-
 Vape-Associated lncRNA Transcript 1 (VALT1) Amplifies the Tumorigenic Effects of e-Cigarette Vapor in Lung Epithelial Cells
Vape-Associated lncRNA Transcript 1 (VALT1) Amplifies the Tumorigenic Effects of e-Cigarette Vapor in Lung Epithelial Cells -
 The Stiff Side of Cancer: How Matrix Mechanics Rewrites Non-Coding RNA Expression Programs
The Stiff Side of Cancer: How Matrix Mechanics Rewrites Non-Coding RNA Expression Programs -
 Comprehensive Schistosoma mansoni Hierarchical Transcriptome Assembly Points to Novel lncRNAs Associated with Sexual Dimorphis
Comprehensive Schistosoma mansoni Hierarchical Transcriptome Assembly Points to Novel lncRNAs Associated with Sexual Dimorphis
Journal Description
Non-Coding RNA
Non-Coding RNA
is an international, peer-reviewed, open access journal on non-coding RNA research dealing with elucidating the structure, function and biology of regulatory non-coding RNAs, and published bimonthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q2 (Genetics and Heredity) / CiteScore - Q1 (Genetics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 34.5 days after submission; acceptance to publication is undertaken in 3.9 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.0 (2024);
5-Year Impact Factor:
4.2 (2024)
Latest Articles
Exploratory Machine Learning Analysis of circRNA-Derived Molecular Features in Autism Spectrum Disorder
Non-Coding RNA 2026, 12(3), 17; https://doi.org/10.3390/ncrna12030017 - 15 May 2026
Abstract
►
Show Figures
Background/Objectives: Autism Spectrum Disorder (ASD) is a set of neurological and neurodevelopmental disorders characterized by difficulties in social communication and interaction, repetitive behaviors, and sensory processing differences. Recent studies have shown that circRNAs play a crucial role in the pathophysiology of ASD. In
[...] Read more.
Background/Objectives: Autism Spectrum Disorder (ASD) is a set of neurological and neurodevelopmental disorders characterized by difficulties in social communication and interaction, repetitive behaviors, and sensory processing differences. Recent studies have shown that circRNAs play a crucial role in the pathophysiology of ASD. In this study, we present an exploratory machine learning framework integrating circRNA sequence features, miRNA interactions, gene targets, and pathway enrichment analysis to investigate ASD-associated molecular signatures. Methods: Differential circRNAs were identified from human peripheral blood datasets, and informative features were selected using attribute-based filtering and Information Gain ranking. Machine learning models were developed using the WEKA platform. Results: The HyperPipes classifier achieved the highest performance (92.5% accuracy under cross-validation). Analysis using an independent ASD gene expression dataset showed consistent discriminative patterns of the derived gene-level signatures across multiple machine learning classifiers. The competitive endogenous RNA network and enriched gene pathways were also analyzed. Conclusions: Overall, this study provides a computational, preliminary framework for analyzing circRNA-associated molecular patterns in ASD. Findings should be interpreted in the context of limited sample size and dataset availability.
Full article
Open AccessArticle
Urinary Exosomal microRNAs as a Novel Approach to Study People with Multiple Sclerosis and Severe Gait Disability: A Preliminary Observation
by
Silvia Grassilli, Andrea Baroni, Marina Pierantoni, Federica Brugnoli, Nicola Lamberti, Sofia Straudi, Fabio Manfredini and Valeria Bertagnolo
Non-Coding RNA 2026, 12(3), 16; https://doi.org/10.3390/ncrna12030016 - 8 May 2026
Abstract
Background: MiRNAs within extracellular vesicles can encompass body barriers, reflecting stage, progression, and response to treatments of various diseases, including multiple sclerosis (MS)—a chronic immune-mediated disease of the central nervous system that causes progressive disability, with highly variable clinical courses. In this context,
[...] Read more.
Background: MiRNAs within extracellular vesicles can encompass body barriers, reflecting stage, progression, and response to treatments of various diseases, including multiple sclerosis (MS)—a chronic immune-mediated disease of the central nervous system that causes progressive disability, with highly variable clinical courses. In this context, urinary exosomal miRNAs could be an appealing source of biomarkers, thanks to their non-invasive and easily repeatable collection. Methods: In this exploratory investigation, we tried to assess if profiling urinary exosomal miRNAs could reveal subtle differences within an apparently homogeneous MS population. The study involved 24 patients with primary or secondary progressive MS, whose urinary exosomes (UEs) were subjected to evaluation of a panel of 87 miRNAs variously correlated with neuroinflammation, cardiovascular functions, and/or involved in MS. Results: We revealed that the examined miRNAs were heterogeneously expressed across the patients, reflecting, as expected, their gender and/or hormonal status. Two miRNAs discriminated against primary or secondary progressive MS, and a panel of 14 commonly upmodulated miRNAs identified patients with longer disease duration and a greater degree of disability. Conclusions: Even if preliminary, these data represent the first relationship between UEs and MS features in humans and suggest that urine could constitute a non-invasive source of exosomal miRNAs, which could prove useful in complementing conventional monitoring to provide a more personalized management of MS patients.
Full article
(This article belongs to the Special Issue ncRNAs in Human Diseases and Therapeutics)
►▼
Show Figures

Figure 1
Open AccessArticle
Hnf1aos1 as a Metabolic Coordinator of Hepatic Lipid Homeostasis and Feedback Control
by
Beshoy Armanios, Jing Jin, Ankit P. Laddha, Le Tra Giang Nguyen, Sherouk M. Tawfik, Neha Mishra, Jose E. Manautou and Xiao-Bo Zhong
Non-Coding RNA 2026, 12(3), 15; https://doi.org/10.3390/ncrna12030015 - 30 Apr 2026
Abstract
►▼
Show Figures
Background: Long noncoding RNAs (lncRNAs) have emerged as critical regulators of hepatic metabolism and disease progression. The hepatocyte nuclear factor 1 alpha antisense 1 (HNF1A-AS1) lncRNA modulates liver-specific transcription factors; however, its physiological role in diet-dependent lipid homeostasis remains poorly defined. Methods: In
[...] Read more.
