-tLip1 pTEF-DGA2-tXPR2 LEU2 homologous downstream sequences | This study |
| H222ΔP ΔL + DGA1 DGA2 ΔSNF1 leu− ura+ (HPDDS) | HPDD, ΔSNF1::URA3 | loxR-URA3-loxP flanked by SNF1 up/down homologous stream sequences | This study |
| Vector Names | Features |
|---|---|
| Cre-recombinase (CR) | Shuttle vector carrying leucine marker, Cre recombinase flanked by TEFin promoter and Xpr2 terminator |
| pGR12 (L) | Shuttle empty vector carrying leucine marker, FBA promoter and lip1 terminator, used for study of leucine biosynthesis |
| POX3 Ura (PU) | Uracil selection marker flanked by POX3 upstream and downstream homologous sequences, used for construction of HP strain |
| LEU2 Ura (LU) | Uracil selection marker flanked by LEU2 upstream and downstream homologous sequences, used for construction of LDD vector |
| SNF1 Ura (SU) | Uracil selection marker flanked by SNF1 upstream and downstream homologous sequences, used for construction of HPDDS strain |
| pGR12 DGA1 (D1) | Single gene centromeric shuttle replicative vector with leucine selection marker, DGA1 gene cloned between FBA promoter and lip1 terminator, used for double gene expression cassette construction |
| pJN44 DGA2 (D2) | Single gene centromeric shuttle replicative vector with leucine selection marker, DGA2 gene cloned between TEFin promoter and xpr2 terminator, used for double gene expression cassette construction |
| DGA1 DGA2 (DD) | Double gene centromeric shuttle replicative vector with leucine selection marker, used for construction of LDD vector |
| LEU2 DGA1 DGA2 (LDD) | Uracil selection marker flanked by LEU2 homologous upstream sequence and combination of double gene expression cassettes and LEU2 homologous downstream sequence, used for construction of HPDD strain |
| No. | Name | Sequence (5′―›3′, Underlined Restriction Site) |
|---|---|---|
| 1 | POX3 up F ApaI | CTATAGGGCCCCTGGGCTGTTCGGTCGA |
| 2 | POX3 up R XbaI | GATCCTCTAGAAGGACGCACAACGCC |
| 3 | POX3 down F SpeI | CTGGACTAGTCGCTCCCATTGGAAACTACGA |
| 4 | POX3 down R NdeI | CCTCACATATGTCTCTTCGCTGTGGTCTAGG |
| 5 | POX3 F Ura | GTCTCTACTTGTAGTTCTGTAGACAGACT |
| 6 | POX3 Ura R | GAAGAATGTATCGTCAAAGTGATCCAAG |
| 7 | POX3 Ura F | TGACTTGTGTATGACTTATTCTCAACTACA |
| 8 | POX3 R Ura | AGATGCGTGATAGATTACTTGGATTTAGT |
| 9 | DGA1 F HindIII | GAGCGAAAGCTTATGACTATCGACTCACAATACTACAAGT |
| 10 | DGA1 R SalI | GTTCAAGTCGACTTACTCAATCATTCGGAACTCTGGG |
| 11 | DGA2 F HindIII | GCAAGGAAGCTTATGGAAGTCCGACGACGA |
| 12 | DGA2 R PstI | ATGCTACTGCAGCTACTGGTTCTGCTTGTAGTTGT |
| 13 | LEU2 up F ApaI | CTATAGGGCCC ACCGGCAAGATCTCGTTAAGACAC |
| 14 | LEU2 up R XbaI | GATCCTCTAGATGTGTGTGGTTGTATGTGTGATGTGG |
| 15 | LEU2 down F SpeI | CTGGACTAGTCTCTATAAAAAGGGCCCAGCCCTG |
| 16 | LEU2 down R NdeI | CCTCACATATG GACAGCCTTGACAACTTGGTTGTTG |
| 17 | LEU2 F Ura | TACAGTTGTAACTATGGTGCTTATCTGGG |
| 18 | LEU2 Ura R | CCTTGGGAACCACCACCGT |
| 19 | LEU2 Ura F | ACTTCCTGGAGGCAGAAGAACTT |
| 20 | LEU2 R Ura | ATAGCAAATTTAGTCGTCGAGAAAGGGTC |
| 21 | SNF1 up F ApaI | CAATTGGGCCCGTGATCAAAGCATGAGATACTGTCAAGG |
| 22 | SNF1 up R XbaI | GATCCTCTAGAGAGGTGGTGGAAGGAGTGGTATGTAGTC |
| 23 | SNF1 down F SpeI | CTGGACTAGT TCATTAATACGTTTCCCTGGTG |
| 24 | SNF1 down R NdeI | CCTCACATATGGGAATTCGTGCAGAAGAACA |
| 25 | SNF1 F Ura | GCGGGAAATCAAGATTGAGA |
| 26 | SNF1 Ura R | CGGTCCATTTCTCACCAACT |
| 27 | SNF1 Ura F | CCTGGAGGCAGAAGAACTTG |
| 28 | SNF1 R Ura | ACTACTGGCGGACTTTGTGG |
| Strain and Culture Medium | Residual Glycerol | DCW (g/L) | Citric Acid (g/L) | Lipid (g/L) |
|---|---|---|---|---|
| HPDDS ura+ leu−, YNB-Ura | 4.93 ± 0.74 a | 7.75 ± 0.89 a | 28.36 ± 4.36 a | 3.6 ± 0.18 a |
| HPDDS ura+ pGR12-leucine, YNB-Leu | 2.84 ± 0.45 b | 6.73 ± 0.24 a | 14.21 ± 1.12 b | 1.99 ± 0.15 b |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abghari, A.; Chen, S. Engineering Yarrowia lipolytica for Enhanced Production of Lipid and Citric Acid. Fermentation 2017, 3, 34. https://doi.org/10.3390/fermentation3030034
Abghari A, Chen S. Engineering Yarrowia lipolytica for Enhanced Production of Lipid and Citric Acid. Fermentation. 2017; 3(3):34. https://doi.org/10.3390/fermentation3030034
Chicago/Turabian StyleAbghari, Ali, and Shulin Chen. 2017. "Engineering Yarrowia lipolytica for Enhanced Production of Lipid and Citric Acid" Fermentation 3, no. 3: 34. https://doi.org/10.3390/fermentation3030034