Microbial Diversity of Marula Wine during Spontaneous Fermentation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Preparation of Marula Wine
2.2. Sampling of the Marula Wine
2.3. Isolation of Bacteria from Marula Wine
2.4. Isolation of Yeasts from the Marula Wine
2.5. Identification of the Microbial Isolates
- a.
- MALDI-TOF Biotyping Technology
- b.
- Next-Generation Sequencing Technology
- DNA Extraction and Amplification of the 16S rDNA and ITS Regions
- Sequence Processing, Operational Taxonomic Units (OTUs) Clustering
2.6. Characterisation of Common Microbiota from Different Marula Wines
3. Results
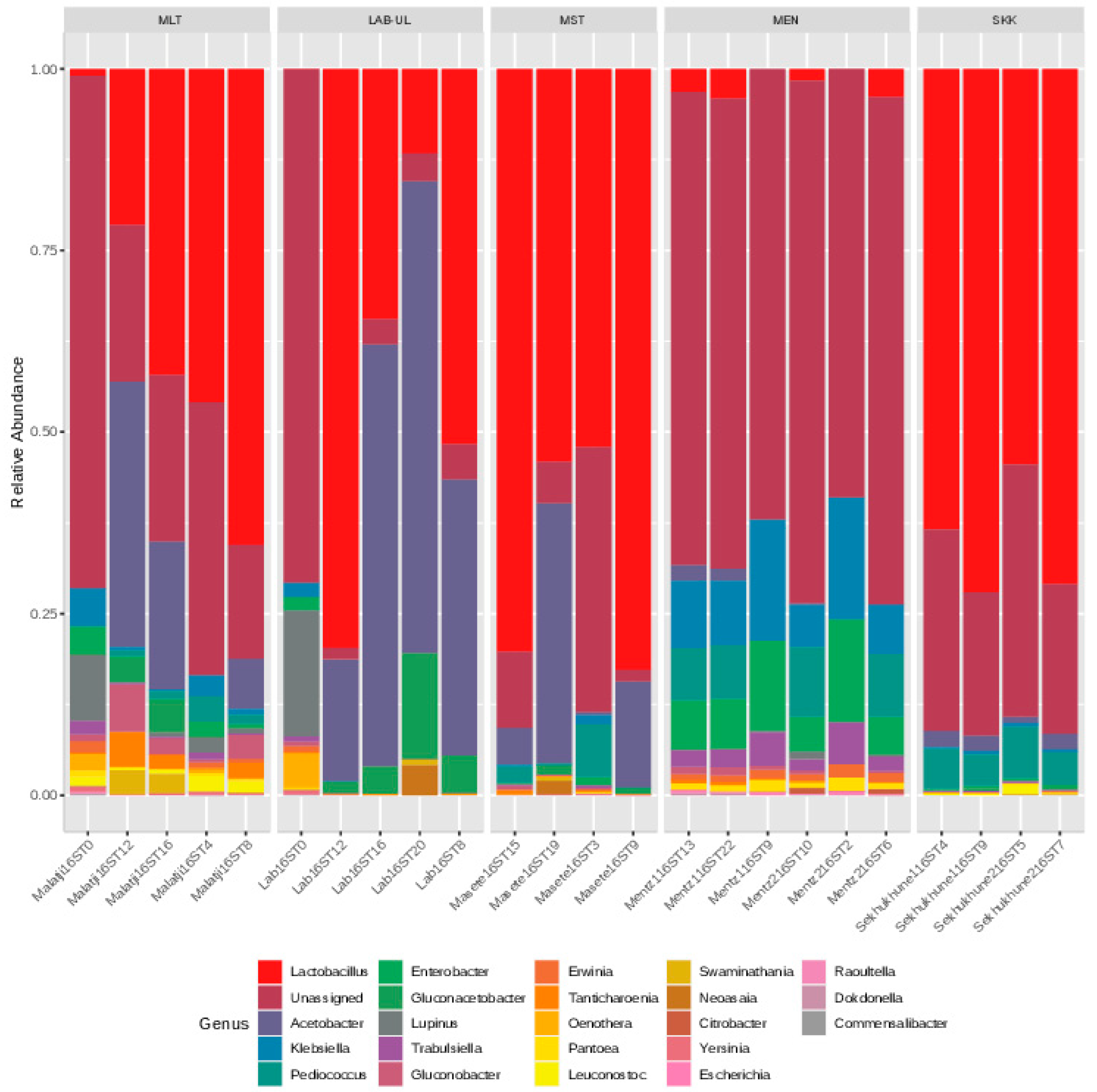
3.1. Bacterial Microbiome in the Marula Wines
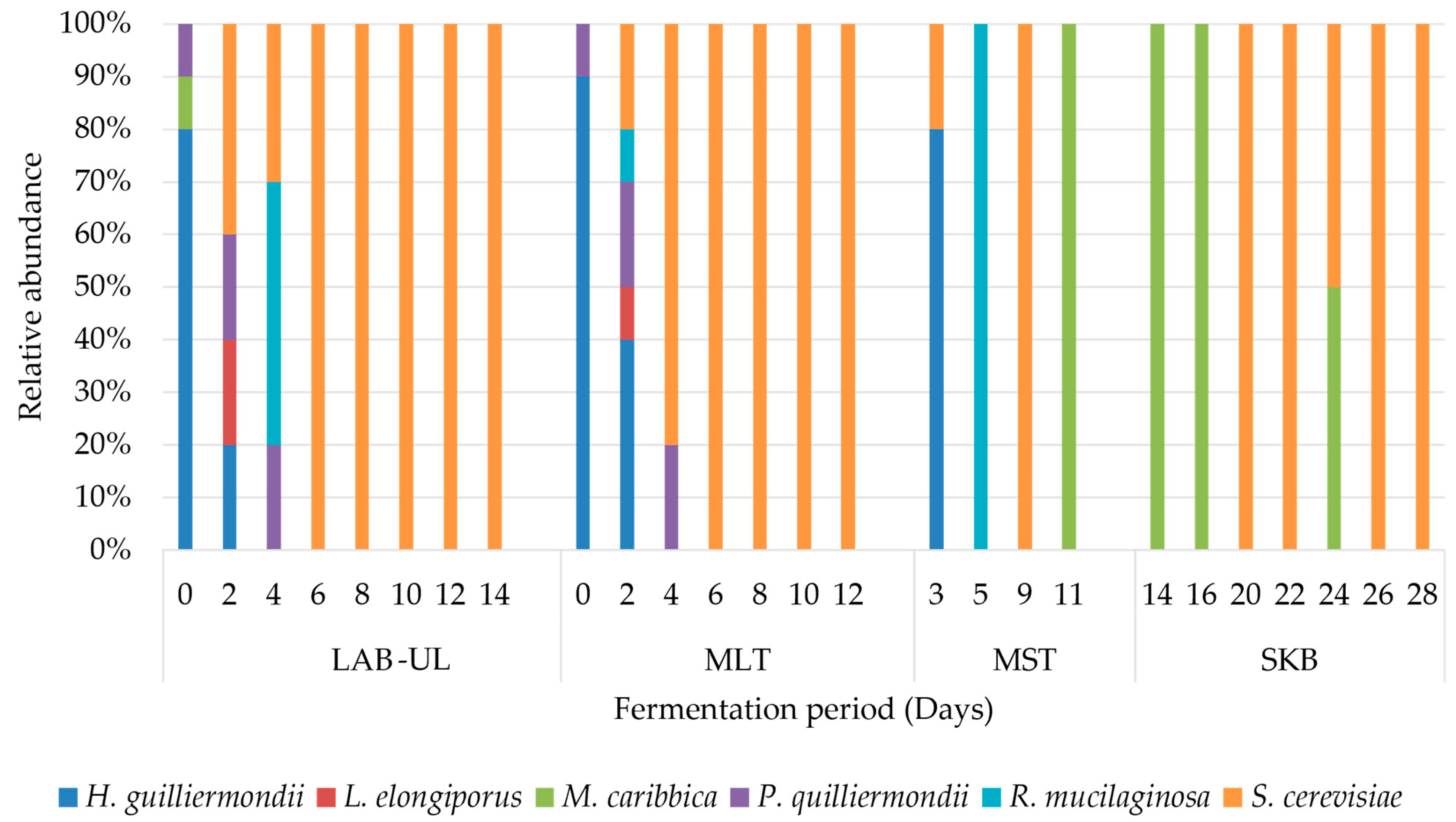
3.2. Yeast Microbiota Present in the Wine during Fermentation
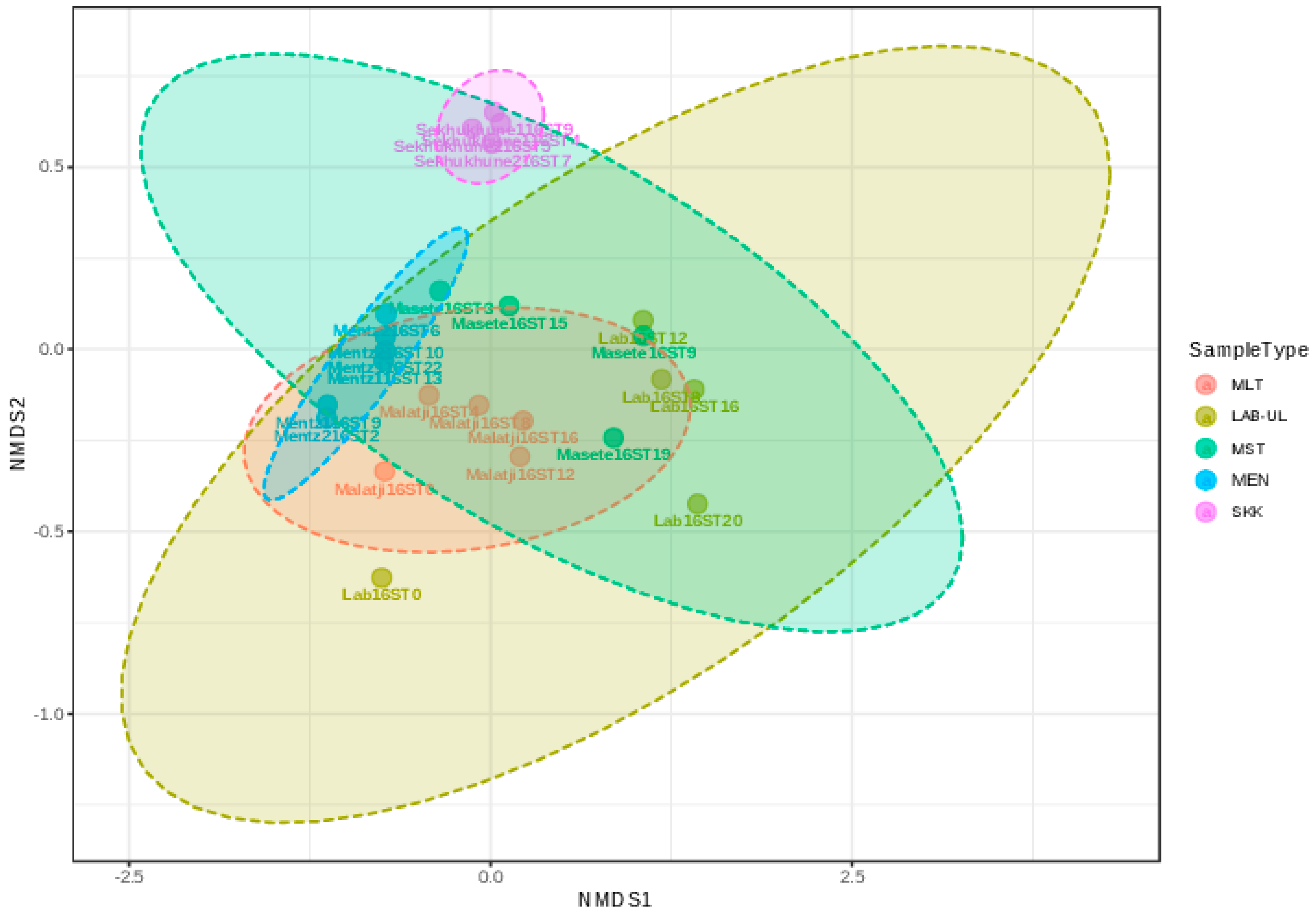
3.3. Microbial Taxonomic Composition of Various Marula Wine during Fermentation
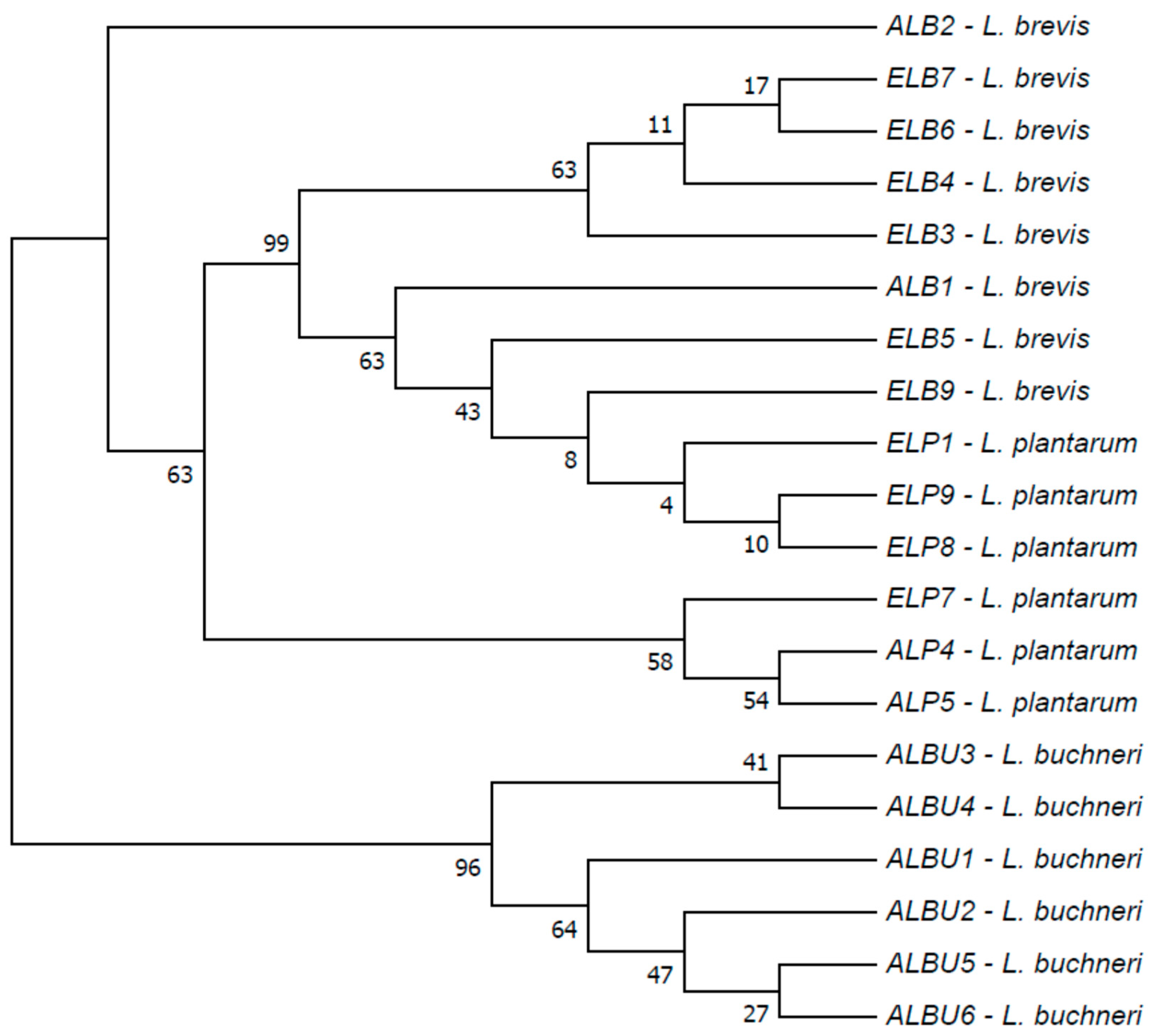
3.4. Characterisation of Common Microbiota from Different Marula Wines
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xaba, P.; Moll, E. Marula: Gardening with traditionally useful indigenous plants. Veld Flora 2011, 97, 76–77. [Google Scholar]
