Targeted Strategies for Degradation of Key Transmembrane Proteins in Cancer
Abstract
:1. Introduction
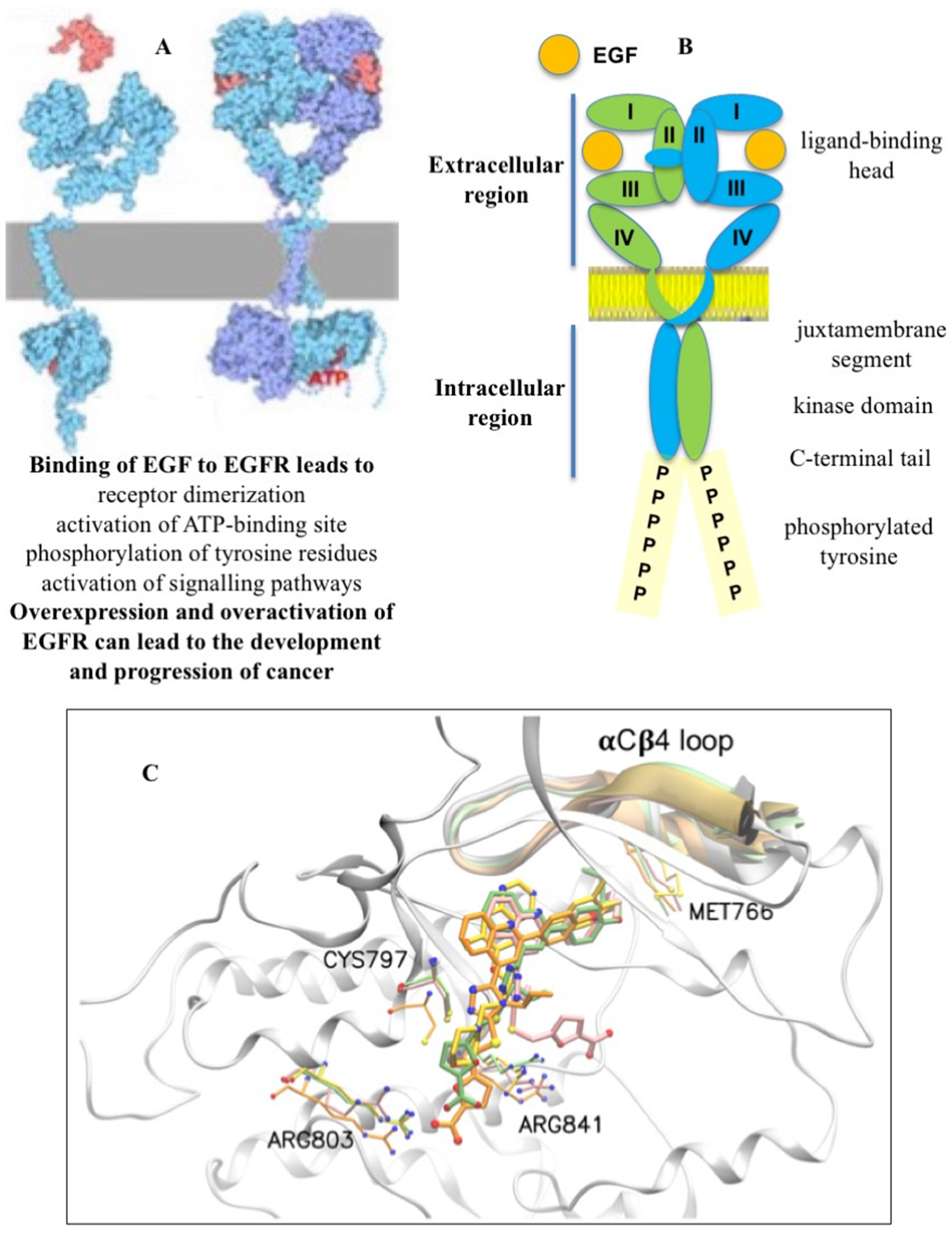
2. Targeted Inhibition of Receptor Tyrosine Kinase EGFR
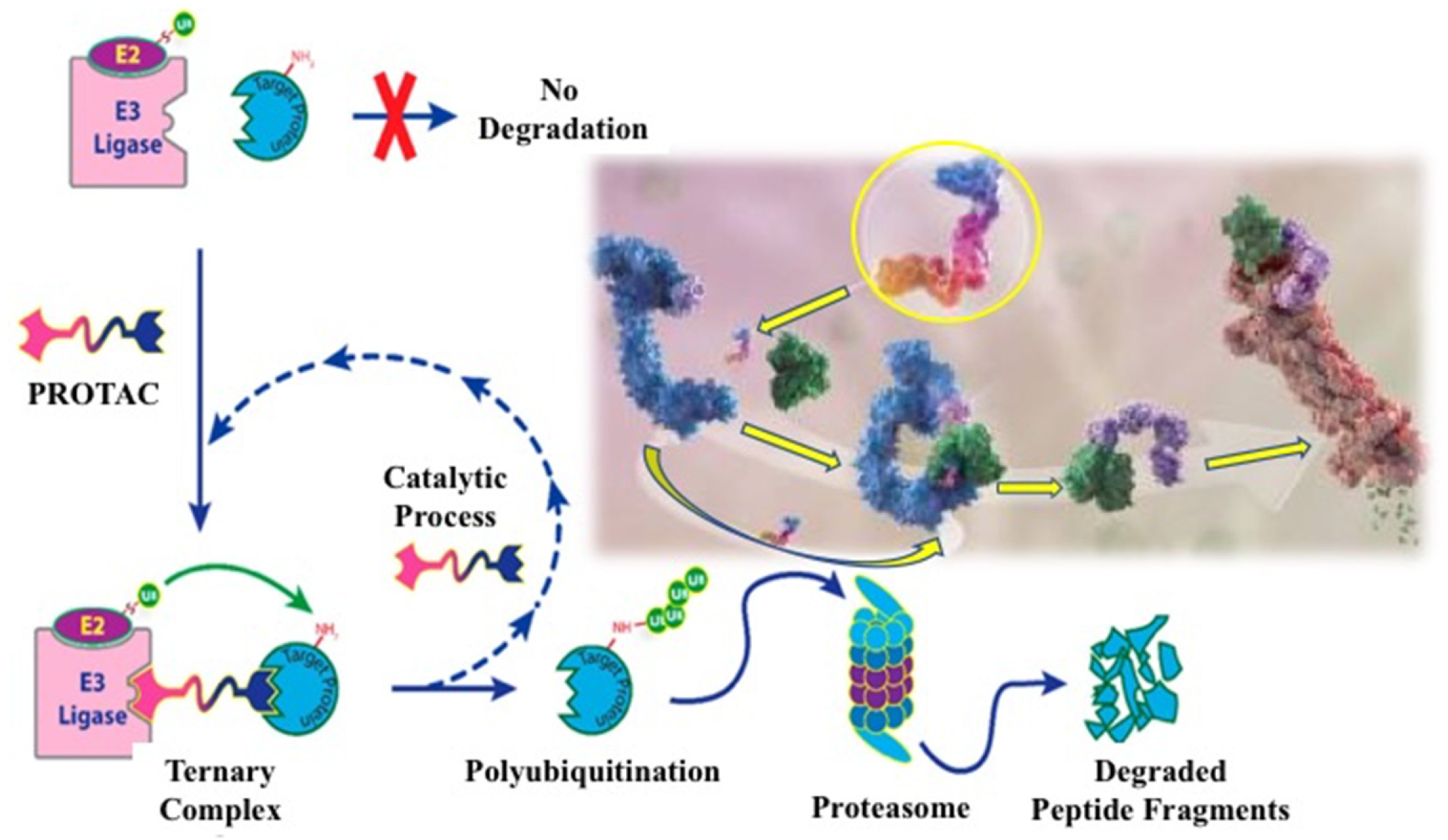
3. EGFR Degradation by PROTAC Technology
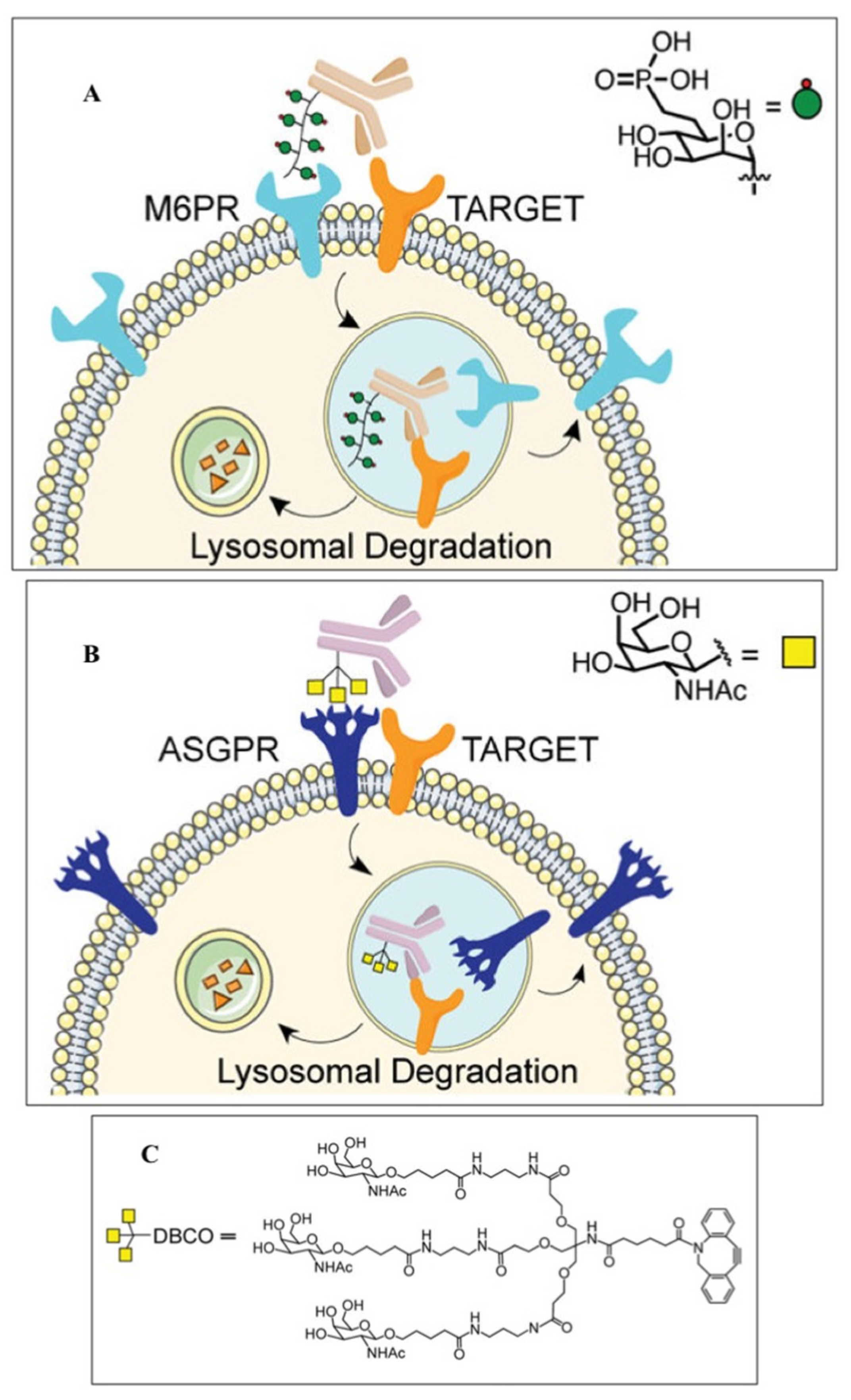
4. Alternative Strategies for EGFR Degradation
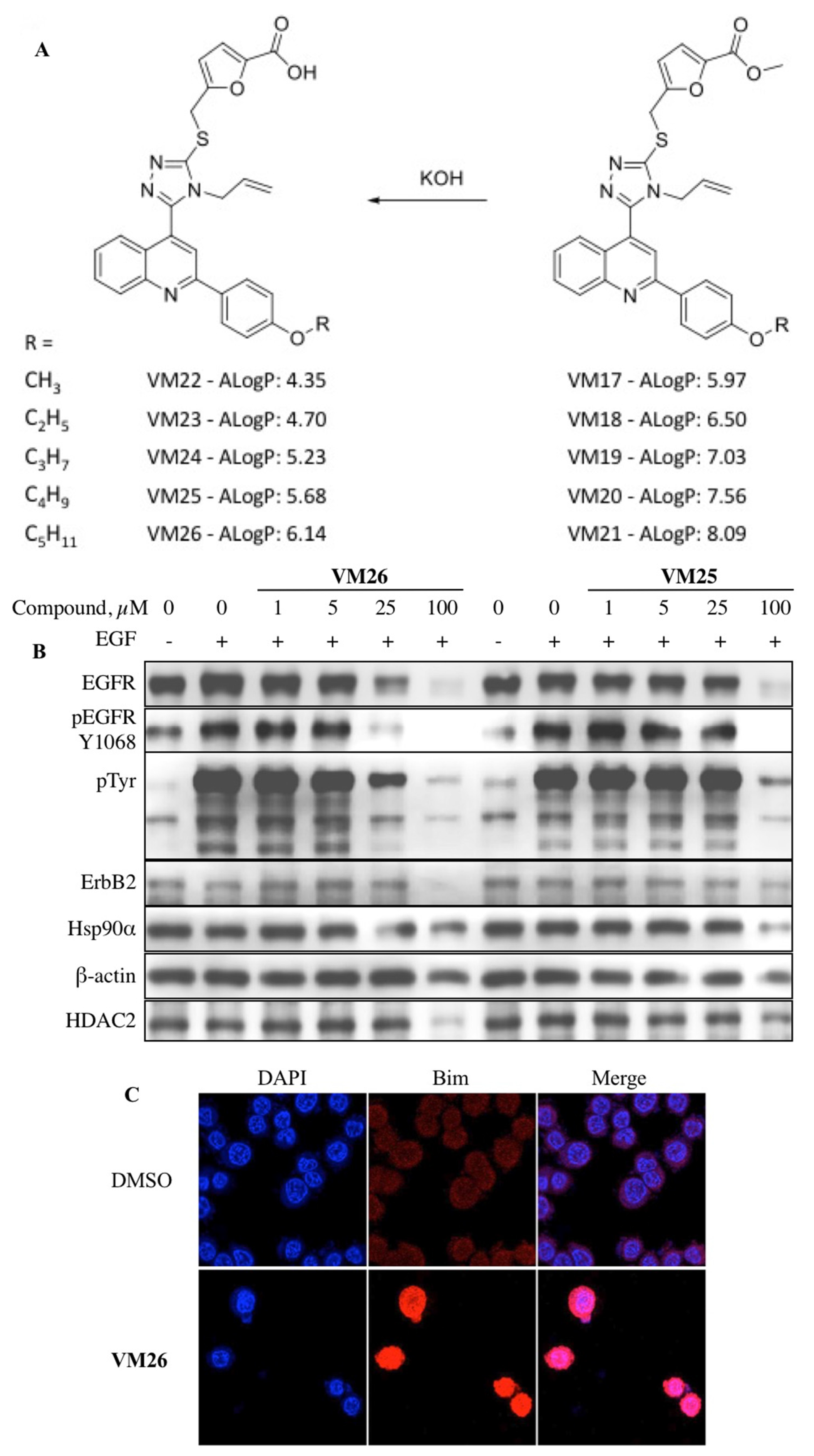
5. New Allosteric Chemicals Bind to EGFR and Lead to Cancer Cell Death
6. New Vision on Cancer Chemotherapy
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Soneji, S.; Beltrán-Sánchez, H.; Sox, H.C. Assessing progress in reducing the burden of cancer mortality, 1985–2005. J. Clin. Oncol. 2014, 32, 444–448. [Google Scholar] [CrossRef] [PubMed]
- Miller, K.D.; Nogueira, L.; Devasia, T.; Mariotto, A.B.; Yabroff, K.R.; Jemal, A.; Kramer, J.; Siegel, R.L. Cancer treatment and survivorship statistics, 2022. CA Cancer J. Clin. 2022, 72, 409–436. [Google Scholar] [CrossRef] [PubMed]
- Willemsen, M.C.; Mons, U.; Fernández, E. Tobacco control in Europe: Progress and key challenges. Tob. Control 2022, 31, 160–163. [Google Scholar] [CrossRef] [PubMed]
- Patel, V.R.; Qasim Hussaini, S.M.; Blaes, A.H.; Morgans, A.K.; Haynes, A.B.; Adamson, A.S.; Gupta, A. Trends in the prevalence of functional limitations among US cancer survivors, 1999–2018. JAMA Oncol. 2023, 9, 1001–1003. [Google Scholar] [CrossRef]
- Jotte, R.M.; Spigel, D.R. Advances in molecular-based personalized non-small-cell lung cancer therapy: Targeting epidermal growth factor receptor and mechanisms of resistance. Cancer Med. 2015, 4, 1621–1632. [Google Scholar] [CrossRef] [PubMed]
- Castañeda, A.M.; Meléndez, C.M.; Uribe, D.; Pedroza-Díaz, J. Synergistic effects of natural compounds and conventional chemotherapeutic agents: Recent insights for the development of cancer treatment strategies. Heliyon 2022, 8, e09519. [Google Scholar] [CrossRef]
- Seshacharyulu, P.; Ponnusamy, M.P.; Haridas, D.; Jain, M.; Ganti, A.K.; Batra, S.K. Targeting the EGFR signaling pathway in cancer therapy. Expert Opin. Ther. Targets 2012, 16, 15–31. [Google Scholar] [CrossRef] [PubMed]
- Patnaik, S.K.; Chandrasekar, M.J.N.; Nagarjuna, P.; Ramamurthi, D.; Swaroop, A.K. Targeting of ErbB1, ErbB2, and their dual targeting using small molecules and natural peptides: Blocking EGFR cell signaling pathways in cancer: A mini-review. Mini Rev. Med. Chem. 2022, 22, 2831–2846. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef]
- Falzone, L.; Salomone, S.; Libra, M. Evolution of cancer pharmacological treatments at the turn of the third millennium. Front. Pharmacol. 2018, 9, 1300. [Google Scholar] [CrossRef]
- Du, Z.; Lovly, C.M. Mechanisms of receptor tyrosine kinase activation in cancer. Mol. Cancer 2018, 17, 58. [Google Scholar] [CrossRef]
- Levantini, E.; Maroni, G.; Del Re, M.; Tenen, D.G. EGFR signaling pathway as therapeutic target in human cancers. Semin. Cancer Biol. 2022, 85, 253–275. [Google Scholar] [CrossRef]
- Pucci, C.; Martinelli, C.; Ciofani, G. Innovative approaches for cancer treatment: Current perspectives and new challenges. Ecancermedicalscience 2019, 13, 961. [Google Scholar] [CrossRef]
- Zhang, C.; Xu, C.; Gao, X.; Yao, Q. Platinum-based drugs for cancer therapy and anti-tumor strategies. Theranostics 2022, 12, 2115–2132. [Google Scholar] [CrossRef]
- Schlessinger, J. Receptor tyrosine kinases: Legacy of the first two decades. Cold Spring Harb. Perspect. Biol. 2014, 6, a008912. [Google Scholar] [CrossRef]
- Sigismund, S.; Avanzato, D.; Lanzetti, L. Emerging functions of the EGFR in cancer. Mol. Oncol. 2018, 12, 3–20. [Google Scholar] [CrossRef]
- Uribe, M.L.; Marrocco, I.; Yarden, Y. EGFR in Cancer: Signaling Mechanisms, Drugs, and Acquired Resistance. Cancers 2021, 13, 2748. [Google Scholar] [CrossRef]
- Dawson, J.P.; Berger, M.B.; Lin, C.C.; Schlessinger, J.; Lemmon, M.A.; Ferguson, K.M. Epidermal growth factor receptor dimerization and activation require ligand-induced conformational changes in the dimer interface. Mol. Cell. Biol. 2005, 25, 7734–7742. [Google Scholar] [CrossRef]
- Stamos, J.; Sliwkowski, M.X.; Eigenbrot, C. Structure of the epidermal growth factor receptor kinase domain alone and in complex with a 4-anilinoquinazoline inhibitor. J. Biol. Chem. 2002, 277, 46265–46272. [Google Scholar] [CrossRef] [PubMed]
