KIR Genotypes Impact Progression to Hepatocellular Carcinoma in Patients with Chronic Hepatitis C Infection
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
2.2. DNA Extraction
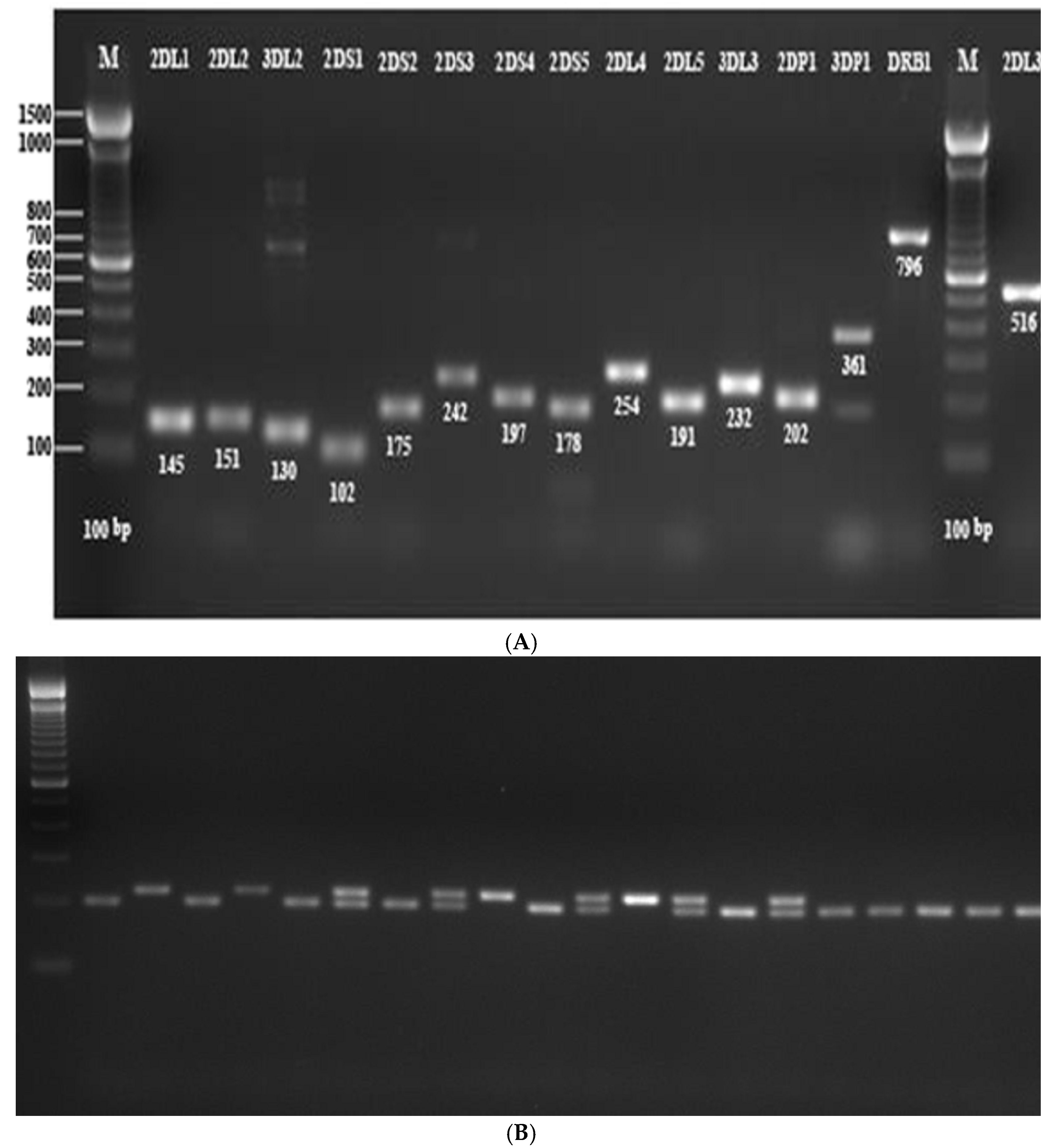
2.3. KIR Genotyping
2.4. KIR2DS4 Analysis
2.5. Histological Assessment of Inflammatory Cells Infiltration in HCC Tumors
2.6. Statistical Data Analysis
3. Results
3.1. Frequencies of KIR Genes and Genotypes in the Studied Populations
3.2. Correlation between the Degree of Inflammatory Infiltration in the Tumor and KIR Genes
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ibrahim, A.S.; Khaled, H.M.; Mikhail, N.N.; Baraka, H.; Kamel, H. Cancer incidence in egypt: Results of the national population-based cancer registry program. J. Cancer Epidemiol. 2014, 2014, 437971. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ministry of Health and Population [Egypt]. El-Zanaty and Associates [Egypt] and ICF International. In 2015 Egypt Health Issues Survey; Ministry of Health: Cairo, Egypt; Population and ICF International: Rockville, MD, USA, 2015. [Google Scholar]
- Kanwal, F.; Khaderi, S.; Singal, A.G.; Marrero, J.A.; Loo, N.; Asrani, S.K.; Amos, C.I.; Thrift, A.P.; Gu, X.; Luster, M.; et al. Risk factors for HCC in contemporary cohorts of patients with cirrhosis. Hepatology 2023, 77, 997–1005. [Google Scholar] [CrossRef]
- Peng, H.; Wisse, E.; Tian, Z. Liver natural killer cells: Subsets and roles in liver immunity. Cell Mol. Immunol. 2016, 13, 328–336. [Google Scholar] [CrossRef] [Green Version]
- Martin, A.M.; Freitas, E.M.; Witt, C.S.; Christiansen, F.T. The genomic organization and evolution of the natural killer immunoglobulin-like receptor (KIR) gene cluster. Immunogenetics 2000, 51, 268–280. [Google Scholar] [CrossRef]
- Trowsdale, J.; Parham, P. Mini-review: Defense strategies and immunity-related genes. Eur. J. Immunol. 2004, 34, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Galarza, F.F.; Christmas, S.; Middleton, D.; Jones, A.R. Allele frequency net: A database and online repository for immune gene frequencies in worldwide populations. Nucleic. Acids. Res. 2011, 39, D913–D919. [Google Scholar] [CrossRef] [Green Version]
- Lanier, L.L.; Corliss, B.C.; Wu, J.; Leong, C.; Phillips, J.H. Immunoreceptor DAP12 bearing a tyrosine-based activation motif is involved in activating NK cells. Nature 1998, 391, 703–707. [Google Scholar] [CrossRef]
- Martin, M.P.; Carrington, M. KIR Locus lymorphisms Genotyping and Disease Association Analysis. In Methods in Molecular Biology; Ewbank, E., Vivier, E., Eds.; Innate immunity; Humana Press: Totowa, NJ, USA, 2008. [Google Scholar] [CrossRef]
