Abstract
The purpose of this paper is to elucidate the roles that microbes may be playing in the rootzone of the medicinal plant Datura inoxia. We hypothesized that the microbes associated with the Datura rootzone would be significantly different than the similar surrounding fields in composition and function. We also hypothesized that rhizospheric and endophytic microbes would be associated with similar metabolic functions to the plant rootzone they inhabited. The methods employed were microbial barcoding, tests of essential oils against antibiotic resistant bacteria and other soil bacterial isolates, 16S Next Generation Sequencing (NGS) metabarcoding, and Whole Genome Shotgun (WGS) taxonomic and functional analyses. A few of the main bacterial genera of interest that were differentially abundant in the Datura root microbiome were Flavobacterium (p = 0.007), Chitinophaga (p = 0.0007), Pedobacter (p = 6 × 10−5), Bradyhizobium (p = 1 × 10−8), and Paenibacillus (p = 1.46 × 10−6). There was significant evidence that the microbes associated with the Datura rootzone had elevated function related to bacterial chalcone synthase (p = 1.49 × 10−3) and permease genes (p < 0.003). There was some evidence that microbial functions in the Datura rootzone provided precursors to important plant bioactive molecules or were beneficial to plant growth. This is important because these compounds are phyto-protective antioxidants and are precursors to many aromatic bioactive compounds that are relevant to human health. In the context of known interactions, and current results, plants and microbes influence the flavonoid biosynthetic pathways of one other, in terms of the regulation of the phenylpropanoid pathway. This is the first study to focus on the microbial ecology of the Datura rootzone. There are possible biopharmaceutical and agricultural applications of the natural interplay that was discovered during this study of the Datura inoxia rhizosphere.
1. Introduction
The Datura plant genus is of interest due to its production of several important cyclic compounds, most notably the tropane alkaloids. Datura produces alkaloids for anti-herbivory including scopolamine and atropine [1]. The plant contains antioxidants such as flavonoids, and phenols. Quercetin is a flavanol with anticancer properties that has free radical scavenging activity. Other compounds may include scopoletin [2], which is a fluorescent coumarin that has antifungal properties against antibiotic resistant strains of Candida [3].
This plant is sometimes referred to as an entheogen due to its use by native people in the United States such as the Chumash. Datura has been used for rituals and ceremonies in Hindu culture [4]. The seeds are highly heterozygous and spontaneous mutations are common in tissue culture [5].
A close relative of Datura inoxia, D. wrightii was recently recognized in rock art paintings by the native Chumash at Pinwheel Cave [6]. Although anthropologists long believed that the rock art paintings depicted a vision or hallucination, the recent discoveries paint a different picture. Rather than depicting a hallucination, the rock art depicts the sacrament itself and reflects the natural cycles of flowering and pollination that are essential to this dynamic outcrossing plant.
Since microbes from the Datura rootzone have been shown to suppress disease in other Solanaceae such as tomato [7], the distinct microbial community could be useful for biocontrol efforts. Recently, a few studies have elucidated the interaction between plants and microbes during secondary metabolite production. It is known that in some species, such as Crotalaria, nodulation is required for alkaloid production [8]. Furthermore, antioxidants have been shown to have antibacterial activity against pathogens [9], and antioxidants produced by soil microbes may have a plant-protective role. Functional analysis of soil metagenomic data has expanded as a field, and is used as a way to consider how plants are influencing the microbiome [10]. The microbiome is equally important in understanding plant secondary metabolism.
Shotgun metagenomics approaches in applied soil science provide the advantage of amplifying sequences from throughout the genome, including coding regions, yielding a high sequencing depth but at a higher cost. 16S metabarcoding is used as a diagnostic tool in clinical sciences and plant ecology, and its forté is identification of bacteria and archaea. However, metagenomics and metabarcoding are culture independent, meaning that no viable accessions are maintained. For that reason, microbial barcoding of streak plates isolated from the soil is also a useful tool that provides material for later wet lab experiments involving plant inoculation or antimicrobial trials, and helps to validate NGS results with tangible materials.
According to Sang et al. (2012), Flavobacterium johnsoniae produced 2,4-di-tert-butylphenol which had biocontrol activity against Phytophthora in Capsicum [11]. Sang’s study references one of a few studies in Solanaceae where Flavobacterium has been shown to suppress disease. Another example is Carrion et al. (2019), where Rhizoctonia solani was suppressed by the endophytic root microbiome [12]. Some of the most important natural products involved were phenazines, polyketides, siderophores, and chitinases. Aryl polyenes, which are structurally similar to carotenoids, were enriched by Flavobacteriaceae; terpenes and resorcinol were enriched by Chitinophaga. Similarly, in 2020 Lucke et al. reported that as biocontrol agents, the secretion of secondary metabolites triggered the induction of systemic resistance in plants [13]. In particular, they pointed out the importance of the chitinase from Flavobacterium and the Polyketide Synthase (PKS) gene cluster from Chitinophaga that is essential for disease suppression.
Del Barrio-Duque et al. (2019) found that 17/19 Mycobacteriaceae strains tested had fungal growth-promoting properties on Serendipita indica in vitro and in tomato; they were plant growth-promoting and helped to alleviate symptoms of Rhizoctonia solani [14]. In this research, Mycolibacterium was in a consortium with Rhizobium and Paenibacillus; in some instances, the isolations worked better than combinations of inoculants. Surprisingly, Burkholderia had a negative impact in vitro on S. indica fungal growth, although it has been shown to be plant growth promoting. An interesting review by Morris et al. 2019 highlights the fact that benzylisoquinoline alkaloid methyltransferases are functionally similar to the cyclopropane methyltransferase from Mycobacterium tuberculosis [15]. Mycobacterium cyclopropanation could contribute to tropane alkaloid biosynthesis in Datura, or provide an intermediate toward the same.
Banuelos-Vazquez reported that endophytes can receive nodulation genes, and in some cases, the ability to fix nitrogen, from Rhizobium etli. Plasmids are transferred to other endophytes in plant roots [16]. Another interesting example is from Kado and Kelly (2006), where they reported a successful protocol for transforming Streptomyces with Agrobacterium [17]. Lucke (2020) also noted the root nodulating capabilities and transfer of plasmids by R. etli and A. tumefaciens, and their ability to induce plant defenses [13].
Furthermore, the plant growth-promoting activities of endophytes may include production of plant growth regulators such as Indole Acetic Acid (IAA), and siderophore production for iron acquisition. Streptomyces spp. within Bruguiera gymnorhiza and Boesenbergia rotunda increased flavonoids and cyclopeptides with anti-HIV and anticancer activities [18]. Endophytic bacteria, as well as fungi, can produce paclitaxel [19]. Within Taxus baccata, Streptomyces produced 0.01–0.02 ng/mL, and Bacillus subtilis produced 1–25 ng/mL [19].
Indeed, as Wu et al. (2021) pointed out, the relationship between medicinal plants and endophytes is a long-term, symbiotic relationship [18]. Endophytes can strongly regulate the synthesis of secondary metabolites in plants. For example, for abiotic stress tolerance, in Pteris vittata, Agrobacterium and Bacillus spp. reduced arsenate to arsenite.
Volatile organic compounds can also help bacteria that are physically separated communicate with one another. Endophytic bacteria can protect plants by quenching quorum sensing molecules. For example, Pseudomonas aeruginosa could degrade 3-hydroxy palmitic acid methyl ester, which is a quorum sensing molecule of Rhizoctonia solanacearum. The inoculation of eggplant with P. aeruginosa reduced bacterial wilt caused by R. solanacearum [18]. In tomato, Bacillus aureus and Serratia nematodiphilia were applied to seeds along with Ralstonia syzigii sub-infection. The treatment under inoculation had increased jasmonic acid concentration in leaves and roots until 12 days post-infection [20].
Antioxidant enzymes are activated by endophytes, such as phenylalanine ammonia lyase, tyrosine ammonia lyase, and polyphenol oxidase (PPO) [20]. Rhizobacterial root colonization significantly impacted phenolic compounds, terpenes, and essential oils in plants. Pseudomonas putida root colonization altered benzoxaninone levels three days after inoculation. Colonization by Rhizobium changed the levels of phenolics, flavonoids, and anthocyanins, in blackberry, which was associated with delayed fungal postharvest growth [20].
In terms of the accumulation of plant secondary metabolites, production of plant sesquiterpenoids was enhanced in Atractyloides macrocephala by Pseudomonas fluorescens. Essential oil production was also increased [20]. According to Sang et al. (2012), endophytes can increase resistance to insects, while the plants provide nutrition and protection [11]. In Nethapodytes fortida and Apodytes dimidiata, two grasses, Fusarium solanii fungi were found to produce campothecin. This quinoline compound was previously taken from the roots of Nethapodytes directly.