- Bae, Y.S.; Kang, S.W.; Seo, M.S.; Baines, I.C.; Tekle, E.; Chock, P.B.; Rhee, S.G. Epidermal growth factor (EGF)-induced generation of hydrogen peroxide. Role in EGF receptor-mediated tyrosine phosphorylation. J. Biol. Chem. 1997, 272, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, C.E.; Truong, T.H.; Garcia, F.J.; Homann, A.; Gupta, V.; Leonard, S.E.; Carroll, K.S. Peroxide-dependent sulfenylation of the EGFR catalytic site enhances kinase activity. Nat. Chem. Biol. 2011, 8, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Carroll, K.S. Chemical biology approaches to study protein cysteine sulfenylation. Biopolymers 2014, 101, 165–172. [Google Scholar] [CrossRef]
- Sakanyan, V.; Angelini, M.; Le Béchec, M.; Lecocq, M.F.; Benaiteau, F.; Rousseau, B.; Gyulkhandanyan, A.; Gyulkhandanyan, L.; Logé, C.; Reiter, E.; et al. Screening and discovery of nitro-benzoxadiazole compounds activating epidermal growth factor receptor (EGFR) in cancer cells. Sci. Rep. 2014, 4, 3977. [Google Scholar] [CrossRef]
- Sakanyan, V.; Hulin, P.; Alves de Sousa, R.; Silva, V.A.; Hambardzumyan, A.; Nedellec, S.; Tomasoni, C.; Logé, C.; Pineau, C.; Roussakis, C.; et al. Activation of EGFR by small compounds through coupling the generation of hydrogen peroxide to stable dimerization of Cu/Zn SOD1. Sci. Rep. 2016, 6, 21088. [Google Scholar] [CrossRef]
- Sakanyan, V.; Benaiteau, F.; Alves de Sousa, R.; Pineau, C.; Artaud, I. Straightforward detection of reactive compound binding to multiple proteins in cancer cells: Towards a better understanding of electrophilic stress. Ann. Clin. Exp. Metabol. 2016, 1, 1006. [Google Scholar]
- Silva, V.A.; Lafont, F.; Benhelli-Mokrani, H.; Breton, M.L.; Hulin, P.; Chabot, T.; Paris, F.; Sakanyan, V.; Fleury, F. Rapid diminution in the level and activity of DNA-dependent protein kinase in cancer cells by a reactive nitro-benzoxadiazole compound. Int. J. Mol. Sci. 2016, 17, 703. [Google Scholar] [CrossRef]
- Sakanyan, V. Reactive Chemicals and Electrophilic Stress in Cancer: A Minireview. High-Throughput 2018, 7, 12. [Google Scholar] [CrossRef]
- Parzych, K.R.; Klionsky, D.J. An overview of autophagy: Morphology, mechanism, and regulation. Antioxid. Redox Signal. 2014, 20, 460–473. [Google Scholar] [CrossRef]
- von Zastrow, M.; Sorkin, A. Mechanisms for Regulating and Organizing Receptor Signaling by Endocytosis. Annu. Rev. Biochem. 2021, 90, 709–737. [Google Scholar] [CrossRef] [PubMed]
- Tomas, A.; Futter, C.E.; Eden, E.R. EGF receptor trafficking: Consequences for signaling and cancer. Trends Cell Biol. 2014, 24, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Caldieri, G.; Malabarba, M.G.; Di Fiore, P.P.; Sigismund, S. EGFR Trafficking in Physiology and Cancer. Prog. Mol. Subcell. Biol. 2018, 57, 235–272. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Sakurai, H. New trend in ligand-induced EGFR trafficking: A dual-mode clathrin-mediated endocytosis model. J. Proteom. 2022, 255, 104503. [Google Scholar] [CrossRef]
- Mendelsohn, J.; Baselga, J. Epidermal growth factor receptor targeting in cancer. Semin. Oncol. 2006, 33, 369–385. [Google Scholar] [CrossRef]
- Cohen, P.; Cross, D.; Jänne, P.A. Kinase drug discovery 20 years after imatinib. Nat. Rev. Drug Discov. 2021, 20, 551–569. [Google Scholar] [CrossRef]
- Red Brewer, M.; Yun, C.H.; Lai, D.; Lemmon, M.A.; Eck, M.J.; Pao, W. Mechanism for activation of mutated epidermal growth factor receptors in lung cancer. Proc. Natl. Acad. Sci. USA 2013, 110, e3595-604, Erratum in Proc. Natl. Acad. Sci. USA 2013, 110, 20344. [Google Scholar] [CrossRef]
- Arter, C.; Trask, L.; Ward, S.; Yeoh, S.; Bayliss, R. Structural features of the protein kinase domain and targeted binding by small-molecule inhibitors. J. Biol. Chem. 2022, 298, 102247. [Google Scholar] [CrossRef]
- Herbst, R.S.; Fukuoka, M.; Baselga, J. Gefitinib—A novel targeted approach to treating cancer. Nat. Rev. Cancer 2004, 4, 956–965. [Google Scholar] [CrossRef]
- Cappuzzo, F.; Ciuleanu, T.; Stelmakh, L.; Cicenas, S.; Szczésna, A.; Juhász, E.; Esteban, E.; Molinier, O.; Brugger, W.; Melezínek, I.; et al. SATURN investigators. Erlotinib as maintenance treatment in advanced non-small-cell lung cancer: A multicentre, randomised, placebo-controlled phase 3 study. Lancet Oncol. 2010, 11, 521–529. [Google Scholar] [CrossRef]
- Jänne, P.A. Challenges of detecting EGFR T790M in gefitinib/erlotinib-resistant tumours. Lung Cancer 2008, 60 (Suppl. 2), S3–S9. [Google Scholar] [CrossRef]
- Shah, R.; Lester, J.F. Tyrosine kinase inhibitors for the yreatment of EGFR mutation-positive Non-Small-Cell Lung Cancer: A clash of the generations. Clin. Lung Cancer 2020, 21, e216–e228. [Google Scholar] [CrossRef]