- Gomez-Lozano, N.; Vilches, C. Genotyping of human killer-cell immunoglobulin-like receptor genes by polymerase chain reaction with sequence-specific primers: An update. Tissue Antigens 2002, 59, 184–193. [Google Scholar] [CrossRef]
- Hayashi, A.; Shibahara, J.; Misumi, K.; Arita, J.; Sakamoto, Y.; Hasegawa, K.; Kokudo, N.; Fukayama, M. Histologic Assessment of Intratumoral Lymphoplasmacytic Infiltration Is Useful in Predicting Prognosis of Patients with Hepatocellular Carcinoma. PLoS ONE 2016, 11, e0155744. [Google Scholar] [CrossRef]
- Littera, R.; Zamboni, F.; Tondolo, V.; Fantola, G.; Chessa, L.; Orru, N.; Sanna, M.; Valentini, D.; Cappai, L.; Mulargia, M.; et al. Absence of activating killer immunoglobulin-like receptor genes combined with hepatitis C viral genotype is predictive of hepatocellular carcinoma. Hum. Immunol. 2013, 74, 1288–1294. [Google Scholar] [CrossRef]
- Castrillon, M.; Marin, N.D.; Karduss-Urueta, A.J.; Velasquez, S.Y.; Alvarez, C.M. Killer-Cell Immunoglobulin-like Receptor Diversity in an Admixed South American Population. Cells 2022, 11, 2776. [Google Scholar] [CrossRef]
- Shen, C.; Ge, Z.; Dong, C.; Wang, C.; Shao, J.; Cai, W.; Huang, P.; Fan, H.; Li, J.; Zhang, Y.; et al. Genetic Variants in KIR/HLA-C Genes Are Associated With the Susceptibility to HCV Infection in a High-Risk Chinese Population. Front. Immunol. 2021, 12, 632353. [Google Scholar] [CrossRef] [PubMed]
- Yindom, L.M.; Mendy, M.; Bodimeade, C.; Chambion, C.; Aka, P.; Whittle, H.C.; Rowland-Jones, S.L.; Walton, R. KIR content genotypes associate with carriage of hepatitis B surface antigen, e antigen and HBV viral load in Gambians. PLoS ONE 2017, 12, e0188307. [Google Scholar] [CrossRef] [Green Version]
- La Nasa, G.; Greco, M.; Littera, R.; Oppi, S.; Celeghini, I.; Caria, R.; Lai, S.; Porcella, R.; Martino, M.; Romano, A.; et al. The favorable role of homozygosity for killer immunoglobulin-like receptor (KIR) A haplotype in patients with advanced-stage classic Hodgkin lymphoma. J. Hematol. Oncol. 2016, 9, 26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barani, S.; Khademi, B.; Ashouri, E.; Ghaderi, A. KIR2DS1, 2DS5, 3DS1 and KIR2DL5 are associated with the risk of head and neck squamous cell carcinoma in Iranians. Hum. Immunol. 2018, 79, 218–223. [Google Scholar] [CrossRef]
- Parham, P.; Moffett, A. Variable NK cell receptors and their MHC class I ligands in immunity, reproduction and human evolution. Nat. Rev. Immunol. 2013, 13, 133–144. [Google Scholar] [CrossRef] [Green Version]