The purpose of this paper is to elucidate the roles that microbes may play in the rootzone of the medicinal plant Datura inoxia. We hypothesized that the microbes associated with the Datura rootzone would be significantly different than the similar surrounding fields in composition and function. We also hypothesized that rhizospheric and endophytic microbes would be associated with similar metabolic functions to the plant rootzone they inhabited. This is the first study to focus on the ecology of the Datura rootzone microbiome, although similar studies have analyzed the rootzones of medicinal plants to uncover how microbes and their host plants interact during secondary metabolite production [21,22].
2. Materials and Methods
Soil samples were collected in triplicate from the field and arboretum at Los Angeles Pierce College. Replicated samples were taken from plant rootzones in the top 3 cm within a 1-foot radius of the plant subjects including Alnus rhombifolia, Datura inoxia, Ethiopian Eragrostis tef (Teff) grass, Opuntia cactus rootzone, and fallow conditions. The samples were collected in sterile cryotubes and were stored at −20 °C.DNA was extracted using the Qiagen (Hilden, Germany) Power Soil DNA kit and sent to Cold Spring Harbor Laboratory for 16S amplification, library preparation, and pooled 16S amplicon NGS on the Illumina (San Diego, CA, USA) MiSeq platform. The 515F (Parada)–806R (Apprill), forward-barcoded primers were used: FWD:GTGYCAGCMGCCGCGGTAA; REV:GGACTACNVGGGTWTCTAAT. A dual indexing strategy was used with these primers following Microbiome Helper’s approach [23,24]. Soil from the same sampling event was sent to Beijing Genomics Institute America, San Jose for logistics and processing, for the replicated Datura, Alnus, and fallow rhizospheric soil samples. These three plant rootzones were selected because Alnus and Datura are both known to produce antioxidants, and the fallow samples served as the reference. It was not possible to sequence the whole metagenome of all 5 categories due to budgetary constraints.
According to the USDA NRCS Web Soil Survey historic data, the fields related to the Datura, Alnus, and fallow samples are all classified identically as Cropley-Urban Land Complex 0–2% slopes, and the reported measurements for pH, texture, cation exchange capacity (CEC), and percent organic matter (% OM) were identical, as shown in Table 1 [25].

Table 1.
Selected physiochemical properties of soil.
Whole genome shotgun sequencing, DNA extraction, and library preparation was subsequently performed by Beijing Genomics Institute on the DNBseq platform. Preliminary trim and QC were carried out using SOAPnuke [26]. MG-RAST (University of Chiago, U.S.) was used to generate functional and taxonomic profiles [27]. Taxonomic identification was performed using RefSeq (NCBI, Bethseda, Maryland, U.S.) and functional profiles were built from Subsystems identifiers. In order to determine which taxa were differentially abundant between categories, DESeq2 (EMBL, Heidelberg, Germany) was used in R [28]. STAMP (Dalhousie University, Nova Scotia, Canada) was used for functional analysis [29].
For soil metabarcoding and 16S barcoding analyses, DNA Subway was used which implements Qiime2 [30,31]. For soil metabarcoding, DNA extraction was performed with the Qiagen PowerSoil kit. Library preparation followed the Earth Microbiome Project 16S Illumina Amplicon protocol [32,33]. Sequencing was performed at Cold Spring Harbor Laboratories. The Greengenes identifier (utilizing 515F/806R primers) was used.
Bacterial isolations were also generated for the Alnus, Datura, and fallow rhizosphere samples in order to provide material for future in vivo plant experiments according to the Soil Science protocols published by St. Clair et al. [34]. Isolates from the soil solutions at 10−3, 10−4, and 10−5 dilutions were plated and grown for three days on various media, summarized in Table 2. Selected isolates were streaked on plates of the same media and colony PCR was performed after three days of isolated growth. For bacterial isolations, DNA was extracted by boiling colonies with chelex beads for 10 min, followed by 16S rRNA amplification. Amplicon sizes were between 500 bp to 1000 bp which allowed for putative identification of the isolates. Sanger sequencing for bacterial isolates was performed by Genewiz. QC was performed and consensus sequences were generated in the Cyverse DNA Subway Blue Line. The Phylip Maximum Likelihood tree was generated in Cyverse. Bacterial putative identifications for the isolates were generated using the EZBioCloud (CJ Bioscience, Seoul, Korea) 16S-based ID application [35]. The putative identification of the isolates with the short sequence length also were expected to reinforce the validity of the barcoding and WGS results, if the results were similar.

Table 2.
Bacterial Strains Isolated and Culture Conditions.
In order to test the response of bacterial isolates in broth culture of plant essential oils, two crude extracts were used. Lemon balm (Melissa officianalis) and Tea tree (Maleleuca alternifolia) essential oils are well-characterized; however, their interaction with rootzone microbes warranted further investigation. Vegetative plant material was harvested from and air dried at room temperature for one week. The samples were handled separately. Soxhlet extraction was carried out for three cycles using reagent ethanol as the solvent. The extracts were subsequently dewaxed by running them through a Buchner funnel and filter paper.
Bacteria were cultured in 1.5 mL tubes of nutrient broth for 48 hours at room temperature. Separate sterile glass tubes filled with 2mL of nutrient broth were inoculated with 100 μL of each bacterial broth culture. The glass tubes were inoculated with 20 μL of either lemon balm oil or tea tree oil from the soxhlet extractions. The clarity of the solution, which was a proxy for bacterial colonization levels, was measured as % transmission at 600 nm. Transmittance at 600nm was measured using a spectrophotometer after culturing at room temperature overnight. This assay was similar to Kryvstova et al.’s biolfilm assay with Arnica montana (2019) [36], although here cuvettes were used rather than microtiter plates. The data was analyzed using the R MASS package (University of Oxford, England) and visualized with ggpubr.
3. Results
3.1. Soil Samples WGS and MG-RAST Taxonomic and Functional Assignments
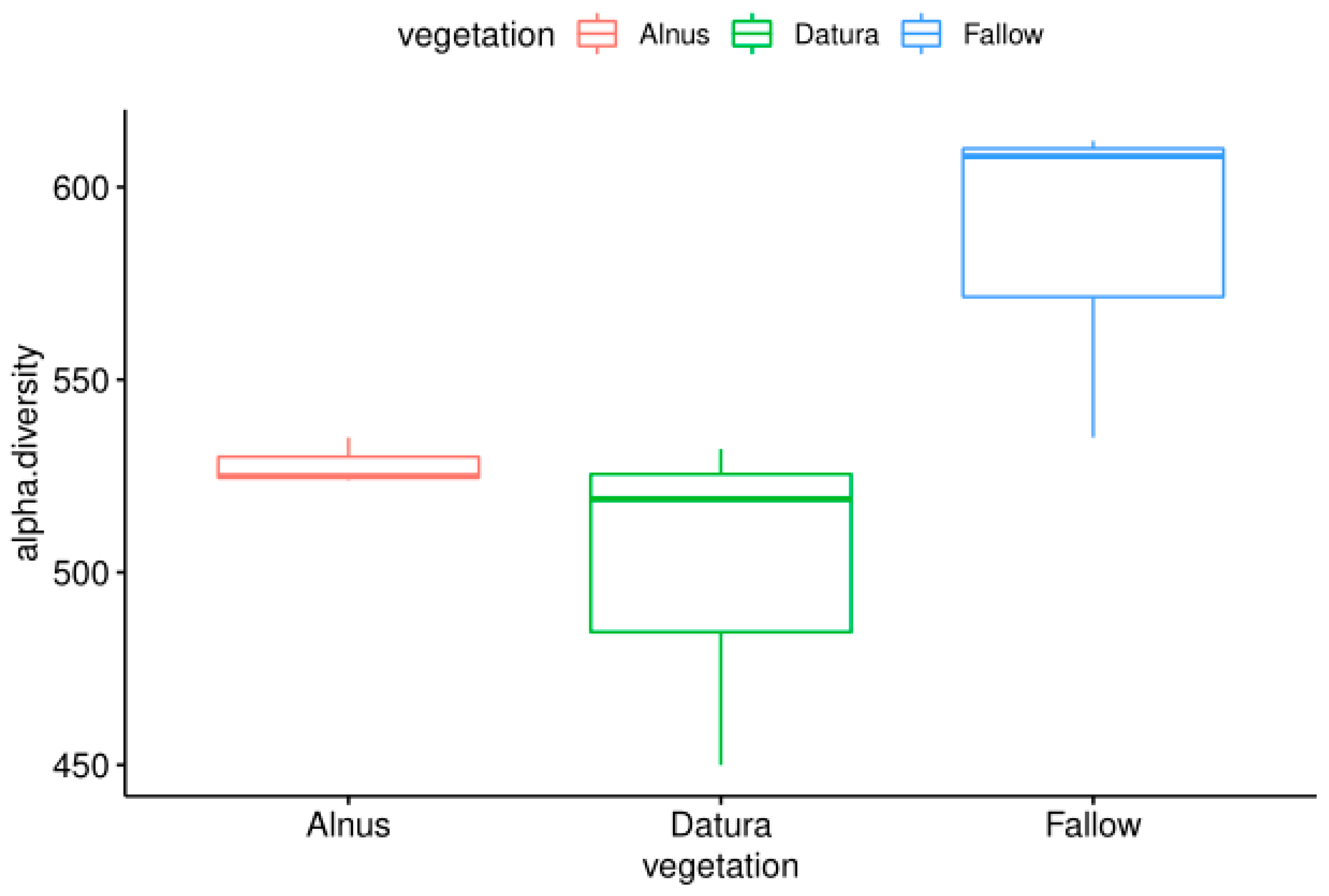
The sequence count ranged from 4,608,913 to 34,551,930 sequences. Interestingly the Fallow field had marginally higher observed alpha diversity overall (p < 0.1), which is contrary to what one might expect considering no plants were growing there. However, it was consistent with the metabarcoding results.