- Hiwilepo-van Hal, P.; Bille, P.G.; Verkerk, R.; Dekker, M. The effect of temperature and time on the quality of naturally fermented marula (Sclerocarya birrea subsp. caffra) juice. LWT-Food Sci. Technol. 2013, 53, 70–75. [Google Scholar] [CrossRef]
- Simatende, P.; Gadaga, T.H.; Nkambule, S.J.; Siwela, M. Methods of preparation of Swazi traditional fermented foods. J. Ethnic Foods 2015, 2, 119–125. [Google Scholar] [CrossRef]
- Nyanga, L.K.; Nout, M.J.; Gadaga, T.H.; Theelen, B.; Boekhout, T.; Zwietering, M.H. Yeasts and lactic acid bacteria microbiota from masau (Ziziphus mauritiana) fruits and their fermented fruit pulp in Zimbabwe. Int. J. Food Microbiol. 2007, 120, 159–166. [Google Scholar] [CrossRef] [PubMed]
- De Vuyst, L.; Weckx, S. The cocoa bean fermentation process: From ecosystem analysis to starter culture development. J. Appl. Microbiol. 2016, 121, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Barata, A.; Malfeito-Ferreira, M.; Loureiro, V. The microbial ecology of wine grape berries. Int. J. Food Microbiol. 2012, 153, 243–259. [Google Scholar] [CrossRef]