- Cariani, E.; Missale, G. KIR/HLA immunogenetic background influences the evolution of hepatocellular carcinoma. Oncoimmunology 2013, 2, e26622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, J.; Heller, G.; Chewning, J.; Kim, S.; Yokoyama, W.M.; Hsu, K.C. Hierarchy of the human natural killer cell response is determined by class and quantity of inhibitory receptors for self-HLA-B and HLA-C ligands. J. Immunol. 2007, 179, 5977–5989. [Google Scholar] [CrossRef] [Green Version]
- Beziat, V.; Traherne, J.A.; Liu, L.L.; Jayaraman, J.; Enqvist, M.; Larsson, S.; Trowsdale, J.; Malmberg, K.J. Influence of KIR gene copy number on natural killer cell education. Blood 2013, 121, 4703–4707. [Google Scholar] [CrossRef] [Green Version]
- Tarek, N.; Le Luduec, J.B.; Gallagher, M.M.; Zheng, J.T.; Venstrom, J.M.; Chamberlain, E.; Modak, S.; Heller, G.; Dupont, B.; Cheung, N.K.V.; et al. Unlicensed NK cells target neuroblastoma following anti-GD2 antibody treatment. J. Clin. Investig. 2012, 122, 3260–3270. [Google Scholar] [CrossRef] [Green Version]
- Tanimine, N.; Ohdan, H. Impact of multiplicity of functional KIR-HLA compound genotypes on hepatocellular carcinoma. Oncoimmunology 2015, 4, e983765. [Google Scholar] [CrossRef] [Green Version]
- Ursu, L.D.; Calenic, B.; Diculescu, M.; Dima, A.; Stoian, I.T.; Constantinescu, I. Clinical and histopathological changes in different KIR gene profiles in chronic HCV Romanian patients. Int. J. Immunogenet. 2021, 48, 16–24. [Google Scholar] [CrossRef]
- Umemura, T.; Joshita, S.; Saito, H.; Wakabayashi, S.I.; Kobayashi, H.; Yamashita, Y.; Sugiura, A.; Yamazaki, T.; Ota, M. Investigation of the Effect of KIR-HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients. Cancers 2021, 13, 3267. [Google Scholar] [CrossRef]
- Tanimine, N.; Tanaka, Y.; Kobayashi, T.; Tashiro, H.; Miki, D.; Imamura, M.; Aikata, H.; Tanaka, J.; Chayama, K.; Ohdan, H. Quantitative Effect of Natural Killer-Cell Licensing on Hepatocellular Carcinoma Recurrence after Curative Hepatectomy. Cancer Immunol. Res. 2014, 2, 1142–1147. [Google Scholar] [CrossRef]
- De Re, V.; Caggiari, L.; De Zorzi, M.; Repetto, O.; Zignego, A.L.; Izzo, F.; Tornesello, M.L.; Buonaguro, F.M.; Mangia, A.; Sansonno, D.; et al. Genetic Diversity of the KIR/HLA System and Susceptibility to Hepatitis C Virus-Related Diseases (vol 10, e0117420, 2015). PLoS ONE 2015, 10, e0128849. [Google Scholar] [CrossRef]
- Parham, P. NK cells lose their inhibition. Science 2004, 305, 786–787. [Google Scholar] [CrossRef] [PubMed]