Background: Long noncoding RNAs (lncRNAs) have emerged as critical regulators of hepatic metabolism and disease progression. The hepatocyte nuclear factor 1 alpha antisense 1 (HNF1A-AS1) lncRNA modulates liver-specific transcription factors; however, its physiological role in diet-dependent lipid homeostasis remains poorly defined. Methods: In this study, we investigated the mouse ortholog, Hnf1a opposite strand 1 (Hnf1aos1), using AAV-mediated knockdown in C57BL/6J mice fed either a chow diet (10% kcal from fat) or a high-fat diet (HFD; 60% kcal from fat) for 12 weeks. Metabolic phenotyping included hepatic lipid quantification, histological analysis, serum biochemistry, and quantitative gene expression profiling. Results: Loss of Hnf1aos1 produced distinct, diet-dependent alterations in hepatic lipid handling. Under chow conditions, knockdown mice exhibited selective hepatic cholesterol accumulation (6.10 ± 2.9 mg/g tissue vs. 3.51 ± 1.1 mg/g in controls), accompanied by dysregulation of cholesterol clearance pathways. In contrast, under HFD conditions, knockdown precipitated severe macrovesicular degeneration, with hepatic triglyceride levels approximately doubled relative to HFD-fed controls (51.72 ± 19.8 mg/g vs. 26.34 ± 11.9 mg/g) and a numerically elevated triglyceride-to-cholesterol ratio (TG:TC ≈ 6.1:1; p = 0.0621, trend). Chow/Kd mice gained significantly less weight than chow-fed controls, whereas HFD/Kd mice exhibited weight gain comparable to HFD controls despite severe hepatic steatosis. This paradoxical phenotype suggests impaired metabolic feedback at the post-transcriptional level, in which compensatory upregulation of Hnf1a mRNA is insufficient to suppress lipid-associated genes such as Cd36, despite profound lipid overload; however, HNF1A protein levels were not directly measured in this study. Conclusion: Collectively, these findings identify Hnf1aos1 as a regulator of hepatic lipid homeostasis whose loss produces a phenotype consistent with inappropriate lipid accumulation during nutrient excess, without defining the underlying molecular mechanism. Our results support a role for Hnf1aos1 in shaping hepatic metabolic plasticity and provide insight into lncRNA-associated MASLD phenotypes.
Full article

Figure 1
Open AccessReview
Circular RNAs in Intestinal Mucosal Homeostasis and Pathologies
by
Hee Kyoung Chung, Lan Xiao and Jian-Ying Wang
Non-Coding RNA 2026, 12(3), 14; https://doi.org/10.3390/ncrna12030014 - 30 Apr 2026
Abstract
The mammalian intestinal epithelium is a rapid self-renewal tissue in the body, serving as a critical interface between the host and the external environment. Maintaining the intestinal epithelium homeostasis requires precise coordination of cellular processes, including proliferation, migration, differentiation, autophagy, and cell-to-cell interaction.
[...] Read more.
The mammalian intestinal epithelium is a rapid self-renewal tissue in the body, serving as a critical interface between the host and the external environment. Maintaining the intestinal epithelium homeostasis requires precise coordination of cellular processes, including proliferation, migration, differentiation, autophagy, and cell-to-cell interaction. An increasing body of evidence has unveiled circular RNAs (circRNAs) as abundant and stable regulatory molecules that play pivotal roles in the intestinal epithelial biology and are intimately involved in many aspects of gut mucosal pathologies. Unlike linear RNAs, circRNAs form covalently closed loop structures through back-splicing events, conferring remarkable stability and resistance to exonucleolytic degradation. circRNAs regulate the growth of the intestinal mucosa, injury-induced epithelial regeneration, and gut barrier function via diverse mechanisms, including interactions with microRNAs and RNA-binding proteins. Deregulated circRNAs are implicated in the pathogenesis of various gut mucosal disorders such as inflammatory bowel disease and malignancies. In this review, we highlight pathobiological functions and mechanisms of intestinal epithelium-enriched circRNAs, particularly circHIPK3, Cdrlas, and circPABPN1, in the epithelium homeostasis and pathologies and also discuss potential clinical application of circRNAs as diagnostic biomarkers and therapeutic targets in patients with critical diseases.
Full article
(This article belongs to the Special Issue Non-Coding RNA: 10th Anniversary)
►▼
Show Figures

Figure 1
Open AccessReview
Neuropathic Pain: Mapping the miRNA Landscape
by
Mario García-Domínguez
Non-Coding RNA 2026, 12(2), 13; https://doi.org/10.3390/ncrna12020013 - 6 Apr 2026
Abstract
►▼
Show Figures
Neuropathic pain represents a complex, prolonged pain state arising from lesions within the somatosensory nervous system. Despite significant advances in elucidating its pathophysiology, current therapeutic approaches remain largely symptomatic and frequently inadequate. MicroRNAs, a class of small non-coding RNAs that regulate gene expression
[...] Read more.
Neuropathic pain represents a complex, prolonged pain state arising from lesions within the somatosensory nervous system. Despite significant advances in elucidating its pathophysiology, current therapeutic approaches remain largely symptomatic and frequently inadequate. MicroRNAs, a class of small non-coding RNAs that regulate gene expression post-transcriptionally, have recently emerged as critical modulators of neuronal excitability, neuroinflammation, and synaptic plasticity, which are crucial processes in the development and maintenance of neuropathic pain. This review summarizes the current evidence linking specific miRNAs to the onset and maintenance of neuropathic pain, with an emphasis on their roles in peripheral and central sensitization. The potential of miRNA-based biomarkers for diagnosis and prognostic evaluation is also highlighted. A thorough understanding of the complex miRNA regulatory networks underlying neuropathic pain could facilitate the development of novel, mechanism-based therapies and ultimately improve clinical outcomes.