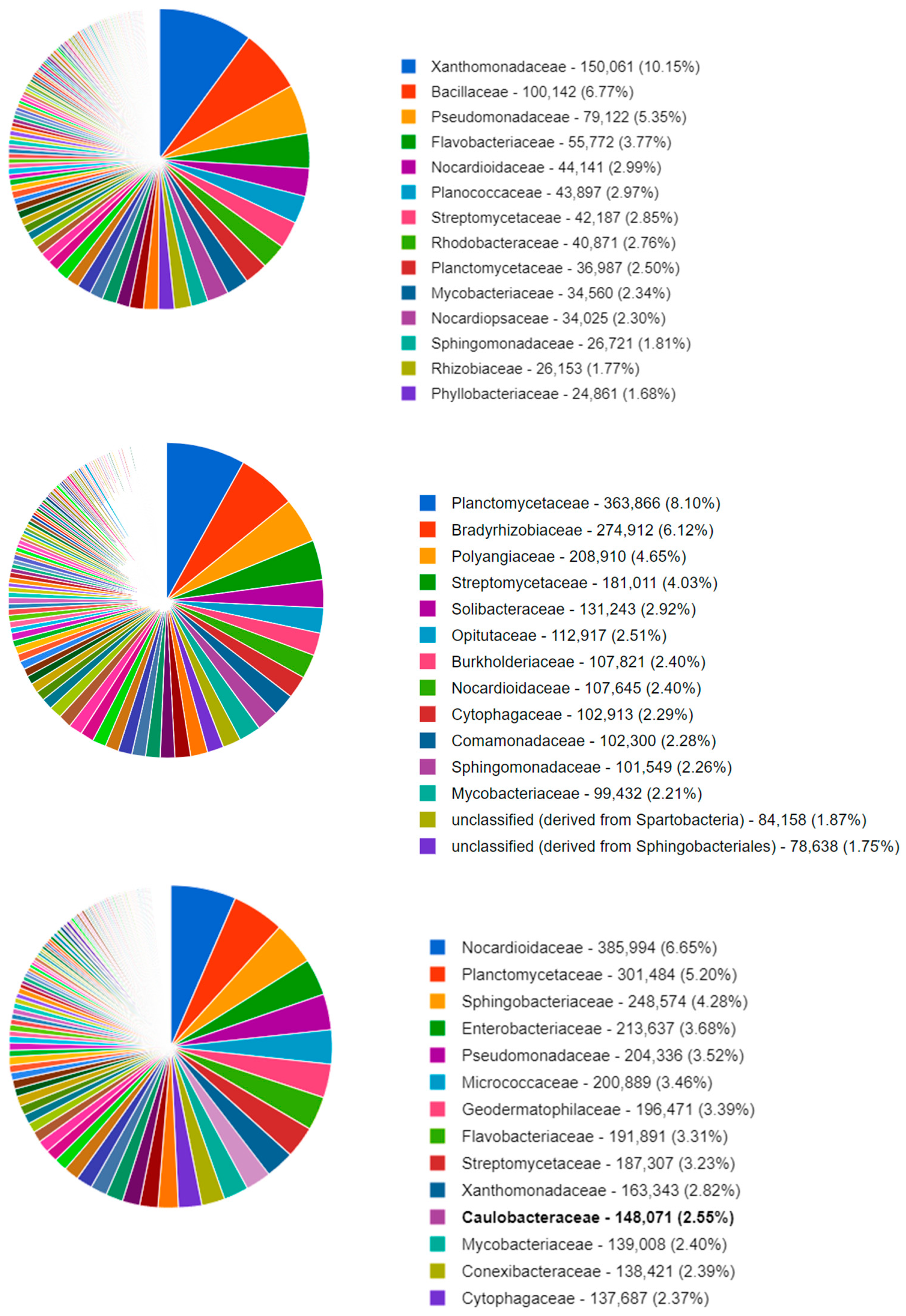
Figure 1 shows the top 14 bacterial families that were identified in a typical soil metagenome associated with the Datura rootzone and a fallow sample from this study. The remaining samples can be visualized in the published study on MG-RAST project mgp98747. All three samples exhibited similar proportions of Mycobacteriaceae and Flavobacteriacea; however, in the Datura rootzone Sphingomonadaceae sequences were more abundant. The Cytophagaceae sequences were represented in similar proportions in the Datura and Alnus associated samples, but were less abundant in the fallow samples.
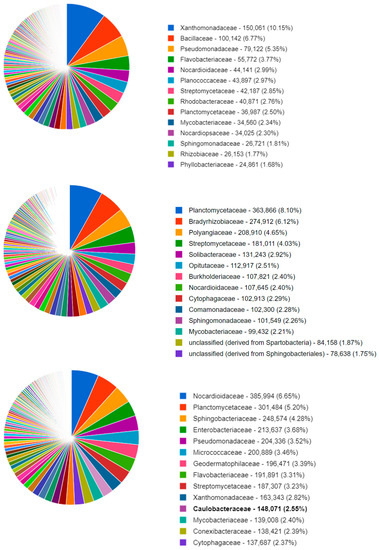
Figure 1.
MG-RAST classification of the NCBI RefSeq identification of the soil organisms at the family level, for a representative sample of fallow (top) Alnus (middle), and Datura (bottom) rootzone soil.
In the Datura rootzone, some of the most abundant genera were Pedobacter and Flavobacterium. Flavobacterium johnsoniae is well-known for its tyrosine ammonia lyase gene, which has been cloned and used in genetic transformations to optimize production of aromatic compounds of pharmaceutical value in microbial expression systems. Some potential applications are production of tropane alkaloids and p-coumaric acid [37,38].
On the other hand, the Datura and Alnus samples had lower species diversity, suggesting that there may be some selection or recruitment occurring in the plant rootzones. The alpha diversity results are visualized in Figure 2. It is also interesting to note that the Alnus rootzone samples had less between group variation.
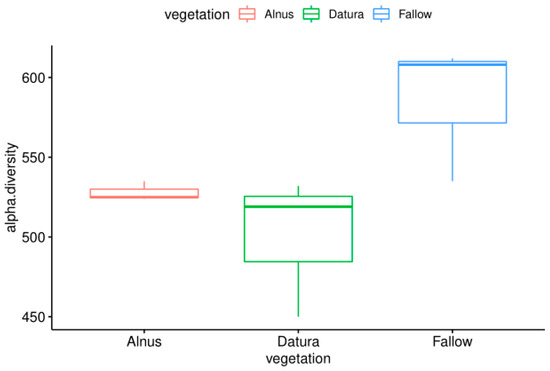
Figure 2.
Comparison of the observed alpha diversity levels from fallow, Alnus, and Datura rootzone-associated soil samples from the Pierce Farm in Los Angeles, California.