- Hirsch, F.R.; Scagliotti, G.V.; Mulshine, J.L.; Kwon, R.; Curran, W.J.; Wu, Y.-L.; Paz-Ares, L. Lung cancer: Current therapies and new targeted treatments. Lancet 2017, 389, 299–311. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Luo, S.; Lin, H.; Yang, H.; Chen, H.; Liao, Z.; Lin, W.; Zheng, W.; Xie, X. Correlation between EGFR mutation status and the incidence of brain metastases in patients with non-small cell lung cancer. J. Thorac. Dis. 2017, 9, 2510–2520. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Franchina, T.; Ricciardi, G.R.R.; Picone, A.; Ferraro, G.; Zanghì, M.; Toscano, G.; Giordano, A.; Adamo, V. A decade of EGFR inhibition in EGFR-mutated non-small-cell lung cancer (NSCLC): Old successes and future perspectives. Oncotarget 2015, 6, 26814–26825. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Boggon, T.J.; Dayaram, T.; Janne, P.A.; Kocher, O.; Meyerson, M.; Johnson, B.E.; Eck, M.J.; Tenen, D.G.; Halmos, B. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 2005, 352, 786–792. [Google Scholar] [CrossRef]
- Pao, W.; Miller, V.A.; Politi, K.A.; Riely, G.J.; Somwar, R.; Zakowski, M.F.; Kris, M.G.; Varmus, H. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med. 2005, 2, e73. [Google Scholar] [CrossRef]
- Godin-Heymann, N.; Ulkus, L.; Brannigan, B.W.; McDermott, U.; Lamb, J.; Maheswaran, S.; Settleman, J.; Haber, D.A. The T790M “gatekeeper” mutation in EGFR mediates resistance to low concentrations of an irreversible EGFR inhibitor. Mol. Cancer Ther. 2008, 7, 874–879. [Google Scholar] [CrossRef]
- Yun, C.H.; Mengwasser, K.E.; Toms, A.V.; Woo, M.S.; Greulich, H.; Wong, K.K.; Meyerson, M.; Eck, M.J. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. USA 2008, 105, 2070–2075. [Google Scholar] [CrossRef]
- Yu, H.A.; Riely, G.J. Second-generation epidermal growth factor receptor tyrosine kinase inhibitors in lung cancers. J. Natl. Compr. Cancer Netw. 2013, 11, 161–169. [Google Scholar] [CrossRef]
- Meng, Y.; Yu, B.; Huang, H.; Peng, Y.; Li, E.; Yao, Y.; Song, C.; Yu, W.; Zhu, K.; Wang, K.; et al. Discovery of dosimertinib, a highly potent, selective, and orally efficacious deuterated EGFR targeting clinical candidate for the treatment of Non-Small-Cell Lung Cancer. J. Med. Chem. 2021, 64, 925–937. [Google Scholar] [CrossRef]
- Liu, Q.; Sabnis, Y.; Zhao, Z.; Zhang, T.; Buhrlage, S.J.; Jones, L.H.; Gray, N.S. Developing irreversible inhibitors of the protein kinase cysteinome. Chem. Biol. 2013, 20, 146–159. [Google Scholar] [CrossRef]
- da Cunha Santos, G.; Shepherd, F.A.; Tsao, M.S. EGFR mutations and lung cancer. Annu. Rev. Pathol. 2011, 6, 49–69. [Google Scholar] [CrossRef]
- Herbst, R.S.; Morgensztern, D.; Boshoff, C. The biology and management of non-small cell lung cancer. Nature 2018, 553, 446–454. [Google Scholar] [CrossRef]
- Huang, F.; Han, X.; Xiao, X.; Zhou, J. Covalent warheads targeting cysteine residue: The promising approach in drug development. Molecules 2022, 27, 7728. [Google Scholar] [CrossRef]
- Zhou, W.; Ercan, D.; Chen, L.; Yun, C.H.; Li, D.; Capelletti, M.; Cortot, A.B.; Chirieac, L.; Iacob, R.E.; Padera, R.; et al. Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature 2009, 462, 1070–1074. [Google Scholar] [CrossRef]
- Finlay, M.R.; Anderton, M.; Ashton, S.; Ballard, P.; Bethel, P.A.; Box, M.R.; Bradbury, R.H.; Brown, S.J.; Butterworth, S.; Campbell, A.; et al. Discovery of a potent and selective EGFR inhibitor (AZD9291) of both sensitizing and T790M resistance mutations that spares the wild type form of the receptor. J. Med. Chem. 2014, 57, 8249–8267. [Google Scholar] [CrossRef]
- Shah, M.P.; Neal, J.W. Targeting acquired and intrinsic resistance mechanisms in Epidermal Growth Factor Receptor mutant Non-Small-Cell Lung Cancer. Drugs 2022, 82, 649–662. [Google Scholar] [CrossRef]
- He, J.; Zhou, Z.; Sun, X.; Yang, Z.; Zheng, P.; Xu, S.; Zhu, W. The new opportunities in medicinal chemistry of fourth-generation EGFR inhibitors to overcome C797S mutation. Eur. J. Med. Chem. 2021, 210, 112995. [Google Scholar] [CrossRef]
- Wang, Z.; Yang, J.J.; Huang, J.; Ye, J.Y.; Zhang, X.C.; Tu, H.Y.; Han-Zhang, H.; Wu, Y.L. Lung adenocarcinoma harboring EGFR T790M and in trans C797S responds to combination therapy of first- and third-generation EGFR TKIs and shifts allelic configuration at resistance. J. Thorac. Oncol. 2017, 12, 1723–1727. [Google Scholar] [CrossRef]
- Jia, Y.; Yun, C.H.; Park, E.; Ercan, D.; Manuia, M.; Juarez, J.; Xu, C.; Rhee, K.; Chen, T.; Zhang, H.; et al. Overcoming EGFR(T790M) and EGFR(C797S) resistance with mutant-selective allosteric inhibitors. Nature 2016, 534, 129–132. [Google Scholar] [CrossRef]
- Wang, S.; Song, Y.; Liu, D. EAI045: The fourth-generation EGFR inhibitor overcoming T790M and C797S resistance. Cancer Lett. 2017, 385, 51–54. [Google Scholar] [CrossRef]
- To, C.; Jang, J.; Chen, T.; Park, E.; Mushajiang, M.; De Clercq, D.J.H.; Xu, M.; Wang, S.; Cameron, M.D.; Heppner, D.E.; et al. Single and dual targeting of mutant EGFR with an allosteric inhibitor. Cancer Discov. 2019, 9, 926–943. [Google Scholar] [CrossRef]