Full article

Figure 1
Open AccessArticle
Paraspeckles Are Associated with the Activation and Nuclear Localization of Unphosphorylated miR-34a
by
Graham H. Read, Kristen McGreevy, Hanny Issawi, Tiffany Yang, Cynthia Tsang, Ihsan A. Turk, Emily Rietdorf, Whitaker Cohn, David W. Salzman, Julian P. Whitelegge and Joanne B. Weidhaas
Non-Coding RNA 2026, 12(2), 12; https://doi.org/10.3390/ncrna12020012 - 31 Mar 2026
Abstract
Background/Objectives: Canonical microRNAs possess a 5′ phosphate required for Argonaute binding and activity. However, prior work identified an unphosphorylated, inactive nuclear pool of the important radiation-responsive microRNA, miR-34, that is rapidly phosphorylated and activated in response to ionizing radiation (IR). Here, we extend
[...] Read more.
Background/Objectives: Canonical microRNAs possess a 5′ phosphate required for Argonaute binding and activity. However, prior work identified an unphosphorylated, inactive nuclear pool of the important radiation-responsive microRNA, miR-34, that is rapidly phosphorylated and activated in response to ionizing radiation (IR). Here, we extend this work and investigate the role of paraspeckles, a phase-separated nuclear sub-compartment, and their association with the localization of unphosphorylated miR-34a. Methods: Mass spectrometry was performed to identify interacting partners of unphosphorylated mir-34. CRISPR-mediated deletion of the paraspeckle NEAT1_2 triple helix motif was performed to create an A549 cell line lacking paraspeckles (dTH). Activity and expression of mir-34a post-irradiation were evaluated by qRT-PCR and luciferase assays comparing dTH and wild-type (WT) A549 cell lines. In situ hybridization (ISH) was performed to evaluate mir-34a localization before and after IR, comparing dTH and WT cell lines. Results: Mass spectrometry identified paraspeckle proteins as significantly enriched interacting partners of unphosphorylated mir-34 mimics. By qRT-PCR and luciferase assays, we found that paraspeckle loss prevented radiation-induced early activation of unphosphorylated mir-34a. We found no difference in radiation-induced transcription of pri-miR-34a, but early processing to pre-miR-34a appeared delayed. ISH confirmed that loss of paraspeckles altered the nuclear localization of miR-34a before and after IR. Conclusions: These data suggest that paraspeckles are associated with nuclear localization and early radiation-responsive activation of unphosphorylated miR-34a. This suggests a coordinated nuclear sequestration of this important miR in its unphosphorylated state to enable an enhanced radiation response.
Full article
(This article belongs to the Section RNA Modifications)
►▼
Show Figures
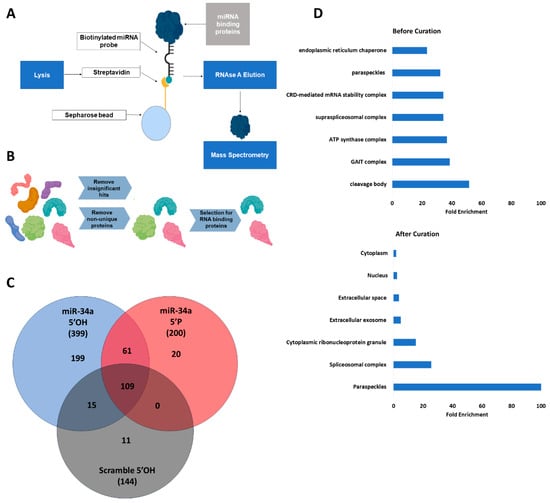
Figure 1
Open AccessReview
Circular and Long Non-Coding RNAs in Cancer Metabolism: Dual Perspective of Biomarkers and Therapeutic Targets
by
Francesca Pia Carbone, Stefania Hanau and Nicoletta Bianchi
Non-Coding RNA 2026, 12(2), 11; https://doi.org/10.3390/ncrna12020011 - 19 Mar 2026
Abstract
Background/Objectives: Metabolic reprogramming is a hallmark of cancer, enabling tumor cells to sustain proliferation, survive under metabolic stress, and develop therapeutic resistance. While oncogenic signaling pathways regulating cancer metabolism have been extensively studied, increasing evidence indicates that non-coding RNAs (ncRNAs) play essential
[...] Read more.
Background/Objectives: Metabolic reprogramming is a hallmark of cancer, enabling tumor cells to sustain proliferation, survive under metabolic stress, and develop therapeutic resistance. While oncogenic signaling pathways regulating cancer metabolism have been extensively studied, increasing evidence indicates that non-coding RNAs (ncRNAs) play essential roles in coordinating metabolic adaptation. This review aims to synthesize current knowledge on long non-coding RNAs (lncRNAs) and circular RNAs (circRNAs) as important but relatively less characterized regulators of cancer metabolic adaptation and discuss their potential as biomarkers and therapeutic targets. Methods: We analyzed their roles across multiple types of cancer, prioritizing studies that integrate ncRNA profiling with metabolomics and mechanistic investigations, with particular attention to their diagnostic, prognostic, and predictive value. Results: LncRNAs and circRNAs regulate major metabolic pathways, including glycolysis, mitochondrial function, glutaminolysis, lipid metabolism, and redox balance. They act through transcriptional and epigenetic mechanisms, protein scaffolding, peptide encoding, and miRNA sponging, frequently converging on key regulators such as HIF-1α, c-Myc, p53, AMPK, and mTOR. However, many reported associations remain largely correlative, with limited integration of quantitative metabolic flux analyses and insufficient validation in physiologically relevant models. Conclusions: Although lncRNAs and circRNAs constitute an important context-dependent regulatory layer linking oncogenic signaling to metabolic reprogramming, future studies should combine ncRNA perturbation with stable isotope tracing, fluxomics, spatial metabolomics, long-read sequencing, and single-cell approaches to define causal and spatially resolved metabolic functions. Such integrative strategies may improve biomarker development and support ncRNA-informed, metabolism-oriented therapeutic interventions.