- Pan, N.; Jiang, W.; Sun, H.; Miao, F.Q.; Qiu, J.; Jin, H.; Xu, J.H.; Shi, Q.; Xie, W.; Zhang, J.Q. KIR and HLA Loci Are Associated with Hepatocellular Carcinoma Development in Patients with Hepatitis B Virus Infection: A Case-Control Study. PLoS ONE 2011, 6, e25682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kawarabayashi, N.; Seki, S.; Hatsuse, K.; Ohkawa, T.; Koike, Y.; Aihara, T.; Habu, Y.; Nakagawa, R.; Ami, K.; Hiraide, H.; et al. Decrease of CD56+T cells and natural killer cells in cirrhotic livers with hepatitis C may be involved in their susceptibility to hepatocellular carcinoma. Hepatology 2000, 32, 962–969. [Google Scholar] [CrossRef] [PubMed]





| Studied Variable | HCC n = 83 | HCV n = 100 | Controls n = 120 | p-Value |
|---|---|---|---|---|
| Age | <0.001 | |||
| Mean ± SD | 56.21 ± 6.73 | 46.22 ± 6.53 | 46.97 ± 10.44 | |
| Median | 57.00 | 47.0 | 47.0 | |
| Min–max | 38.0–75.00 | 35.0–60.00 | 21.0–76.00 | |
| Gender | 0.001 | |||
| Female | 13 (15.7) | 21 (21.0) | 45 (37.5) | |
| Male | 70 (84.3) | 79 (79.0) | 75 (62.5) |
| Univariate Analysis | Multivariate Analysis * | |||
|---|---|---|---|---|
| p-Value | OR (95% CI) | Adjusted p-Value | Adjusted OR (95% CI) | |
|
KIR AA haplotype vs. KIR Bx haplotypes | Between HCC and healthy controls | |||
| 0.003 | 0.24 (0.09–0.62) | <0.001 | 0.12 (0.04–0.39) | |
| Between HCC and HCV groups | ||||
| 0.001 | 0.21 (0.08–0.54) | 0.03 | 0.26 (0.08–0.88) | |
| Genotype ID | The Study Groups | p-Values | |||||
|---|---|---|---|---|---|---|---|
| HCC (n = 83) | Chronic HCV (n = 100) | Healthy Control (n = 119) | P1 HCC vs. HCV | P2 HCC vs. Healthy Control | P3 HCV vs. Healthy Control | P4 among the Three Groups | |
| 1 | 6 (7.2) | 27 (27.0) | 29 (24.4) | 0.0005 | 0.001 | 0.65 | 0.001 |
| 6 | 16 (19.3) | 8 (8.0) | 8 (6.7) | 0.02 | 0.006 | 0.79 | 0.01 |
| 5 | 10 (12.0) | 20 (20.0) | 14 (11.8) | 0.14 | 0.95 | 0.09 | 0.16 |
| 2 | 8 (9.6) | 7 (7.0) | 11 (9.2) | 0.51 | 0.92 | 0.54 | 0.77 |
| 4 | 7 (8.4) | 7 (7.0) | 8 (6.7) | 0.71 | 0.64 | 0.93 | 0.89 |
| 71 | 5 (6.0) | 5 (5.0) | 6 (5.0) | 0.76 | 0.76 | 0.98 | 0.94 |
| 19 | 4 (4.8) | 3 (3.0) | 1 (0.8) | 0.52 | 0.73 | 0.33 | 0.21 |
| 7 | 4 (4.8) | 2 (2.0) | 6 (5.0) | 0.28 | 0.94 | 0.29 | 0.46 |
| 73 | 2 (2.4) | 6 (6.0) | 1 (0.8) | 0.23 | 0.36 | 0.04 | 0.07 |
| 3 | 2 (2.4) | 4 (4.0) | 7 (5.9) | 0.53 | 0.23 | 0.57 | 0.48 |
| 9 | 2 (2.4) | 1 (1.0) | 2 (1.7) | 0.45 | 0.71 | 0.99 | 0.75 |
| 90 | 2 (2.4) | 1 (1.0) | 1 (0.8) | 0.45 | 0.36 | 1 | 0.59 |
| 93 | 2 (2.4) | 0 | 0 | 0.2 | 0.16 | 1 | NA |
| 81 | 2 (2.4) | 0 | 1 (0.8) | 0.2 | 0.36 | 1 | NA |