- Jang, J.; To, C.; De Clercq, D.J.H.; Park, E.; Ponthier, C.M.; Shin, B.H.; Mushajiang, M.; Nowak, R.P.; Fischer, E.S.; Eck, M.J.; et al. Mutant-selective allosteric EGFR degraders are effective against a broad range of drug-resistant mutations. Angew. Chem. Int. Ed. Engl. 2020, 59, 14481–14489. [Google Scholar] [CrossRef]
- Dong, H.; Ye, X.; Zhu, Y.; Shen, H.; Shen, H.; Chen, W.; Ji, M.; Zheng, M.; Wang, K.; Cai, Z.; et al. Discovery of potent and wild-type-sparing fourth-generation EGFR Inhibitors for treatment of osimertinib-resistance NSCLC. J. Med. Chem. 2023, 66, 6849–6868. [Google Scholar] [CrossRef]
- Marasco, M.; Misale, S. Resistance is futile with fourth-generation EGFR inhibitors. Nat. Cancer 2022, 3, 381–383. [Google Scholar] [CrossRef]
- Xu, L.; Xu, B.; Wang, J.; Gao, Y.; He, X.; Xie, T.; Ye, X.Y. Recent advances of novel fourth generation EGFR inhibitors in overcoming C797S mutation of lung cancer therapy. Eur. J. Med. Chem. 2023, 245 Pt 1, 114900. [Google Scholar] [CrossRef]
- Chirnomas, D.; Hornberger, K.R.; Crews, C.M. Protein degraders enter the clinic—A new approach to cancer therapy. Nat. Rev. Clin. Oncol. 2023, 20, 265–278. [Google Scholar] [CrossRef]
- Sakamoto, K.M.; Kim, K.B.; Kumagai, A.; Mercurio, F.; Crews, C.M.; Deshaies, R.J. Protacs: Chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl. Acad. Sci. USA 2001, 98, 8554–8559. [Google Scholar] [CrossRef]
- Sakamoto, K.M.; Kim, K.B.; Verma, R.; Ransick, A.; Stein, B.; Crews, C.M.; Deshaies, R.J. Development of Protacs to target cancer-promoting proteins for ubiquitination and degradation. Mol. Cell. Proteom. 2003, 2, 1350–1358. [Google Scholar] [CrossRef]
- Bondeson, D.; Mares, A.; Smith, I.; Ko, E.; Campos, S.; Miah, A.H.; Mulholland, K.E.; Routly, N.; Buckley, D.L.; Gustafson, J.L.; et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 2015, 11, 611–617. [Google Scholar] [CrossRef]
- Dale, B.; Cheng, M.; Park, K.S.; Kaniskan, H.Ü.; Xiong, Y.; Jin, J. Advancing targeted protein degradation for cancer therapy. Nat. Rev. Cancer 2021, 21, 638–654. [Google Scholar] [CrossRef] [PubMed]
- Békés, M.; Langley, D.R.; Crews, C.M. PROTAC targeted protein degraders: The past is prologue. Nat. Rev. Drug Discov. 2022, 21, 181–200. [Google Scholar] [CrossRef] [PubMed]
- Hipp, M.S.; Kasturi, P.; Hartl, F.U. The proteostasis network and its decline in ageing. Nat. Rev. Mol. Cell Biol. 2019, 20, 421–435. [Google Scholar] [CrossRef]
- Li, X.; Pu, W.; Zheng, Q.; Ai, M.; Chen, S.; Peng, Y. Proteolysis-targeting chimeras (PROTACs) in cancer therapy. Mol. Cancer 2022, 21, 99. [Google Scholar] [CrossRef]
- Yau, R.; Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 2016, 18, 579–586. [Google Scholar] [CrossRef] [PubMed]
- Bard, J.A.M.; Goodall, E.A.; Greene, E.R.; Jonsson, E.; Dong, K.C.; Martin, A. Structure and function of the 26S proteasome. Annu. Rev. Biochem. 2018, 87, 697–724. [Google Scholar] [CrossRef]
- Wolska-Washer, A.; Smolewski, P. Targeting protein degradation pathways in tumors: Focusing on their role in hematological malignancies. Cancers 2022, 14, 3778. [Google Scholar] [CrossRef]
- Ishida, T.; Ciulli, A. E3 ligase ligands for PROTACs: How they were found and How to discover new ones. SLAS Discov. 2021, 26, 484–502. [Google Scholar] [CrossRef]
- Li, M.; Zhi, Y.; Liu, B.; Yao, Q. Advancing strategies for proteolysis-targeting chimera design. J. Med. Chem. 2023, 66, 2308–2329. [Google Scholar] [CrossRef]
- Smith, B.E.; Wang, S.L.; Jaime-Figueroa, S.; Harbin, A.; Wang, J.; Hamman, B.D.; Crews, C.M. Differential PROTAC substrate specificity dictated by orientation of recruited E3 ligase. Nat. Commun. 2019, 10, 131. [Google Scholar] [CrossRef] [PubMed]
- Kofink, C.; Trainor, N.; Mair, B.; Wohrle, S.; Wurm, M.; Mischerikow, N.; Roy, M.J.; Bader, G.; Greb, P.; Garavel, G.; et al. A selective and orally bioavailable VHL-recruiting PROTAC achieves SMARCA2 degradation in vivo. Nat. Commun. 2022, 13, 5969. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Huang, Y.; Huang, J.; He, R.; Huang, M.; Lu, X.; Xu, Y.; Zhou, F.; Zhang, Z.; Ding, K. Discovery of the first examples of threonine tyrosine kinase PROTAC degraders. J. Med. Chem. 2022, 65, 2313–2328. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Hu, B.; Wang, M.; Xu, F.; Miao, B.; Yang, C.-Y.; Wang, M.; Liu, Z.; Hayes, D.F.; Chinnaswamy, K.; et al. Discovery of ERD-308 as a highly potent proteolysis targeting chimera (PROTAC) degrader of estrogen receptor (ER). J. Med. Chem. 2019, 62, 1420–1442. [Google Scholar] [CrossRef] [PubMed]