Full article
(This article belongs to the Special Issue Non-coding RNA as Biomarker in Cancer)
►▼
Show Figures

Figure 1
Open AccessArticle
Vape-Associated lncRNA Transcript 1 (VALT1) Amplifies the Tumorigenic Effects of e-Cigarette Vapor in Lung Epithelial Cells
by
Daniel Angelo R. Mirador, Jose Lorenzo M. Ferrer, Kim Denyse Hao Lin and Reynaldo L. Garcia
Non-Coding RNA 2026, 12(2), 10; https://doi.org/10.3390/ncrna12020010 - 16 Mar 2026
Abstract
Background/Objectives: Lung cancer remains a major global health burden, largely driven by cigarette use. Although electronic cigarettes (e-cigarettes) are viewed as safer alternatives due to their reduced chemical load, growing evidence shows their vapor can disrupt cellular transcriptomes, including long noncoding RNAs
[...] Read more.
Background/Objectives: Lung cancer remains a major global health burden, largely driven by cigarette use. Although electronic cigarettes (e-cigarettes) are viewed as safer alternatives due to their reduced chemical load, growing evidence shows their vapor can disrupt cellular transcriptomes, including long noncoding RNAs (lncRNAs). In this study, we examined the regulation and function of vape-associated lncRNA transcript 1 (VALT1), a novel transcript upregulated in the oral transcriptomes of e-cigarette users and similarly elevated in non-small-cell lung cancer (NSCLC) tumors. Methods: Publicly available RNA-seq datasets were analyzed, and VALT1 was identified as an e-cigarette-responsive lncRNA. Its dose-dependent induction by e-cigarette smoke extract (eCSE) and cytoplasmic localization were confirmed via RT-qPCR. Its effects on cancer-associated phenotypes including proliferation, ROS detoxification, resistance to apoptosis, migration, cytoskeletal disorganization, and nuclear remodeling were assessed through overexpression and siRNA-mediated knockdown in A549 and BEAS-2B cells. Results: Acute eCSE exposure induced a biphasic, dose-dependent increase in VALT1 expression, accompanied by enhanced proliferation, ROS detoxification, apoptosis resistance, migration, cytoskeletal disorganization, and nuclear remodeling in A549 cells. VALT1 overexpression reproduced these phenotypes in both cell lines without eCSE treatment, whereas knockdown attenuated them. VALT1 promoted survival under cytotoxic stress in A549 but not BEAS-2B cells. Conclusions: These findings support an active role for VALT1 as an e-cigarette vapor-upregulated transcript that contributes to its phenotypic readout and enhances cellular survival under extracellular chemical stress—thereby aggravating tumorigenic phenotypes even in the absence of mutations that contribute to malignant transformation.
Full article
(This article belongs to the Section Long Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessArticle
Comprehensive Schistosoma mansoni Hierarchical Transcriptome Assembly Points to Novel lncRNAs Associated with Sexual Dimorphism
by
Caio Felipe Freire, Thalles Souza-Lopes, Murilo Sena Amaral, Ana Carolina Tahira and Sergio Verjovski-Almeida
Non-Coding RNA 2026, 12(2), 9; https://doi.org/10.3390/ncrna12020009 - 12 Mar 2026
Abstract
Background/Objectives: Schistosomiasis is a neglected tropical disease affecting >200 million people worldwide. Praziquantel is the sole recommended drug against Schistosoma mansoni; however, it lacks activity against juvenile forms and cannot prevent reinfection. Thus, there is an urgent need to identify novel therapeutic
[...] Read more.
Background/Objectives: Schistosomiasis is a neglected tropical disease affecting >200 million people worldwide. Praziquantel is the sole recommended drug against Schistosoma mansoni; however, it lacks activity against juvenile forms and cannot prevent reinfection. Thus, there is an urgent need to identify novel therapeutic targets. Long noncoding RNAs (lncRNAs) are known to regulate various biological processes in S. mansoni, including parasite pairing and fertility; therefore, screening for novel lncRNAs could reveal new potential targets. Methods: We compiled all publicly available RNA-seq data from the Sequence Read Archive (SRA) and performed a hierarchical transcriptome assembly using the multi-sample assembler Ryūtō, combined with version 10 of the S. mansoni genome. We applied HOMER for peak-calling and identification of histone marks and used weighted gene co-expression network analysis (WGCNA) to infer putative functions of lncRNAs in sexual dimorphism. Results: Using a robust pipeline, we identified 10,170 novel lncRNA genes comprising 16,990 novel lncRNA transcripts, including 8783 intergenic, 7918 antisense, and 289 intronic lncRNA transcripts. Most (78.7%) have histone regulatory marks (H3K4me3, H3K27me3, H3K27ac, or H4K20me1) near their transcription start sites, indicating potential expression regulation. Comparing male and female samples, we identified 1991 differentially expressed genes (FDR < 5%, |log2FC| ≥ 1.5), including 296 known lncRNAs and 339 novel lncRNAs. WGCNA identified hub lncRNAs within co-expression modules, and Gene Ontology enrichment analyses (FDR ≤ 5%) suggest that these lncRNAs are involved in cell differentiation and morphogenesis pathways. Conclusions: We provide a comprehensive catalog of S. mansoni lncRNAs. These findings offer opportunities to discover potential new therapeutic targets, advancing the future development of anti-schistosome therapies.
Full article
(This article belongs to the Section Long Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessReview
Short and Long Non-Coding RNAs in Renal Cell Carcinoma
by
Monia Cecati, Valentina Pozzi, Valentina Schiavoni, Giuseppina Barrasso, Veronica Pompei, Daniela Marzioni, Nicoletta Bonci, Stefania Fumarola, Andrea Ballini, Davide Sartini and Roberto Campagna
Non-Coding RNA 2026, 12(2), 8; https://doi.org/10.3390/ncrna12020008 - 27 Feb 2026
Abstract
Renal cell carcinoma (RCC) represents the most frequent kidney malignancy and remains a major clinical challenge due to its often silent onset, high metastatic potential, and limited responsiveness to conventional chemotherapy. Increasing evidence indicates that non-coding RNAs (ncRNAs), including microRNAs (miRNAs), long non-coding
[...] Read more.
