Abstract
Photoperiod is a pivotal regulatory factor in the flowering of Camellia oleifera Abel. (C. oleifera). GIGANTEA (GI) serves as a pivotal regulator, not only orchestrating the intricate circadian rhythm but also governing photoperiod-dependent flowering. In order to explore the function of GI in C. oleifera (CoGI), we obtained a CoGI gene-coding sequence and analyzed a CoGI protein sequence using bioinformatics. Furthermore, we conducted a spatiotemporal expression analysis of CoGI. And a yeast two-hybridization assay was used to screen the interacting proteins of CoGI. Evolutionary analysis revealed high conservation of the CoGI protein, which clustered with the GI protein from Camellia sinensis (CsGI) on a common evolutionary branch. The expression of CoGI was different in each part, and a tissue expression analysis revealed that the relative expression level of the CoGI gene is highest in the leaves of C. oleifera, while it is at its lowest in the seed coats. Transgenic Arabidopsis thaliana (Arabidopsis) overexpressing CoGI exhibited early flowering under long-day conditions. In addition, the yeast two-hybrid library screening revealed interactions between seven C. oleifera proteins and CoGI: CoACR9, CoLAO, CoDExH12-like, CoIT1K-like, CoUPF0481, CoIDM3, and CoAt4g27190-like. The findings demonstrated that CoGI is crucial to C. oleifera’s flowering.
1. Introduction
Camellia oleifera Abel. (C. oleifera), a unique woody oil tree species endemic to China, is primarily found in the Yangtze River basin and the hilly regions south of China [1,2]. It is a late self-incompatibility plant. The flowering periods of different varieties are different, and pollination is difficult to complete. Moreover, its flowering period coincides with autumn and winter, a time when low temperatures and unpredictable weather conditions hinder flowering and pollination. Consequently, in practical production, there are more flowers and less fruit, a low fruit setting rate, and a low yield phenomenon [3,4]. Hence, investigating the flowering mechanism of C. oleifera holds immense significance for enhancing its yield.
Photoperiod, the phenomenon that plants regulate their flowering time by sensing the duration of daylight and darkness, is a crucial factor influencing plants flowering and fruiting [5]. The circadian clock, which functions as an endogenous timer that detects changes in day length during the photoperiodic response, is a primary pathway for photoperiodic flowering induction [6,7,8,9]. The GI gene, situated between the circadian clock and flowering-regulating genes, serves as a circadian clock output gene [9,10]. In Arabidopsis thaliana (Arabidopsis), the CONSTANS (CO) gene plays an essential role in regulating flowering. It induces flowering in plants by promoting the expression of FLOWERING LOCUS T (FT), thereby [11]. As one of the flowering regulatory factors, GI regulates plants’ flowering by modulating the expression of the CO and FT genes under long-day conditions. GI stands as an upstream gene of CO and FT, making it significantly important in regulating plant physiological rhythms and the flowering process [12,13,14,15].
In Arabidopsis, the ZTL protein regulates the distribution and expression peak of the GI protein between the nucleus and cytoplasm through its N-terminal LOV domain, thus affecting the regulation of the biological rhythm and flowering time of Arabidopsis [16]. Through an analysis of the structure and function of CmGI in chrysanthemums, it was found that CmGI could be regulated in response to biological clock signals in the photoperiodic flowering pathway of chrysanthemums [17]. In rice, OsGI regulates the expression of three-quarters of genes associated with the circadian rhythm [18]. Related studies on the circadian rhythm of Norwegian spruce showed that the functions of PaCCA1, PaGI, and PaZTL were similar to those of Arabidopsis homologous isomerins [19]. These results indicate that the function of GI in regulating plant circadian rhythm may be highly conserved in different types of plants. Therefore, we speculate that the CoGI gene is the key gene regulating the flowering time in C. oleifera in response to the circadian rhythm. In this study, the CDS sequence of the CoGI gene of C. oleifera was obtained by screening the transcriptome sequencing results. After obtaining the full-length sequence, its function and potential interacting proteins were studied to verify its function in C. oleifera and speculate the possible biosynthesis pathway.
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
C. oleifera trees were planted at the Central South University of Forestry and Technology (112.994587° E, 28.131841° N), while Arabidopsis (Columbia, Col-0) cultivation took place in the greenhouse of the university’s Key Laboratory of Economic Forest. Seeds were initially sterilized and planted on a 1/2 MS (50 mg/L Kanamycin) solid medium in order to screen transgenic Arabidopsis overexpressing CoGI. Then, the solid media were transferred to the growth chamber following a vernalization period of 2 days at 4 °C in the dark. After two weeks, the germinated seedlings were transplanted into pots containing a substrate mixed with nutrient soil and perlite 3:2 for further cultivation in the greenhouse. The environmental conditions in both the greenhouse and growth chamber were 16 h of light and 8 h of darkness at 22 °C. The light source was an LED lamp with the same temperature day and night. The cultivation of Arabidopsis was performed using a small, plastic, square basin with a diameter of 8 cm and a height of 8 cm. The soil humidity was not strictly controlled, but was monitored through manual observation and a small number of watering.
2.2. RNA Extraction and cDNA Synthesis
C. oleifera leaves were ground in liquid nitrogen. An EZ-10 DNAaway RNA mini-preps kit (BBI, Shanghai, China) was utilized to extract RNA and a HiScript II 1st Strand cDNA Synthesis Kit (+gDNA wiper, removing the residual genomic DNA to ensure the reliability of subsequent quantitative results) (Vazyme, Nanjing, China) was then used to synthesize cDNA from the isolated RNA. Samples of the RNA and cDNA were kept for later usage at −80 °C in a freezer.
2.3. Clone and Vector Construction of CoGI
Primers (Table A1) for CoGI were designed using Primer Premier 5 in order to clone the CDS fragment of the CoGI gene from the cDNA of C. oleifera leaves. The PCR product was then detected, recovered, and purified via gel electrophoresis before being ligated to the pCAMBIA-2300-GFP vector. The vector that was created was converted into competent E. Coli DH5α competent cells, which were then plated on LB (50 mg/L Kana) within the sterile workbench. Individual colonies were chosen for PCR identification after a period of overnight incubation at 37 °C [20,21]. Positive strains were then cultured in an LB (50 mg/L Kanamycin) liquid medium and sent to Beijing TsingKe Biotech Co., Ltd. (Beijing, China) for sequencing. The obtained CDS sequence was aligned with the CoGI CDS sequence using DNAMAN 9.0 software to confirm the cloning of the target gene before proceeding with subsequent experiments.
2.4. Bioinformatics Analysis of CoGI
DNAMAN 9.0 software was utilized to meticulously compare the sequencing outcomes with the CDS sequence of CoGI [21]. The basic physicochemical properties of the CoGI protein were analyzed using the Expasy tool, ProtParam (https://www.expasy.org/, accessed on 10 August 2023), and the hydrophilicity and hydrophobicity of the protein were further analyzed using its ProtScale tool. The conserved domains and gene families of the CoGI proteins were analyzed using InterPro (https://www.ebi.ac.uk/interpro/, accessed on 10 August 2023). SOPMA (http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_sopma.html, accessed on 10 August 2023) was used to predict the secondary structure of CoGI protein, and Expasy’s SWISS-MODEL tool was used to calculate the tertiary structure model of the CoGI protein. NCBI was used (https://www.ncbi.nlm.nih.gov/, accessed on 10 August 2023) to search the GIs protein sequences, and multiple comparisons were conducted using DNAMAN 9.0. Meanwhile, the conserved motifs of the protein sequences were analyzed on MEME (https://meme-suite.org/meme/index.html, accessed on 10 August 2023). Phylogenetic trees were constructed with precision using MEGA 11 and TBtools-II v2.010 [21,22].
2.5. Expression Analysis of CoGI
From 31 July to 30 October, the C. oleifera flower buds were picked at a 15-day interval [23]. Mid-September saw the collection of C. oleifera’s roots, stems, leaves, flowers, and fruits. Flowers were separated into petals and stamens, while fruits were divided into seed coats and kernels. Moreover, samples were taken every two hours on a day when the weather was good. After being placed in liquid nitrogen, these materials were kept in a freezer at −80 °C in order to extract total RNA and synthesize cDNA. Glyceraldehyde-3-phosphate dehydrogenase (CoGAPDH) was used as the internal reference gene in the qRT-PCR experiment for CoGI expression level, which was conducted using ChamQ Universal SYBR qPCR Master Mix (Vazyme, Nanjing, China) [24]. Table A1 and Table A3 provide specifics on the primers and reaction conditions for qRT-PCR, respectively. Three technical duplicates of each qRT-PCR reaction were run on the CFX96 real-time PCR system (Bio-Rad, Hercules, CA, USA). Using the 2−∆∆CT method, collecting one set of sample data as a 1 fold contrast group, relative expression levels were determined.
2.6. Transformation and Screening of Arabidopsis
The plasmid pCAMBIA-2300-GFP-CoGI, once constructed, was extracted and effectively converted via electroporation into Agrobacterium tumefaciens AGL0. Subsequently, the floral dip technique was utilized to introduce this plasmid into Arabidopsis (Col-0) [25]. After being sterilized, the collected T0 seeds were planted on the 1/2 MS (50 mg/L Kanamycin) solid medium for screening. Following 2 days of vernalization at 4 °C, the solid mediums were moved to the growth chamber and left there for 2 weeks. After two weeks, T0 seedlings exhibiting four leaves and elongated roots were carefully selected from the solid medium and transplanted into pots. These seedlings were then moved to a greenhouse for further growth. Three Arabidopsis lines that had Kanamycin resistance and phenotypes that were obviously distinct from Col-0 were carefully selected, and the seeds collected were designated as the T1 generation.
2.7. Flowering Phenotypic Observation of Overexpression of CoGI in Transgenic Arabidopsis
The CoGI-overexpressing transgenic Arabidopsis plants underwent rigorous screening up to the T2 generation, and the seeds were then collected for phenotypic observation. Prior to sowing, the Arabidopsis seeds were thoroughly disinfected. The Col-0 seeds were planted on the 1/2 MS solid medium, whereas the T2 seedlings were placed on the 1/2 MS (50 mg/L Kanamycin) solid medium. After a fortnight of growth, seedlings with four leaves and elongated roots were carefully chosen from the solid medium and transplanted into pots for further cultivation. The flowering time and the count of leaves at flowering were meticulously observed and recorded. The collected phenotypic data were subsequently analyzed using GraphPad Prism 9.
2.8. The Autoactivation Activity Test of CoGI
Using the primers described in Table A1, the CoGI fragment was successfully cloned into the pGBKT7 vector (CoGI-BD). This experimental setup included four groups: the blank control group (pGBKT7 + pGADT7), the positive control group (pGBKT7-53 + pGADT7-T), the negative control group (pGBKT7-Lam + pGADT7-T), and the experimental group (CoGI-BD + pGADT7). All of these groups underwent transformation into the yeast strain AH109. After selecting yeast colonies from each group, they were resuspended in sterile water. Subsequently, 3 μL of each suspension was transferred onto an SD/-Trp/-Leu/-His (TDO) + 3-AT solid medium containing a gradient of 3-AT concentrations ranging from 0 to 60 mM/L. Following the drying, sealing, and inverted culturing of the solid media for 3–5 days, the lowest 3-AT concentration that corresponded to a plate with no colony growth was identified. This concentration was then selected for further library screening procedures [26].
2.9. Screen of Yeast Two-Hybrid Library
Yeast two-hybrid screening utilized the mating approach, enabling the screening of protein-protein interactions using various selective media [27,28]. CoGI-BD was turned into yeast AH109 and grown in an SD/-Trp liquid medium at 28 °C, 200 rpm shaker until OD600 reached 0.8. The yeast cells were then resuspended in the SD/-Trp liquid media at a density above 1 × 108/mL and mixed with Y187 library yeast, which was constructed from the cDNA of mixed samples of C. oleifera bud and leaves. This mixture was further cultured in a 2 × YPDA liquid medium at 28 °C, 200 rpm shaker for 16–20 h, until heterozygous “clover” shapes were observed under a microscope. The yeast cells were resuspended in sterile water and plated on TDO solid medium with 3-AT for 4–7 days. All yeast colonies were resuspended and transferred to both TDO + 3-AT and SD/-Trp/-Leu-His-Ade (QDO) + 3-AT + X-α-Gal solid medium. Yeast Plasmid Extraction Kit (Coolaber, Beijing, China) was utilized to extract the plasmids from blue colonies that had been chosen and cultured in the SD/-Trp-leu liquid medium for an entire night. After that, the plasmids were introduced into E. coli DH5α in order to be sequenced.
After sequencing, NCBI blastp was used to evaluate protein sequences, and the results were compiled. Each fragment’s associated plasmid was taken out of E. coli DH5α and introduced into yeast AH109 with CoGI-BD. Verification was performed on TDO + 3-AT and QDO + 3-AT + X-α-gal solid media [29,30].
