1. Introduction
Ocypode Weber, 1795 (Brachyura: Ocypodidae) [
1] is the popularly known ghost crab, with 21 recognized species and it is one of the most abundant crabs of the family Ocypodidae Rafinesque, 1815.
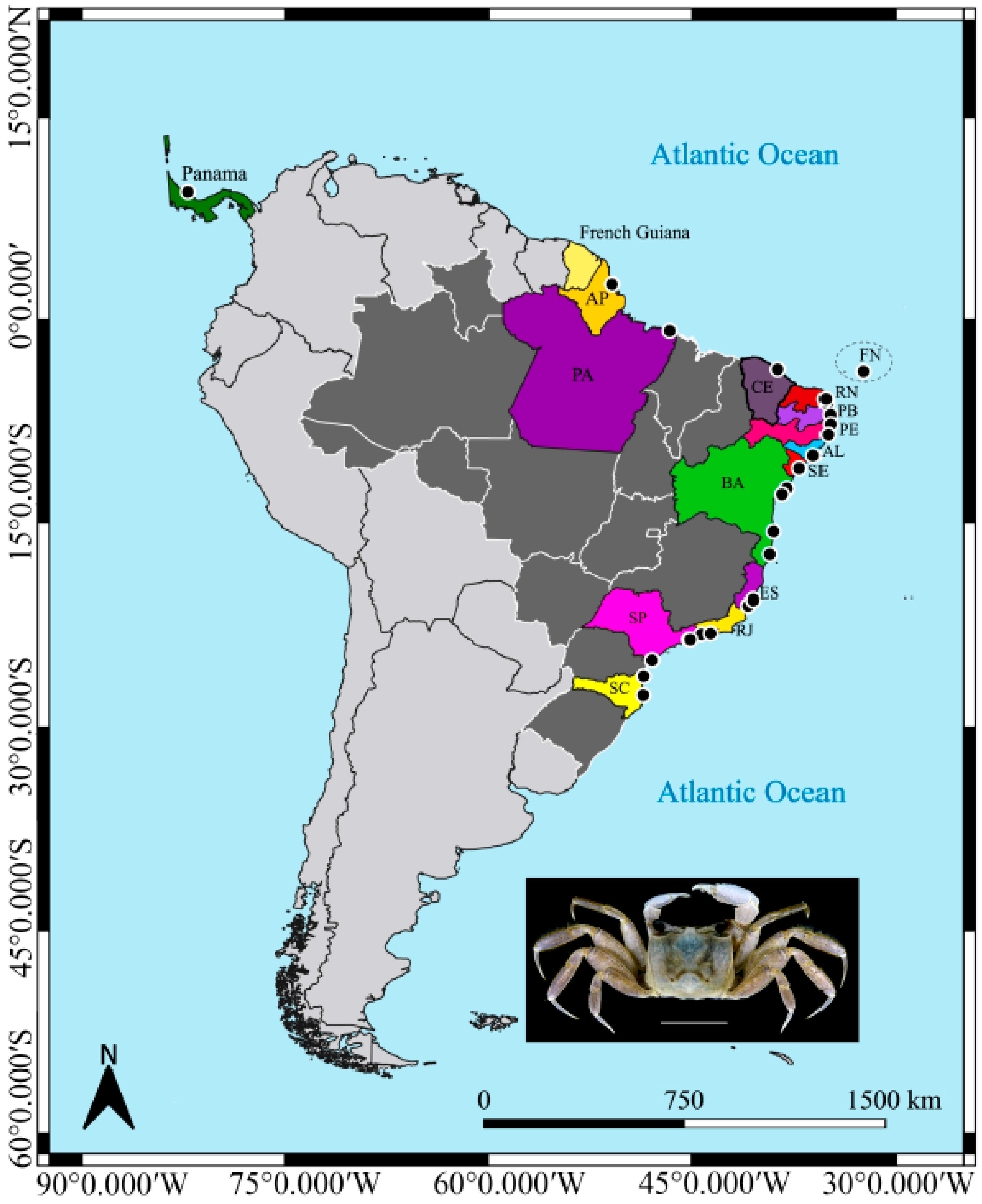
Ocypode quadrata (Fabricius, 1787) is the only species of the genus that occurs across the western Atlantic region, from the USA (Massachusetts) to Brazil (Rio Grande do Sul), Central America, and Antilles [
2,
3,
4] (
Figure 1). This species is semiterrestrial and typically inhabits the entire supralittoral zone, including dunes, sandy beaches, and vegetated areas, also being found in the upper midlittoral zone, where individual burrows are flooded during high tides [
2,
5]. This crab can provide essential information about the negative impact on beach ecosystems and contribute in efforts towards the conservation of beach biodiversity, since the threat of human impact is to have an irreversible effect on this environment [
6,
7]. The widespread distribution of
O. quadrata, along with its semiterrestrial habitat, its long planktonic larval phases, and its potential as a tool for impact on beach ecosystems, make this species an interesting example to understand different patterns of genetic differentiation, and population structure, mainly through the study of genetic variability.
