First Description of the Mitogenome and Phylogeny of Culicinae Species from the Amazon Region
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection and Total DNA Extraction
2.2. Quantification of Total Extracted DNA, Construction of the Genomic Library, and Sequencing
2.3. Pre-Processing of Sequenced Products
2.4. Genomic Assembly
2.5. Analyses of the Obtained Mitochondrial Sequences
2.6. Phylogenetics Analysis
3. Results and Discussion
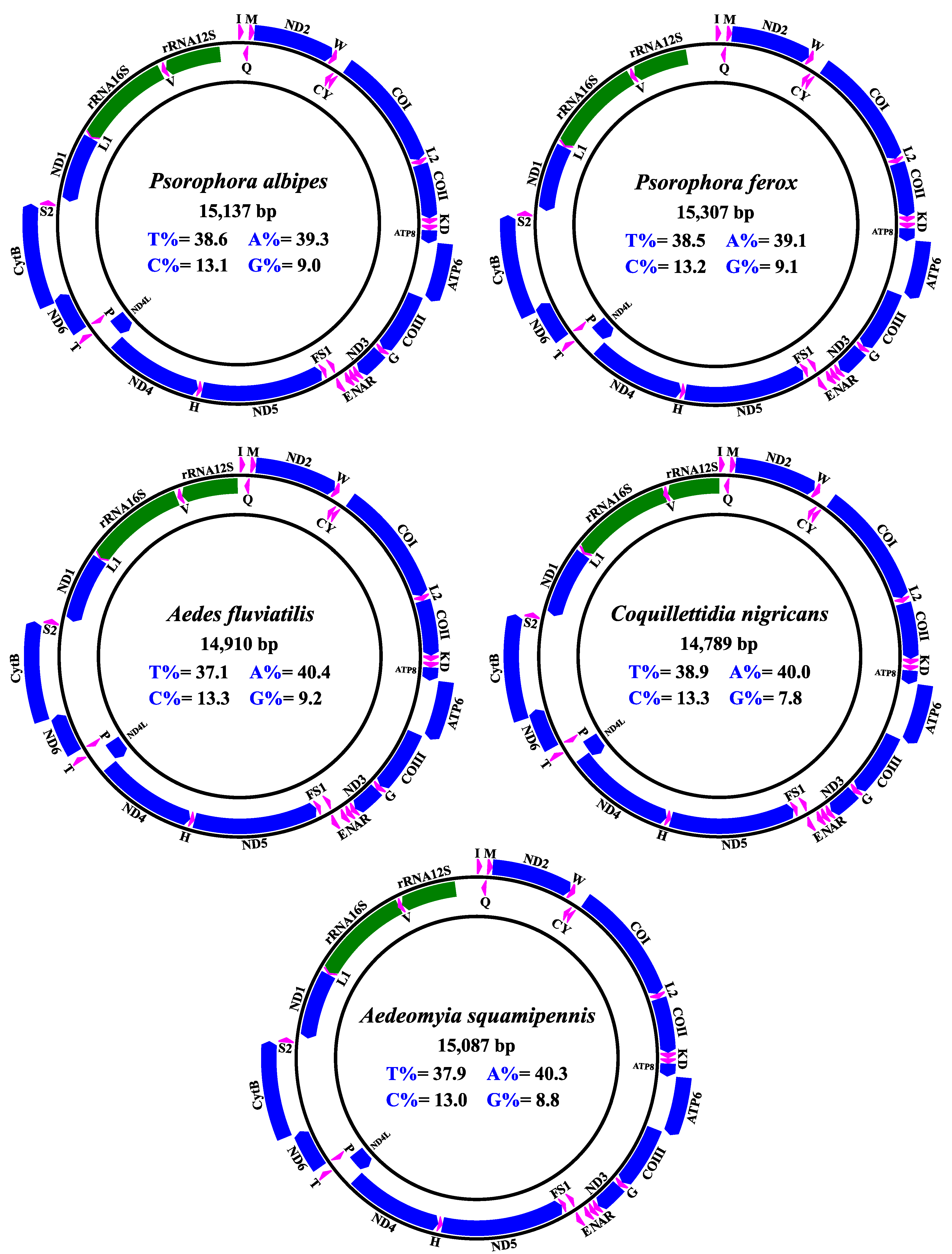
3.1. Organization, Structure and Content of the Mitogenomes Obtained
3.2. Description of Protein-Coding Genes (PCGs)
3.3. Evolutive Analysis
3.4. Phylogenetic Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Harbach, R.E. Mosquito Taxonomic Inventory. 2013. Available online: https://www.mosquito-taxonomic-inventory.info/ (accessed on 3 November 2021).