3.2. Functional Analysis in STAMP
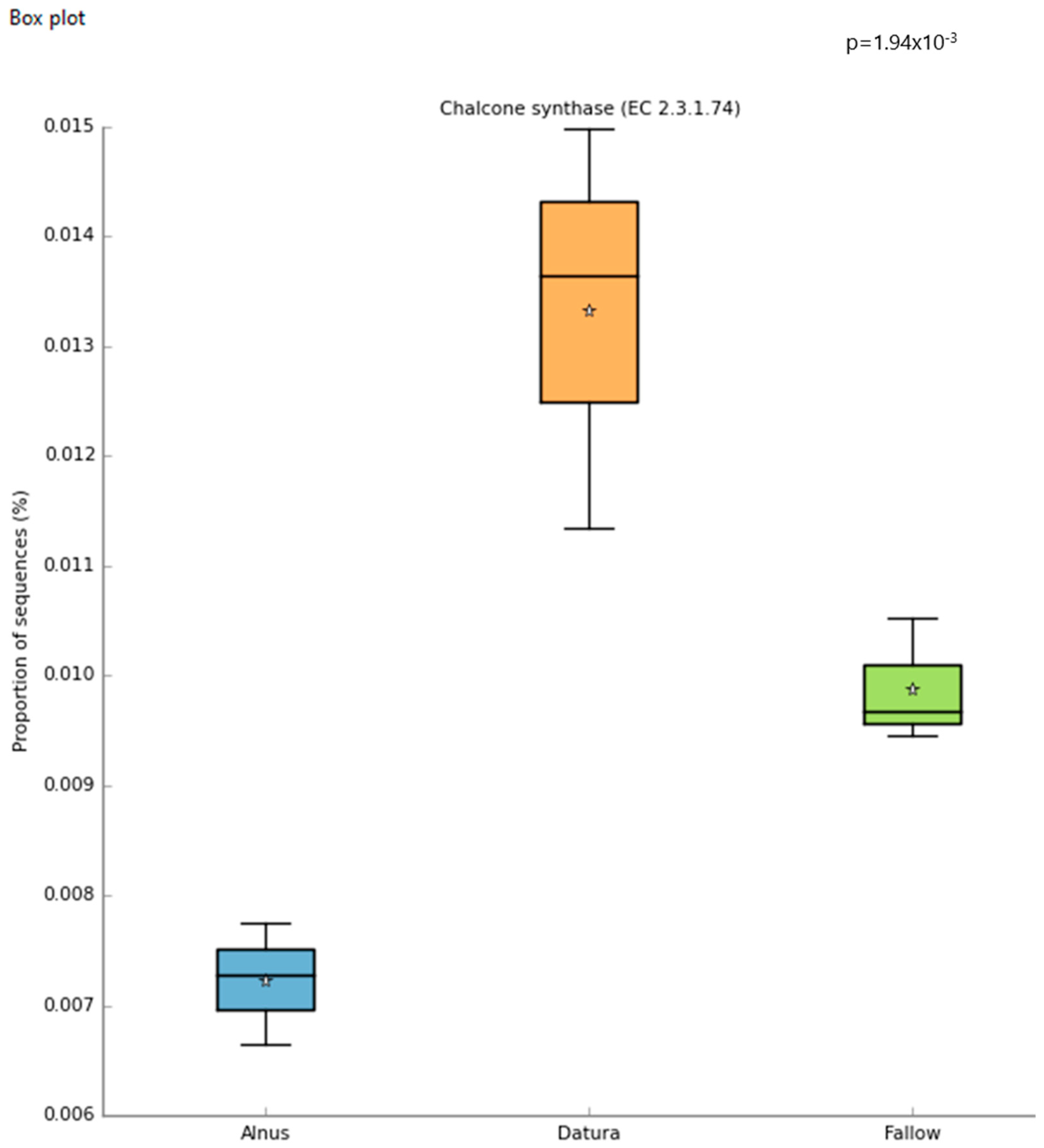
Differential abundance of chalcone synthase genes was detected in the Datura rootzone based on STAMP analysis of the MG-RAST generated Subsystems metagenomic functional profile (Figure 3). Using a cutoff of alpha = 0.05, the following were significantly higher in the Datura-associated soil samples: Alpha-xylosidase (p < 0.017), chalcone synthase (p < 0.002) (Figure 3), putrescine utilization (p = 0.05), Aromatic amino acid transport protein AroP (p < 0.003), polyols ABC transporter permease component (p < 0.003), Quinate permease (p = 0.002), ubiquinone biosynthesis enzyme COQ 7 (p < 7 × 10−3), Heavy metal sensor histidine kinase (p < 6.4 × 10−3), flavin reductase family (p = 0.022), para-hydroxybenzoate- polyprenyl transferase (p < 0.004), Siderophore achromobactin ABC transporter (p < 0.0021), NADPH quinone oxidoreductase 2 (p < 0.0021), vanillate O-demthylase oxygenase subunit (p < 0.0014), Phenylacetic acid degradation protein PaaN (p < 0.0013), clavulanic acid biosynthesis (p = 0.027), cobalamin synthesis (p < 0.005), and MAP kinase pathways (p < 0.0045).
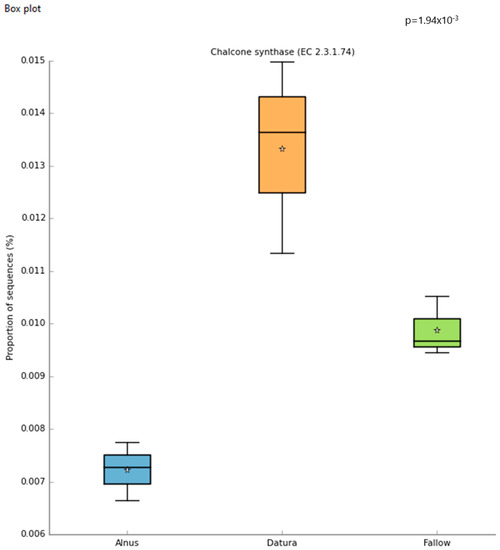
Figure 3.
Boxplots of the MG-RAST generated Subsystems metagenomic functional profile give insights into the community functions that are most significant. The star symbol represents the mean proportion of sequences for each sample group.
On the other hand, unlike the Datura samples, the Na+ translocating NADH-quinone reductase subunit was differentially abundant in Marquis C (p < 7 × 10−4), which was the fallow field near the compost pile. Gram positive cell wall components were differentially abundant in Marquis C (p < 0.008), although Gram negative cell wall components were equally likely to be present in all samples (p = 0.244). The CRISPR associated protein TM1812 was enriched in the fallow field near the compost pile (p < 0.0014). Phenlypropanoid compound degradation functional genes were most abundant in the Alnus samples (p < 6.4 × 10−5). Tocopherol biosynthesis genes were also differentially abundant in the Alnus rhizospere-associated soil (p < 0.004).
There was enrichment of cell wall degradation and depletion of Gram-positive bacteria, stress response, siderophores, catabolism of tyrosine, enrichment for aromatic compounds and permeases, production and reduction of flavonoids, antibiotics and cobalamin, and degradation of oxidized products in the Datura rootzone. The presence of differentially abundant permease components for essential oils and quinates suggests that there is a secretion pathway that would allow these compounds to enter plant roots and contribute to flavonoid, alkaloid and terpenoid biosynthesis in plant hosts.
3.3. Purple Line Analysis and Comparison
Demultiplexed sequence counts for the DNA Subway Purple Line paired end data set ranged from 5472 to 3.63 million, prior to alpha rarefaction. The lowest number of reads were for one of the Datura rootzone soil samples and a tilled fallow sample nearby in Marquis C. The highest number of reads were from Field 17, a fallow field tilled with cow patties and Marquis C near the compost pile. Indeed, in Field 17 also had a trend toward the highest alpha diversity, followed by Marquis C. There appeared to be more within group variation reflected by the 16S workflow compared to WGS.
Enrichment of Pedobacter matched up with MG-RAST results, and with Mycobacterium which may be related to alkaloid biosynthesis. Pedobacter represented >32% of the reads for one of the Datura rootzone samples. Blastococcus sequences were abundant in all three Datura rootzone soil samples. Streptomyces, Rubrobacter and Mycobacterium were abundant in one of the Datura samples, but were not detected in the other two. There appeared to be a similar drought tolerant entourage of bacteria associated with Datura, Teffgrass, and the cactus-associated soil samples, based on the Principal Coordinates Analysis (DNA Subway Purple Line Public project 7102).
Some unexpected results were detected by the isolation of bacterial colonies and analysis on the Blue Line. Four Chryseobacterium accessions were isolated from the soil that did not show up on 16S amplicon sequencing for the same soil samples. However, the Chryseobacterium genus was detected with DNBseq WGS (see Table 3). Since the WGS results are expected to be the most reliable, it seems likely that the Chryseobacterium putative identifications were in fact valid at least to the genus level, although that genus was not detected in the Datura-associated samples with metabarcoding in this instance. However, these results should not be interpreted to indicate novel species were isolates without further investigation, since the sequence lengths were too short to make a reliable identification at the species level. The Blue Line isolates were amplified using the same primers used for metabarcoding in this study and therefore have similar sequence lengths. In fact, the WGS results on the MG-RAST pipeline should be similarly considered reliable to the genus level.

Table 3.
The 34 genera with both high negative log fold change values, and significant adjusted p-values (p < 0.001) representing those associated with the Datura rootzone are shown.