Renal cell carcinoma (RCC) represents the most frequent kidney malignancy and remains a major clinical challenge due to its often silent onset, high metastatic potential, and limited responsiveness to conventional chemotherapy. Increasing evidence indicates that non-coding RNAs (ncRNAs), including microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and circular RNAs (circRNAs), are key regulators of RCC tumorigenesis, progression, and therapy resistance. Rather than providing a purely descriptive overview, this review focuses on emerging mechanistic paradigms through which ncRNAs actively shape tumor behavior and therapeutic response in RCC. This review summarizes current knowledge on the biological and clinical relevance of ncRNAs in RCC, highlighting their dual roles as oncogenic drivers or tumor suppressors through the modulation of pathways involved in proliferation, apoptosis, angiogenesis, invasion, immune evasion, metabolic reprogramming, and ferroptosis. Particular emphasis is placed on mechanistically defined ncRNA regulatory axes controlling ferroptosis, autophagy, metabolic reprogramming, and immune escape, as well as on ncRNA-mediated intercellular communication via extracellular vesicles, which promotes the dissemination of resistance to targeted therapies. The review also addresses ncRNA-based diagnostic and prognostic applications, including miRNA signatures capable of discriminating RCC subtypes and circulating ncRNAs as minimally invasive biomarkers. Moreover, the manuscript discusses ncRNA-mediated mechanisms of resistance to targeted therapies such as sunitinib, sorafenib, and axitinib, emphasizing regulatory networks involving miRNA targets, lncRNA–miRNA sponging, RNA-binding proteins, extracellular vesicle transfer, and epigenetic modulation. Emerging therapeutic opportunities are also addressed, including strategies aimed at inhibiting oncogenic ncRNAs or restoring tumor-suppressive ncRNAs to enhance drug sensitivity and improve patient stratification.
Full article
(This article belongs to the Section Clinical Applications of Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessReview
The Stiff Side of Cancer: How Matrix Mechanics Rewrites Non-Coding RNA Expression Programs
by
Alma D. Campos-Parra, Jonathan Puente-Rivera, César López-Camarillo, Stephanie I. Nuñez-Olvera, Nereyda Hernández Nava, Gabriela Alvarado Macias and Macrina Beatriz Silva-Cázares
Non-Coding RNA 2026, 12(1), 7; https://doi.org/10.3390/ncrna12010007 - 18 Feb 2026
Abstract
Extracellular matrix (ECM) stiffening is a defining biophysical feature of solid tumors that reshape gene regulation through mechanotransduction. Increased collagen crosslinking and stromal remodeling enhance integrin engagement, focal-adhesion signaling and force transmission to the nucleus, where key hubs such as lysyl oxidase (LOX),
[...] Read more.
Extracellular matrix (ECM) stiffening is a defining biophysical feature of solid tumors that reshape gene regulation through mechanotransduction. Increased collagen crosslinking and stromal remodeling enhance integrin engagement, focal-adhesion signaling and force transmission to the nucleus, where key hubs such as lysyl oxidase (LOX), focal adhesion kinase (FAK) and the Hippo co-activators YAP1 and TAZ (WWTR1) promote proliferation, invasion, stemness and therapy resistance. Here, we synthesize evidence that quantitative changes in matrix stiffness remodel the miRNome and lncRNome in both tumor and stromal compartments, including extracellular vesicle cargo that reprograms metastatic niches. To address heterogeneity in experimental support, we classify mechanosensitive ncRNAs into studies directly validated by stiffness manipulation (e.g., tunable hydrogels/AFM) versus indirect associations based on mechanosensitive signaling, and we summarize physiological versus pathophysiological stiffness ranges across tissues discussed. We further review competing endogenous RNA (ceRNA) networks converging on mechanotransduction nodes and ECM remodeling enzymes, and discuss translational opportunities and challenges, including targeting mechanosensitive ncRNAs, combining ncRNA modulation with anti-stiffening strategies, delivery barriers in dense tumors, and the potential of circulating/exosomal ncRNAs as biomarkers. Overall, integrating ECM mechanics with ncRNA regulatory circuits provides a framework to identify feed-forward loops sustaining aggressive phenotypes in rigid microenvironments and highlights priorities for validation in physiologically relevant models.
Full article
(This article belongs to the Section Long Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessArticle
Computational Analysis of Telomerase RNA Evolution in Caenorhabditis Species
by
Christopher Klapproth, Franziska Reinhardt, Peter F. Stadler and Sven Findeiß
Non-Coding RNA 2026, 12(1), 6; https://doi.org/10.3390/ncrna12010006 - 11 Feb 2026
Abstract
Background/Objectives: The telomerase RNA (TR) is an indispensable part of the telomerase protein complex responsible for telomere elongation in most eukaryotic species. Although the telomere terminal repeat sequence (TTAGGC)n in Caenorhabditis elegans has been known for years, a telomerase RNA gene was
[...] Read more.
Background/Objectives: The telomerase RNA (TR) is an indispensable part of the telomerase protein complex responsible for telomere elongation in most eukaryotic species. Although the telomere terminal repeat sequence (TTAGGC)n in Caenorhabditis elegans has been known for years, a telomerase RNA gene was not identified in the entire phylum of Nematoda until recently. Methods: In this exploratory study, we employ a combination of different approaches to identify likely telomerase RNA candidates among putative non-coding transcripts. Results: A detailed analysis of our prime candidate shows compelling evidence that it encodes the missing RNA element of the telomerase complex, which is notably located in an intron of the coding gene nmy-2. Using nmy-2 homologs in other nematodes as anchors, we annotate the conserved TR gene in 21 Caenorhabditis species. We furthermore show that the intronic localization of the TR gene is conserved in two distinct branching groups of the Caenorhabditis phylogeny and demonstrate that this property likely emerged from a single point of origin. Conclusions: While the intronic TR represents a very interesting evolutionary adaption that seems to have been successful in the Elegans and Japonica groups, the question regarding the macroscopic nematode TR evolution remains.