3. Results
3.1. Cloning of the CoGI Coding Sequence and Analysis of the Protein
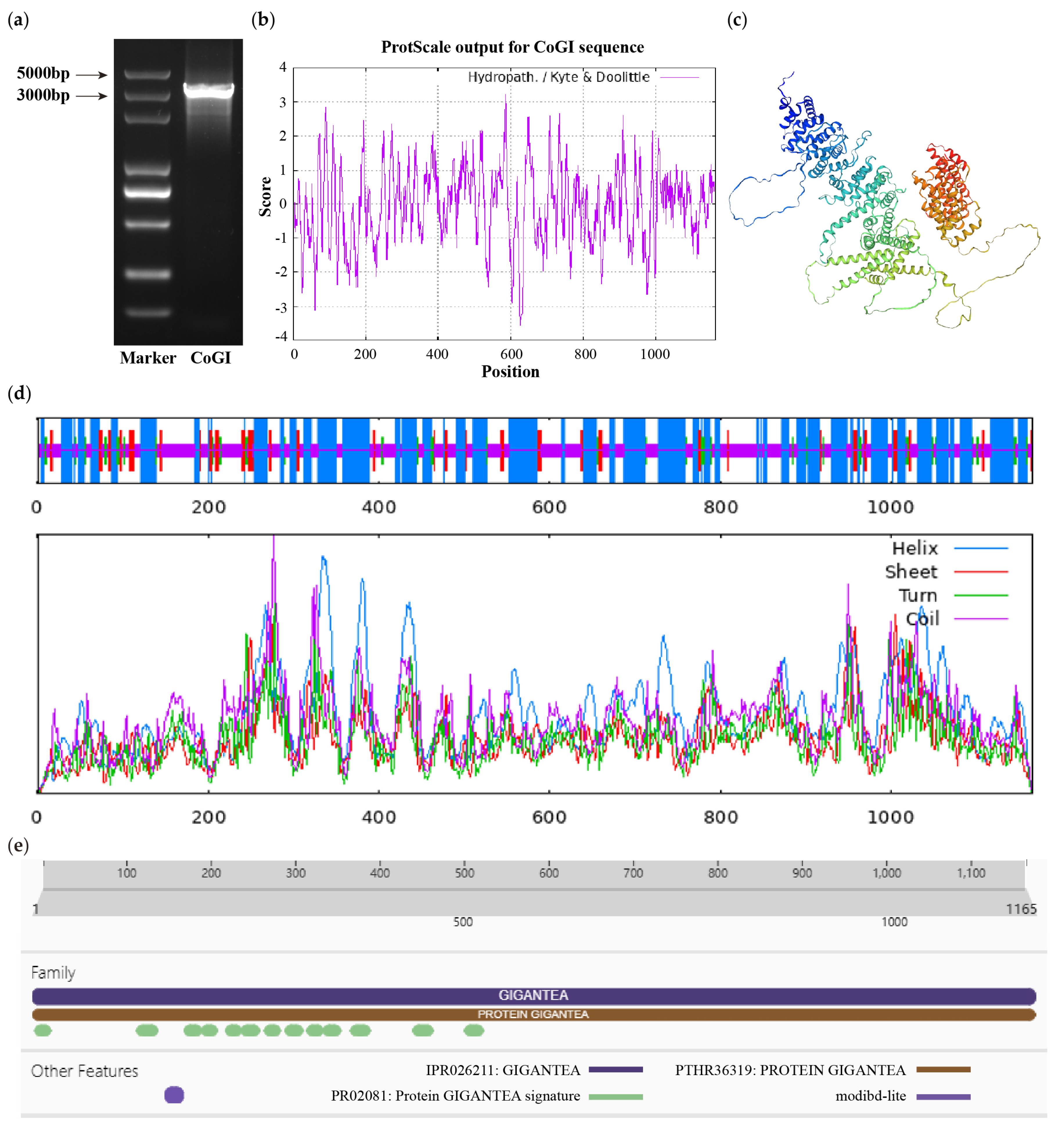
From the cDNA of C. oleifera ‘Huashuo’, the CDS sequence of CoGI was cloned (Figure 1a). The sequence results were consistent with the projected size, with a total length of 3498 bp encoding 1165 amino acids. The CoGI protein, the molecular formula of which was predicted to be C5702H9042N1564O1685S53, had a relative molecular weight of 128.17 kD, a theoretical isoelectric point of 6.06, a hydrophilic of −0.029 and an instability index of 53.32 (Figure 1b). It is generally believed that negative numbers represent hydrophilicity and positive numbers represent hydrophobicity. Proteins with an instability index less than 40 are stable, while proteins with an instability index greater than 40 are unstable. Therefore, we speculate that it is an unstable hydrophilic protein. The secondary structure mainly contains four structures: α-helix, random coil, extended strand, and β-turn, accounting for 47.55%, 40.09%, 7.90% and 4.46%, respectively (Figure 1d), which is basically consistent with the tertiary structure model (Figure 1c). Its protein sequence domain was predicted to belong to the GIGANTEA gene family (Figure 1e).
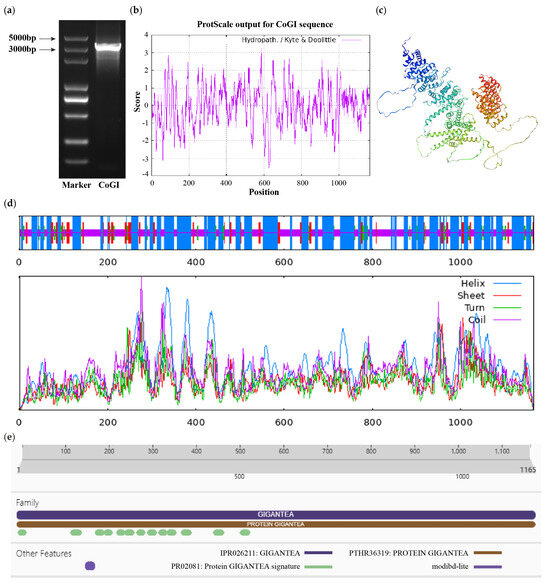
Figure 1.
CDS fragment detection and bioinformatics analysis of CoGI protein. Notes: (a) CDS fragment detection of CoGI; (b) hydrophilicity and hydrophobicity analysis of CoGI protein, negative number is hydrophilic, positive number is hydrophobic; (c) tertiary structure prediction of CoGI protein; (d) secondary structure prediction of CoGI protein; (e) conserved domains of CoGI protein.
3.2. Homology Analysis and Phylogenetic Tree Construction of CoGI Protein from C. oleifera
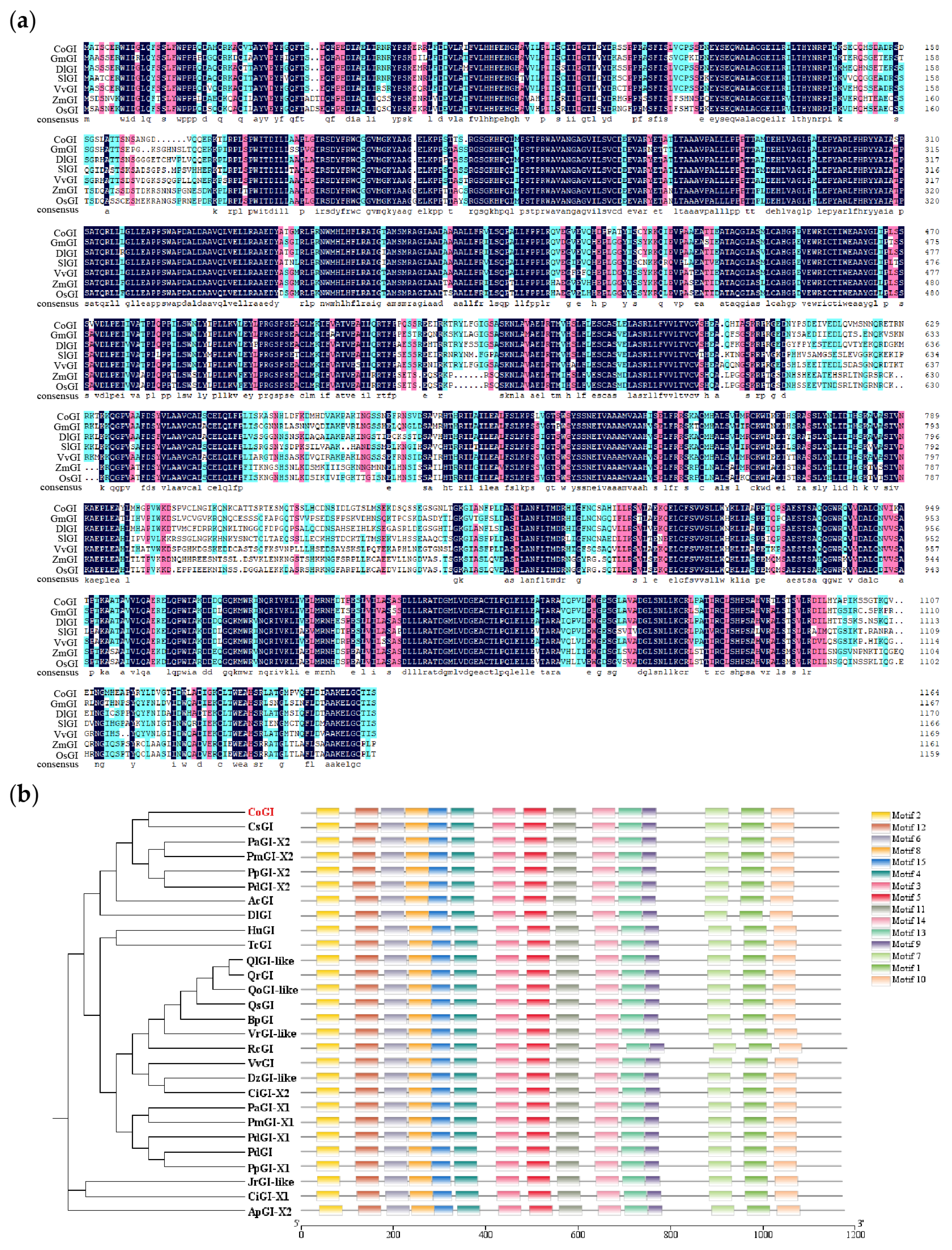
The CoGI protein sequence of C. oleifera was compared with the GI protein sequences of Glycine max (GmGI), Dimocarpus longan (DlGI), Solanum lycopersicum (SlGI), Vitis vinifera (VvGI), Zea mays (ZmGI), and Oryza sativa (OsGI). The results showed that the GI protein sequence changed little and was highly conserved (Figure 2a). The degree of sequence composition similarity between C. oleifera and the six species was 79.24%, 83.04%, 77.64%, 83.46%, 69.37%, 69.79%. NCBI was used to search and select 27 GIs protein sequences of 21 species for constructing a phylogenetic tree. The results showed that CsGI and CoGI protein were in the same branch (Figure 2b).
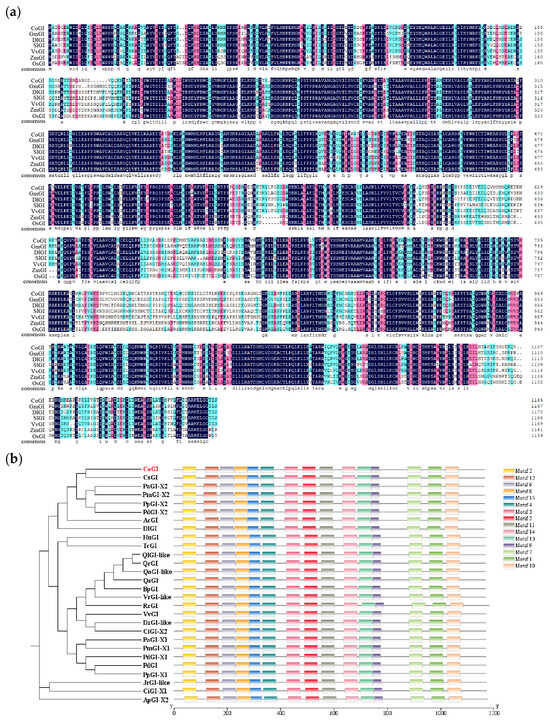
Figure 2.
Multiple sequence alignment and phylogenetic tree. Notes: (a) Multiple sequence alignment, the GI protein sequences are from Glycine max, Dimocarpus longan, Solanum lycopersicum, Vitis vinifera, Zea mays, and Oryza sativa. Highlight homology level:  = 100%,
= 100%,  ≥ 75%,
≥ 75%,  > 50%; (b) phylogenetic tree of GI protein family from C. oleifera and other species.
> 50%; (b) phylogenetic tree of GI protein family from C. oleifera and other species.
 = 100%,
= 100%,  ≥ 75%,
≥ 75%,  > 50%; (b) phylogenetic tree of GI protein family from C. oleifera and other species.
> 50%; (b) phylogenetic tree of GI protein family from C. oleifera and other species.