- Maicas, S. The role of yeasts in fermentation processes. Microorganism 2020, 8, 1142. [Google Scholar] [CrossRef] [PubMed]
- Bartowsky, E.J. Bacterial spoilage of wine and approaches to minimize it. Lett. Appl. Microbiol. 2009, 48, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Bleve, G.; Tufariello, M.; Vetrano, C.; Mita, G.; Grieco, F. Simultaneous alcoholic and malolactic fermentations by Saccharomyces cerevisiae and Oenococcus oeni cells co-immobilized in alginate beads. Front. Microbiol. 2016, 7, 943. [Google Scholar] [CrossRef]
- Langer, E.M. Molecular Ferment: The Rise and Proliferation of Yeast Model Organism Research; University of California: San Francisco, CA, USA, 2016. [Google Scholar]
- Liu, W.; Li, H.; Jiang, D.; Zhang, Y.; Zhang, S.; Sun, S. Effect of Saccharomyces cerevisiae, Torulaspora delbrueckii and malolactic fermentation on fermentation kinetics and sensory property of black raspberry wines. Food Microbiol. 2020, 91, 103551. [Google Scholar] [CrossRef]
- Bartowsky, E.J.; Henschke, P.A. Acetic acid bacteria spoilage of bottled red wine a review. Int. J. Food Microbiol. 2008, 125, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Juvonen, R.; Virkajarv, I.V.; Priha, O.; Laitila, A. Microbiological Spoilage and Safety Risks in Non-Beer Beverages; VTT Tiedotteita—Research Notes: Espoo, Finland, 2011; pp. 29–34. [Google Scholar]
- Gadaga, T.H.; Mutukumira, A.N.; Narvhus, J.A.; Feresu, S.B. A review of traditional fermented foods and beverages of Zimbabwe. Int. J. Food Microbiol. 1999, 53, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Nwonwu, F.O. The socio-economic and economic relevance of the marula tree and its sustainable use in South Africa. Afr. Insight 2006, 36, 249–265. [Google Scholar]
- Kutyauripo, J.; Parawira, W.; Tinofa, S.; Kudita, I.; Ndengu, C. Investigation of shelf-life extension of sorghum beer (Chibuku) by removing the second conversion of malt. Int. J. Food Microbiol. 2009, 129, 271–276. [Google Scholar] [CrossRef]
- Bottari, B. Culture Independent Approach for the Evaluation of Presence and Activity of Microorganisms in Food. Doctoral Thesis, Università degli Studi di Parma, Dipartimento di Sanità Pubblica, Italy, 2009. Available online: https://hdl.handle.net/1889/1024 (accessed on 5 March 2023).