- Harbach, R.E. The Culicidae (Diptera): A Review of Taxonomy, Classification and Phylogeny. Zootaxa 2007, 1668, 591–638. [Google Scholar] [CrossRef] [Green Version]
- Harbach, R.E.; Kitching, I.J. Phylogeny and Classification of the Culicidae (Diptera). Syst. Entomol. 1998, 23, 327–370. [Google Scholar] [CrossRef]
- Forattini, O.P. Culicidologia Médica: Identificação, Biologia, Epidemiologia; EDUSP: São Paulo, Brazil, 2002. [Google Scholar]
- Hao, Y.J.; Zou, Y.L.; Ding, Y.R.; Xu, W.Y.; Yan, Z.T.; Li, X.D.; Fu, W.B.; Li, T.J.; Chen, B. Complete Mitochondrial Genomes of Anopheles stephensi and An. dirus and Comparative Evolutionary Mitochondriomics of 50 Mosquitoes. Sci. Rep. 2017, 7, 7666. [Google Scholar] [CrossRef]
- Sallum, M.A.M.; Schultz, T.R.; Foster, P.G.; Aronstein, K.; Wirtz, R.A.; Wilkerson, R.C. Phylogeny of Anophelinae (Diptera: Culicidae) Based on Nuclear Ribosomal and Mitochondrial DNA Sequences. Syst. Entomol. 2002, 27, 361–382. [Google Scholar] [CrossRef] [Green Version]
- Hay, S.I.; Sinka, M.E.; Okara, R.M.; Kabaria, C.W.; Mbithi, P.M.; Tago, C.C.; Benz, D.; Gething, P.W.; Howes, R.E.; Patil, A.P.; et al. Developing Global Maps of the Dominant Anopheles Vectors of Human Malaria. PLoS Med. 2010, 7, e1000209. [Google Scholar] [CrossRef] [PubMed]
- Nanfack Minkeu, F.; Vernick, K.D. A Systematic Review of the Natural Virome of Anopheles mosquitoes. Viruses 2018, 10, 222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foster, P.G.; De Oliveira, T.M.P.; Bergo, E.S.; Conn, J.; Sant’Ana, D.C.; Nagaki, S.S.; Nihei, S.; Lamas, C.E.; Gonzalez, C.; Moreira, C.C.; et al. Phylogeny of Anophelinae Using Mitochondrial Protein Coding Genes. R. Soc. Open Sci. 2017, 4, 170758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Judd, D.D. Review of the Systematics and Phylogenetic Relationships of the Sabethini (Diptera: Culicidae). Syst. Entomol. 1996, 21, 129–150. [Google Scholar] [CrossRef]
- Reidenbach, K.R.; Cook, S.; Bertone, M.A.; Harbach, R.E.; Wiegmann, B.M.; Besansky, N.J. Phylogenetic Analysis and Temporal Diversification of Mosquitoes (Diptera: Culicidae) Based on Nuclear Genes and Morphology. BMC Evol. Biol. 2009, 9, 298. [Google Scholar] [CrossRef] [Green Version]
- Lorenz, C.; Alves, J.M.P.; Foster, P.G.; Sallum, M.A.M.; Suesdek, L. First Record of Translocation in Culicidae (Diptera) Mitogenomes: Evidence from the Tribe Sabethini. BMC Genom. 2019, 20, 721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laurito, M.; de Oliveira, T.M.P.; Almirón, W.R.; Sallum, M.A.M. COI Barcode versus Morphological Identification of Culex (Culex) (Diptera: Culicidae) Species: A Case Study Using Samples from Argentina and Brazil. Mem. Inst. Oswaldo Cruz 2013, 108, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Laurito, M.; Ayala, A.M.; Almirón, W.R.; Gardenal, C.N. Molecular Identification of Two Culex (Culex) Species of the Neotropical Region (Diptera: Culicidae). PLoS ONE 2017, 12, e0173052. [Google Scholar] [CrossRef]