The ecological characteristics of this species were widely studied [
5,
8,
9,
10,
11,
12,
13,
14], as well as its capability of survival in the intertidal areas [
9], which facilitates its collection and monitoring. Furthermore, it can be used as a bioindicator for human actions on several beaches in the Americas [
11,
15,
16].
Ocypode quadrata has planktonic larvae, five zoeal stages, and a megalopa that stays 60 days in the water column [
17,
18]; the morphology of the megalopa is probably adaptive for survival through postponed metamorphosis for more than 34 days into the first crab stage [
19]. Those larval features can facilitate high dispersal capacity, which can be related to genetic structure, as well as observed in populations that are not panmictic but are very widely dispersed [
20].
All these characteristics draw attention to
O. quadrata being a potential model for several studies, as was the case in other regional genetic population studies [
6,
21] and in systematics and taxonomy [
1]. Despite this well-developed knowledge on biology and ecology of
O. quadrata, the information on genetic variability with this taxon is still scanty. In this way, molecular analyses using the genetic barcoding can help in the characterization of the genetic variability among the populations, detecting homogeneity or structuring, and improving the knowledge of the dispersion mechanism of this species.
Studies of genetic variability and using the barcoding approach can be seen not only in different groups of Decapoda (e.g., [
22,
23,
24,
25,
26]), but also in many other organisms (e.g., [
27,
28,
29,
30]).
Some species are widely spread due to their planktonic larval characteristics and, along with potential barriers to gene flow, such as isolation by distance, sea currents, or changes in salinity caused by the outflow of the mouth of the Amazon River into the Atlantic Ocean [
31,
32,
33,
34,
35,
36], are a promising model in studies, for example, of genetic variability. Our study is going to increase the number of DNA sequences from two genes’ fragments (COI and 16S), amplifying the sample area of this species distribution, different from what has been previously studied. From this, our study tests the hypothesis of genetic structure in
O. quadrata across the western Atlantic coast of America among populations of the Caribbean Sea and Northeast and Southeast Brazil influenced by the Amazon River plume.
2. Materials and Methods
Specimens of
O. quadrata were obtained from donations and loans and are deposited in the following collections: Coleção de Crustáceos do Departamento de Biologia (CCDB), Faculdade de Filosofia, Ciências e Letras de Ribeirão Preto–Universidade de São Paulo (FFCLRP/USP), Ribeirão Preto, Brazil; Museu de Zoologia, Universidade de São Paulo (MZUSP), São Paulo, Brazil; Coleção de Crustáceos da Universidade Federal do Rio Grande do Sul (UFRGS), Porto Alegre, Brazil. The specimens examined are listed in
Table 1.
Genomic DNA was extracted from muscle samples from pereiopods with Chelex
® (Chelating Ion Exchange Resin; [
41]). The final concentration of extracted DNA was measured using a spectrophotometer (NanoDrop 2000/2000c) to calculate the amount of DNA to be used in the PCR reaction.