3.3. Expression Analysis of CoGI in C. oleifera
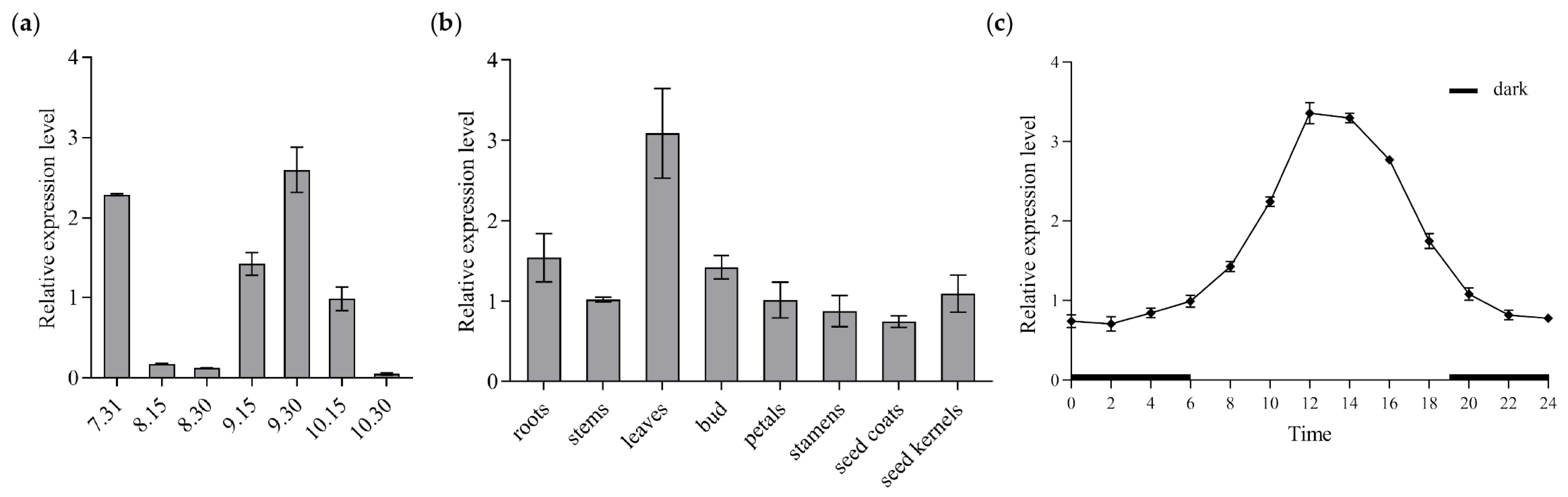
Flower buds in seven different stages were collected every 15 d for CoGI expression analysis, and qRT-PCR was conducted using cDNA templates derived from samples of different periods. The results (Figure 3a) indicated that CoGI expression reached its peak in late September. To investigate the CoGI gene’s expression pattern across various tissues of C. oleifera, qRT-PCR was conducted using cDNA templates derived from roots, stems, leaves, buds, petals, stamens, seed coats, and seed kernels. The findings (Figure 3b) revealed that leaves had the highest level of CoGI expression whereas seed coats had the lowest level. By analyzing the expression of CoGI in leaves during the day and night, it was found that the expression of CoGI in leaves showed a circadian rhythm change within 1 day under the long-day condition. At night, the expression level was relatively low, from 0:00 to 6:00 and 20:00 to 24:00. The expression level began to rise after dawn at 6:00, reached a peak at 12:00, and then declined (Figure 3c).
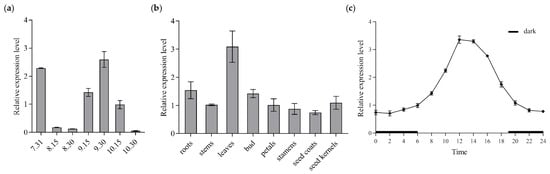
Figure 3.
Relative expression level of CoGI in C. oleifera. Notes: (a) Relative expression level of CoGI in different stage flower buds; (b) relative expression level of CoGI in different tissues; (c) relative expression level of CoGI in leaves at different time in one day. Data are means ± SE.
3.4. Heterologous Overexpression of CoGI in Arabidopsis
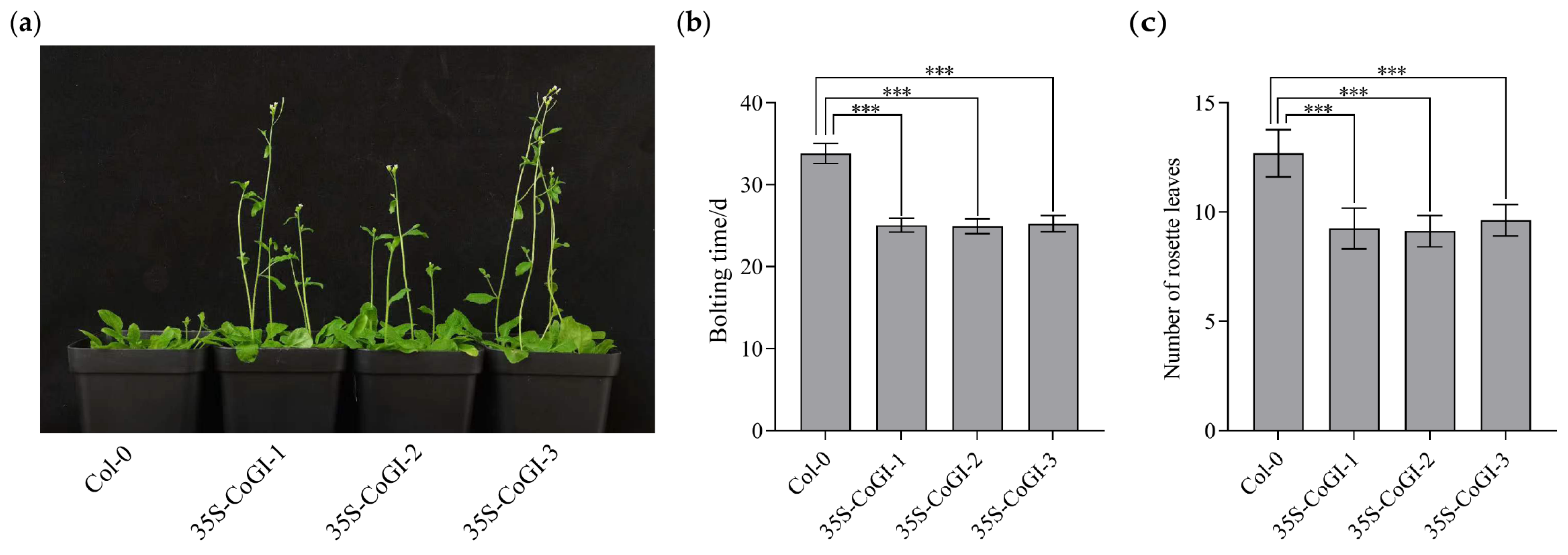
In order to verify the function of the CoGI gene in Arabidopsis, we introduced the constructed overexpression vector of the CoGI gene into Col-0 through Agrobacterium tumefaciens AGL0, and observed its phenotype. The result (Figure 4a) showed that compared with Col-0 plants, overexpressed strains had an earlier flowering time. The flowering time and number of rosette leaves were calculated and mapped. As shown in Figure 4b,c, compared with Col-0 plants, CoGI overexpressing plants had earlier flowering time and fewer rosette leaves at flowering time. The flowering times of three CoGI overexpressing lines under long sunshine conditions were 8.8, 8.9 and 8.6 days earlier than that of Col-0 plants, respectively. The number of rosette leaves decreased by 3.4, 3.6 and 3.1, respectively, showing an obvious early flowering phenotype.
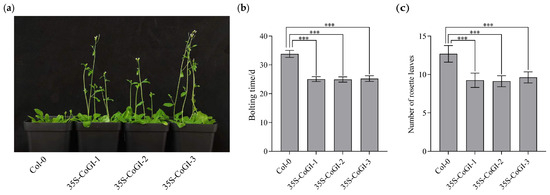
Figure 4.
Observation of the overexpression CoGI transgenic Arabidopsis. Notes: (a) Flowering observation; (b) bolting time; (c) number of leaves at bolting time. The data are means ± SD. p-values were determined using Tukey’s HSD test. ns, no statistically significant difference; *** p < 0.001.
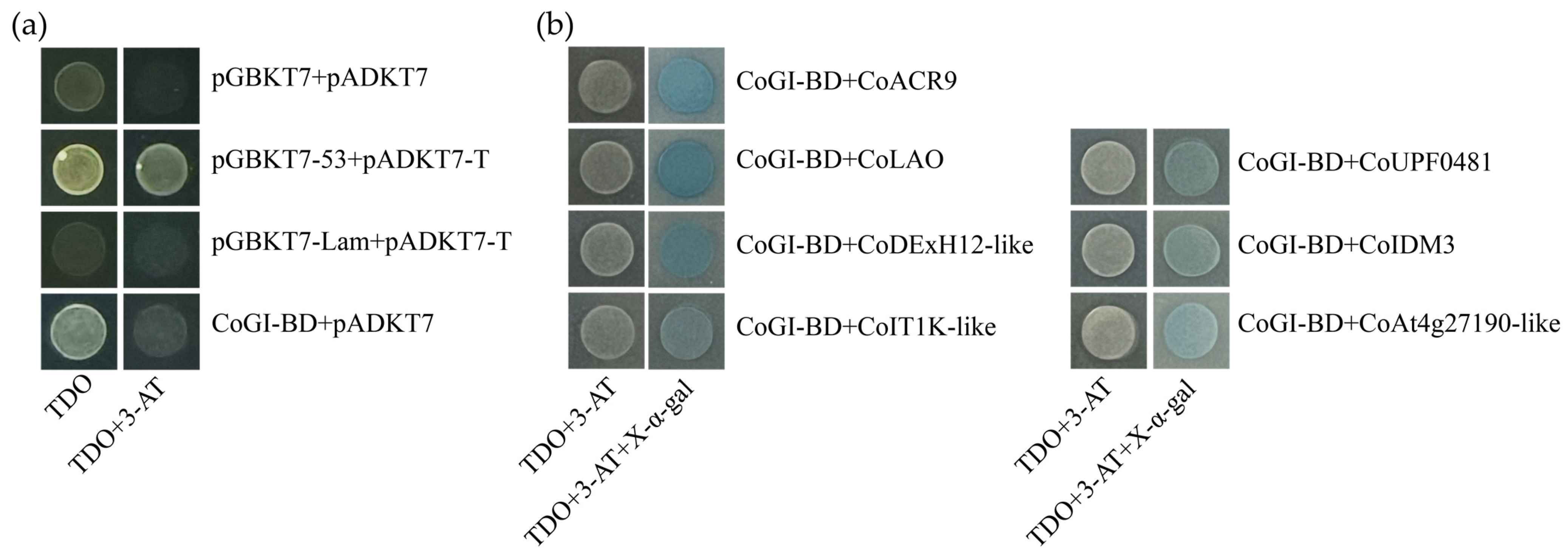
3.5. Autoactivation Activity Test and Yeast Two-Hybrid Screening of CoGI
When the 20 mM 3-AT was added, CoGI showed autoactivation that could be inhibited (Figure 5a). From yeast two-hybrid library screening, eight yeast strains were sequenced, yielding eight gene fragments. The eight fragments were translated and retrieved using the BLASTP tool. Subsequently, seven plasmids that matched known protein sequences were taken out of E. coli DH5α and transformed into yeast AH109 with CoGI-BD (Figure 5b). The specific information of the seven proteins is shown in Table 1, and the bioinformatics analysis of the proteins is illustrated in Figure A1, Figure A2, Figure A3, Figure A4, Figure A5, Figure A6 and Figure A7.
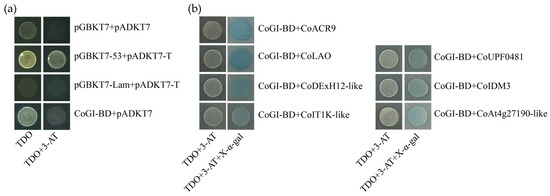
Figure 5.
Autoactivation activity test and yeast two-hybrid screening of CoGI. Notes: (a) Detection of autoactivation activity of CoGI protein; (b) validation of interaction between seven proteins and CoGI.

Table 1.
List of seven proteins that interacted with CoGI protein.
4. Discussion
The GI protein has no known functional domain, and it is a higher plant-specific nuclear protein [31]. In this study, we cloned the CDS sequence of CoGI and conducted a translation as well as a bioinformatics analysis of its encoded protein. We found that the CoGI protein sequence was more than 80% similar to other species, indicating that GI was highly conserved among different species, which was consistent with previous studies.
It was found that GI genes have tissue and organ specificity in different species. In Arabidopsis, AtGI exhibits elevated expression levels in leaves, flowers, and stems, with higher expression observed in stems compared to roots [32]. In Glycine max, the expression of GmGI was the highest in leaves and flower buds under a long-day condition. However, under short-day conditions, GmGI expression peaked in roots and leaves [33]. BnGI was expressed in all tissues of Brassica napus, with significantly higher levels in rapeseed compared to other tissues [34]. In Dioscorea esculenta, DeGI expression was higher in leaves and roots, but lower in stems [35]. In Solanum tuberosum, StGI was highly expressed in roots, stolons, and sepals [36]. In Citrus reticulata Blanco, CrGI expression peaked in leaves, followed by stems [37]. Similarly, in Juglans regia, JrGI expression was significantly higher in leaves compared to leaf buds and flower buds [38]. In this study, CoGI expression in different tissues of C. oleifera was analyzed and CoGI was found to be highest in leaves. Previous studies have shown that the circadian clock components control leaf senescence in Arabidopsis, and GI positively regulates leaf senescence [14,39,40,41]. In addition, the highest expression of GI in plant leaves may also be because leaves are the most sensitive organs for sensing photoperiods and responding to biorhythmic clock signals.