3.4. Differential Abundance Analysis in DESeq2
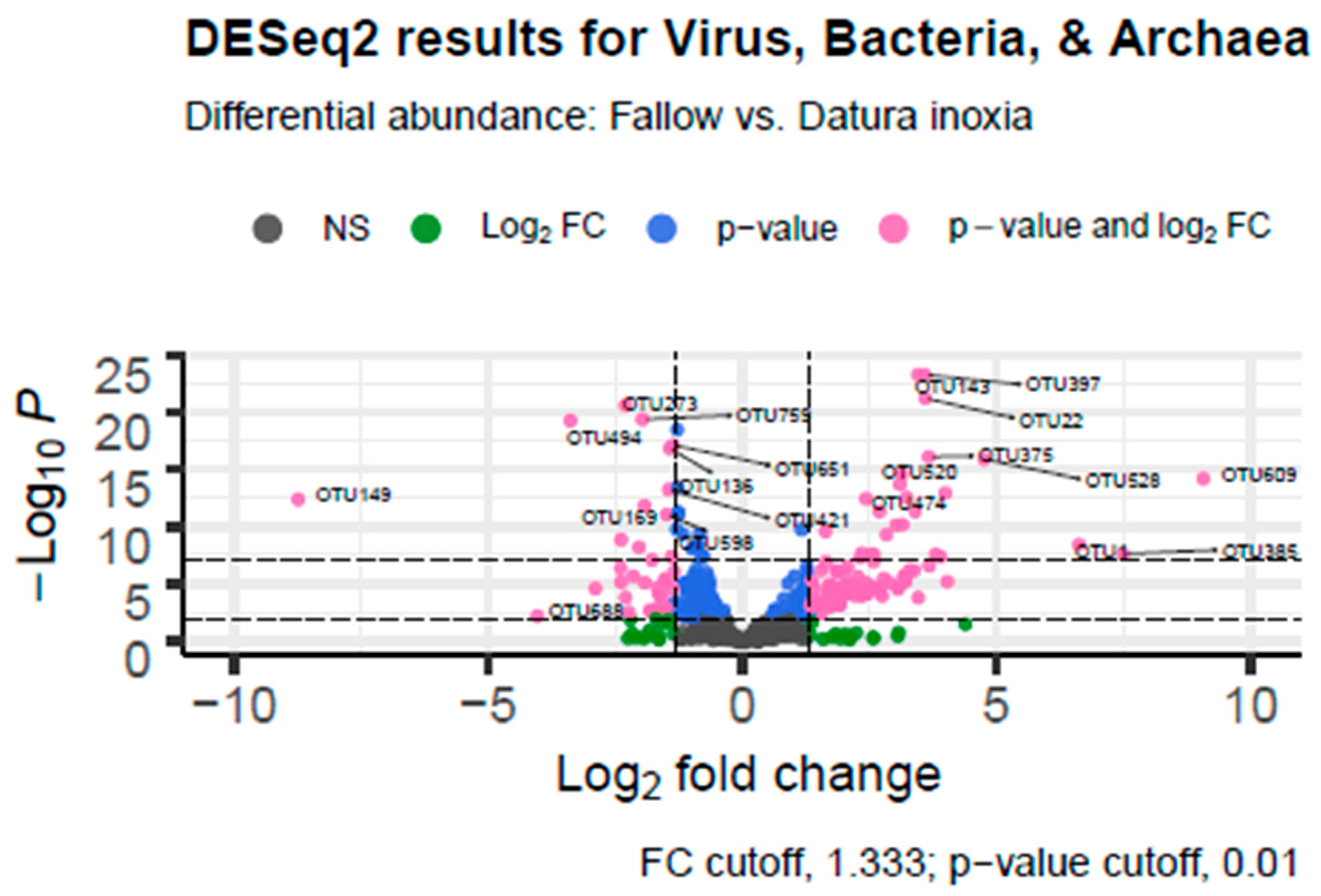
Differential Abundance Analysis in R showed that there were a large number of taxa that were differentially expressed in the fallow (positive log fold change) versus Datura (negative log fold change) samples. There were 223 virus, archaea, or bacterial genera that were differentially abundant between the Datura rootzone samples and the Fallow samples from Marquis Field near the compost pile. There were 140 species that were associated with the Datura rootzone and 83 species that were associated with the fallow samples, based on adjusted p-value < 0.001. Of those, 124 genera were also significant based on the log fold change cutoff of 1.33, 34 were associated with the Datura rootzone, and the remaining genera were associated with the fallow samples The results are summarized in Table 3 and visualized in Figure 4.
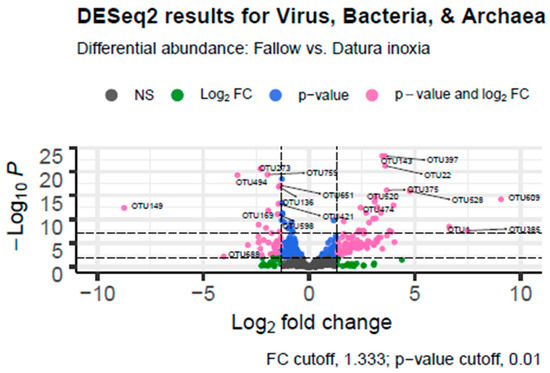
Figure 4.
Differential Abundance Analysis Volcano Graph. DESeq.
Among the more notable results were differentially abundant bacteria in the Datura rootzone including Flavobacterium, Chryseobacterium, and unclassified members of the Flavobacteriaceae. Chitinophaga, Mucilaginibacter, and Pedobacter of the closely related Sphingobacteriaceae were also differentially associated with the Datura rootzone. These genera are associated with degradation of polysaccharides or chitin. There was also evidence that Verrucomicrobium that are typically associated with Flavobacterium as well as unclassified Verrucomicrobium were significantly associated with the Datura rootzone. These genera are known for hydrolysis of xylans. Dyadobacter of the Cytophagaceae, which has polysaccharide and amino acid degrading functions, was present in differentially large quantities; this species lives in glaciers and was formerly classified as a member Flavobacteriales. Stigmatella, which is responsible for breaking down insoluble debris, was also significantly associated with the Datura rootzone.
Herbasperillum and Oxalobacter from the Oxalobacteriaceae were differentially abundant, along with Janthinobacterium, which produces violacein, an antifungal, antiprotist, antibacterial, and anti-tumor compound [39]. Interestingly, sequences from the plant pathogen Erwinia were also present in significantly high numbers, as well as the poultry disease Riemerella, and the infectious Serratia which is consistent with the animal bedding that is spread from the compost pile to all of the adjacent fields. Variovorax is involved in disrupting quorum sensing and has swarming motility, and was differentially abundant in the Datura rootzone. Bradyrhizobium and Azorhizobium abundance in the Datura rootzone was significant based on adjusted p-value (p < 0.01) but not based on log fold change values.
3.5. Microbial Barcoding and Essential Oil Trial
The Phylip Maximum Likelihood phylogenetic tree indicated that there are four main clusters of bacteria that were isolated. The first cluster includes isolates related to Pseudomonas sp. The second cluster consists of two groupings; one group is related to Massilia sp. and Achromobacter sp., while the second group is related to Chryseobacterium sp. The third cluster consisted of Bacillus sp. and Microbacteriaceae. Interestingly, one of the isolates was putatively identified as Priestia megaterium, which is used as an herbicide in organic production [40]. The fourth cluster is made up of mostly pathogenic species such as Erwinia, Pantoea, and Klebsiella. Most of the pathogenic strains were associated with the Arboretum samples. Stenotrophomonas was used as the outgroup. There were four Chryseobacterium isolates that were detected with barcoding, which were not detected with 16S NGS metabarcoding. The ascertainment of the Achromobacter isolate agrees with NGS results. The Xanthomonas isolation agrees with the WGS results, since Xanthomonadaceae was the top family detected in the WGS samples for the Datura inoxia rootzone. There were several accessions with low similarity and high completeness which may warrant further investigation in order to more accurately identify the species. This could be achieved with a larger amplicon size or WGS of the isolates.