- Yokoo, H.; Shibata, N.; Naganuma, M.; Murakami, Y.; Fujii, K.; Ito, T.; Aritake, K.; Naito, M.; Demizu, Y. Development of a hematopoietic prostaglandin D synthase-degradation inducer. ACS Med. Chem. Lett. 2021, 12, 236–241. [Google Scholar] [CrossRef]
- Khan, S.; Zhang, X.; Lv, D.; Zhang, Q.; He, Y.; Zhang, P.; Liu, X.; Thummuri, D.; Yuan, Y.; Wiegand, J.S.; et al. A selective BCL-XL PROTAC degrader achieves safe and potent antitumor activity. Nat. Med. 2019, 25, 1938–1947. [Google Scholar] [CrossRef]
- Li, X.; Song, Y. Proteolysis-targeting chimera (PROTAC) for targeted protein degradation and cancer therapy. J. Hematol. Oncol. 2020, 13, 50. [Google Scholar] [CrossRef]
- Zhao, Q.; Lan, T.; Su, S.; Rao, Y. Induction of apoptosis in MDAMB-231 breast cancer cells by a PARP1-targeting PROTAC small molecule. Chem. Commun. 2019, 55, 369–372. [Google Scholar] [CrossRef]
- Yan, W.; Pan, B.; Shao, J.; Lin, H.; Li, H. Feasible column chromatography-free, multi-gram scale synthetic process of VH032 amine, which could enable rapid PROTAC library construction. ACS Omega 2022, 7, 26015–26020. [Google Scholar] [CrossRef]
- Burslem, G.M.; Smith, B.E.; Lai, A.C.; Jaime-Figueroa, S.; McQuaid, D.C.; Bondeson, D.P.; Toure, M.; Dong, H.; Qian, Y.; Wang, J.; et al. The advantages of targeted protein degradation over inhibition: An RTK case study. Cell Chem. Biol. 2018, 25, 67–77.e3. [Google Scholar] [CrossRef]
- Jones, L.H.; Mitchell, C.A.; Loberg, L.; Pavkovic, M.; Rao, M.; Roberts, R.; Stamp, K.; Volak, L.; Wittwer, M.B.; Pettit, S. Targeted protein degraders: A call for collective action to advance safety assessment. Nat. Rev. Drug Discov. 2022, 21, 401–402. [Google Scholar] [CrossRef] [PubMed]
- Zeng, S.; Huang, W.; Zheng, X.; Cheng, L.; Zhang, Z.; Wang, J.; Shen, Z. Proteolysis targeting chimera (PROTAC) in drug discovery paradigm: Recent progress and future challenges. Eur. J. Med. Chem. 2021, 210, 112981. [Google Scholar] [CrossRef] [PubMed]
- Ottis, P.; Crews, C.M. Proteolysis-targeting chimeras: Induced protein degradation as a therapeutic strategy. ACS Chem. Biol. 2017, 12, 892–898. [Google Scholar] [CrossRef] [PubMed]
- Bondeson, D.P.; Smith, B.E.; Burslem, G.M.; Buhimschi, A.D.; Hines, J.; Jaime-Figueroa, S.; Wang, J.; Hamman, B.D.; Ishchenko, A.; Crews, C.M. Lessons in PROTAC design from selective degradation with a promiscuous warhead. Cell Chem. Biol. 2018, 25, 78–87.e5. [Google Scholar] [CrossRef]
- Cromm, P.M.; Samarasinghe, K.T.G.; Hines, J.; Crews, C.M. Addressing kinase-independent functions of Fak via PROTAC mediated degradation. J. Am. Chem. Soc. 2018, 140, 17019–17026. [Google Scholar] [CrossRef]
- Burslem, G.M.; Crews, C.M. Proteolysis-targeting chimeras as therapeutics and tools for biological discovery. Cell 2020, 181, 102–114. [Google Scholar] [CrossRef]
- Cheng, M.; Yu, X.; Lu, K.; Xie, L.; Wang, L.; Meng, F.; Han, X.; Chen, X.; Liu, J.; Xiong, Y.; et al. Discovery of potent and selective epidermal growth factor receptor (EGFR) bifunctional small-molecule degraders. J. Med. Chem. 2020, 63, 1216–1232. [Google Scholar] [CrossRef]
- He, K.; Zhang, Z.; Wang, W.; Zheng, X.; Wang, X.; Zhang, X. Discovery and biological evaluation of proteolysis targeting chimeras (PROTACs) as EGFR degraders based on osimertinib and lenalidomide. Bioorg. Med. Chem. Lett. 2020, 30, 127167. [Google Scholar] [CrossRef]
- Qu, X.; Liu, H.; Song, X.; Sun, N.; Zhong, H.; Qiu, X.; Yang, X.; Jiang, B. Effective degradation of EGFRL858R+T790M mutant proteins by CRBN-based PROTACs through both proteosome and autophagy/lysosome degradation systems. Eur. J. Med. Chem. 2021, 218, 113328. [Google Scholar] [CrossRef]
- Zhang, X.; Xu, F.; Tong, L.; Zhang, T.; Xie, H.; Lu, X.; Ren, X.; Ding, K. Design and synthesis of selective degraders of EGFRL858R/T790M mutant. Eur. J. Med. Chem. 2020, 192, 112199. [Google Scholar] [CrossRef]
- Zhao, H.-Y.; Yang, X.-Y.; Lei, H.; Xi, X.-X.; Lu, S.-M.; Zhang, J.-J.; Xin, M.; Zhang, S.-Q. Discovery of potent small molecule PROTACs targeting mutant EGFR. Eur. J. Med. Chem. 2020, 208, 112781. [Google Scholar] [CrossRef]
- Kabeya, Y.; Mizushima, N.; Ueno, T.; Yamamoto, A.; Kirisako, T.; Noda, T.; Kominami, E.; Ohsumi, Y.; Yoshimori, T. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 2000, 19, 5720–5728. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, H.-Y.; Xi, X.-X.; Liu, Y.-J.; Xin, M.; Mao, S.; Zhang, J.-J.; Lu, A.X.; Zhang, S.-Q. Discovery of potent epidermal growth factor receptor (EGFR) degraders by proteolysis targeting chimera (PROTAC). Eur. J. Med. Chem. 2020, 189, 112061. [Google Scholar] [CrossRef]