GI has many functions and effects in regulating flowering time. In the study of Arabidopsis, it was discovered that AtGI can interact with FLAVIN-BINDING, KELCH REPEAT, and F-BOX 1 (FKF1) proteins to form the ATGI-FKF1 complex under long-day conditions, degrade CDF1 protein on CO promoter, and release the inhibition of CO. Thus, it can promote the expression of its target gene FT and promote the flowering of Arabidopsis [42,43,44,45,46]. However, under short-day conditions, GI can enhance the expression of FT without up-regulating CO expression, thus promoting the flowering of Arabidopsis [47]. Moreover, in the relevant studies on the long-day monocotyledonous plant Brachypodiam distachyon, it was found that the function of BdGI to induce flowering was the same as that in Arabidopsis [48]. In this study, we transferred the CoGI into Col-0 Arabidopsis plants to obtain transgenic CoGI-overexpressing Arabidopsis plants. It was found that transgenic plants exhibited significantly earlier flowering time compared to Col-0 Arabidopsis plants, and the number of rosette leaves was significantly reduced during bolting.
Although many proteins interacting with GI genes have been reported in Arabidopsis, it is not clear which proteins CoGI binds to and influences the expression in C. oleifera. Therefore, we conducted yeast two-hybrid experiments to find the proteins binding to CoGI and explore the response mechanism of CoGI in the C. oleifera flower formation from the perspective of genetic regulation. One of them, the ACR9 gene encodes a protein containing ACT domain repeats. In rice, the distribution and accumulation of the OsACR9 protein show dynamic changes in different tissues and developmental stages, especially in the parts where nitrogen N-metabolism is active. For example, the accumulation of OsACR9 protein in rice root cells increases significantly after the application of ammonium salt [49]. These results indicate that it is involved in the regulation of N-metabolism. In Arabidopsis, acr9 mutants are highly sensitive to glucose, especially in early seedling development, root growth, anthocyanin accumulation, and significantly increased expression of sugar response genes, suggesting that ACR9 is a negative regulatory component in glucose signaling pathways. At the same time, acr9 mutants are also hypersensitive to Abscisic Acid (ABA), suggesting that ACR9 may also play an inhibitory role in the ABA signaling pathway and may mediate the interaction between C/N signals [50]. At4g27190 is a protein with potential disease resistance in plants. In a study of the durian plant, a large number of resistance gene analogs (RGAs) were identified through the mining and analysis of the reference genome of the durian “Musang King”, among which 135 copies of the disease resistance protein encoded by At4g27190 were found in the durian genome. Suggesting that it may play an important role in durian defense responses to a variety of pathogens [51]. In comparative proteomic studies during lychee flowering, homologous proteins encoded by At4g27190 were specifically expressed in the phloem exudate at the start-up stage of flower buds, and were associated with signal transduction, hormone-mediated signaling pathways, plant response to ABA and other biological processes [52]. GI is not only related to glucose metabolism in plants, but also plays a key role in the photoperiodic signal response to drought stress. GI can also activate FT/TSF and SOC1 genes in response to ABA, resulting in early flowering.
The UPF0481 protein has been shown to have many possible functions in plants, including stress response, tissue culture, and regeneration processes. In the study of Elsholshia haichuensis, the UPF0481 protein was differentially expressed between copper-resistant and non-copper-resistant populations, indicating that this protein may play a role in plant response to heavy metals (such as copper) stress [53]. In the study of eucalyptus, the homologous gene of this protein showed dynamic expression changes during callus development. This suggests that it also plays a role in dedifferentiation, redifferentiation and developmental regulation of plant cells [54]. IDM3 protein plays multiple roles in plant DNA methylation regulation. It is not only a core component of the ROS1-mediated DNA demethylation pathway, but also a key factor involved in histone acetylation regulation. In Arabidopsis, IDM3 interacts with proteins such as IDM1, IDM2, and MBD7, and this complex can trigger histone acetylation modification targeting specific methylation regions, thus initiating the recruitment of demethylases such as ROS1 and achieving DNA demethylation [55,56]. In Camellia sinensis, CsIDM3 also interacts with MBD family members CsMBD5 and CsMBD16, suggesting that IDM family members may regulate the epigenetic silencing of specific methylation regions by interacting with MBD proteins [57]. LAO is a copper-containing oxidase in plant cells, mainly located in the cytoplasm or near the cell wall, which participates in terminal oxidation reactions, and can catalyze the oxidation of ascorbic acid (vitamin C) to dehydroascorbic acid, a process that is crucial for material metabolism in plants, especially playing a central role in regulating the REDOX state outside the plastid [58]. As an antioxidant and signaling molecule, ascorbic acid concentration fluctuates under strong light environment, which may be related to plant adaptation to light intensity change, regulation of the photoprotection mechanism and coping with oxidative stress. Studies have shown that under intense light conditions, ascorbic acid concentration in plant leaves changes regularly during the day, and is closely related to the establishment of flowering time, defense response to stress, and expression regulation of genes related to plant hormone signaling [59,60,61]. The above three proteins are involved in plant stress response, DNA methylation, and antioxidant synthesis, etc. Although no studies have proven that CoGI has a direct relationship with these three enzymes, GI genes play an important role in plant stress response. In Arabidopsis, the activity of DNA MENTHYLTRAN-SFERASE 1 (MET1) is coordinated with the oscillation of the biological clock, which affects the methylation state of genes related to circadian rhythms, and thus regulates the expression of these genes [62]. Moreover, the daily regular variation of ascorbic acid concentration in plant leaves indicates that it may be influenced by circadian rhythms.
At present, there are few reports about DExH12-like and IT1K-like in plants, and the functions of these two plant proteins and the biological processes involved in them are limited. Therefore, their mechanisms of action and regulatory pathways were not discussed in depth in this study.
5. Conclusions
We performed a comprehensive analysis of CoGI in this study. Through phylogenetic tree construction, we discovered a close relationship between CoGI and CsGI. Expression analysis demonstrated that CoGI expression was highest in the leaves of C. oleifera and lowest in the seed coats. Meanwhile, transgenic Arabidopsis plants overexpressing CoGI showed an early flowering phenotype under long-day conditions. In addition, a total of seven protein fragments that can interact with CoGI in yeast were screened in a yeast two-hybrid experiment. The functions which the genes were mostly related to signal transduction. Overall, our results deepen our comprehension of CoGI and offer promising avenues for future investigations on the role of GI genes in plants growth and development.
Author Contributions
Conceptualization, J.Y. and J.L.; methodology, J.Y. and L.J.; software, L.J.; validation, L.J. and J.Y.; formal analysis, L.J., S.R., Q.L. and L.Z.; investigation, L.J., S.R., Q.L. and L.Z.; resources, J.Y. and J.L.; data curation, L.J., S.R., Q.L. and L.Z.; writing—original draft preparation, L.J.; writing—review and editing, S.R., Q.L., L.Z. and J.Y.; visualization, J.Y.; supervision, J.Y. and J.L.; project administration, J.Y. and J.L.; funding acquisition, J.Y. and J.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by National Natural Science Foundation of China, grant number 32371930; China Postdoctoral Science Foundation, grant number 2021M703653; Natural Science Foundation of Hunan Province, grant number 2023JJ41041; the Research Foundation of Education Bureau of Hunan Province, grant number 22B0281.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding authors.
Conflicts of Interest
The authors declare no conflicts of interest.
Appendix A

Table A1.
The list of Primer.
Table A1.
The list of Primer.
| Primer Name | Primer Sequence (5′ to 3′) |
|---|---|
| CoGI-2300-F | GGTACCCGGGGATCCATGGCTACTTCATGCGAAAGG |
| CoGI-2300-R | TCCTCTAGAGGATCCAATTGATATGGTACATCCTAATTCC |
| CoGI-qPCR-F | TCAACAAAGCCGAACCA |
| CoGI-qPCR-R | CAGCAGGGAAACAACAGAG |
| GAPDH-F | CTACTGGAGTTTTCACCGA |
| GAPDH-R | TAAGACCCTCAACAATGCC |
| CoGI-BD-F | AATTCCCGGGGATCCATGGCTACTTCATGCGAAAGG |
| CoGI-BD-R | CAGGTCGACGGATCCAATTGATATGGTACATCCTAATTCC |

Table A2.
The reaction condition of PCR.
Table A2.
The reaction condition of PCR.
| Step | Temperature/°C | Time | Cycles |
|---|---|---|---|
| Predenaturation | 95 | 3 min | 35 |
| Denaturation | 95 | 15 s | |
| Annealing | 58 | 15 s | |
| Extension | 72 | 2 min | |
| Complete extension | 72 | 5 min | |
| Finish | 22 | Forever |

Table A3.
The reaction condition of qRT-PCR.
Table A3.
The reaction condition of qRT-PCR.
| Step | Temperature/°C | Time | Cycles |
|---|---|---|---|
| Predenaturation | 95 | 5 min | 45 |
| Denaturation | 95 | 30 s | |
| Annealing | 58 | 30 s | |
| Extension | 72 | 30 s |

Table A4.
The information of the proteins for multiple sequence alignment.
Table A4.
The information of the proteins for multiple sequence alignment.
| Protein | Accession | Species |
|---|---|---|
| GmGI | NP_001341719.1 | Glycine max |
| DlGI | AII99806.1 | Dimocarpus longan |
| SlGI | XP_004237832.1 | Solanum lycopersicum |
| VvGI | XP_010665062.1 | Vitis vinifera |
| ZmGI | NP_001288586.1 | Zea mays |
| OsGI | NP_001396355.1 | Oryza sativa |

Table A5.
Information of the proteins for the phylogenetic tree.
Table A5.
Information of the proteins for the phylogenetic tree.
| Protein | Accession | Species | Protein | Accession | Species |
|---|---|---|---|---|---|
| CsGI | XP_028106519.1 | Camellia sinensis | QoGI-like | XP_030952170.1 | Quercus lobata |
| ClGI | KAI8004298.1 | Camellia lanceoleosa | QlGI-like | XP_030952183.1 | |
| RcGI | XP_024166287.1 | Rosa chinensis | CiGI-X1 | XP_042962978.1 | Carya illinoinensis |
| VvGI | XP_059590911.1 | Vitisvinifera | CiGI-X2 | XP_042962979.1 | |
| DlGI | XP_052190440.1 | Diospyros lotus | PdGI | AJC01622.1 | Prunus dulcis |
| AcGI | PSS17443.1 | Actinidia chinensis | PdGI-X1 | XP_034196939.1 | |
| QrGI | XP_050269337.1 | Quercus robur | PdGI-X2 | XP_034196940.1 | |
| DzGI-like | XP_022767964.1 | Durio zibethinus | PmGI-X1 | XP_008237481.1 | Prunus mume |
| TcGI | XP_017981038.1 | Theobroma cacao | PmGI-X2 | XP_016650762.1 | |
| JrGI-like | XP_018846578.1 | Juglans regia | PaGI-X1 | XP_021818210.1 | Prunus avium |
| QsGI | XP_023907641.2 | Quercus suber | PaGI-X2 | XP_021818214.1 | |
| VrGI-like | XP_034674793.1 | Vitis riparia | PpGI-X1 | XP_007199688.2 | Prunus persica |
| HuGI | XP_021285158.1 | Herrania umbratica | PpGI-X2 | XP_020426395.1 | |
| BpGI | ALL25874.1 | Betula platyphylla | ApGI-X2 | XP_027367357.1 | Abrus precatorius |
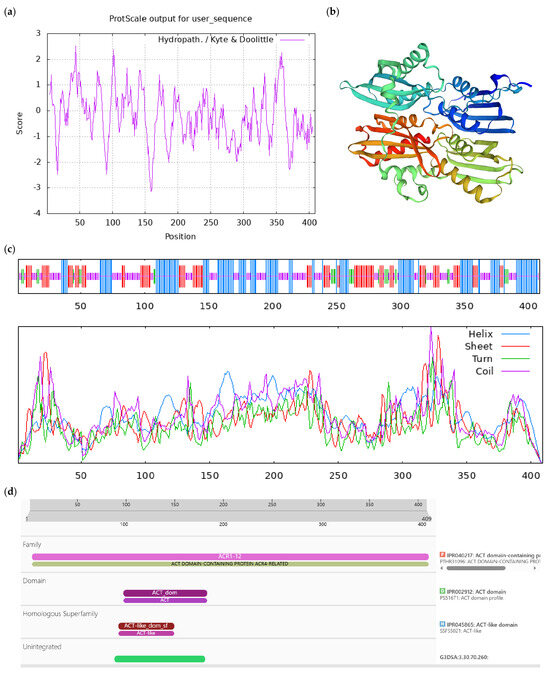
Figure A1.
Bioinformatics analysis of CoACR9 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoACR9 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoACR9 protein; (c) secondary structure prediction of CoACR9 protein; (d) conserved domains of CoACR9 protein.
Figure A1.
Bioinformatics analysis of CoACR9 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoACR9 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoACR9 protein; (c) secondary structure prediction of CoACR9 protein; (d) conserved domains of CoACR9 protein.