Full article
(This article belongs to the Section Evolution of Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessArticle
Extracellular Small RNAs in Human Milk: Molecular Profiles, Stability and Fragment-Specific Responses in Cell-Based Assays
by
Clara Claus, Carla Borini Etichetti, Bruno Costa, Julieta B. Grosso, Juan Pablo Tosar, Uciel Chorostecki and Silvana V. Spinelli
Non-Coding RNA 2026, 12(1), 5; https://doi.org/10.3390/ncrna12010005 - 9 Feb 2026
Abstract
Background/Objectives: Human milk is a complex biological fluid containing not only macro- and micronutrients but also diverse bioactive molecules, including extracellular RNAs. Although RNA has been detected in milk for decades, only a subset of RNA species has been characterized in detail, and
[...] Read more.
Background/Objectives: Human milk is a complex biological fluid containing not only macro- and micronutrients but also diverse bioactive molecules, including extracellular RNAs. Although RNA has been detected in milk for decades, only a subset of RNA species has been characterized in detail, and abundant families such as tRNA-, yRNA-, and rRNA-derived fragments remain underexplored. This study aimed to define the composition, fragmentation patterns, stability, and exploratory functional activity of these highly abundant RNAs in human milk. Methods: We performed small RNA sequencing on skim milk samples and analyzed the resulting profiles in comparison with publicly available milk and biofluid datasets. RNA stability assays, Northern blotting, and RT-qPCR were conducted to validate RNA abundance and degradation kinetics. Extracellular vesicles (EVs) and non-vesicular fractions were analyzed to determine the subcellular distribution of RNA species. Exploratory functional assays using synthetic RNA fragments were carried out to assess their ability to modulate cellular responses in vitro. Results: Human milk was found to be highly enriched in small RNA fragments derived from tRNA, yRNA, and rRNA, dominated by a limited set of discrete sequences. These profiles were highly reproducible across independent datasets and distinct biofluids. Orthologal validation assays confirmed their abundance and stability, with RNA levels exceeding those of serum by over two orders of magnitude. Full-length transcripts were enriched in EVs, whereas shorter fragments predominated in the non-vesicular fraction. Synthetic milk-derived exRNAs showed detectable pro-survival activity under stress conditions in vitro. Conclusions: This study reveals that human milk carries a limited set of highly abundant stable sRNA molecules, primarily derived from tRNAs, yRNAs, and rRNAs. These findings provide new insights into the RNA cargo of human milk and offer preliminary evidence that selected sRNA fragments can modulate cellular stress responses in in vitro models.
Full article
(This article belongs to the Section Small Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessReview
Mechanisms at the Intersection of lncRNA and m6A Biology
by
Samuel J. Gonzalez, Edgardo Linares, Allison M. Porman Swain and Aaron M. Johnson
Non-Coding RNA 2026, 12(1), 4; https://doi.org/10.3390/ncrna12010004 - 31 Jan 2026
Abstract
This review provides a thorough survey of long noncoding RNAs that bear the RNA modification N6-methyladenosine (m6A) and current work to understand the resulting mechanistic and biological consequences. We give an overview of lncRNA and m6A biology first, describing the writers, erasers, and
[...] Read more.
This review provides a thorough survey of long noncoding RNAs that bear the RNA modification N6-methyladenosine (m6A) and current work to understand the resulting mechanistic and biological consequences. We give an overview of lncRNA and m6A biology first, describing the writers, erasers, and readers of m6A and their targeting of lncRNAs. Next, we give an in-depth review of the field of nuclear lncRNAs that regulate chromatin and their regulation via m6A. We then describe the growing appreciation of liquid–liquid phase separation properties in lncRNA and m6A biology. Finally, we cover examples of cytoplasmic lncRNAs regulated by m6A. Overall, this review aims to emphasize how epitranscriptomics influences noncoding RNA mechanisms to provide additional layers of regulation, integrated into downstream biological processes.
Full article
(This article belongs to the Section Long Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessArticle
Hypercholesterolemia Impairs the Expression of Angiogenic MicroRNAs in Extracellular Vesicles Within Ischemic Skeletal Muscles
by
Nozha Raguema, Sylvie Dussault, Kevin Sawaya, Michel Desjarlais, Eric Boilard, Sylvain Chemtob and Alain Rivard
Non-Coding RNA 2026, 12(1), 3; https://doi.org/10.3390/ncrna12010003 - 26 Jan 2026
Abstract
►▼
Show Figures
Background/Objectives: In severe peripheral artery disease (PAD) with limb ischemia, hypercholesterolemia (HC) is associated with impaired neovascularization. Extracellular vesicles (EVs) are present within ischemic muscles, and they contain microRNAs (miRs) involved in several biological functions, including angiogenesis and neovascularization. Methods: We
[...] Read more.
Background/Objectives: In severe peripheral artery disease (PAD) with limb ischemia, hypercholesterolemia (HC) is associated with impaired neovascularization. Extracellular vesicles (EVs) are present within ischemic muscles, and they contain microRNAs (miRs) involved in several biological functions, including angiogenesis and neovascularization. Methods: We used a mouse model of PAD and compared the response to hindlimb ischemia in hypercholesterolemic ApoE−/− vs. normocholesterolemic mice. Next-generation sequencing (NGS) was used to perform full miR expression profiling in ischemic skeletal muscles and in EVs of varying sizes—large EVs (lEVs) and small EVs (sEVs)—within these muscles. Results: We identified several miRs with potential pro-angiogenic effects (angiomiRs) that are reduced by HC in lEVs (Let-7b-5p, miR-151-3p, Let-7c-5p) or sEVs (miR-21a-5p, miR-196b-5p, miR-340-5p). As proof of principle, we showed that the overexpression of Let-7b-5p in lEVs, or miR-21a-5p in sEVs, can significantly increase the angiogenic capacity of these EVs in vitro. HC also impaired the enrichment of specific angiomiRs in lEVs (miR-100-5p), sEVs (miR-142a-3p), or in both lEVs and sEVs (miR-146b-5p). In silico approaches, including the prediction of miR targets, pathway unions, and gene unions, identified the resulting predictive effects of HC-modulated miRs in EVs on processes with key roles in the modulation of angiogenesis and neovascularization, such as the regulation of the actin cytoskeleton and focal adhesion and the HIF-1, MAPK, AMPK, and PI3K-Akt signaling pathways. Conclusions: Our results constitute an important first step towards the identification of specific miRs that could be targeted to improve EV angiogenic function in hypercholesterolemic conditions and reduce tissue ischemia in patients with severe PAD.