- Zhao, H.-Y.; Wang, H.P.; Mao, Y.Z.; Zhang, H.; Xin, M.; Xi, X.-X.; Lei, H.; Mao, S.; Li, D.H.; Zhang, S.-Q. Discovery of potent PROTACs targeting EGFR mutants through the optimization of covalent EGFR ligands. J. Med. Chem. 2022, 65, 4709–4726. [Google Scholar] [CrossRef]
- Ricordel, C.; Friboulet, L.; Facchinetti, F.; Soria, J.C. Molecular mechanisms of acquired resistance to third-generation EGFR-TKIs in EGFR T790M-mutant lung cancer. Ann. Oncol. 2018, 29 (Suppl. 1), 128–137. [Google Scholar] [CrossRef]
- Wang, S.H.; Tsui, S.T.; Liu, C.; Song, Y.P.; Liu, D.L. EGFR C797S mutation mediates resistance to third-generation inhibitors in T790M-positive non-small cell lung cancer. J. Hematol. Oncol. 2016, 9, 59. [Google Scholar] [CrossRef]
- Zhang, H.; Xie, R.; AI-Furas, H.; Li, Y.; Wu, Q.; Li, J.; Xu, F.; Xu, T. Design, synthesis, and biological evaluation of novel EGFR PROTACs targeting del19/T790M/C797S mutation. ACS Med. Chem. Lett. 2022, 13, 278–283. [Google Scholar] [CrossRef]
- Chen, Y.; Tandon, I.; Heelan, W.; Wang, Y.X.; Tang, W.P.; Hu, Q.Y. Proteolysis-targeting chimera (PROTAC) delivery system: Advancing protein degraders towards clinical translation. Chem. Soc. Rev. 2022, 51, 5330–5350. [Google Scholar] [CrossRef]
- Raina, K.; Lu, J.; Qian, Y.; Altieri, M.; Gordon, D.; Rossi, A.M.K.; Wang, J.; Chen, X.; Dong, H.Q.; Siu, K.; et al. PROTAC-induced BET protein degradation as a therapy for castration-resistant prostate cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 7124–7129. [Google Scholar] [CrossRef]
- Moreau, K.; Coen, M.; Zhang, A.X.; Pachl, F.; Castaldi, M.P.; Dahl, G.; Boyd, H.; Scott, C.; Newham, P. Proteolysis-targeting chimeras in drug development: A safety perspective. Brit. J. Pharmacol. 2020, 177, 1709–1718. [Google Scholar] [CrossRef]
- Hong, K.B.; An, H. Degrader-antibody conjugates: Emerging new modality. J. Med. Chem. 2023, 66, 140–148. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Crews, C.M. PROTACs: Past, present and future. Chem. Soc. Rev. 2022, 51, 5214–5236. [Google Scholar] [CrossRef] [PubMed]
- Dragovich, P.S. Degrader-antibody conjugates. Chem. Soc. Rev. 2022, 51, 3886–3897. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Huang, L.; Zhou, Y.; Xue, T.; Chen, Z.; Han, G. Near-infrared-light activatable nanoparticles for deep-tissue-penetrating wireless optogenetics. Adv. Healthc. Mater. 2019, 8, e1801132. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Y.; Song, G.; He, Y.; Zhang, X.; Liu, Y.; Ju, H. A DNA-azobenzene nanopump fueled by upconversion luminescence for controllable intracellular drug release. Angew. Chem. Int. Ed. Engl. 2019, 58, 18207–18211. [Google Scholar] [CrossRef]
- He, Q.; Zhou, L.; Yu, D.; Zhu, R.; Chen, Y.; Song, M.; Liu, X.; Liao, Y.; Ding, T.; Fan, W.; et al. Near-infrared-activatable PROTAC nanocages for controllable target protein degradation and on-demand antitumor therapy. J. Med. Chem. 2023, 66, 10458–10472. [Google Scholar] [CrossRef]
- Schirrmacher, V. From chemotherapy to biological therapy: A review of novel concepts to reduce the side effects of systemic cancer treatment (Review). Int. J. Oncol. 2019, 54, 407–419. [Google Scholar] [CrossRef]
- Pirker, R. EGFR-directed monoclonal antibodies in non-small cell lung cancer. Target. Oncol. 2013, 8, 47–53. [Google Scholar] [CrossRef]
- Tabasinezhad, M.; Omidinia, E.; Talebkhan, Y.; Omrani, M.D.; Mahboudi, F.; Ghaedi, H.; Wenzel, W. The effects of somatic mutations on EGFR interaction with anti-EGFR monoclonal antibodies: Implication for acquired resistance. Proteins 2020, 88, 3–14. [Google Scholar] [CrossRef]
- Capdevila, J.; Elez, E.; Macarulla, T.; Ramos, F.J.; Ruiz-Echarri, M.; Tabernero, J. Anti-epidermal growth factor receptor monoclonal antibodies in cancer treatment. Cancer Treat. Rev. 2009, 35, 354–363. [Google Scholar] [CrossRef]
- Ermondi, G.; Garcia-Jimenez, D.; Caron, G. PROTACs and building blocks: The 2D chemical space in very early drug discovery. Molecules 2021, 26, 672. [Google Scholar] [CrossRef] [PubMed]
- Weng, G.; Cai, X.; Cao, D.; Du, H.; Shen, C.; Deng, Y.; He, Q.; Yang, B.; Li, D.; Hou, T. PROTAC-DB 2.0: An updated database of PROTACs. Nucleic Acids Res. 2023, 51, D1367–D1372. [Google Scholar] [CrossRef] [PubMed]
- O’Brien Laramy, M.N.; Luthra, S.; Brown, M.F.; Bartlett, D.W. Delivering on the promise of protein degraders. Nat. Rev. Drug Discov. 2023, 22, 410–427. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; de Camargo Correia, G.S.; Wang, J.; Manochakian, R.; Zhao, Y.; Lou, Y. Emerging targeted therapies in advanced Non-Small-Cell Lung Cancer. Cancers 2023, 15, 2899. [Google Scholar] [CrossRef] [PubMed]
- Banik, S.M.; Pedram, K.; Wisnovsky, S.; Ahn, G.; Riley, N.M.; Bertozzi, C.R. Lysosome- targeting chimaeras for degradation of extracellular proteins. Nature 2020, 584, 291–297. [Google Scholar] [CrossRef]