- Chan-Chable, R.J.; Martínez-Arce, A.; Mis-Avila, P.C.; Ortega-Morales, A.I. DNA Barcodes and Evidence of Cryptic Diversity of Anthropophagous Mosquitoes in Quintana Roo, Mexico. Ecol. Evol. 2019, 9, 4692–4705. [Google Scholar] [CrossRef] [Green Version]
- DeSalle, R.; Egan, M.G.; Siddall, M. The Unholy Trinity: Taxonomy, Species Delimitation and DNA Barcoding. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1905–1916. [Google Scholar] [CrossRef] [Green Version]
- Tabachnick, W.J. Nature, Nurture and Evolution of Intra-Species Variation in Mosquito Arbovirus Transmission Competence. Int. J. Environ. Res. Public Health 2013, 10, 249–277. [Google Scholar] [CrossRef] [Green Version]
- Barzon, L.; Lavezzo, E.; Militello, V.; Toppo, S.; Palù, G. Applications of Next-Generation Sequencing Technologies to Diagnostic Virology. Int. J. Mol. Sci. 2011, 12, 7861–7884. [Google Scholar] [CrossRef] [Green Version]
- Boore, J.L. Animal Mitochondrial Genomes. Nucleic Acids Res. 1999, 27, 1767–1780. [Google Scholar] [CrossRef] [Green Version]
- Aragão, A.O.; Nunes Neto, J.P.; Cruz, A.C.R.; Casseb, S.M.M.; Cardoso, J.F.; da Silva, S.P.; Ishikawa, E.A.Y. Description and Phylogeny of the Mitochondrial Genome of Sabethes chloropterus, Sabethes glaucodaemon and Sabethes belisarioi (Diptera: Culicidae). Genomics 2019, 111, 607–611. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Wang, Y.; Li, X.Y.; Peng, H.; Ma, Y.J. Sequencing and Analysis of the Complete Mitochondrial Genome in Anopheles sinensis (Diptera: Culicidae). Infect. Dis. Poverty 2017, 6, 1–6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Demari-Silva, B.; Foster, P.G.; de Oliveira, T.M.P.; Bergo, E.S.; Sanabani, S.S.; Pessôa, R.; Sallum, M.A.M. Mitochondrial Genomes and Comparative Analyses of Culex camposi, Culex coronator, Culex usquatus and Culex usquatissimus (Diptera:CUlicidae), Members of the Coronator Group. BMC Genom. 2015, 16, 831. [Google Scholar] [CrossRef] [Green Version]
- Da Silva, F.S.; Cruz, A.C.R.; de Almeida Medeiros, D.B.; Silva, S.P.; Nunes, M.R.T.; Martins, L.C.; Chiang, J.O.; da Silva Lemos, P.; Cunha, G.M.; Araujo, R.F.; et al. Mitochondrial Genome Sequencing and Phylogeny of Haemagogus albomaculatus, Haemagogus leucocelaenus, Haemagogus spegazzinii, and Haemagogus tropicalis (Diptera: Culicidae). Sci. Rep. 2020, 10, 16948. [Google Scholar] [CrossRef]
- Ye, F.; Liu, T.; King, S.D.; You, P. Mitochondrial Genomes of Two Phlebotomine Sand Flies, Phlebotomus chinensis and Phlebotomus papatasi (Diptera: Nematocera), the First Representatives from the Family Psychodidae. Parasites Vectors 2015, 8, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, N.; Liang, A.P.; Bu, C.P. A Molecular Phylogeny of Hemiptera Inferred from Mitochondrial Genome Sequences. PLoS ONE 2012, 7, e48778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pita, S.; Panzera, F.; Vela, J.; Mora, P.; Palomeque, T.; Lorite, P. Complete Mitochondrial Genome of Triatoma Infestans (Hemiptera, Reduviidae, Triatominae), Main Vector of Chagas Disease. Infect. Genet. Evol. 2017, 54, 158–163. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S. FastQC v.0.11.9. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 16 September 2021).