We used a two-locus mitochondrial approach that included partial sequences of 16S rRNA (16S) and cytochrome oxidase I (COI), which were found to be valuable in other marine decapod population studies [
23,
26,
33,
35,
36,
42]. Fragments of 600 bp (16S) and 700 bp (COI) were amplified by means of a polymerase chain reaction (PCR), using the universal primers LCO1490 (5′-GGT CAA ATC ATA AAG ATA TTG-3′) and HCO2198 (5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′) [
43], and 16Sar (5′-CGC CTG TTT ATC AAA AAC AT-3′) and 16Sbr (5′-CCG GTC TGA ACT CAG ATC ACG T-3′) [
44]. PCR procedures followed [
45,
46,
47] with adjustments in annealing temperatures according to the specific primers, decreasing until a 5 °C melting temperature. PCRs were performed in a total volume of 25 μL, which included 4.5 μL of Millipore-filtered water, 5 μL of betaine (5 M), 4 μL of dNTP (10 μM), 3 μL of MgCl
2 (25 μM), 3 μL of a 10× PCR Taq buffer with (NH
4)
2SO
4, 1 μL of each primer (10 μM for 16S and 20 μM for COI), 0.5 μL of 1.5 U Taq DNA polymerase. We followed the thermal cycles of initial denaturing for 4 min at 94 °C; pairing for 40 cycles (50 s at 94 °C, 70 s at 45/48 °C, and 1 min at 72 °C); final extension for 10 min at 72 °C [
1]. PCR products were purified using a Sure Clean Plus
® kit (Bioline, Tauton, MA, USA) and were sequenced with the ABI 3730 XL DNA Analyzer
® (Applied Biosystems, 850 Lincoln Centre Drive, Foster City, CA 94404, USA).
All sequences were confirmed by sequencing forward and reverse strands. The consensus sequence for both strands was obtained using the computational program Geneious v.2021.2 [
48]. Sequences were aligned in MAFFT v.7 [
49] using default parameters. We used the program GBlocks v.091b [
50,
51] to locate and exclude ambiguous areas of the alignment or each locus, using relaxed gap selection criteria (allowed gap positions = all). Alignments were concatenated in the software Geneious v.2021. 2. All sequences were submitted to GenBank under specific accession numbers (COI: OR354404-OR354434; 16S: OR352135-OR352150; see
Table 1). Genetic vouchers used in molecular analyses are deposited in the collection of the CCDB/FFCLRP/USP or at the institution of origin.
Samples were grouped according to geographical distribution in Brazilian States, and countries for those localities with fewer individuals (USA, Mexico, Panama, and French Guiana). Genetic distances were calculated to determine intra- and interspecific variation rates with the software MEGA v. 11.0.10 [
52], using the p-distance method; distance values are in percentages.
Phylogenetic hypotheses were proposed using Maximum Likelihood (ML) [
53] performed on the W-IQ-TREE online platform using models selected by the program based on the data provided [
54]. For the concatenate ML tree, we chose just specimens with both DNA fragments (16S and COI), since the other samples have just one sequence, which would configure a lot of missing data (
Table 1); this analysis aimed to contextualize
O. quadrata within the genus. The best-fit models according to BIC were GTR+F+I+G4 for concatenate genes; branch support was evaluated by ultrafast bootstrapping (1000 pseudoreplicates); bootstrap values > 60% are shown in all trees.
GenBank sequences from
Ocypode africana De Man, 1881;
Ocypode ceratophthalmus (Pallas, 1772);
Ocypode macrocera H. Milne Edwards, 1837;
Ocypode nobilii De Man, 1902;
Ocypode cordimana Latreille, 1818;
Ocypode kuhlii De Haan, 1835;
Ocypode rotundata Miers, 1882;
Ocypode ryderi Kingsley, 1880;
Ocypode sinensis Dai & Yang in Song & Yang, 1985;
Ocypode stimpisoni Ortmann, 1897;
Ocypode occidentalis Stimpson, 1860; and
Ocypode gaudichaudii H. Milne Edwards & Lucas, 1843 were analyzed as sister groups and
Afruca tangeri (Eydoux, 1835) as an outgroup (
Table 1), following the phylogeny proposal for Ocypodidae [
1].