Full article

Figure 1
Open AccessReview
The Role of microRNAs as Potential Biomarkers in Diffuse Large B-Cell Lymphoma
by
Eirini Panteli, Epameinondas Koumpis, Vasileios Georgoulis, Georgios Petros Barakos, Evangelos Kolettas, Panagiotis Kanavaros, Alexandra Papoudou-Bai and Eleftheria Hatzimichael
Non-Coding RNA 2026, 12(1), 2; https://doi.org/10.3390/ncrna12010002 - 7 Jan 2026
Abstract
►▼
Show Figures
Diffuse large B-cell lymphoma (DLBCL) is the most common and clinically aggressive subtype of non-Hodgkin lymphoma (NHL). While novel therapies such as rituximab and polatuzumab vedotin have led to improved outcomes, approximately 35% of patients eventually develop relapsed or refractory disease. MicroRNAs (miRNAs),
[...] Read more.
Diffuse large B-cell lymphoma (DLBCL) is the most common and clinically aggressive subtype of non-Hodgkin lymphoma (NHL). While novel therapies such as rituximab and polatuzumab vedotin have led to improved outcomes, approximately 35% of patients eventually develop relapsed or refractory disease. MicroRNAs (miRNAs), a class of endogenous single-stranded RNAs approximately 22 nucleotides in length, play a pivotal role in the regulation of gene expression at the post-transcriptional level through interactions with complementary target RNAs and contribute significantly to the development, progression, and treatment response of DLBCL. Oncogenic miRNAs, such as miR-155, miR-21, and the miR-17–92 cluster, promote proliferation, survival, immune evasion, and therapy resistance by modulating pathways including PI3K/AKT, NF-κB, and MYC. Conversely, tumor-suppressive miRNAs such as miR-34a, miR-144, miR-181a, and miR-124-3p inhibit oncogene activity and enhance apoptosis, with their loss often associated with adverse outcomes. Among these, miR-155 and miR-21 are particularly well studied, playing central roles in both tumor progression and remodeling of the tumor microenvironment. This review summarizes current evidence on the biological and clinical relevance of miRNAs in DLBCL, emphasizing their diagnostic and prognostic potential.
Full article

Figure 1
Open AccessArticle
Functional and Molecular Characterization of Extracellular Vesicles Enriched in Exosomes Released by Bone Marrow Mesenchymal Stromal Cells Exposed to IFNγ in Combination with Autophagy Modulators Tamoxifen or Chloroquine
by
Vladimir Beljanski, Maria J. Moreno Hollweg, Renee Potens, Tanner Blaylock, Andres B. Irausquin, Nikhila Paleati and Lubov Nathanson
Non-Coding RNA 2026, 12(1), 1; https://doi.org/10.3390/ncrna12010001 - 24 Dec 2025
Cited by 1
Abstract
►▼
Show Figures
Background/Objectives: Bone marrow mesenchymal stromal cells (MSCs) are therapeutic cells that adopt an immunomodulatory phenotype when exposed to pro-inflammatory cytokines. Recent research efforts uncovered that many therapeutic benefits of MSCs can be attributed to the secretion of extracellular vesicles (EVs) such as
[...] Read more.
Background/Objectives: Bone marrow mesenchymal stromal cells (MSCs) are therapeutic cells that adopt an immunomodulatory phenotype when exposed to pro-inflammatory cytokines. Recent research efforts uncovered that many therapeutic benefits of MSCs can be attributed to the secretion of extracellular vesicles (EVs) such as exosomes, small membrane vesicles of endocytic origin present in the cellular secretome. EVs’ formation and release are impacted by the autophagy pathway, which recycles proteins and organelles via lysosomal degradation. Methods: To evaluate how modulation of autophagy affects properties of MSC EVs enriched in exosomes under pro-inflammatory conditions, we treated the cells with either tamoxifen (TX) or chloroquine (CQ), two drugs known to stimulate or inhibit autophagy, respectively, together with IFNγ. MSC EVs enriched in exosomes were then purified from serum-free media, and their immunoregulatory properties were evaluated ex vivo using activated CD4 T cells; small RNA sequencing was also conducted to determine EVs’ microRNA content. Results: Our data indicate that MSCs treated with CQ + IFNγ yield EVs that possess somewhat higher capacity to decrease T cell proliferation compared to other EVs. Small RNA sequencing revealed that, although similar microRNAs were found in EVs isolated from all treated cells, the treatments exerted more effect on the levels of multiple microRNAs that are known to regulate either cancer or inflammation-related biological pathways in target cells. Conclusions: Overall, we conclude that the co-treatment of MSCs with TX or CQ in the presence of pro-inflammatory cytokine IFNγ has the potential to modulate microRNA content of EVs, potentially affecting biological properties of such EVs and their effect on target cells.
Full article

Figure 1
Open AccessArticle
Transposable Element-Derived miR-28-5p and miR-708-5p: Exploring Potential Roles in Lung Cancer
by
Sergiu Chira, Cornelia Braicu, Stefan Strilciuc, George A. Calin and Ioana Berindan-Neagoe
Non-Coding RNA 2025, 11(6), 81; https://doi.org/10.3390/ncrna11060081 - 18 Dec 2025
Abstract
►▼
Show Figures
Background: Transposable elements are normally silenced by epigenetic mechanisms; however, during malignant transformation, epigenetic alterations enable transposons to produce functional molecules like miRNAs. Among these, LINE-2 (L2) elements can generate miRNAs capable of regulating key genes, including tumor suppressors. Two L2-derived miRNAs, miR-28
[...] Read more.