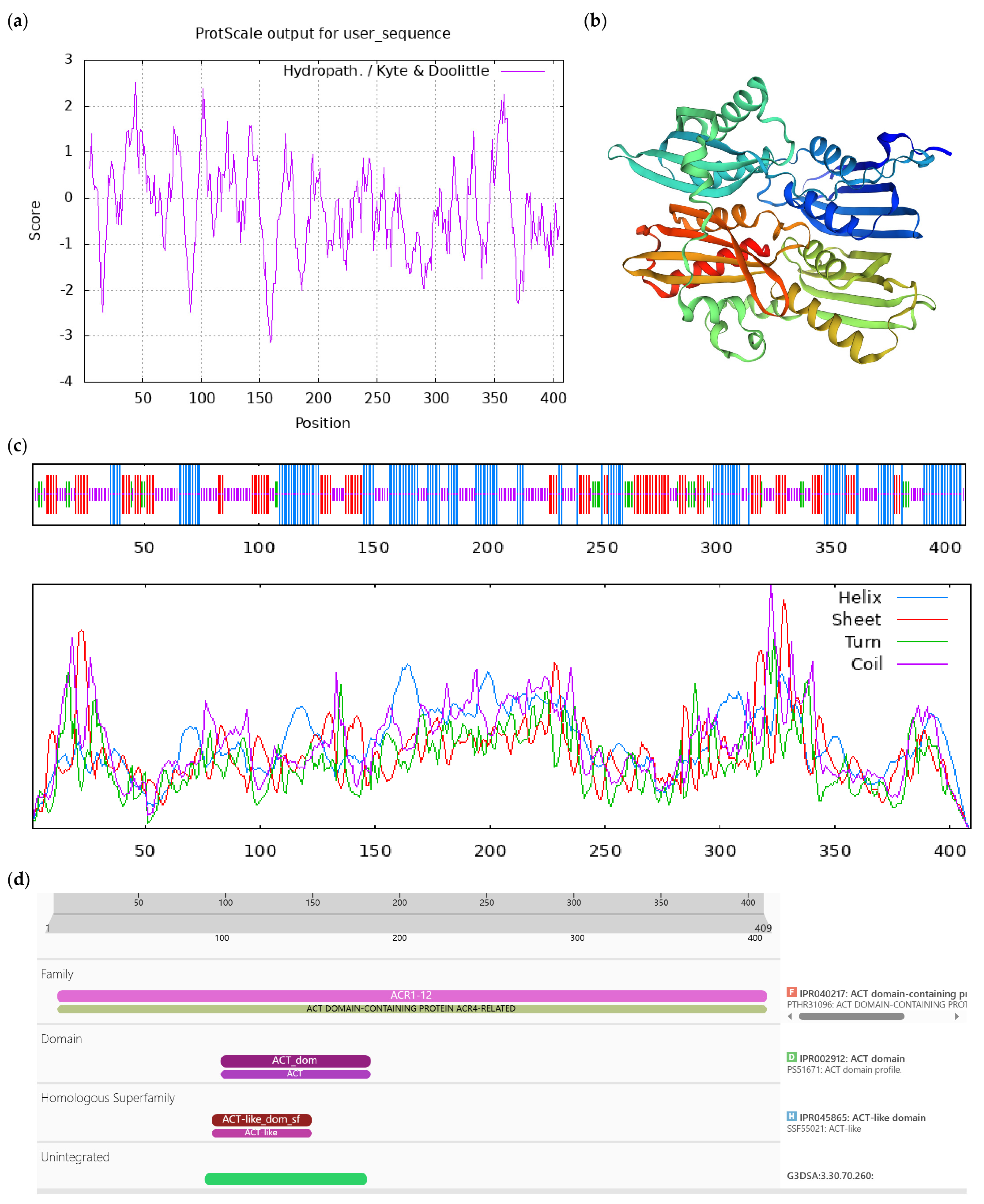
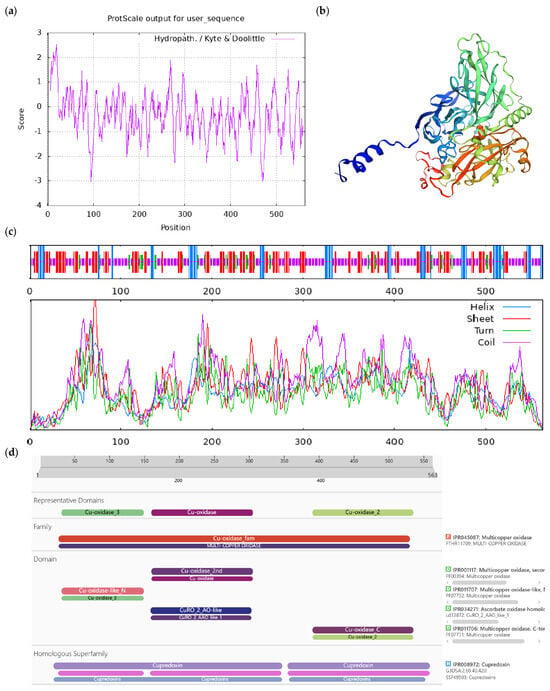
Figure A2.
Bioinformatics analysis of CoLAO protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoLAO protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoLAO protein; (c) secondary structure prediction of CoLAO protein; (d) conserved domains of CoLAO protein.
Figure A2.
Bioinformatics analysis of CoLAO protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoLAO protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoLAO protein; (c) secondary structure prediction of CoLAO protein; (d) conserved domains of CoLAO protein.
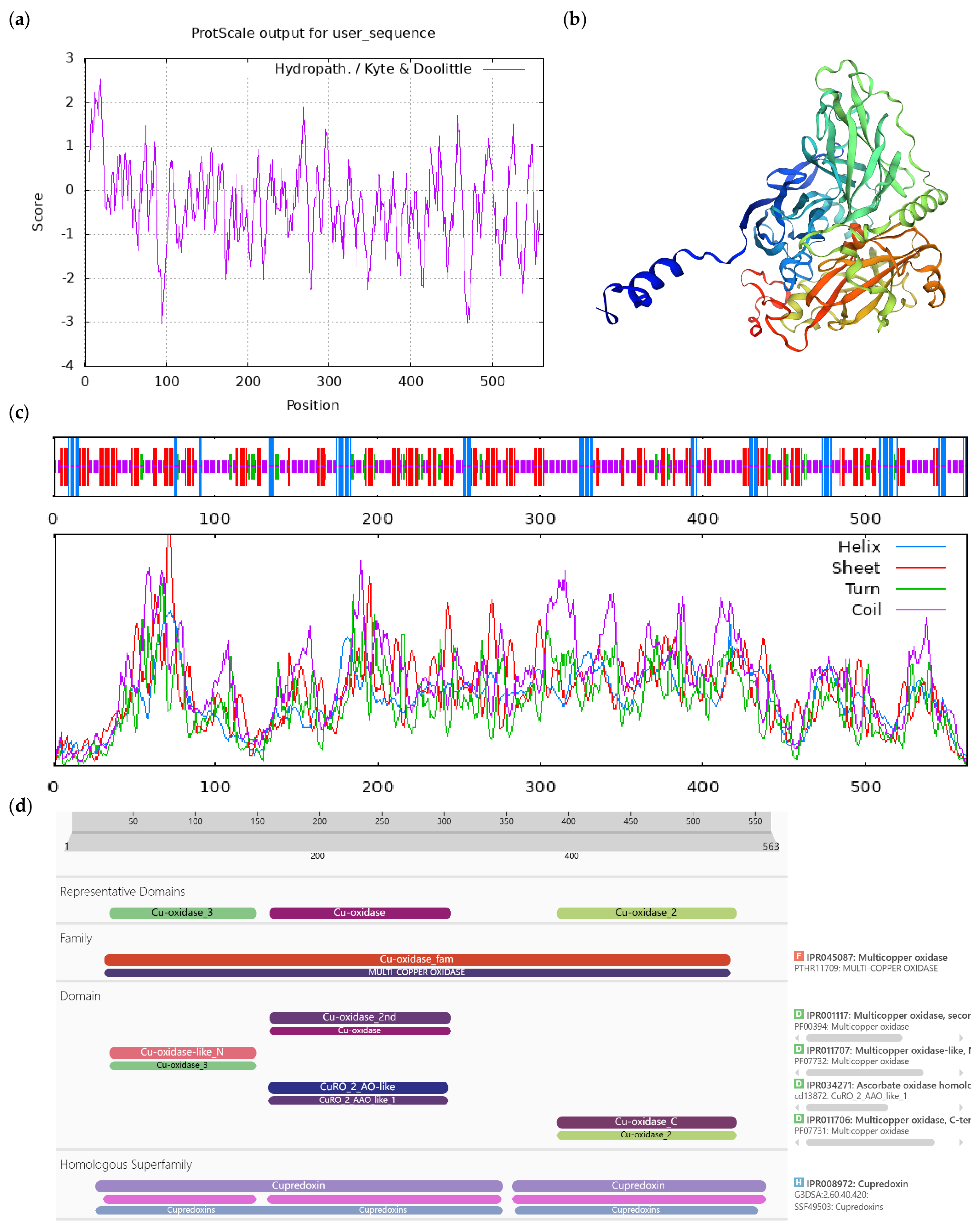
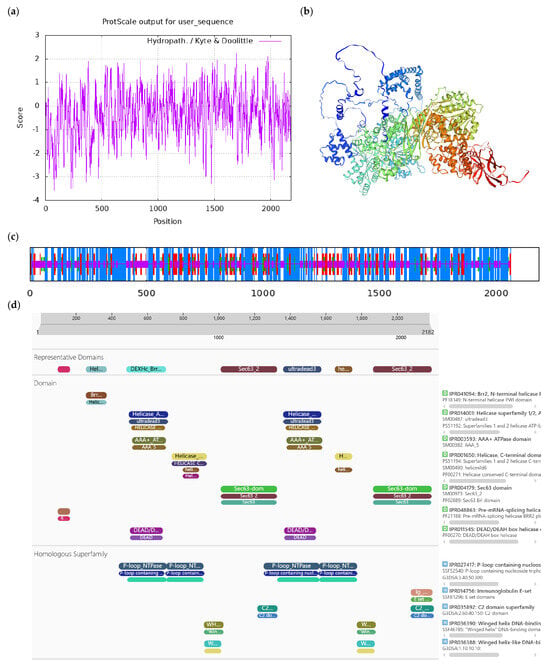
Figure A3.
Bioinformatics analysis of CoDExH12-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoDExH12-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoDExH12-like protein; (c) secondary structure prediction of CoDExH12-like protein; (d) conserved domains of CoDExH12-like protein.
Figure A3.
Bioinformatics analysis of CoDExH12-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoDExH12-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoDExH12-like protein; (c) secondary structure prediction of CoDExH12-like protein; (d) conserved domains of CoDExH12-like protein.
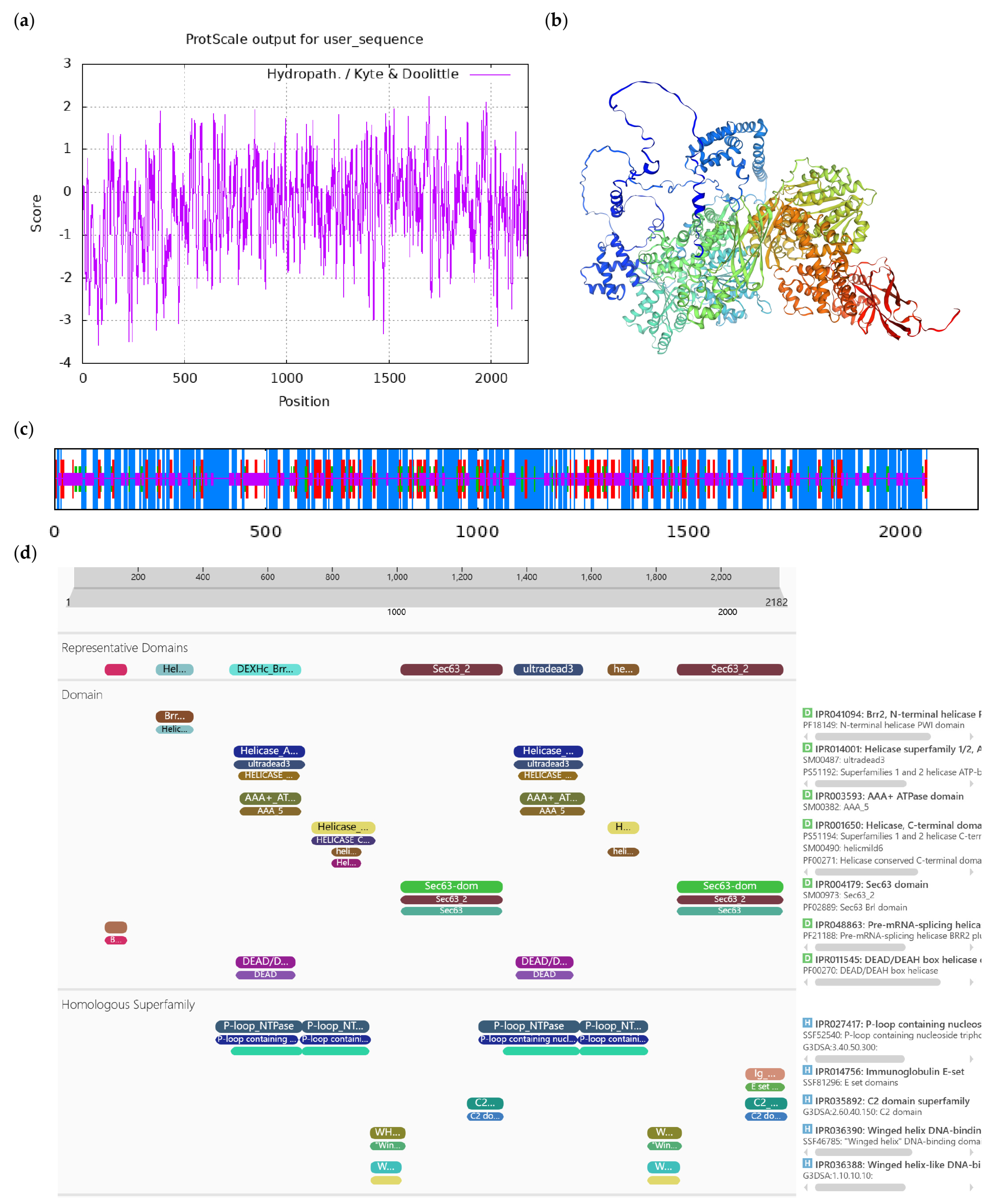
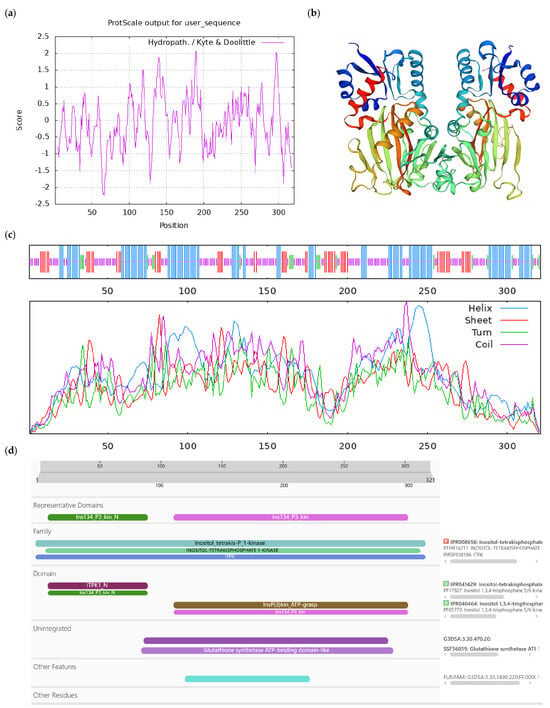
Figure A4.