Population parameters of genetic variability within each locality are represented by the number of haplotypes (h), segregating sites (S), haplotype diversity (Hd), nucleotide diversity (π), and average number of nucleotide differences (k), estimated in DnaSP 6.12.1 [
55]. The haplotype networks were constructed with PopART v. 4.8.4 [
56] using the statistical parsimony method TCS Network [
57]. A linear cross-mark and the total number of mutational steps represent the links among lineages; the number in the circle represents the frequency of each haplotype, and the smallest circles indicate only one haplotype. The black dots show missing haplotypes and the connections among haplotypes indicate one mutation step. The black dots or the average vectors are considered hypothetical haplotypes (sequences), generated by the program to connect the sampled haplotypes [
58]. The analysis of molecular variance (AMOVA) and the pairwise fixation index (F
ST) were evaluated in Arlequin 3.5 [
59], with 10,000 permutations, to calculate the variance within localities, and between Brazilian States, the USA, Mexico, Panama, and French Guiana.
The neutrality tests Tajima’s D [
60] and Fu’s F
s [
61] were performed in DnaSP 6.12.1 [
55] to explain observed patterns in genetic variation within populations of
O. quadrata considering the interaction between genetic drift and mutation, including other parameters like nucleotide heterozygosity and the number of segregating sites in a DNA sequences’ dataset; these tests are useful to detect whether mutations were neutral or under the influence of selection, and also for detecting population growth. In addition, pairwise mismatch distribution was analyzed to test occurrence of contraction or population expansion [
62], assay demographic expansion, and detect stability. The sample distribution of pairwise differences will usually deviate from the distribution expected. From this, a variety of shapes can be found, including bimodal and trimodal distributions; this wide variety of distributions is correlated to the pairwise differences and can be because of the history of coalescent events in a single sample of genes [
63]. Furthermore, if the observed distribution of pairwise differences is close to a Poisson distribution, that means it is consistent with the hypothesis that the population has been growing exponentially in size [
63]; the continued exponential growth suggests a sudden burst of population growth [
62]. If the population decreases in size, distributions are initially L-shaped, and then converge quickly to an equilibrium; if the populations are in equilibrium, the theoretical curves are free of waves [
62], or a population that has been growing presents mismatch distributions that are smooth and have a peak [
64].
3. Results
The present study contributes with 50 new sequences of
O. quadrata, 16 sequences from 16S and 34 from COI, another 86 sequences (74 from 16S, 12 for COI) from the same species taken from GenBank, and 24 sequences from other species of the genus (
Table 1).
The final alignment of the COI fragment consists of 636 base pairs (bp) and the alignment of the 16S rRNA fragment consists of 511 bp. The intraspecific divergence for
Ocypode quadrata varied from 0 to 13.9% and 0 to 6.4% for COI and 16S, respectively, whereas interspecific values ranged from 12.1 to 21.7% for COI (
Table S1) and 6.8 to 14.0% for 16S (
Table S2), with these values corresponding to the divergence between sequences of
O. africana and
O. ceratophthalmus. Genetic distances among localities ranged from 0.4 to 19.7% (
Table 2) for COI and 0.0 to 6.4% (
Table 3) for 16S.
The concatenated phylogram suggested the monophyly of
O. quadrata with bootstrap values of 100 (
Figure 2), with an external branch consisting of a specimen from Rio Grande do Norte, and a subclade with specimens from different localities (bootstrap: 78%), configuring no pattern dividing groups that could reveal genetic structure.
The genetic variability of
Ocypode quadrata could be evaluated by the number of segregating sites (S) with 121 and 33 for COI and 16S, respectively; a nucleotide diversity (π) of 0.0373 (COI) and 0.01108 (16S), with k values of 20.853 (COI) and 5.243 (16S); and haplotype diversity (Hd) of 0.9707 for COI and 0.6014 for 16S (
Table 4).