Background: Transposable elements are normally silenced by epigenetic mechanisms; however, during malignant transformation, epigenetic alterations enable transposons to produce functional molecules like miRNAs. Among these, LINE-2 (L2) elements can generate miRNAs capable of regulating key genes, including tumor suppressors. Two L2-derived miRNAs, miR-28 and miR-708, have been linked to lung cancer, yet the mechanisms underlying their dysregulation remain poorly understood. Our study reveals how genomic context contributes to aberrant gene expression through comprehensive bioinformatic analyses. Methods: Using bioinformatics analysis, we evaluated the expression of miR-28 and miR-708 in lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC) datasets from TCGA. Further, we assessed the expression and methylation status of miR-28 and miR-708 host genes, LPP and TENM4, respectively, using computational tools. Finaly, we searched for potential candidate tumor suppressor genes targeted by miR-28 and miR-708, which are downregulated in LUAD and LUSC. Results: We found that intragenic L2-derived miR-28 and miR-708 are significantly upregulated in LUAD and LUSC. While TENM4 gene also displays a marked increase in expression in LUAD and LUSC, in tumor versus normal tissue, this difference is less obvious for the LPP gene. We suggest that such dysregulations in expression might be linked to specific methylation patterns of their genomic locations. Furthermore, we emphasize that miR-28 and miR-708 might contribute to lung cancer pathogenesis by targeting key tumor suppressor genes. Conclusions: Alterations in the methylation status of L2-miRNAs genomic loci might result in elevated levels of miRNAs and subsequent targeting of tumor suppressor genes with potential implications in lung cancer pathogenesis.
Full article

Figure 1
Open AccessReview
MicroRNAs in Breast Cancer Bone Metastasis Formation and Progression: An Overview on Recent Progress in This Research Field
by
Margherita Puppo
Non-Coding RNA 2025, 11(6), 80; https://doi.org/10.3390/ncrna11060080 - 18 Dec 2025
Abstract
Bone metastasis is a common and severe complication in advanced stages of breast cancer (BC) that is characterised by limited treatment options and poor patient prognosis. MicroRNAs (miRNAs) are a large class of regulatory small non-coding RNAs (ncRNAs) expressed by cells. Moreover, miRNAs
[...] Read more.
Bone metastasis is a common and severe complication in advanced stages of breast cancer (BC) that is characterised by limited treatment options and poor patient prognosis. MicroRNAs (miRNAs) are a large class of regulatory small non-coding RNAs (ncRNAs) expressed by cells. Moreover, miRNAs can be released by cells into the blood and lymphatic streams, acting as distant cell-to-cell communicators. Of note, miRNAs have pivotal roles in the metastatic progression of BC to bone. This review summarises the most recent findings on miRNAs and their mRNA targets in driving BC bone metastasis. Furthermore, the potential clinical uses of miRNAs as future therapeutic targets/agents or biomarkers for BC bone metastasis are discussed.
Full article
(This article belongs to the Section Small Non-Coding RNA)
►▼
Show Figures

Figure 1
Open AccessReview
CARINH, an Interferon-Induced LncRNA in Cancer and Inflammation
by
Morgane Gourvest and Coen van Solingen
Non-Coding RNA 2025, 11(6), 79; https://doi.org/10.3390/ncrna11060079 - 21 Nov 2025
Abstract
►▼
Show Figures
CARINH is an intriguing long noncoding RNA whose unique regulatory functions intersect the seemingly distinct processes of innate immunity and cancer development. Notably, CARINH is conserved across species, offering powerful experimental models for uncovering its mechanistic roles and physiological functions across diverse biological
[...] Read more.
CARINH is an intriguing long noncoding RNA whose unique regulatory functions intersect the seemingly distinct processes of innate immunity and cancer development. Notably, CARINH is conserved across species, offering powerful experimental models for uncovering its mechanistic roles and physiological functions across diverse biological contexts. Stimulated by interferons and viral infections, CARINH stands out as a key player in the body’s antiviral defense mechanisms. Additionally, its dysregulation has been implicated in autoimmune disorders such as psoriasis, asthma, and inflammatory bowel disease, underscoring its broader role in maintaining immune homeostasis. Furthermore, alterations in CARINH expression have been connected to cancer progression, highlighting its dual role in immune response and tumor suppression. In this review, we delve into CARINH’s pivotal function in modulating interferon responses and influencing cancer development, with a focus on the molecular pathways that regulate its expression and contribute to its diverse roles. Understanding these pathways is crucial for evaluating CARINH’s significance as a biomarker and therapeutic target, potentially leading to groundbreaking advancements in medical research and treatment strategies.
Full article

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Cancers, Cells, Diseases, Genes, IJMS, ncRNA, Sci
Alternative Splicing in RNA Metabolism, Tissue Homeostasis and Human Disease: From Mechanisms to Therapeutics and Clinical Applications
Topic Editors: Christos K. Kontos, Dongyu JiaDeadline: 15 January 2027

Conferences
Special Issues
Special Issue in
ncRNA
Women’s Special Issue Series: Worldwide Advances in Non-Coding RNA Research
Guest Editors: Ling Yang, Barbara PardiniDeadline: 30 June 2026
Special Issue in
ncRNA
Non-Coding RNA: 10th Anniversary
Guest Editor: George A. CalinDeadline: 1 September 2026
Special Issue in
ncRNA
The Importance of Non-Coding RNAs in Epithelial Cancers—2nd Edition
Guest Editor: Leandro CastellanoDeadline: 21 September 2026
Special Issue in
ncRNA
ncRNAs in Human Diseases and Therapeutics
Guest Editors: Sujay Paul, Neil RenwickDeadline: 20 October 2026
Topical Collections
Topical Collection in
ncRNA
Regulatory RNAs in Cardiovascular Development and Disease
Collection Editors: Yvan Devaux, Francisco J. Enguita, Andrea Caporali
Topical Collection in
ncRNA
The Non-Coding RNA Journal Club: Highlights on Recent Papers
Collection Editor: George A. Calin