- Li, Z.; Zhu, C.; Ding, Y.; Fei, Y.; Lu, B. ATTEC: A potential new approach to target proteinopathies. Autophagy 2020, 16, 185–187. [Google Scholar] [CrossRef]
- Takahashi, D.; Arimoto, H. Targeting selective autophagy by AUTAC degraders. Autophagy 2020, 16, 765–766. [Google Scholar] [CrossRef]
- Ji, C.H.; Lee, M.J.; Kim, H.Y.; Heo, A.J.; Park, D.Y.; Kim, Y.K.; Kim, B.Y.; Kwon, Y.T. Targeted protein degradation via the autophagy-lysosome system: AUTOTAC (AUTOphagy-TArgeting Chimera). Autophagy 2022, 18, 2259–2262. [Google Scholar] [CrossRef]
- Itoh, Y.; Ishikawa, M.; Naito, M.; Hashimoto, Y. Protein knockdown using methyl bestatin-ligand hybrid molecules: Design and synthesis of inducers of ubiquitination-mediated degradation of cellular retinoic acid-binding proteins. J. Am. Chem. Soc. 2010, 132, 5820–5826. [Google Scholar] [CrossRef]
- Ahn, G.; Banik, S.M.; Bertozzi, C.R. Degradation from the outside in: Targeting extracellular and membrane proteins for degradation through the endolysosomal pathway. Cell Chem. Biol. 2021, 28, 1072–1080. [Google Scholar] [CrossRef]
- Hong, D.; Zhou, B.; Zhang, B.; Ren, H.; Zhu, L.; Zheng, G.; Ge, M.; Ge, J. Recent advances in the development of EGFR degraders: PROTACs and LYTACs. Eur. J. Med. Chem. 2022, 239, 114533. [Google Scholar] [CrossRef] [PubMed]
- Maity, P.; Chatterjee, J.; Patil, K.T.; Arora, S.; Katiyar, M.K.; Kumar, M.; Samarbakhsh, A.; Joshi, G.; Bhutani, P.; Chugh, M.; et al. Targeting the epidermal growth factor receptor with molecular degraders: State-of-the-art and future opportunities. J. Med. Chem. 2023, 66, 3135–3172. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, Q.; Xie, X.; Peng, C.; Pang, Q.; Liu, B.; Han, B. Application of novel degraders employing autophagy for expediting medicinal research. J. Med. Chem. 2023, 66, 1700–1711. [Google Scholar] [CrossRef] [PubMed]
- Ahn, G.; Banik, S.M.; Miller, C.L.; Riley, N.M.; Cochran, J.R.; Bertozzi, C.R. LYTACs that engage the asialoglycoprotein receptor for targeted protein degradation. Nat. Chem. Biol. 2021, 17, 937–946. [Google Scholar] [CrossRef]
- Iradyan, M.; Iradyan, N.; Hulin, P.; Hambardzumyan, A.; Gyulkhandanyan, A.; Alves de Sousa, R.; Hessani, A.; Roussakis, C.; Bollot, G.; Bauvais, C.; et al. Targeting degradation of EGFR through the allosteric site leads to cancer cell detachment-promoted death. Cancers 2019, 11, 1094. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Smith, I.E.; Reis-Filho, J.S. Triple-negative breast cancer. N. Engl. J. Med. 2010, 363, 1938–1948. [Google Scholar] [CrossRef]
- Tanida, I.; Ueno, T.; Kominami, E. LC3 conjugation system in mammalian autophagy. Int. J. Biochem. Cell. Biol. 2004, 36, 2503–2518. [Google Scholar] [CrossRef]
- Taipale, M.; Jarosz, D.F.; Lindquist, S. HSP90 at the hub of protein homeostasis: Emerging mechanistic insights. Nat. Rev. Mol. Cell Biol. 2010, 11, 515–528. [Google Scholar] [CrossRef]
- Miyata, Y.; Nakamoto, H.; Neckers, L. The therapeutic target Hsp90 and cancer hallmarks. Curr. Pharm. Des. 2013, 19, 347–365. [Google Scholar] [CrossRef]
- Citri, A.; Harari, D.; Shohat, G.; Ramakrishnan, P.; Gan, J.; Lavi, S.; Eisenstein, M.; Kimchi, A.; Wallach, D.; Pietrokovski, S.; et al. Hsp90 recognizes a common surface on client kinases. J. Biol. Chem. 2006, 281, 14361–14369. [Google Scholar] [CrossRef]
- Pick, E.; Kluger, Y.; Giltnane, J.M.; Moeder, C.; Camp, R.L.; Rimm, D.L.; Kluger, H.M. High HSP90 expression is associated with decreased survival in breast cancer. Cancer Res. 2007, 67, 2932–2937. [Google Scholar] [CrossRef]
- Sakanyan, V.A.; Iradyan, M.A.; Iradyan, N.S. Development of targeted EGFR degradation for cancer treatment. Nat. Acad. Sci. Armen. Rep. 2022, 122, 218–227. [Google Scholar] [CrossRef]
- Paoli, P.; Giannoni, E.; Chiarugi, P. Anoikis molecular pathways and its role in cancer progression. Biochim. Biophys. Acta 2013, 1833, 3481–3498. [Google Scholar] [CrossRef]
- Vlahakis, A.; Debnath, J. The Interconnections between autophagy and integrin-mediated cell adhesion. J. Mol. Biol. 2017, 429, 515–530. [Google Scholar] [CrossRef]
- Reginato, M.J.; Mills, K.R.; Paulus, J.K.; Lynch, D.K.; Sgroi, D.C.; Debnath, J.; Muthuswamy, S.K.; Brugge, J.S. Integrins and EGFR coordinately regulate the pro-apoptotic protein Bim to prevent anoikis. Nat. Cell Biol. 2003, 5, 733–740. [Google Scholar] [CrossRef] [PubMed]
- Chi, X.; Nguyen, D.; Pemberton, J.M.; Osterlund, E.J.; Liu, Q.; Brahmbhatt, H.; Zhang, Z.; Lin, J.; Leber, B.; Andrews, D.W. The carboxyl-terminal sequence of bim enables bax activation and killing of unprimed cells. eLife 2020, 9, e44525. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, L.; Strasser, A.; O’Reilly, L.A.; Hausmann, G.; Adams, J.M.; Cory, S.; Huang, D.C. Bim: A novel member of the Bcl-2 family that promotes apoptosis. EMBO J. 1998, 17, 384–395. [Google Scholar] [CrossRef] [PubMed]