For the COI fragment (34 new sequences, 12 from GenBank), the number of haplotypes (Hap) was 33, with 1 highly frequent haplotype (Hap_9) with seven individuals from four different localities, linked with several unique haplotypes (Hap_10, Hap_17, Hap_18, Hap_20; Hap_26; Hap_27), and 2 less frequent haplotypes (Hap_12, Hap_22). One individual from the USA (Hap_1) with other Mexican haplotypes were separated by more than 20 mutation steps from the Brazilian haplotypes (
Figure 3).
For the 16S fragment (16 new sequences, 74 from GenBank), the number of haplotypes was 24; the most common was Hap_2 with 56 specimens from 14 localities, linked with 11 unique haplotypes (Hap_3, Hap_6, Hap_7, Hap_10, Hap_11, Hap_13, Hap_16, Hap_20, Hap_22, Hap_23, Hap_24), and 4 were less frequent (Hap_4, Hap_9, Hap_12, Hap_14, Hap_18). One individual from the USA was separated by 17 mutation steps from three Brazilian haplotypes (Hap_4, Hap_14, Hap_18) (
Figure 4). Within Brazilian States, among Brazil, Panama, and French Guiana, both haplotype networks for
O. quadrata fit into the star-shaped pattern, suggesting a recent demographic expansion, with no apparent population genetic structure; mainly for the COI gene, the USA and Mexico populations may be a new lineage.
AMOVA results also did not detect any genetic structure among populations (within Brazilian States; among Brazil, Panama, and French Guiana), with
p = 0.00168 for the COI fragment, and
p = 0.21465 for 16S (
Table 5). Most of the F
ST values were not significant, with a few exceptions of the comparison between localities (
Table 6). The result of the neutrality test Tajima’s D was −0.69521 and Fu’s Fs was −3.770, which were not significant (
p > 0.10) for the COI gene; and Tajima’s D was −2.45254 and the Fu’s Fs statistic was −25.390, which were significant (
p < 0.001), for 16S. For the COI fragment, evolution in
O. quadrata occurs by genetic drift and mutation, emphasizing the lack of evidence of selection for these populations. For 16S, these significant values demonstrated that populations of
O. quadrata do not reach the equilibrium, for example, if they experienced a bottleneck recently [
60].
Mismatch distribution graphics revealed multimodal distributions, which means it deviated from the expected, and pronounced waves with rough crests, which suggest coalescent events and ancient population explosion. Populations possibly passed through contraction and expansion events, which is consistent with the neutrality test (
Figure 5A,B), and a third peak suggests multiple genetic clusters (
Figure 5A), such as the USA plus Mexico separated from the other localities.
4. Discussion
In the present study, the initial hypothesis of genetic structure in
O. quadrata, supported by its wide geographic distribution, was refuted. Other marine Decapoda presented the same pattern, lack of genetic structure, for short and long distances such as the crab
Ucides cordatus Linnaeus, 1763 [
42]; the mangrove crab
Sesarma rectum Randall, 1840 [
33]; the swimming crab
Callinectes danae Smith, 1869 [
35,
36]; the western Atlantic hermit crab
Clibanarius antillensis Stimpson, 1859 [
26]; and the shrimp
Artemesia longinaris [
23]. Therefore, the wide distribution cannot be considered as a predictive character for this condition in marine organisms. Furthermore, since the barriers to gene flow in these environments influence each organism in a singular way, patterns of genetic structure are not easily established [
65,
66]. The USA and Mexican specimens presented higher values of genetic distances and many mutation steps at the haplotype networks, suggesting signs of a new lineage. Like those observed herein, for the orange claw hermit crab
Calcinus tibicen (Herbst, 1791), results suggested two genetically well-defined groups, North and South Atlantic, but no morphological pattern for each genetic group was observed [
34].