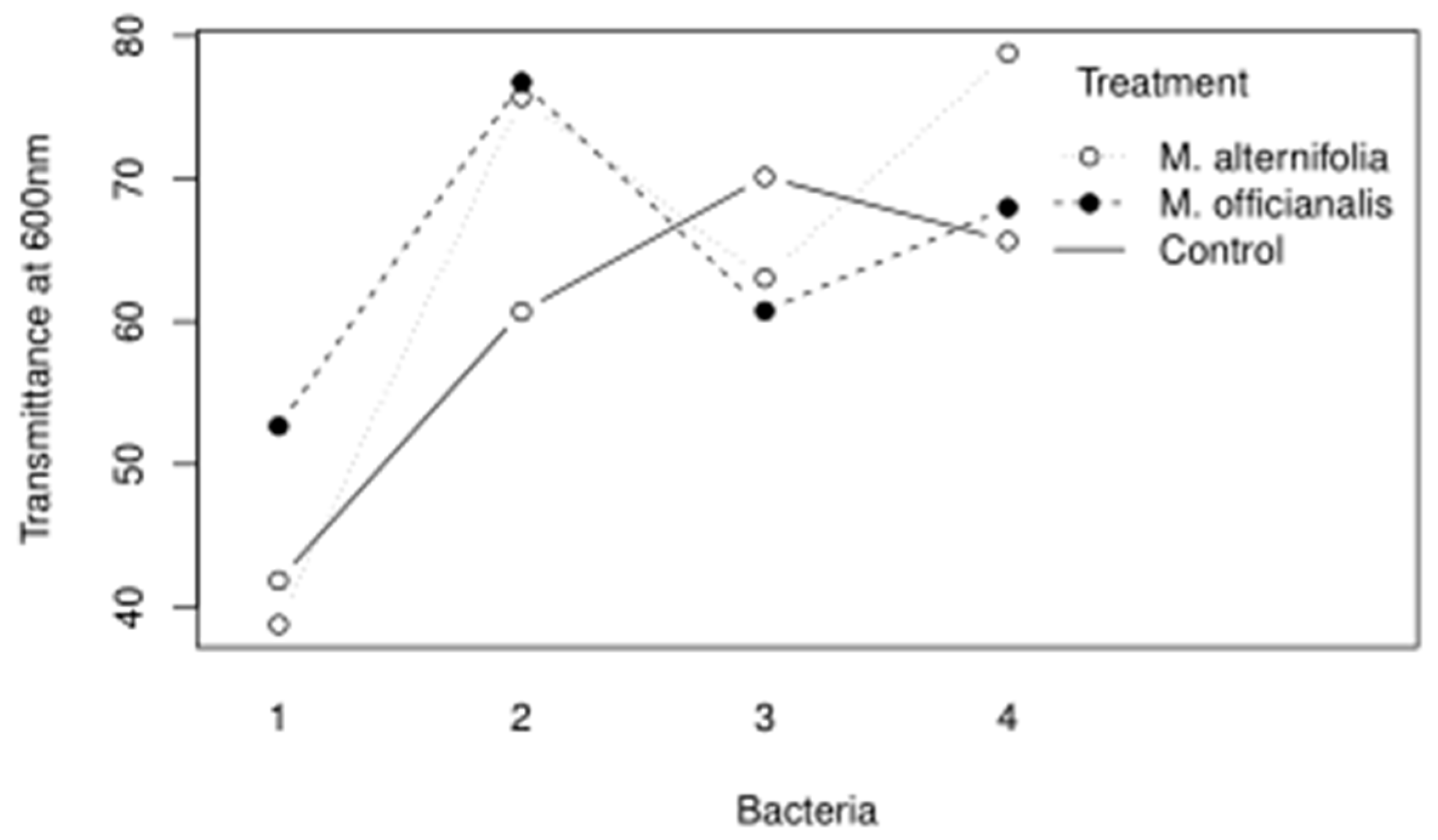
The main findings of the essential oil trials indicate that the isolate putatively identified as Pseudomonas mucoides was not susceptible to penicillin, and not susceptible to lemon balm essential oils. In retrospect, since P. mucoides is a Gram-negative bacterium, it would not be expected to be susceptible to penicillin. Bacillus pseudomycoides was resistant to penicillin but grew even more with lemon balm and tea tree essential oils. It is interesting because B. pseudomycoides was tolerant of high alcohol content, possibly since it is a lactic acid fermenter. Priestia qingshengii was not susceptible to penicillin and grew less when lemon balm or tea tree essential oils were added. Leuthyella okanaganae was resistant to streptomycin; however, growth was controlled significantly when lemon balm or tea tree essential oils were added and the least growth occurred when the tea tree treatment was used. Interestingly, there was an interaction between the essential oil used and the bacterial accession it was tested against, as shown in Figure 5.
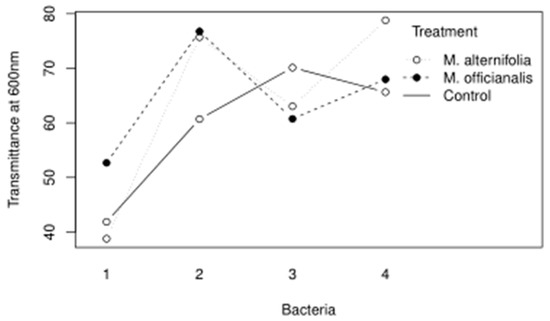
Figure 5.
The interaction plot for the Bacteria and Treatment variables of the essential oil trial are shown.
It is interesting because Luna et al. 2007 found that several Bacillus pseudomycoides strains were susceptible to beta-lactams, however the strain ascertained in this study was resistant to penicillin [41]. Bacillus pseudomycoides is related to Bacillus anthracis and Bacillus cereus; these species are notorious for being antibiotic resistant but B. pseudomycoides and B. cereus are particularly resistant to clindamycin [41].
Pseudomonas is also notoriously resistant to antibiotics, although its inclusion here was somewhat serendipitous since it was not selected with an antibiotic that would be typically used to combat Gram negative bacteria. It was also found that tea tree and lemon balm oils were ineffective against the P. mycoides isolate.
4. Discussion
Cytophagaceae are known to degrade polysaccharides [42]. Pedobacter inoculation has been shown to significantly increase the antioxidant content of strawberries [43]. Closely related species are the Flavobacterium, which are famous for producing flavonoids including quinones and they play a role in disease suppression in soils [44]. Flavonoid and alkaloid production gene ontologies are closely related [45,46]. Rhizoctonia solani is suppressed by Flavobacterium [47]. This is believed to be related to its chitinase activity which is also of interest for energy production from biomass [48].
As Fadiji 2020 pointed out, plant protective endophytes stimulate plant secondary metabolite production while inducing plant resistance to pathogens [49]. In addition, endophytic fungi produce strong antioxidants. In Taxus cuspidata paclitaxel production was elevated, and in Euphorbia pekinensis there was elevated terpene production in response to the Fusarium E5 elicitor; these are two examples from the article. Some examples of protection provided by the endophytes include antibiotics, competition for resources with pathogens, and lytic enzymes such as chitinases that break down fungi and plant cell walls.
The chitinase activity provides a possible answer to how endophytes and their products are transported into the plant. Furthermore, xylanase and other cell wall degrading enzymes such as chitinases and cellobio-hydrolase were detected in the metagenome of several endophytes [20]. These lytic enzymes have the dual function of giving microbes access to plant cells, ducts, and intercellular spaces for colonization and transport, and the enzymes can also break down fungi cell walls to assist the plant host while fighting off infections.
Our preliminary evidence shows that the functional hits for Chalcone synthase in the microbes of the Datura rootzone are differentially abundant in contrast to the fallow sample. Chalcone synthase is involved in antioxidant production in bacteria and plants, and is homologous with polyketide synthase, which is part of the alkaloid production pathway in plants [50,51]. There appears to be synergy between the plant and microbe functions in terms of secondary metabolite production in the rootzone.
Genetic Engineering and the Flavonoid Biosynthesis Pathway
Another way to look at this is that microbes may be engineered to produce the plant medicinal compounds directly [52]. Chemical synthesis of aromatic compounds uses benzene, toluene, and xylene as starting materials; these materials are derived from petroleum. Microorganisms are a renewable source of plant-derived secondary compounds. These compounds include phenolic acids, flavonoids, stilbenoids, coumarin, and conjugates of the same. A pathway of interest for the engineering of microbes for the production of plant metabolites, which occurs in nature, is the pathway by which tyrosine is converted to p-coumaric acid by tyrosine ammonia lyase and later converted to quercetin.
Yonekura et al. 2019 shed light on the flavonoid biosynthesis pathway [45]. In the plant kingdom, flavonoids are widely distributed in all subclasses, except for hornworts. This suggests that chalcone synthase, the first enzyme in the pathway, was evolved many times in parallel. If the enzyme was universal, then it must have been lost by many species along the way, since not all species within the subclasses have it [45]. The first two enzymes on the biosynthetic pathway are chalcone synthase and chalcone isomerase, which are believed to also have evolved first from a common ancestor involved in lipid metabolism.
There are 9000 flavonoid compounds defined strictly as:
- Structural derivatives of phenyl-substituted propylbenzenes with a C15 backbone,
- Phenyl-substituted propylbenzene derivatives with a C16 skeleton, or
- Phenyl-substituted propylbenzenes condensed with C6-C3 lignan precursors to form flavonolignans.
More broadly, we can also include chalcones and dihydrochalcones, anthocyanins, and aurones [45]. It is believed that defense against UV radiation, and plant hormone regulation were the first flavonoid functions. Flavonoid accumulation could help with harsh field conditions in Datura. They are structurally related to carotenoids, which are part of the accessory light harvesting complex in plants. Interestingly, UV-B radiation of mosses increased flavonoid content, which supports this notion [45].
Chalcone synthase (CHS) is derived from beta-ketoacyl ACP synthase. It is in the Type III polyketide synthase (PKS) gene family. CHS may have evolved before chalcone isomerase (CHI), since CHS catalyzes the first step and the step catalyzed by CHI may happen spontaneously [45]. P-coumaroyl CoA is transformed to naringenin chalcone by CHS, then to naringenin by CHI. Arabidopsis has four Type III PKS genes and one functional CHS.
CHS is promiscuous in the sense that it can use many substrates, although it cannot function on bulky substrates [45]. Flavobacterium johnsoniae has a chalcone synthase gene according to the Uniprot database [53], which may be working synergistically with Datura to increase flavonoid content in the plant, supply acyl compounds to the plant for downstream production for plant secondary metabolites, or have a modified function that is specific to its plant host. It is worth mentioning that Datura, Streptomyces, and Agrobacterium also have this gene. In vitro, CHS from Huperzia serrata can produce other polyketides besides flavonoids, like aromatic tricyclic pyridoisoindoles [45].