| 18 | 1 (1.2) | 2 (2.0) | 0 | 0.67 | 0.41 | 0.2 | NA |
| 8 | 1 (1.2) | 0 | 2 (1.7) | 0.2 | 0.78 | 0.5 | NA |
| 80 | 1 (1.2) | 0 | 0 | 0.2 | 0.41 | 1 | NA |
| 28 | 1 (1.2) | 1 (1.0) | 0 | 0.89 | 0.2 | 0.45 | NA |
| 64 | 1 (1.2) | 1 (1.0) | 0 | 0.89 | 0.2 | 0.45 | NA |
| 228 | 1 (1.2) | 1 (1.0) | 0 | 0.89 | 0.2 | 0.45 | NA |
| 106 | 1 (1.2) | 0 | 0 | 0.2 | 0.2 | 1 | NA |
| 167 | 1 (1.2) | 0 | 0 | 0.2 | 0.2 | 1 | NA |
| 294 | 1 (1.2) | 0 | 0 | 0.2 | 0.2 | 1 | NA |
| 30 | 1 (1.2) | 0 | 0 | 0.2 | 0.2 | 1 | NA |
| 21 | 1 (1.2) | 0 | 1 (0.8) | 0.2 | 0.79 | 0.99 | NA |
| 69 | 0 | 1 (1.0) | 2 (1.7) | 1 | 0.51 | 0.99 | NA |
| 70 | 0 | 1 (1.0) | 2 (1.7) | 1 | 0.51 | 1 | NA |
| 11 | 0 | 1 (1.0) | 2 (1.7) | 1 | 0.51 | 0.99 | NA |
| 175 | 0 | 1 (1.0) | 0 | 1 | 1 | 0.45 | NA |
| 27 | 0 | 0 | 2 (1.7) | 1 | 0.51 | 0.5 | NA |
| 12 | 0 | 0 | 2 (1.7) | 1 | 0.51 | 0.5 | NA |
| 382 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 331 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 159 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 14 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 20 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 72 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 13 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 35 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 75 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 118 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| 68 | 0 | 0 | 1 (0.8) | 1 | 1 | 1 | NA |
| KIR AA haplotypes | 6 (7.2) | 27 (27.0) | 29 (24.2) | 0.0005 | 0.001 | 0.65 | 0.001 |
| KIR Bx haplotypes | 77 (92.8) | 73 (73.0) | 91 (75.8) | 0.0005 | 0.001 | 0.63 | 0.001 |
| 2DS4 full length | 29 (34.9) | 14 (14.0) | 18 (15.13) | 0.0008 | 0.001 | 0.82 | 0.0004 |
| 2DS4 variant (22 bp deletion) | 38 (45.7) | 55 (55.0) | 68 (57.14) | 0.21 | 0.11 | 0.75 | 0.25 |
| 2DS4 Heterozygous | 9 (10.8) | 29 (29.0) | 24 (20.17) | 0.002 | 0.07 | 0.12 | 0.01 |
| 2DS4 Absence | 7 (8.4) | 2 (2.0) | 9 (7.56) | 0.04 | 0.82 | 0.06 | 0.11 |
| KIR Gene | HCC (n = 83) | Chronic HCV (n = 100) | Healthy Control (n = 120) | P1 HCC vs. HCV | P2 HCC vs. Healthy Control | P3 HCV vs. Healthy Control | P4 among the Three Groups |
|---|---|---|---|---|---|---|---|
| 2DL1 | 82 (98.8) | 100 (100.0) | 119 (99.1) | 1 | 1 | 1 | 1 |
| 2DL2 | 66 (79.51) | 64 (64.0) | 68 (56.7) | 0.18 | 0.01 | 0.33 | 0.03 |
| 2DL3 | 70 (84.34%) | 86 (86.0) | 109 (90.8) | 0.75 | 0.15 | 0.26 | 0.33 |
| 3DL1 | 79 (95.18) | 98 (98.0) | 112 (93.3) | 0.28 | 0.58 | 0.18 | 0.25 |
| 3DL2 | 83 (100.0) | 100 (100.0) | 120 (100.0) | NA | NA | NA | NA |