- Calmin, G.; Lefort, F.; Belbahri, L. Multi-Loci Sequence Typing (MLST) for two lacto-acid bacteria (LAB) species: Pediococcus áparvulus and P. ádamnosus. Mol. Biotechnol. 2008, 40, 170–179. [Google Scholar] [CrossRef]
- Dušková, M.; Šedo, O.; Kšicová, K.; Zdráhal, Z.; Karpíšková, R. Identification of Lactobacilli isolated from food by genotypic methods and MALDI-TOF MS. Int. J. Food Microbiol. 2012, 159, 107–114. [Google Scholar] [CrossRef]
- Carvalho, B.F.; Ávila, C.L.S.; Bernardes, T.F.; Pereira, M.N.; Santos, C.; Schwan, R.F. Fermentation profile and identification of lactic acid bacteria and yeasts of rehydrated corn kernel silage. J. Appl. Microbiol. 2017, 122, 589–600. [Google Scholar] [CrossRef]
- Rodrigues, N.P.A.; Garcia, E.F.; De Souza, E.L. Selection of Lactic Acid Bacteria with Promising Probiotic Aptitudes from Fruit and Ability to Survive in Different Food Matrices. Braz. J. Microbiol. 2021, 52, 2257–2269. [Google Scholar] [CrossRef]
- Bruker Daltonik, G. Bruker Guide to MALDI Sample Preparation. 2015. Available online: https://researchservices.pitt.edu/sites/default/files/Bruker_Guide%20for%20MALDI_Sample_Preparation.pdf (accessed on 10 July 2016).