Evolutionarily, CHS on the flavonoid biosynthetic pathway may have paved the way for biosynthesis of medicinal secondary metabolites by land plants. It expanded the chemical variety of the metabolites plants produced over time [45]. Type III PKS can also synthesize oxidosqualane and terpenes in other specialized metabolisms, for example. It is not known why plants and their endophytes have evolved this important set of enzymes, but it warrants further investigation.
Many medicinal plants produce economically important compounds, and synthetic production is not possible because the pathways are poorly understood. Further elucidating the interaction between microbes and wild plants could also shed light on resilience mechanisms, which could be used for crop improvement. For example, it has been shown that beneficial microbes prime plant defenses, giving a competitive advantage to organic farming [54]. Furthermore, understanding how these plants adapt to their environment will empower weed control efforts, including biocontrol.
5. Conclusions
We hypothesized that the microbes associated with the Datura rootzone would be significantly different than the similar surrounding fields in composition and function. We found strong evidence to support this hypothesis, based on significant Log fold change values and adjusted p-values. The main results indicated that Flavobacterium, Pedobacter and Chitinophaga were differentially abundant in the Datura rootzone when compared with the fallow condition. We also hypothesized that rhizospheric and endophytic microbes would be associated with similar metabolic functions to the plant rootzone they inhabited. There was some evidence to support this notion, based on experimental evidence and statistical analysis of the functional profiles. One of the soil bacterial isolates, putatively identified as Bacillus pseudomycoides, grew more when treated with lemon balm and tea tree essential oils. There were several functions associated with flavonoid and aromatic compound production, as well as cell wall degradation, that were differentially abundant in the Datura rootzone, compared to the fallow condition. Further research focusing on metabolomic profiles of Datura plants grown in the presence and absence of rootzone bacteria will be useful to reveal to what degree rootzone bacteria influence secondary metabolite production.
Author Contributions
Conceptualization, S.S.; formal analysis, S.S.; investigation, S.S., K.P., A.L.B.; writing—original draft preparation, S.S.; writing—review and editing, K.P., A.L.B. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by training and resources from the National Science Foundation IUSE grant, NSF DUE-1821657 awarded to Bruce Nash.in part to support Community College faculty. This material is based upon work supported by the National Science Foundation under Grant Nos. DBI-0735191, DBI-1265383, and DBI-1743442 which supported iPlant Collaborative and DNA Subway development.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
WGS profiles, metadata and genomic data are freely available on MG-RAST under public project mgp98747. DNA Subway Purple Line metabarcoding results are available on Cyverse under public project 7102. Raw fastq files are available at: https://de.cyverse.org/data/ds/iplant/home/shared/iplant_DNA_subway/metabarcoding_data/210611_M00970/St_Claire (accessed on 24 June 2021). DNA Subway Blue line barcoding results and trace files are available under public project 238876 on Cyverse.
Acknowledgments
Thank you to Maryam Saraylou for advice on cultivation dependent methods. Thank you to Daila Melendez, Rosealie Vicuna, and Karu Smith for assistance with plant material extraction. Thank you to Bruce Nash for materials. Thank you to Jay R. Reichman for technical writing assistance.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Sonia, P.; Siddiqui, A.A.; Dwivedi, J.; Sonia, V. Pharmacological properties of Datura stramonium L. as a potential medicinal tree: An overview. Asian Pac. J. Trop. Biomed. 2012, 2, 1002–1008. [Google Scholar] [CrossRef] [Green Version]
- Han, X.L.; Wang, H.; Zhang, Z.H.; Tan, Y.; Wang, J.H. Study on chemical constituents in seeds of Datura metel from Xinjiang. Zhong yao cai = Zhongyaocai = J. Chin. Med. Mater. 2015, 38, 1646–1648. [Google Scholar]
- Lemos Ari, S.O.; Florêncio Jônatas, R.; Pinto Nícolas, C.C.; Campos Lara, M.; Silva Thiago, P.; Grazul Richard, M.; Pinto Priscila, F.; Tavares Guilherme, D.; Elita, S.; Apolônio Ana Carolina, M.; et al. Antifungal Activity of the Natural Coumarin Scopoletin Against Planktonic Cells and Biofilms from a Multidrug-Resistant Candida tropicalis Strain. Front. Microbiol. 2020, 11, 1525. Available online: https://www.frontiersin.org/article/10.3389/fmicb.2020.01525 (accessed on 9 December 2021). [CrossRef] [PubMed]
- Bulcsu, S. Datura rituals in the vajramahabhairava-tantra. Acta Orient. Acad. Sci. Hung. 1994, 47, 409–416. Available online: www.jstor.org/stable/23658487 (accessed on 18 July 2021).
- Rajewski, A.; Carter-House, D.; Stajich, J.; Litt, A. Datura genome reveals duplications of psychoactive alkaloid biosynthetic genes and high mutation rate following tissue culture. BMC Genom. 2021, 22, 201. [Google Scholar] [CrossRef]
- Robinson, D.W.; Brown, K.; McMenemy, M.; Dennany, L.; Baker, M.J.; Allan, P.; Cartwright, C.; Bernard, J.; Sturt, F.; Kotoula, E.; et al. Datura quids at Pinwheel Cave, California, provide unambiguous confirmation of the ingestion of hallucinogens at a rock art site. Proc. Natl. Acad. Sci. USA 2020, 117, 31026–31037. [Google Scholar] [CrossRef] [PubMed]
- Rania, A.B.A.; Jabnoun-Khiareddine, H.; Nefzi, A.; Mokni-Tlili, S.; Daami-Remadi, M. Endophytic bacteria from Datura metel for plant growth promotion and bioprotection against Fusarium wilt in tomato. Biocontrol Sci. Technol. 2016, 26, 1139–1165. [Google Scholar] [CrossRef]
- Irmer, S.; Podzun, N.; Langel, D.; Heidemann, F.; Kaltenegger, E.; Schemmerling, B.; Geilfus, C.M.; Zörb, C.; Ober, D. New aspect of plant–rhizobia interaction: Alkaloid biosynthesis in Crotalaria depends on nodulation. Proc. Natl. Acad. Sci. USA 2015, 112, 4164–4169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhuyar, P.; Rahim, M.H.; Sundararaju, S.; Maniam, G.P.; Govindan, N. Antioxidant and antibacterial activity of red seaweed Kappaphycus alvarezii against pathogenic bacteria. Glob. J. Environ. Sci. Manag. 2019, 6, 47–58. [Google Scholar]
- Xia, Y.; Dong, M.; Yu, L.; Kong, L.; Seviour, R.; Kong, Y. Compositional and Functional Profiling of the Rhizosphere Microbiomes of the Invasive Weed Ageratina adenophora and Native Plants. PeerJ 2021, 9, E10844. [Google Scholar] [CrossRef] [PubMed]
- Sang, M.K.; Kim, K.D. The volatile-producing Flavobacterium johnsoniae strain GSE09 shows biocontrol activity against Phytophthora capsici in pepper. J. Appl. Microbiol. 2012, 113, 383–398. [Google Scholar] [CrossRef]
- Carrión, V.J.; Perez-Jaramillo, J.; Cordovez, V.; Tracanna, V.; De Hollander, M.; Ruiz-Buck, D.; Mendes, L.W.; Van Ijcken, W.F.J.; Gomez-Exposito, R.; Elsayed, S.S.; et al. Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 2019, 366, 606–612. [Google Scholar] [CrossRef] [PubMed]
- Miriam, L.; Gabriel, C.M.; Asaf, L. The Role of Secretion Systems, Effectors, and Secondary Metabolites of Beneficial Rhizobacteria in Interactions with Plants and Microbes. Front. Plant Sci. 2020, 11, 1718. Available online: https://www.frontiersin.org/article/10.3389/fpls.2020.589416 (accessed on 26 July 2021).