There were low intraspecific genetic distances when comparing interspecific values between
O. quadrata from Brazilian, Panama, and French Guiana specimens, giving evidence of a lack of structuring, as well as in other marine crabs, like
Ocypode species, in which the intraspecific variation for homogeneous populations was around 0.52 ± 0.11%, whereas the values between populations of other genetically structured species were 10.14 ± 0.46% [
67]. Considering that individuals of the same species are not identical to each other and may present transversions or random transitions in their genetic material, as well as being subject to environmental factors with potential for phenotypic changes [
68], we can infer that this population group of
O. quadrata characterizes a lack of genetic structure, since such minimal variation will be present even in species with constant gene flow. Furthermore, we suggest that genetic variability in Brazil seems to be high and populations are diverse, as observed in the different unique haplotypes. In addition, as previously suggested [
6], the USA and Mexican specimens are separated from the other localities and may present a new lineage. However, we cannot infer with a few sequences and without observing morphological characters.
Results among distinct localities within Brazil along with Panama and French Guiana also suggested the absence of genetic separation, evidenced by low genetic distance values, and by the polytomies in the phylogenetic tree and the star-shaped haplotype network, like those observed for other populations of
O. quadrata [
6,
21]. Herein, it is plausible to infer that the absence of a geographic pattern in the composition of the formed subgroups is a good indicator of high panmixia, which is a relevant fact, since the pattern found in most living species tends to be the structuring at some level for distant geographical localities [
25,
69,
70]. Considering the broad extension of the Brazilian coast, it is possible that these crabs have a high dispersion capacity, facilitating gene flow even to distant populations, including the populations of Panama and French Guiana, maintaining genetic homogeneity. This dispersion may be attributed to larval characteristics, as five zoeal stages and a megalopa, and their long period of permanence in the plankton [
17,
18,
19].
Different from the other localities, the individual from the USA together with the Mexican population were separated by many mutation steps from the Brazilian with Panama and French Guiana populations, possibly representing a distinct lineage. This aspect has already been subtly pointed out, suggesting an existing barrier to gene flow in the tropical–temperate transition between the Caribbean and the east coast of the USA [
6]. The megalopae of
O. quadrata have been reported ≈190 km beyond any adult populations in the USA coast, and can tolerate temperature fluctuations, which means that megalopae can migrate for long distances along the coast and might survive long enough to burrow into a sandy beach [
71]. In the present study, these characteristics can explain the separation of the USA and Mexico populations from the Brazilian and the other populations. Furthermore, we suggested that the sea currents of the North Atlantic Current and Gulf Stream [
72] are influencing the larval dispersion of
O. quadrata in long distances and can be responsible for such northern and southern distinct groups. In the present study, we expanded the number of specimens and localities, mainly in Brazil, which increase the number of mutation steps and reinforce the idea of a distinct entity. However, although outside the scope of this study, it would be interesting to analyze morphological characters of USA specimens along with the taxonomical history of
O. quadrata and related species.
In
Ocypode quadrata, the nucleotide diversity and the haplotype diversity were like other brachyuran species, including the congener species
O. ceratophthalmus (Pallas, 1772), which also did not show significant genetic variation among the Japanese populations, but showed a slight difference between the Philippine populations, separated by 30 to 1200 km [
67], and three genetically distinct geographical groups, corresponding to the east, west, and central region of the species’ range in the Indo-Pacific region [
73]. This small difference between populations is possibly a result of recent population expansion or dispersion [
66,
67], along with gene flow.
Regarding population size changes, our results indicated that the population of
O. quadrata from Brazil, Panama, and French Guina did not show recent abrupt expansions or a bottleneck, which is corroborated by the genetic balance between these populations because of the same demographic balance [
74,
75], which can also be noted in neutrality data and the star-like haplotype network. Another possibility, a star-like haplotype network with a great quantity of rare haplotypes, could also suggest an insufficient sampling of intraspecific genetic variation since
O. quadrata has a wide distribution. Thus, no demographic change was drastic enough to promote lineage divergence between Brazilian, Panama, and French Guiana populations.