- White, T.J.; Bruns, T.; Lee, S.J.W.T.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Elsevier: Amsterdam, The Netherlands, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Lane, D.J. 16S/23S rRNA Sequencing. In Nucleic Acid Techniques in Bacterial Systematic; Stackebrandt, E., Goodfellow, M., Eds.; John Wiley and Sons: New York, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Masella, A.P.; Bartram, A.K.; Truszkowski, J.; Brown, D.G.; Neufeld, J.D. PANDAseq: Paired-End Assembler for Illumina Sequences. BMC Bioinform. 2012, 13, 31. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and Clustering Orders of Magnitude Faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Ashenafi, M. A review on the microbiology of indigenous fermented foods and beverages of Ethiopia. Ethiop. J. Biol. Sci. 2006, 5, 189–245. [Google Scholar]
- Walker, G.M.; Stewart, G.G. Saccharomyces cerevisiae in the production of fermented beverages. Beverages 2016, 2, 30. [Google Scholar] [CrossRef]
- Motlhanka, K.; Zhou, N.; Lebani, K. Microbial and chemical diversity of traditional non-cereal based alcoholic beverages of Sub-Saharan Africa. Beverages 2018, 4, 36. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Bamforth, C.W. The microbiology of malting and brewing. Microbiol. Mol. Biol. Rev. 2013, 77, 157–172. [Google Scholar] [CrossRef]
- Felšöciová, S.; Kowalczewski, P.Ł.; Krajčovič, T.; Dráb, Š.; Kačániová, M. Quantitative and Qualitative Composition of Bacterial Communities of Malting Barley Grain and Malt during Long-Term Storage. Agronomy 2020, 10, 1301. [Google Scholar] [CrossRef]
- Nemo, R.; Bacha, K. Microbial, Physicochemical and Proximate Analysis of Selected Ethiopian Traditional Fermented Beverages. Lebensm.-Wiss. Technol. 2020, 131, 109713. [Google Scholar] [CrossRef]
- Schutte, L.M. Isolation and Identification of the Microbial Consortium Present in Fermented Milks from Sub-Saharan Africa. Masters of Science in Food Science, Stellenbosch University. 2013. Available online: https://scholar.sun.ac.za/bitstream/handle/10019.1/80020/schutte_isolation_2013.pdf?sequence=2 (accessed on 21 July 2016).
- Dlamini, N.R.; Dube, S. Studies on the physico-chemical, nutritional and microbiological changes during the traditional preparation of Marula wine in Gwanda, Zimbabwe. Nutr. Food Sci. 2008, 38, 61–69. [Google Scholar] [CrossRef]
- Costantini, A.; Garcia-Moruno, E.; Moreno-Arribas, M.V. Biochemical Transformations Produced by Malolactic Fermentation. In Wine Chemistry and Biochemistry; Springer: New York, NY, USA, 2008; pp. 27–57. [Google Scholar] [CrossRef]
- Berbegal, C.; Peña, N.; Russo, P.; Grieco, F.; Pardo, I.; Ferrer, S.; Spano, G.; Capozzi, V. Technological properties of Lactobacillus plantarum strains isolated from grape must fermentation. Food Microbiol. 2016, 57, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Raj, T.; Chandrasekhar, K.; Kumar, A.N.; Kim, S.H. Recent biotechnological trends in lactic acid bacterial fermentation for food processing industries. Syst. Microbiol. Biomanuf. 2021, 2, 14–40. [Google Scholar] [CrossRef]
- Bangar, S.P.; Suri, S.; Trif, M.; Özogul, F. Organic Acids Production from Lactic Acid Bacteria: A Preservation Approach. Food Biosci. 2022, 46, 101615. [Google Scholar] [CrossRef]