- Wilfling, F.; Weber, A.; Potthoff, S.; Vögtle, F.N.; Meisinger, C.; Paschen, S.A.; Häcker, G. BH3-only proteins are tail-anchored in the outer mitochondrial membrane and can initiate the activation of bax. Cell Death Differ. 2012, 19, 1328–1336. [Google Scholar] [CrossRef] [PubMed]
- Gogada, R.; Yadav, N.; Liu, J.; Tang, S.; Zhang, D.; Schneider, A.; Seshadri, A.; Sun, L.; Aldaz, C.M.; Tang, D.G.; et al. Bim, a proapoptotic protein, up-regulated via transcription factor E2F1-dependent mechanism, functions as a prosurvival molecule in cancer. J. Biol. Chem. 2013, 288, 368–381. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Letai, A.; Sarosiek, K. Regulation of apoptosis in health and disease: The balancing act of BCL-2 family proteins. Nat. Rev. Mol. Cell Biol. 2019, 20, 175–193. [Google Scholar] [CrossRef]
- Tong, J.; Taylor, P.; Moran, M.F. Proteomic analysis of the epidermal growth factor receptor (EGFR) interactome and post-translational modifications associated with receptor endocytosis in response to EGF and stress. Mol. Cell. Proteom. 2014, 13, 1644–1658. [Google Scholar] [CrossRef]
- Galluzzi, L.; Vitale, I.; Aaronson, S.A.; Abrams, J.M.; Adam, D.; Agostinis, P.; Alnemri, E.S.; Altucci, L.; Amelio, I.; Andrews, D.W. Molecular mechanisms of cell death: Recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. [Google Scholar] [CrossRef]
- Tang, D.; Kang, R.; Berghe, T.V.; Vandenabeele, P.; Kroemer, G. The molecular machinery of regulated cell death. Cell Res. 2019, 29, 347–364. [Google Scholar] [CrossRef] [PubMed]
- Yan, G.; Elbadawi, M.; Efferth, T. Multiple cell death modalities and their key features (Review). World Acad. Sci. J. 2020, 2, 39–48. [Google Scholar] [CrossRef]
- Le Gall, M.; Chambard, J.C.; Breittmayer, J.P.; Grall, D.; Pouyssegur, J.; Obberghen-Schilling, E. The p42/p44 MAP kinase pathway prevents apoptosis induced by anchorage and serum removal. Mol. Biol. Cell 2000, 11, 1103–1112. [Google Scholar] [CrossRef]
- Quadros, M.R.; Peruzzi, F.; Kari, C.; Rodeck, U. Complex regulation of signal transducers and activators of transcription 3 activation in normal and malignant keratinocytes. Cancer Res. 2004, 64, 3934–3939. [Google Scholar] [CrossRef]
- Chamberlain, P.P.; Hamann, L.G. Development of targeted protein degradation therapeutics. Nat. Chem. Biol. 2019, 15, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Iradyan, M.A.; Iradyan, N.S.; Hambardzumyn, A.A.; Panosyan, H.A.; Tamazyan, R.A.; Ayvazyan, A.G.; Hovhannisyan, G.S.; Alves de sousa, R.; Sakanyan, V.A. Selective N-, S-alkylation of 4-allyl-3-[2-(4-alkoxyphenyl)-quinolin-4-yl]-4,5-dihydro-1h-1,2,4-triazole-5-thiones with substituted benzylchlorides. synthesis, docking analysis and cytotoxic action. Chem. J. Arm. 2018, 71, 389–406. [Google Scholar]
- Iradyan, M.A.; Iradyan, N.S.; Hambardzumyan, A.A.; Minasyan, N.S.; Roussakis, C.; Sakanyan, V.A. Synthesis of furfuryl derivatives of 4-allyl-1-(4-hydroxy-3-nitrobenzyl)-3-[2- (4-alkoxyphenyl)-quinolin-4-yl]-4,5-dihydro-1H-1,2,4-triazole-5-thions and their toxicity in cancer cells. Chem. J. Arm. 2018, 71, 559–570. [Google Scholar]
- Iradyan, M.A.; Iradyan, N.S.; Hambardzumyan, A.A.; Nersesyan, L.E.; Aharonyan, A.S.; Danielyan, I.S.; Muradyan, R.E.; Paronikyan, R.V.; Stepanyan, G.M. Docking analysis and some biological properties of furfuryl derivatives of 4-allyl-5-[2-(4-alkoxyphenyl) quinolin-4-yl]-4H-1,2,4-triazol-3-thiol. Biol. J. Arm. 2018, 70, 100–107. [Google Scholar]
- Available online: https://www.creative-biolabs.com/blog/protac/protac-reviews/pros-and-cons-protac-technology/ (accessed on 4 July 2023).
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sakanyan, V.; Iradyan, N.; Alves de Sousa, R. Targeted Strategies for Degradation of Key Transmembrane Proteins in Cancer. BioTech 2023, 12, 57. https://doi.org/10.3390/biotech12030057
Sakanyan V, Iradyan N, Alves de Sousa R. Targeted Strategies for Degradation of Key Transmembrane Proteins in Cancer. BioTech. 2023; 12(3):57. https://doi.org/10.3390/biotech12030057
Chicago/Turabian StyleSakanyan, Vehary, Nina Iradyan, and Rodolphe Alves de Sousa. 2023. "Targeted Strategies for Degradation of Key Transmembrane Proteins in Cancer" BioTech 12, no. 3: 57. https://doi.org/10.3390/biotech12030057
APA StyleSakanyan, V., Iradyan, N., & Alves de Sousa, R. (2023). Targeted Strategies for Degradation of Key Transmembrane Proteins in Cancer. BioTech, 12(3), 57. https://doi.org/10.3390/biotech12030057