Bioinformatics analysis of CoIT1K-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoIT1K-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoIT1K-like protein; (c) secondary structure prediction of CoIT1K-like protein; (d) conserved domains of CoIT1K-like protein.
Figure A4.
Bioinformatics analysis of CoIT1K-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoIT1K-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoIT1K-like protein; (c) secondary structure prediction of CoIT1K-like protein; (d) conserved domains of CoIT1K-like protein.
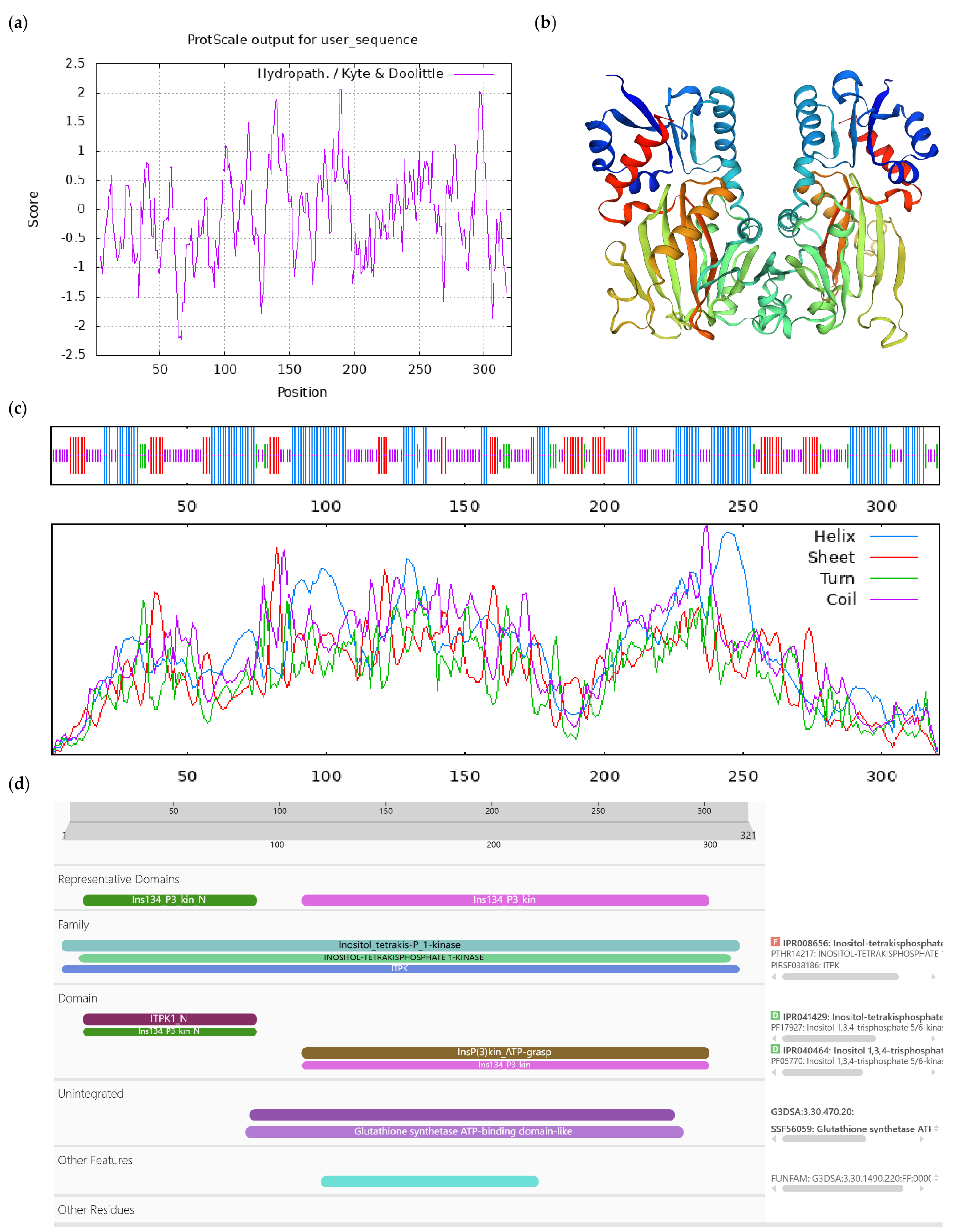
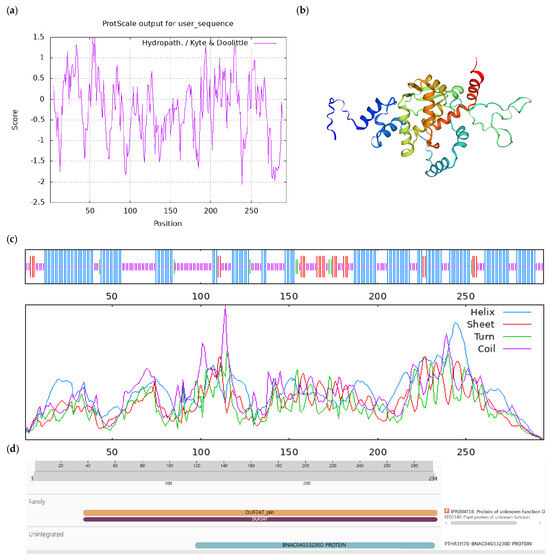
Figure A5.
Bioinformatics analysis of CoUPF0481 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoUPF0481 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoUPF0481 protein; (c) secondary structure prediction of CoUPF0481 protein; (d) conserved domains of CoUPF0481 protein.
Figure A5.
Bioinformatics analysis of CoUPF0481 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoUPF0481 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoUPF0481 protein; (c) secondary structure prediction of CoUPF0481 protein; (d) conserved domains of CoUPF0481 protein.
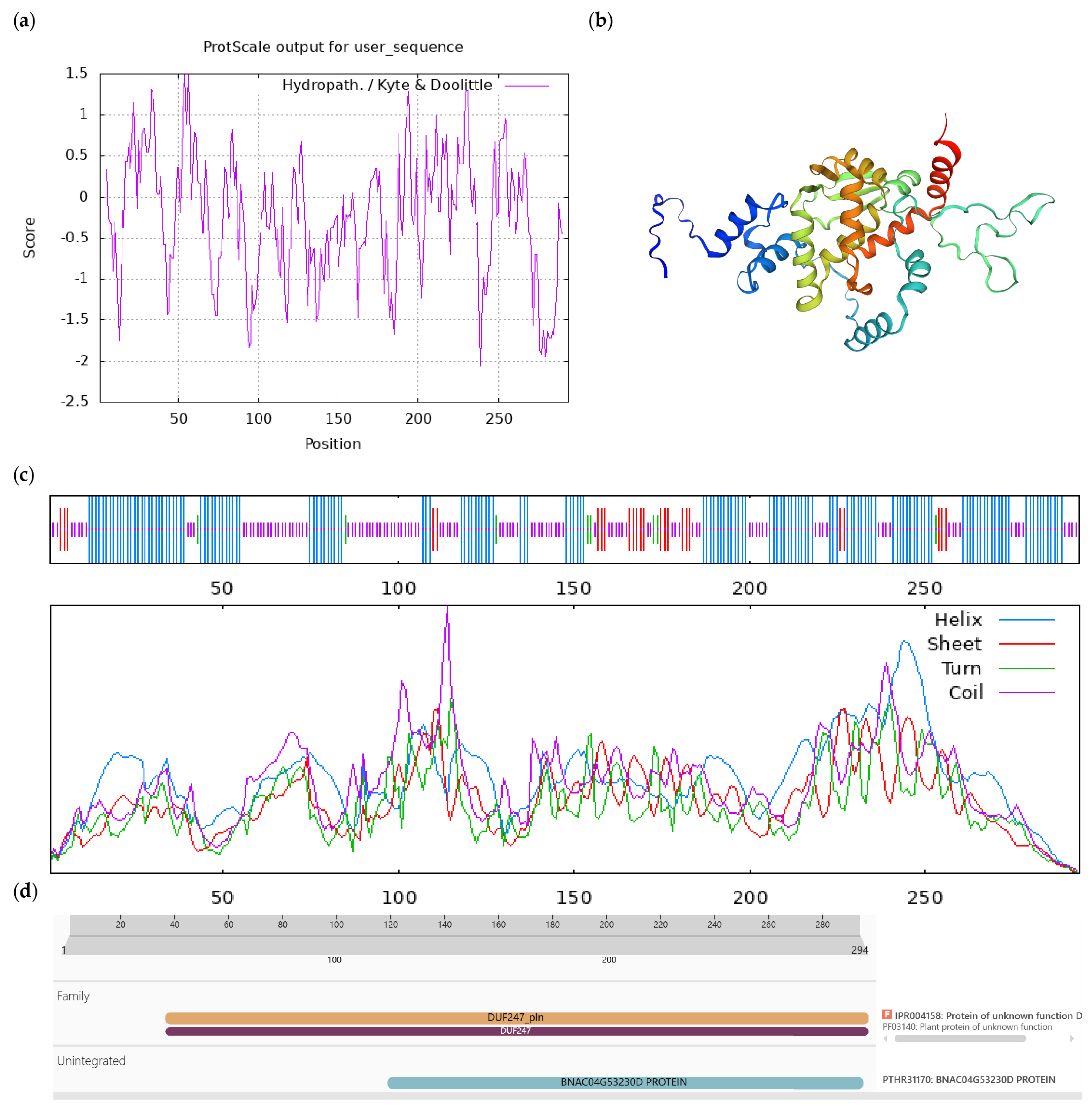
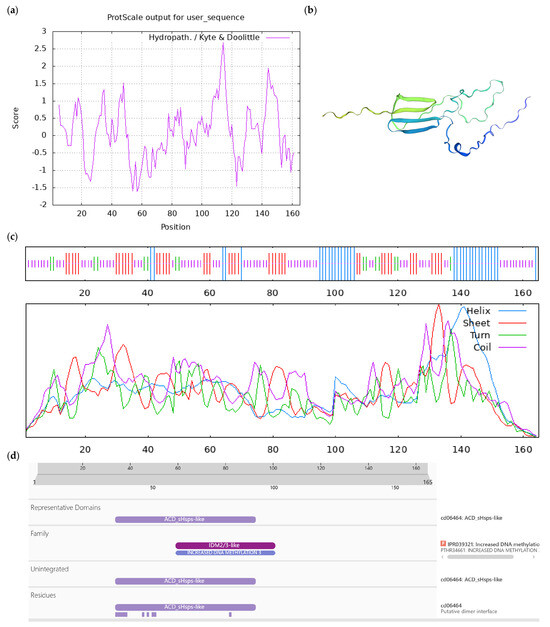
Figure A6.
Bioinformatics analysis of CoIDM3 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoIDM3 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoIDM3 protein; (c) secondary structure prediction of CoIDM3 protein; (d) conserved domains of CoIDM3 protein.
Figure A6.
Bioinformatics analysis of CoIDM3 protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoIDM3 protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoIDM3 protein; (c) secondary structure prediction of CoIDM3 protein; (d) conserved domains of CoIDM3 protein.