Possibly, USA and Mexican populations have been exposed to events of strong selective pressure and/or genetic drift, changing their size abruptly, which makes certain individuals more efficient in transmitting their genes than others [
62,
76]. Genetic structure and many mutational steps were also observed for
Ilyoplax pusilla (De Haan, 1835) [
77] and in the swimming crab
Callinectes ornatus Ordway, 1863 [
78]. These steps indicate modifications by transversions or transitions in the nitrogenous bases, in relation to the previous haplotype, increasing the genetic distance between them. Along the generations, this process can facilitate the genetic structure within populations and posteriorly, the speciation [
68]. A population expansion can be indicated by the smoothness of the mismatch distribution, which is not affected by population structure, whereas mean sequence divergence increases in a pooled sample from highly isolated subpopulations [
64]. The increase in sequence divergence may be the case of the USA and Mexican populations. Different from
O. quadrata, the unimodal distribution observed in the mismatch distributions’ graphics was related to demographic and spatial expansion, suggesting a recent bottleneck effect or sudden population expansion for the crab species
S. rectum [
33], the occurrence of a recent demographic expansion during the evolutionary history for another crab,
U. cordatus [
15], and demographic expansion for the hermit crab
C. antillensis [
26]. As well as in the present study for populations of
O. quadrata from the USA and Mexico, the haplotype network and long branches in the phylogram indicate some groups that could present haplotypic structure [
6,
77], despite the low number of mutational steps. However, the low number of mutational steps within Brazilian populations of
O. quadrata was attributed to non-exclusive continuous gene flow and/or genetic variations to each individual, in which some subgroups do not necessarily indicate a genetic separation.
Possible explanations for the high gene flow between populations of geographically distant organisms can be related to the type of occupied habitat [
79], its ability to disperse [
80,
81], and dependency of the life-history strategies [
82]. In marine animals, water temperature is an important variable that is influenced by the currents and that can even determine their survival and subsequent reproductive success of individuals in its respective environment. The semiterrestrial
Ocypode quadrata is exposed to temperatures that vary between about 16 °C in the north of the United States, 25 °C in the north of Brazil, and 20 °C in the south [
18,
83]. At temperatures lower than 16 °C, the organism can survive, but in an inactive state, and can remain in its sealed burrow for almost three months of the year [
18], which may have an influence on its wide distribution.
As individuals of
O. quadrata inhabit supralittoral and intertidal zones, their ability to disperse is mainly attributed to larvae [
84]. There are six larval stages, five zoea and a megalopa, that stay in the water column for 60 days [
17,
18]. It is also described as a unique characteristic of this group, when in the early stages of development, that the megalopa can regulate its development cycle and postpone the metamorphosis until 34 days [
19], which further increases its dispersive success, preventing maturation/settlement in not-favorable environments. Those larval features allow us to attribute a high dispersal capacity to the larvae, which explain the lack of genetic structuring in this species. It is like those observed, along the Brazilian coast, in the fiddler crab,
Uca maracoani (Latreille, 1803), a semiterrestrial coastal species with an extensive marine pelagic larval duration [
66,
85];
Minuca panema (Coelho, 1972) [
86], and the eastern Pacific
Emerita analoga (Stimpson, 1857), an intertidal sandy beach crab with an especially wide latitudinal distribution and a long pelagic larval phase, whose populations are not panmictic but are very widely dispersed and approaching genetic homogeneity [
20].
Marine currents may also be facilitators of the dispersion of
O. quadrata from southern Brazil to North America. In Brazil, the three main currents are the North Brazil Current (NBC), the Brazil Current, and the Malvinas Current [
87], the first two being cold and the last one warm. The NBC is the strongest one, allowing the dispersion of larvae from Rio Grande Norte (Brazil) to Central America [
88]. Similar examples were observed for other marine decapods along the western Atlantic, including the tiny shrimp
Hippolyte obliquimanus Dana, 1852 [
32]; the fiddler crab
Minuca rapax (Smith, 1870) (as
Uca rapax) [
89]; and the orange claw hermit crab
C. tibicen [
34].
Alternatively, the long-distance larval dispersal can be a consequence of the so-called steppingstone process, which is common for distinct animals, including marine ones [
33,
90], and can be applicable in
O. quadrata. It consists of the connection between fragments of habitats by a given species, enabling the connection between distant regions, providing gene flow from the occupation of these fragments and subsequent dispersal [
90,
91].