- del Barrio-Duque, A.; Ley, J.; Samad, A.; Antonielli, L.; Sessitsch, A.; Compant, S. Beneficial Endophytic Bacteria-Serendipita indica Interaction for Crop Enhancement and Resistance to Phytopathogens. Front. Microbiol. 2019, 10, 2888. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.S.; Facchini, P.J. Molecular Origins of Functional Diversity in Benzylisoquinoline Alkaloid Methyltransferases. Front. Plant Sci. 2019, 10, 1058. [Google Scholar] [CrossRef] [Green Version]
- Alfredo, B.L.; Daniel, C.; Susana, R.; Laura, C.L.; Rosana, S.; Castellani Lucas, G.; Torres, T.G.; Susana, B. Transfer of the Symbiotic Plasmid of Rhizobium etli CFN42 to Endophytic Bacteria Inside Nodules. Front. Microbiol. 2020, 11, 1752. [Google Scholar]
- Kado, C.I.; Kelly, B. Transformation of Actinomycetes. Agrobacterium Protoc. 2006, 2, 395–402. [Google Scholar]
- Wu, W.; Chen, W.; Liu, S.; Wu, J.; Zhu, Y.; Qin, L.; Zhu, B. Beneficial Relationships between Endophytic Bacteria and Medicinal Plants. Front. Plant Sci. 2021, 12, 646146. [Google Scholar] [CrossRef]
- Flores-Bustamante, Z.R.; Rivera-Orduna, F.N.; Martínez-Cárdenas, A.; Flores-Cotera, L.B. Microbial Paclitaxel: Advances and Perspectives. J. Antibiot. 2010, 63, 460–467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oukala, N.; Aissat, K.; Pastor, V. Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress. Plants 2021, 10, 1012. [Google Scholar] [CrossRef]
- Chen, H.; Wu, H.; Yan, B.; Zhao, H.; Liu, F.; Zhang, H.; Sheng, Q.; Miao, F.; Liang, Z. Core Microbiome of Medicinal Plant Salvia miltiorrhiza Seed: A Rich Reservoir of Beneficial Microbes for Secondary Metabolism? Int. J. Mol. Sci. 2018, 19, 672. [Google Scholar] [CrossRef] [Green Version]
- You, C.; Qin, D.; Wang, Y.; Lan, W.; Li, Y.; Yu, B.; Peng, Y.; Xu, J.; Dong, J. Plant Triterpenoids Regulate Endophyte Community to Promote Medicinal Plant Schisandra sphenanthera Growth and Metabolites Accumulation. J. Fungi 2021, 7, 788. [Google Scholar] [CrossRef]
- Comeau, A.M.; Douglas, G.M.; Langille, M.G.I. Microbiome Helper: A Custom and Streamlined Workflow for Microbiome Research. mSystems 2017, 2, e00127-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ul-Hasan, S.; Bowers, R.M.; Figueroa-Montiel, A.; Licea-Navarro, A.F.; Beman, J.M.; Woyke, T. Community ecology across bacteria, archaea and microbial eukaryotes in the sediment and seawater of coastal Puerto Nuevo, Baja California. PLoS ONE 2019, 14, e0212355. [Google Scholar] [CrossRef] [Green Version]
- Soil Survey Staff, Natural Resources Conservation Service, United States Department of Agriculture. Web Soil Survey. Available online: http://websoilsurvey.sc.egov.usda.gov/ (accessed on 14 October 2021).
- Chen, Y.; Chen, Y.; Shi, C.; Huang, Z.; Zhang, Y.; Li, S.; Li, Y.; Ye, J.; Yu, C.; Li, Z.; et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 2018, 7, gix120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyer, F.; Paarmann, D.; D’Souza, M.; Olson, R.; Glass, E.M.; Kubal, M. The metagenomics RAST server—A public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform. 2008, 9, 386. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef] [Green Version]
- Hilgert, U.; McKay, S.; Khalfan, M.; Williams, J.; Ghiban, C.; Micklos, D. DNA Subway: Making Genome Analysis Egalitarian. In Proceedings of the 2014 Annual Conference on Extreme Science and Engineering Discovery Environment, Atlanta, GA, USA, 13–18 July 2014; Association for Computing Machinery: New York, NY, USA, 2014; Volume 70, pp. 1–3. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4516–4522. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef] [Green Version]
- Clair, S.S.; Saraylou, M.; Maine, E. Introduction to Soil Science Lab Manual v1. Protocols.io 2019. [Google Scholar] [CrossRef]
- Yoon, S.H.; Ha, S.M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar] [CrossRef]
- Kryvtsova, M.V.; Trush, K.; Koščová, J.; Eftimova, J. Antimicrobial, antioxidant and some biochemical properties of arnica montanal. Bull Probl. Biol. Med. 2019, 3, 268. [Google Scholar] [CrossRef]
- Eudes, A.; Mouille, M.; Robinson, D.S.; Benites, V.T.; Wang, G.; Roux, L.; Tsai, Y.-L.; Baidoo, E.E.K.; Chiu, T.-Y.; Heazlewood, J.L.; et al. Exploiting members of the BAHD acyltransferase family to synthesize multiple hydroxycinnamate and benzoate conjugates in yeast. Microb. Cell Factories 2016, 15, 198. [Google Scholar] [CrossRef] [Green Version]
- Jendresen, C.B.; Stahlhut, S.G.; Li, M.J.; Gaspar, P.; Siedler, S.; Foörster, J.; Maury, J.; Borodina, I.; Nielsen, A.T. Highly Active and Specific Tyrosine Ammonia-Lyases from Diverse Origins Enable Enhanced Production of Aromatic Compounds in Bacteria and Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2015, 81, 4458–4476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Friedrich, I.; Hollensteiner, J.; Schneider, D.; Poehlein, A.; Hertel, R.; Daniel, R. First Complete Genome Sequences of Janthinobacterium lividum EIF1 and EIF2 and Their Comparative Genome Analysis. Genome Biol. Evol. 2020, 12, 1782–1788. [Google Scholar] [CrossRef]
- Dara, S.K. Biopesticides: Categories and use strategies for IPM and IRM. E-J. Entomol. Biolog. 2021. Available online: https://ucanr.edu/blogs/blogcore/postdetail.cfm?postnum=46134 (accessed on 29 December 2021).
- Luna, V.A.; King, D.S.; Gulledge, J.; Cannons, A.C.; Amuso, P.T.; Cattani, J. Susceptibility of Bacillus anthracis, Bacillus cereus, Bacillus mycoides, Bacillus pseudomycoides and Bacillus thuringiensis to 24 antimicrobials using Sensititre automated microbroth dilution and Etest agar gradient diffusion methods. J. Antimicrob. Chemother. 2007, 60, 555–567. [Google Scholar] [CrossRef] [Green Version]
- Baek, K.; Choi, A. Draft genome sequence of the polysaccharide degrading bacterium Cytophagaceae strain SJW1-29. Korean J. Microbiol. 2020, 56, 422–425. [Google Scholar] [CrossRef]
- Maria, M.; Mucha, Â.; Ferreira, H.; Gonçalves, B.; Bacelar, E.; Marques, G. Comparative study of plant growth-promoting bacteria on the physiology, growth, and fruit quality of strawberry. J. Sci. Food Agric. 2019, 99, 5341–5349. [Google Scholar]
- Kolton, M.; Erlacher, A.; Berg, G.; Cytryn, E. Ch 9: The Flavobacterium genus in the Plant Holobiont: Ecological, Physiological, and Applicative Insights. In Microbial Models from Environmental to Industrial Sustainabilty; Springer: Singapore, 2016. [Google Scholar]
- Yonekura-Sakakibara, K.; Higashi, Y.; Nakabayashi, R. The Origin and Evolution of Plant Flavonoid Metabolism. Front. Plant Sci. 2019, 10, 943. [Google Scholar] [CrossRef] [Green Version]
- Abe, I. Biosynthesis of medicinally important plant metabolites by unusual type III polyketide synthases. J. Nat. Med. 2020, 74, 639–646. [Google Scholar] [CrossRef]
- Schlatter, D.; Kinkel, L.; Thomashow, L.; Weller, D.; Paulitz, T. Disease Suppressive Soils: New Insights from the Soil Microbiome. Phytopathol. Rev. 2017, 107, 1284–1297. [Google Scholar] [CrossRef] [Green Version]
- Nordberg, H.; Cantor, M.; Dusheyko, S.; Hua, S.; Poliakov, A.; Shabalov, I.; Smirnova, T.; Grigoriev, I.V.; Dubchak, I. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 2014, 42, D26–D31. [Google Scholar] [CrossRef] [PubMed]
- Fadiji, A.E.; Babalola, O.O. Elucidating Mechanisms of Endophytes Used in Plant Protection and Other Bioactivities with Multifunctional Prospects. Front. Bioeng. Biotechnol. 2020, 8, 467. [Google Scholar] [CrossRef]
- Bedewitz, M.A.; Jones, A.D.; D’Auria, J.C.; Barry, C.S. Tropinone synthesis via an atypical polyketide synthase and P450 mediated cyclization. Nat. Commun. 2018, 9, 5281. [Google Scholar] [CrossRef] [Green Version]
- Huang, J.P.; Fang, C.; Ma, X.; Wang, L.; Yang, J.; Luo, J.; Huang, S.X. Tropane alkaloids biosynthesis involves an unusual type III polyketide synthase and non-enzymatic condensation. Nat. Commun. 2019, 10, 4036. [Google Scholar] [CrossRef] [Green Version]
- Huccetogullari, D.; Luo, Z.W.; Lee, S.Y. Metabolic engineering of microorganisms for production of aromatic compounds. Microb. Cell Factories 2019, 18, 41. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D1. [Google Scholar]
- Krey Karol, L.; Nabity Paul, D.; Blubaugh Carmen, K.; Zhen, F.; Van Leuven James, T. Organic Farming Sharpens Plant Defenses in the Field. Front. Sustain. Food Syst. 2020, 4, 97. Available online: https://www.frontiersin.org/article/10.3389/fsufs.2020.00097 (accessed on 29 December 2021). [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).