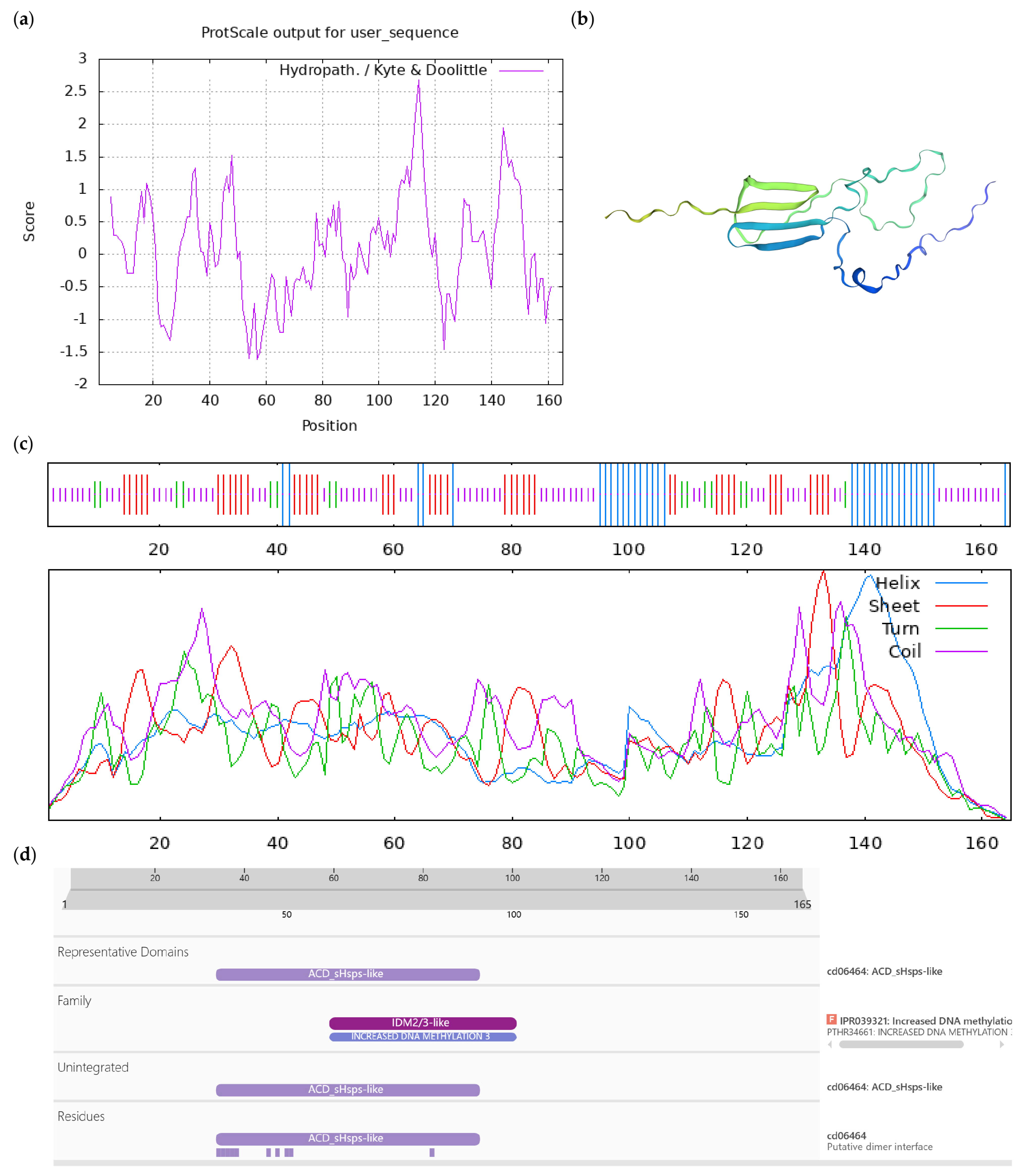
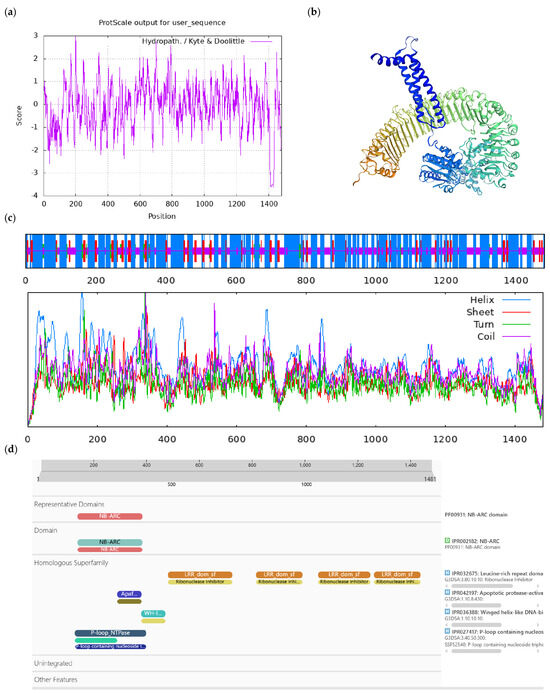
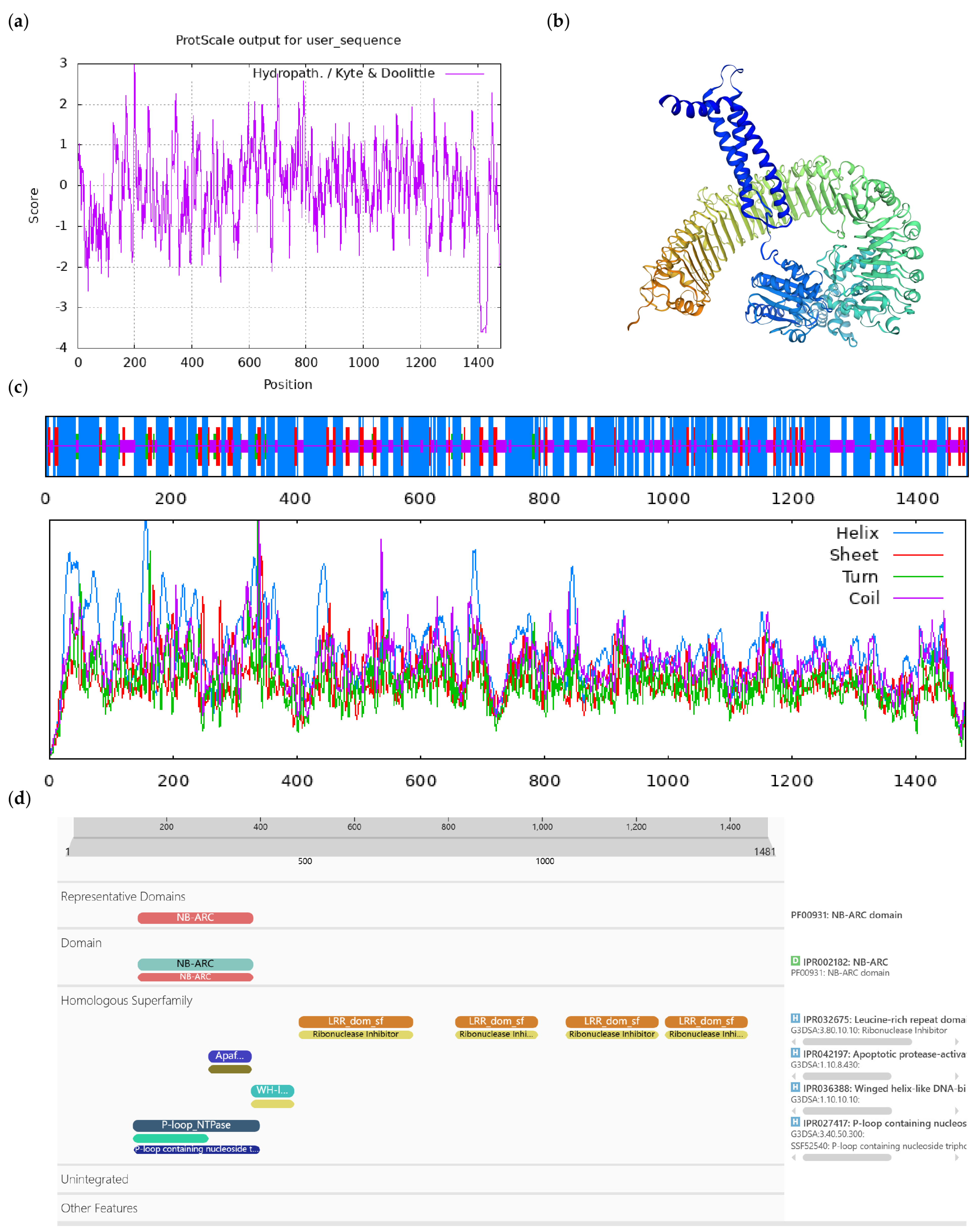
Figure A7.
Bioinformatics analysis of CoAt4g27190-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoAt4g27190-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoAt4g27190-like protein; (c) secondary structure prediction of CoAt4g27190-like protein; (d) conserved domains of CoAt4g27190-like protein.
Figure A7.
Bioinformatics analysis of CoAt4g27190-like protein. Notes: (a) Hydrophilicity and hydrophobicity analysis of CoAt4g27190-like protein, negative number is hydrophilic, positive number is hydrophobic; (b) tertiary structure prediction of CoAt4g27190-like protein; (c) secondary structure prediction of CoAt4g27190-like protein; (d) conserved domains of CoAt4g27190-like protein.
References
- Wang, R.; He, Z.; Zhang, Y.; Zhang, Z.; Wang, X.; Chen, Y. Physiological and Molecular Responses of Camellia oleifera Seedlings to Varied Nitrogen Sources. Horticulturae 2023, 9, 1243. [Google Scholar] [CrossRef]
- Luo, J.; Wei, S.; Zhou, X.; Tian, Y.; Chen, Y.; Song, Q.; Chen, L. Nutrient contents in the organs and soil of young and mature Camellia oleifera C. Abel forests in China. Bangladesh J. Bot. 2022, 51, 359–369. [Google Scholar]
- Wu, L.L.; Li, J.A.; Gu, Y.Y.; Zhang, F.H.; Gu, L.; Tan, X.F.; Shi, M.W. Effect of Chilling Temperature on Chlorophyll Florescence, Leaf Anatomical Structure, and Physiological and Biochemical Characteristics of Two Camellia oleifera Cultivars. Int. J. Agric. Biol. 2020, 23, 777–785. [Google Scholar]
- Zhang, Y.Q.; Guo, Q.Q.; Luo, S.Q.; Pan, J.W.; Yao, S.; Gao, C.; Guo, Y.Y.; Wang, G. Light Regimes Regulate Leaf and Twigs Traits of Camellia oleifera (Abel) in Pinus massoniana Plantation Understory. Forests 2022, 13, 918. [Google Scholar] [CrossRef]
- Hoong-Yeet, Y. Solar rhythm in the regulation of photoperiodic flowering of long-day and short-day plants. J. Exp. Bot. 2013, 64, 2643–2652. [Google Scholar] [CrossRef]
- Chow, B.Y.; Kay, S.A. Global approaches for telling time: Omics and the Arabidopsis circadian clock. Semin. Cell Dev. Biol. 2013, 24, 383–392. [Google Scholar] [CrossRef]
- Suarez-Lopez, P.; Wheatley, K.; Robson, F.; Onouchi, H.; Valverde, F.; Coupland, G. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 2001, 410, 1116–1120. [Google Scholar] [CrossRef] [PubMed]
- Andrés, F.; Coupland, G. The genetic basis of flowering responses to seasonal cues. Nature reviews. Genetics 2012, 13, 627–639. [Google Scholar] [CrossRef]
- Shuai, M.M. Evolution Mechanism of Photoperiodic Floral Genes GIGANTEA and CONSTANS. Master’s Thesis, Zhejiang A&F University, Hangzhou, China, 2018. [Google Scholar]
- Oliverio, K.A.; Crepy, M.; Martin-Tryon, E.L.; Milich, R.; Harmer, S.L.; Putterill, J.; Yanovsky, M.J.; Casal, J.J. GIGANTEA regulates phytochrome A-mediated photomorphogenesis independently of its role in the circadian clock. Plant Physiol. 2007, 144, 495–502. [Google Scholar] [CrossRef][Green Version]
- Qi, X.H.; Wu, D.T.; Li, G.Z.; Zhao, J.L.; Li, M.L. Research progress of flower formation regulation pathway of Arabidopsis. J. Shanxi Agric. Univ. (Nat. Sci. Ed.) 2018, 38, 36. [Google Scholar] [CrossRef]
- Nohales, M.A.; Kay, S.A. Molecular mechanisms at the core of the plant circadian oscillator. Nat. Struct. Mol. Biol. 2016, 23, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- Usman, M.A.; Amanda, D.; Jon, S.D.; Marcel, Q. Photoperiod sensing of the circadian clock is controlled by EARLY FLOWERING 3 and GIGANTEA. Plant J. Cell Mol. Biol. 2020, 101, 1397–1410. [Google Scholar] [CrossRef]
- Kim, H.; Park, S.J.; Kim, Y.; Nam, H.G. Subcellular Localization of GIGANTEA Regulates the Timing of Leaf Senescence and Flowering in Arabidopsis. Front. Plant Sci. 2020, 11, 589707. [Google Scholar] [CrossRef]
- Liang, Q.Z.; Song, K.H.; Lu, M.S.; Dai, T.; Yang, J.; Wan, J.X.; Li, L.; Chen, J.J.; Zhan, R.L.; Wang, S.B. Transcriptome and Metabolome Analyses Reveal the Involvement of Multiple Pathways in Flowering Intensity in Mango. Front. Plant Sci. 2022, 13, 933923. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Geng, R.; Gallenstein, R.A.; Somers, D.E. The F-box protein ZEITLUPE controls stability and nucleocytoplasmic partitioning of GIGANTEA. Development 2013, 140, 4060–4069. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Wang, X.F.; Zheng, C.S.; Xing, S.Y.; Shu, H.R. Full-length cDNA cloning, sequence information and quantitative expression analysis of CmGI (GIGANTEA) output gene from Chrysanthemum circadian clock. Chin. J. Agric. Sci. 2012, 45, 2690–2703. [Google Scholar]
- Izawa, T.; Mihara, M.; Suzuki, Y.; Gupta, M.; Itoh, H.; Nagano, A.J.; Motoyama, R.; Sawada, Y.; Yano, M.; Hirai, M.Y.; et al. Os-GIGANTEA confers robust diurnal rhythms on the global transcriptome of rice in the field. Plant Cell 2011, 23, 1741–1755. [Google Scholar] [CrossRef]
- Karlgren, A.; Gyllenstrand, N.; Källman, T.; Lagercrantz, U. Conserved function of core clock proteins in the gymnosperm Norway spruce (Picea abies L. Karst). PLoS ONE 2013, 8, e60110. [Google Scholar] [CrossRef]
- Yan, J.D. Molecular Genetics and Biochemical Analysis of Arabidopsis FKF1 Controlling Flowering by Regulating Gibberellin Signaling. Ph.D. Thesis, Hunan University, Changsha, China, 2020. [Google Scholar] [CrossRef]
- Ren, S.S. Cloning and Functional Study of Blue Light Receptor Gene CoZTL in Camellia oleifolia. Master’s Thesis, Central South University of Forestry and Technology, Changsha, China, 2023. [Google Scholar] [CrossRef]
- You, L.N. Screening and Functional Analysis of VfbZIP and VfWRKY Genes Involved in Oil Synthesis of Alnia oleifera. Master’s Thesis, Central South University of Forestry and Technology, Changsha, China, 2023. [Google Scholar] [CrossRef]
- Wang, X.N. Research on Phenology and Blossom Biology of Camellia oleifera Abel. Master’s Thesis, Central South University of Forestry and Technology, Changsha, China, 2011. [Google Scholar]
- Tai, Y.; Wei, C.; Yang, H.; Zhang, L.; Chen, Q.; Deng, W.; Wei, S.; Zhang, J.; Fang, C.; Ho, C.; et al. Transcriptomic and phytochemical analysis of the biosynthesis of characteristic constituents in tea (Camellia sinensis) compared with oil tea (Camellia oleifera). BMC Plant Biol. 2015, 15, 190. [Google Scholar] [CrossRef]
- Wiktorek-Smagur, A.; Hnatuszko-Konka, K.; Kononowicz, A.K. Flower bud dipping or vacuum infiltration-two methods of Arabidopsis thaliana transformation. Russ. J. Plant Physiol. 2009, 56, 560–568. [Google Scholar] [CrossRef]
- Ren, S.S.; Juan, L.M.; He, J.C.; Liu, Q.; Yan, J.D.; Li, J.A. Expression Analysis and Interaction Protein Screening of CoZTL in Camellia oleifera Abel. Horticulturae 2023, 9, 833. [Google Scholar] [CrossRef]
- Makuch, L. Yeast Two-Hybrid Screen. Methods Enzymol. 2014, 539, 31–51. [Google Scholar] [CrossRef] [PubMed]
- Soellick, T.R.; Uhrig, J.F. Development of an optimized interaction-mating protocol for large-scale yeast two-hybrid analyses. Genome Biol. 2001, 2, research0052.0051. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.Q.; Ling, H.E.; Jun, W.; Cheng, J. Screening of hepatocyte proteins binding with C-terminally truncated surface antigen middle protein of hepatitis B virus (MHBst167) by a yeast two-hybrid system. Mol. Med. Rep. 2014, 10, 1259–1263. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Wang, T.; Che, J.J.; Yu, F.; Liu, H.L.; Zuo, Z.Y.; Yang, Z.H.; Fan, H.D. Screening and identification of proteins interacting with IL-24 by the yeast two-hybrid screen, Co-IP, and FRET assays. Anti-Cancer Drugs 2016, 27, 318–327. [Google Scholar] [CrossRef] [PubMed]