- Krueger, F. Trim Galore v.0.6.5. Available online: https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ (accessed on 16 September 2021).
- Li, D.; Liu, C.-M.; Luo, R.; Sadakane, K.; Lam, T.-W. MEGAHIT: An Ultra-Fast Single-Node Solution for Large and Com- Plex Metagenomics Assembly via Succinct de Bruijn Graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef] [Green Version]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [Green Version]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and Sensitive Protein Alignment Using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef]
- Huson, D.H.; Auch, A.F.; Qi, J.; Schuster, S.C. MEGAN Analysis of Metagenomic Data. Genome Res. 2007, 17, 377–386. [Google Scholar] [CrossRef] [Green Version]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Bernt, M.; Donath, A.; Jühling, F.; Externbrink, F.; Florentz, C.; Fritzsch, G.; Pütz, J.; Middendorf, M.; Stadler, P.F. MITOS: Improved de Novo Metazoan Mitochondrial Genome Annotation. Mol. Phylogenet. Evol. 2013, 69, 313–319. [Google Scholar] [CrossRef]
- Stothard, P.; Wishart, D.S. Circular Genome Visualization and Exploration Using CGView. Bioinformatics 2005, 21, 537–539. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Perna, N.T.; Kocher, T.D. Patterns of Nucleotide Composition at Fourfold Degenerate Sites of Animal Mitochondrial Genomes. J. Mol. Evol. 1995, 41, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, B.; Wittelsbürger, U.; Ramos-Onsins, S.E.; Lercher, M.J. PopGenome: An Efficient Swiss Army Knife for Population Genomic Analyses in R. Mol. Biol. Evol. 2014, 31, 1929–1936. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Yang, Z. PAML 4: Phylogenetic Analysis by Maximum Likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian Inference of Phylogenetic Trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [Green Version]
- Rambaut, A. Figtree v1.4.4. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 1 November 2021).
- Inkscape. Available online: https://inkscape.org/pt-br/ (accessed on 1 November 2021).