- Kersters, K.; Lisdiyanti, P.; Komagata, K.; Swings, J. The family acetobacteraceae: The genera Acetobacter, Acidomonas, Asaia, Gluconacetobacter, Gluconobacter, and Kozakia. Prokaryotes 2006, 5, 163–200. [Google Scholar] [CrossRef]
- Nielsen, D.S.; Teniola, O.D.; Ban-Koffi, L.; Owusu, M.; Andersson, T.S.; Holzapfel, W.H. The microbiology of Ghanaian cocoa fermentations analysed using culture-dependent and culture-independent methods. Int. J. Food Microbiol. 2007, 114, 168–186. [Google Scholar] [CrossRef]
- Jackson, R.S. Wine Science: Principles and Applications; Elsevier Science Publishing: London, UK, 2008. [Google Scholar]
- Joyeux, A.; Lafon-Lafourcade, S.; Ribéreau-Gayon, P. Evolution of acetic acid bacteria during fermentation and storage of wine. Appl. Environ. Microbiol. 1984, 48, 153–156. [Google Scholar] [CrossRef]
- Fleet, G.H. Yeast and wine flavour. Int. J. Food Microbiol. 2003, 86, 11–22. [Google Scholar] [CrossRef]
- Moreira, N.; Mendes, F.; de Pinho, P.G.; Hogg, T.; Vasconcelos, I. Heavy sulphur compounds, higher alcohols and esters production profile of Hanseniaspora uvarum and Hanseniaspora guilliermondii grown as pure and mixed cultures in grape must. Int. J. Food Microbiol. 2008, 124, 231–238. [Google Scholar] [CrossRef]
- García, M.; Esteve-Zarzoso, B.; Arroyo, T. Non-Saccharomyces yeasts: Biotechnological role for wine production. In Grape and Wine Biotechnology; Books on Demand: Norderstedt, Germany, 2016; Volume 10, p. 64957. [Google Scholar] [CrossRef]
- Pretorius, I.S. Tailoring wine yeast for the new millennium: Novel approaches to the ancient art of winemaking. Yeast 2000, 16, 675–729. [Google Scholar] [CrossRef] [PubMed]
- Parapouli, M.; Vasileiadis, A.; Afendra, A.S.; Hatziloukas, E. Saccharomyces cerevisiae and its industrial applications. AIMS Microbiol. 2020, 6, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Aldrete-Tapia, J.A.; Escalante-Minakata, P.; Miranda-Castilleja, D.E.; Hernández-Iturriaga, M. Fermentation conditions for yeast selection and effect of yeast–bacterial interaction in developing a starter culture for tequila fermentation. J. Food Sci. 2022, 87, 5089–5098. [Google Scholar] [CrossRef] [PubMed]





| Strain ID | Strain Code | Strain Origin (as Origin of the Wine) |
|---|---|---|
| L. plantarum | ELP1, ELP8, ELP9, | LAB-UL |
| L. plantarum | ELP7 | MLT |
| L. plantarum | ALP4 | MOSHIRA * |
| L. plantarum | ALP5 | DENILTON * |
| L. brevis | ALB1 | MOSHIRA * |
| L. brevis | ALB2 | DENILTON * |
| L. brevis | ELB3, ELB4, ELB5, ELB6, ELB7, ELB9 | LAB-UL |
| L. buchneri | ALBU1, ALBU2, ALBU5, ALBU6 | MOSHIRA * |
| L. buchneri | ALBU3, ALBU4 | DENILTON * |
| S. cerevisiae | EL0DG4, MWRS5, MGS1, EL2DS1, MCS5, MLG1, MRW1, MRWR1, MWRS4, MRW3 | LAB-UL |
| S. cerevisiae | ES20DG, EM6RW1 | MLT |
| S. cerevisiae | ES22DG | SKB |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maluleke, E.; Lekganyane, M.A.; Moganedi, K.L.M. Microbial Diversity of Marula Wine during Spontaneous Fermentation. Fermentation 2023, 9, 862. https://doi.org/10.3390/fermentation9100862
Maluleke E, Lekganyane MA, Moganedi KLM. Microbial Diversity of Marula Wine during Spontaneous Fermentation. Fermentation. 2023; 9(10):862. https://doi.org/10.3390/fermentation9100862
Chicago/Turabian StyleMaluleke, Evelyn, Maleho Annastasia Lekganyane, and Kgabo L. Maureen Moganedi. 2023. "Microbial Diversity of Marula Wine during Spontaneous Fermentation" Fermentation 9, no. 10: 862. https://doi.org/10.3390/fermentation9100862