| 2DL5 | 66 (79.51) | 63 (63.0) | 80 (66.6) | 0.01 | 0.04 | 0.57 | 0.04 |
| 3DL3 | 83 (100.0) | 100 (100.0) | 120 (100.0) | NA | NA | NA | NA |
| 2DS1 | 48 (57.83) | 36 (36.0) | 54 (45.0) | 0.01 | 0.2 | 0.22 | 0.07 |
| 2DS2 | 58 (69.88) | 57 (57.0) | 67 (55.8) | 0.07 | 0.04 | 0.86 | 0.09 |
| 2DS3 | 48 (57.83) | 46 (46.0) | 50 (41.6) | 0.11 | 0.02 | 0.58 | 0.07 |
| 2DS4 | 76 (91.57) | 98 (98.0) | 111 (92.5) | 0.04 | 0.8 | 0.06 | 0.12 |
| 2DS5 | 42 (50.60) | 33 (33.0) | 44 (36.6) | 0.01 | 0.04 | 0.67 | 0.03 |
| 3DS1 | 45 (54.22) | 34 (34.0) | 53 (44.1) | 0.01 | 0.33 | 0.13 | 0.06 |
| 2DL4 | 83 (100.0) | 100 (100.0) | 120 (100.0) | NA | NA | NA | NA |
| 3DP1 | 83(100.0) | 100 (100.0) | 120 (100.0) | NA | NA | NA | NA |
| 2DP1 | 82(98.80) | 100 (100.0) | 119 (99.1) | 1 | 1 | 1 | 1 |
| Genotype ID | Frequency in Tumors | p-Value | |
|---|---|---|---|
| With Infiltration (n = 21) | Without Infiltration (n = 26) | ||
| 1 | 2 (9.52) | 0 (0.00) | 0.19 |
| 2 | 1 (4.76) | 3 (11.54) | 0.4 |
| 4 | 3 (14.28) | 1 (3.85) | 0.2 |
| 5 | 2 (9.52) | 3 (11.54) | 0.82 |
| 6 | 2 (9.52) | 8 (30.77) | 0.07 |
| 7 | 2 (9.52) | 1 (3.85) | 0.42 |
| 8 | 0 (0.00) | 1 (3.85) | 1 |
| 18 | 1 (4.76) | 0 (0.00) | 0.45 |
| 19 | 1 (4.76) | 1 (3.85) | 0.8 |
| 21 | 0 (0.00) | 1 (3.85) | 1 |
| 28 | 0 (0.00) | 1 (3.85) | 1 |
| 30 | 1 (4.76) | 0 (0.00) | 0.45 |
| 71 | 2 (9.52) | 0 (0.00) | 0.19 |
| 73 | 0 (0.00) | 1 (3.85) | 1 |
| 81 | 0 (0.00) | 2 (7.69) | 0.49 |
| 90 | 0 (0.00) | 1 (3.85) | 1 |
| 93 | 0 (0.00) | 1 (3.85) | 1 |
| 167 | 1 (4.76) | 0 (0.00) | 0.45 |
| 294 | 1 (4.76) | 0 (0.00) | 0.45 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdelmaguid, W.; Maher, D.; Kohla, M.A.S.; Ezzat, S.; Moaz, I.; Abdel-Mageed, W.S.; El-Halfawy, K.A.; Abdel-Rahman, M.H. KIR Genotypes Impact Progression to Hepatocellular Carcinoma in Patients with Chronic Hepatitis C Infection. Livers 2023, 3, 354-368. https://doi.org/10.3390/livers3030027
Abdelmaguid W, Maher D, Kohla MAS, Ezzat S, Moaz I, Abdel-Mageed WS, El-Halfawy KA, Abdel-Rahman MH. KIR Genotypes Impact Progression to Hepatocellular Carcinoma in Patients with Chronic Hepatitis C Infection. Livers. 2023; 3(3):354-368. https://doi.org/10.3390/livers3030027
Chicago/Turabian StyleAbdelmaguid, Waleed, Doha Maher, Mohamed A. S. Kohla, Sameera Ezzat, Inas Moaz, Wael S. Abdel-Mageed, Khalil A. El-Halfawy, and Mohamed H. Abdel-Rahman. 2023. "KIR Genotypes Impact Progression to Hepatocellular Carcinoma in Patients with Chronic Hepatitis C Infection" Livers 3, no. 3: 354-368. https://doi.org/10.3390/livers3030027