- Sun, L.; Li, T.J.; Fu, W.B.; Yan, Z.T.; Si, F.L.; Zhang, Y.J.; Mao, Q.M.; Demari-Silva, B.; Chen, B. The Complete Mt Genomes of Lutzia halifaxia, Lt. fuscanus and Culex pallidothorax (Diptera: Culicidae) and Comparative Analysis of 16 Culex and Lutzia Mt Genome Sequences. Parasites Vectors 2019, 12, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinez-Villegas, L.; Assis-Geraldo, J.; Koerich, L.B.; Collier, T.C.; Lee, Y.; Main, B.J.; Rodrigues, N.B.; Orfano, A.S.; Pires, A.C.A.M.; Campolina, T.B.; et al. Characterization of the Complete Mitogenome of Anopheles Aquasalis, and Phylogenetic Divergences among Anopheles from Diverse Geographic Zones. PLoS ONE 2019, 14, e219523. [Google Scholar] [CrossRef] [Green Version]
- Luo, Q.C.; Hao, Y.J.; Meng, F.; Li, T.J.; Ding, Y.R.; Hua, Y.Q.; Chen, B. The Mitochondrial Genomes of Culex tritaeniorhynchus and Culex pipiens Pallens (Diptera: Culicidae) and Comparison Analysis with Two Other Culex Species. Parasites Vectors 2016, 9, 406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, D.X.; Hewitt, G.M. Insect Mitochondrial Control Region: A Review of Its Structure, Evolution and Usefulness in Evolutionary Studies. Biochem. Syst. Ecol. 1997, 25, 99–120. [Google Scholar] [CrossRef]
- Zhang, S.K.; Shu, J.P.; Wang, Y.D.; Liu, Y.N.; Peng, H.; Zhang, W.; Wang, H.J. The Complete Mitochondrial Genomes of Two Sibling Species of Camellia weevils (Coleoptera: Curculionidae) and Patterns of Curculionini Speciation. Sci. Rep. 2019, 9, 3412. [Google Scholar] [CrossRef] [PubMed]
- Behura, S.K.; Lobo, N.F.; Haas, B.; deBruyn, B.; Lovin, D.D.; Shumway, M.F.; Puiu, D.; Romero-Severson, J.; Nene, V.; Severson, D.W. Complete Sequences of Mitochondria Genomes of Aedes aegypti and Culex quinquefasciatus and Comparative Analysis of Mitochondrial DNA Fragments Inserted in the Nuclear Genomes. Insect Biochem. Mol. Biol. 2011, 41, 770–777. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oliveira, T.M.P.; Foster, P.G.; Bergo, E.S.; Nagaki, S.S.; Sanabani, S.S.; Marinotti, O.; Marinotti, P.N.; Sallum, M.A.M. Mitochondrial Genomes of Anopheles (Kerteszia) (Diptera: Culicidae) from the Atlantic Forest, Brazil. J. Med. Entomol. 2016, 53, 790–797. [Google Scholar] [CrossRef]
- Timmis, J.N.; Ayliff, M.A.; Huang, C.Y.; Martin, W. Endosymbiotic Gene Transfer: Organelle Genomes Forge Eukaryotic Chromosomes. Nat. Rev. Genet. 2004, 5, 123–135. [Google Scholar] [CrossRef]
- Lemos, P.d.S.; Monteiro, H.A.d.O.; Castro, F.C.; de Lima, C.P.S.; da Silva, D.E.A.; de Vasconcelos, J.M.; de Oliveira, L.F.; da Silva, S.P.; Cardoso, J.F.; Vianez Júnior, J.L.d.S.G.; et al. Characterization of Mitochondrial Genome of Haemagogus janthinomys (Diptera: Culicidae). Mitochondrial DNA Part A DNA Mapp. Seq. Anal. 2017, 28, 50–51. [Google Scholar] [CrossRef] [PubMed]
- Kambhampati, S.; Smith, P.T. PCR Primers for the Amplification of Four Insect Mitochondrial Gene Fragments. Insect Mol. Biol. 1995, 4, 233–236. [Google Scholar] [CrossRef]
- Xu, J.; Fonseca, D.M. One-Way Sequencing of Multiple Amplicons from Tandem Repetitive Mitochondrial DNA Control Region. Mitochondrial DNA 2011, 22, 155–158. [Google Scholar] [CrossRef]
- Wang, G.; Li, C.; Guo, X.; Xing, D.; Dong, Y.; Wang, Z.; Zhang, Y.; Liu, M.; Zheng, Z.; Zhang, H.; et al. Identifying the Main Mosquito Species in China Based on DNA Barcoding. PLoS ONE 2012, 7, e47051. [Google Scholar] [CrossRef] [PubMed]