- Kundu, P.; Sahu, R. GIGANTEA confers susceptibility to plants during spot blotch attack by regulating salicylic acid signalling pathway. Plant Physiol. Biochem. 2021, 167, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.G.; Seo, P.J. Dependence and independence of the root clock on the shoot clock in Arabidopsis. Genes Genom. 2018, 40, 1063–1068. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Zhang, X.M.; Hu, R.B.; Wu, F.Q.; Ma, J.H.; Meng, Y.; Fu, Y.F. Identification and molecular characterization of FKF1 and GI homologous genes in soybean. PLoS ONE 2013, 8, e79036. [Google Scholar] [CrossRef]
- Zhou, X.C.; Han, H.Q.; Chen, H.Y.; Liu, Y. Cloning and expression analysis of biorhythm clock output gene BnGI in Brassica napus. J. Shanghai Jiao Tong Univ. (Agric. Sci. Ed.) 2014, 32, 5–11. [Google Scholar]
- Tang, W.; Yan, H.; Su, Z.X.; Park, S.C.; Liu, Y.J.; Zhang, Y.J.; Wang, X.; Kou, M.; Ma, D.F.; Kwak, S.S.; et al. Cloning and characterization of a novel GIGANTEA gene in sweet potato. Plant Physiol. Biochem. 2017, 116, 27–35. [Google Scholar] [CrossRef]
- Karsai, R.F.; Odgerel, K.; Jose, J.; Bánfalvi, Z. In Silico Characterization and Expression Analysis of GIGANTEA Genes in Potato. Biochem. Genet. 2022, 60, 2137–2154. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.F. Expression Analysis and Functional Verification of ClGI Gene in Citrus limon. Master’s Thesis, Huazhong Agricultural University, Wuhan, China, 2019. [Google Scholar] [CrossRef]
- Yuan, X.; Liu, J.M.; Guo, C.H.; Kang, C.; Zhang, Z.G.; Quan, S.W.; Niu, J.X. Cloning and expression analysis of JrGI gene in walnut. Chin. J. Bioeng. 2023, 39, 640–652. [Google Scholar] [CrossRef]
- Song, Y.; Jiang, Y.; Kuai, B.; Li, L. CIRCADIAN CLOCK-ASSOCIATED 1 Inhibits Leaf Senescence in Arabidopsis. Front. Plant Sci. 2018, 9, 280. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Kim, H.J.; Vu, Q.T.; Jung, S.; McClung, C.R.; Hong, S.; Nam, H.G. Circadian control of ORE1 by PRR9 positively regulates leaf senescence in Arabidopsis. Proc. Natl. Acad. Sci. USA 2018, 115, 8448–8453. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.Y.; Wang, Y.; Wei, H.; Li, N.; Tian, W.W.; Chong, K.; Wang, L. Circadian Evening Complex Represses Jasmonate-Induced Leaf Senescence in Arabidopsis. Mol. Plant 2018, 11, 326–337. [Google Scholar] [CrossRef] [PubMed]
- Takato, I.; Thomas, F.S.; Frank, G.H.; Lindsey, A.H.; Steve, A.K. FKF1 F-Box Protein Mediates Cyclic Degradation of a Repressor of CONSTANS in Arabidopsis. Science 2005, 309, 293–297. [Google Scholar] [CrossRef]
- Kim, W.Y.; Fujiwara, S.; Suh, S.S.; Kim, J.; Kim, Y.; Han, L.Q.; David, K.; Putterill, J.; Nam, H.G.; Somers, D.E. ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature 2007, 449, 356–360. [Google Scholar] [CrossRef] [PubMed]
- Mariko, S.; Dmitri, A.N.; Steve, A.K.; Takato, I. FKF1 and GIGANTEA Complex Formation is Required for Day-Length Measurement in Arabidopsis. Science 2007, 318, 261–265. [Google Scholar] [CrossRef]
- Fabio, F.; Kishore, C.S.P.; Lionel, G.; Nicolas, S.; Mark, R.; José, A.J.; George, C. Arabidopsis DOF Transcription Factors Act Redundantly to Reduce CONSTANS Expression and Are Essential for a Photoperiodic Flowering Response. Dev. Cell 2009, 17, 75–86. [Google Scholar] [CrossRef]
- Tang, W.; Liu, Y.J.; Zhang, Y.G.; Wang, X.; Hou, M.; Yan, H.; Ma, D.F.; Li, Q. Research progress on physiological functions of plant GI genes. Mol. Plant Breed. 2014, 12, 1044–1049. [Google Scholar] [CrossRef]
- Sawa, M.; Kay, S.A. GIGANTEA directly activates Flowering Locus T in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2011, 108, 11698–11703. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.Y.; Lee, S.M.; Seo, P.J.; Yang, M.S.; Park, C.M. Identification and molecular characterization of a Brachypodium distachyon GIGANTEA gene: Functional conservation in monocot and dicot plants. Plant Mol. Biol. 2010, 72, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Kudo, T.; Kawai, A.; Yamaya, T.; Hayakawa, T. Cellular distribution of ACT domain repeat protein 9, a nuclear localizing protein, in rice (Oryza sativa). Physiol. Plant. 2008, 133, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Liao, H.S.; Chen, Y.J.; Hsieh, W.Y.; Li, Y.C.; Hsieh, M.H. Arabidopsis ACT DOMAIN REPEAT9 represses glucose signaling pathways. Plant Physiol. 2023, 192, 1532–1547. [Google Scholar] [CrossRef] [PubMed]
- Cortaga, C.Q.; Latina, R.A.; Habunal, R.R.; Lantican, D.V. Identification and characterization of genome-wide resistance gene analogs (RGAs) of durian (Durio zibethinus L.). J. Genet. Eng. Biotechnol. 2022, 20, 29. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.J.; Chen, H.B.; Hu, Z.Q.; Lu, X.Y.; Wang, H.Y.; Liu, H.; Zhou, B.Y. Comparative proteomics of phloem exudates reveals long-distance signals potentially involved in Litchi chinensis flowering. Biol. Plant. 2020, 64, 220–224. [Google Scholar] [CrossRef]
- Liu, R.X. Molecular Mechanism of Differential Expression of Transcriptome and Cell Wall Invertase Genes in Different Cupr-resistant Populations of Elsholtzia Haizhou. Ph.D. Thesis, Wuhan University, Wuhan, China, 2023. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, J.J.; Li, C.G.; Chen, S.K.; Tang, Q.L.; Xiao, Y.F.; Zhong, L.X.; Chen, Y.Y.; Chen, B. Gene expression programs during callus development in tissue culture of two Eucalyptus species. BMC Plant Biol. 2022, 22, 1. [Google Scholar] [CrossRef]
- Miao, W.; Dai, J.; Wang, Y.T.; Wang, Q.Q.; Lu, C.; La, Y.M.; Niu, J.Y.; Tan, F.; Zhou, S.X.; Wu, Y.F.; et al. Roles of IDM3 and SDJ1/2/3 in Establishment and/or Maintenance of DNA Methylation in Arabidopsis. Plant Cell Physiol. 2021, 62, 1409–1422. [Google Scholar] [CrossRef] [PubMed]
- Duan, C.G.; Wang, X.G.; Xie, S.J.; Pan, L.; Miki, D.; Tang, K.; Hsu, C.C.; Lei, M.G.; Zhong, Y.L.; Hou, Y.; et al. A pair of transposon-derived proteins function in a histone acetyltransferase complex for active DNA demethylation. Cell Res. 2017, 27, 226–240. [Google Scholar] [CrossRef]
- Lang, Z.B.; Lei, M.G.; Wang, X.G.; Tang, K.; Miki, D.; Zhang, H.M.; Mangrauthia, S.K.; Liu, W.S.; Nie, W.F.; Ma, G.J.; et al. The Methyl-CpG-Binding Protein MBD7 Facilitates Active DNA Deme-thylation to Limit DNA Hyper-Methylation and Transcriptional Gene Silencing. Mol. Cell 2015, 57, 971–983. [Google Scholar] [CrossRef]
- Guo, Y.; Zhu, J.; Xu, Z.C.; Zhang, S.C. Research progress of plant ascorbate oxidase. Chin. Agric. Sci. Bull. 2008, 3, 196–199. [Google Scholar]
- Dowdle, J.; Ishikawa, T.; Gatzek, S.; Rolinski, S.; Smirnoff, N. Two Genes in Arabidopsis Tha-liana Encoding GDP-L-Galactose Phosphorylase Are Required for Ascor-bate Biosynthesis and Seedling Viability. Plant J. Cell Mol. Biol. 2007, 52, 673–689. [Google Scholar] [CrossRef] [PubMed]
- Aniqa, A.; Nudrat, A.A.; Muhammad, A. Influence of natural and synthetic vitamin C (ascorbic acid) on primary and secondary metabolites and associated metabolism in quinoa (Chenopodium quinoa Willd.) plants under water deficit regimes. Plant Physiol. Biochem. 2018, 123, 192–203. [Google Scholar] [CrossRef]
- Li, C. Cloning and Low Temperature Response Mechanism of Tobacco Ascorbate Oxidase Promoter. Master’s Thesis, Henan Agricultural University, Zhengzhou, China, 2021. [Google Scholar] [CrossRef]
- Kim, M.; Ohr, H.; Lee, J.W.; Hyun, Y.; Fischer, R.L.; Choi, Y. Temporal and spatial downregulation of Arabidopsis MET1 activity results in global DNA hypomethylation and developmental defects. Mol. Cells 2008, 26, 611–615. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).