- Makhawi, A.M.; Liu, X.B.; Yang, S.R.; Liu, Q.Y. Genetic Variations of ND5 Gene of MtDNA in Populations of Anopheles sinensis (Diptera: Culicidae) Malaria Vector in China. Parasites Vectors 2013, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Pu, D.Q.; Liu, H.L.; Gong, Y.Y.; Ji, P.C.; Li, Y.J.; Mou, F.S.; Wei, S.J. Mitochondrial Genomes of the Hoverflies Episyrphus balteatus and Eupeodes corollae (Diptera: Syrphidae), with a Phylogenetic Analysis of Muscomorpha. Sci. Rep. 2017, 7, 44300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Zhang, Y.; Hu, K.; Zhao, Y.; Lin, R.; Li, Y.; Huang, Z.; Zhang, X.; Geng, X.; Ding, J. Mitochondrial Genome Characteristics of Two Sphingidae Insects (Psilogramma increta and Macroglossum stellatarum) and Implications for Their Phylogeny. Int. J. Biol. Macromol. 2018, 113, 592–600. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.J.; Li, D.F.; Li, H.; Yang, M.F.; Dai, R.H. Structural and Phylogenetic Implications of the Complete Mitochondrial Genome of Ledra Auditura. Sci. Rep. 2019, 9, 15746. [Google Scholar] [CrossRef]
- Krzywinski, J.; Li, C.; Morris, M.; Conn, J.E.; Lima, J.B.; Povoa, M.M.; Wilkerson, R.C. Analysis of the Evolutionary Forces Shaping Mitochondrial Genomes of a Neotropical Malaria Vector Complex. Mol. Phylogenet. Evol. 2011, 58, 469–477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological Identifications through DNA Barcodes. Proc. R. Soc. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laurito, M.; Almirón, W.R. Phylogenetic Relationships of Culex (Culex) Species (Diptera, Culicidae) from Argentina Based on Morphological Characters. Zootaxa 2013, 3652, 117–155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cywinska, A.; Hunter, F.F.; Hebert, P.D.N. Identifying Canadian Mosquito Species through DNA Barcodes. Med. Vet. Entomol. 2006, 20, 413–424. [Google Scholar] [CrossRef] [PubMed]
- Murugan, K.; Vadivalagan, C.; Karthika, P.; Panneerselvam, C.; Paulpandi, M.; Subramaniam, J.; Wei, H.; Aziz, A.T.; Alsalhi, M.S.; Devanesan, S.; et al. DNA Barcoding and Molecular Evolution of Mosquito Vectors of Medical and Veterinary Importance. Parasitol. Res. 2016, 115, 107–121. [Google Scholar] [CrossRef] [PubMed]
- Gonz, R. Molecular Tracing with Mitochondrial ND5 of the Invasive Mosquito Aedes (Stegomyia) Albopictus (Skuse) in Northern South America. J. Entomol. Zool. Stud. 2016, 1, 32–39. [Google Scholar]
- Hess, P.N.; De Moraes Russo, C.A. An Empirical Test of the Midpoint Rooting Method. Biol. J. Linn. Soc. 2007, 92, 669–674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harbach, R.E.; Kitching, I.J. The Phylogeny of Anophelinae Revisited: Inferences about the Origin and Classification of Anopheles (Diptera: Culicidae). Zool. Scr. 2016, 45, 34–47. [Google Scholar] [CrossRef] [Green Version]
- Harbach, R.E. The Phylogeny and Classification of Anopheles. In Anopheles mosquitoes—New Insights into Malaria Vectors, Sylvie Manguin; InTech: Rijeka, Croatia, 2013. [Google Scholar] [CrossRef] [Green Version]
- Peng, X.Y.; Zhou, P.; Duan, X.Y.; Qian, Z.Q. The Mitochondrial Genomes of Twelve Anopheles mosquitoes (Diptera: Culicidae) and Their Phylogenetic Implications. Conserv. Genet. Resour. 2016, 8, 387–390. [Google Scholar] [CrossRef]
- Silva, A.F.; Machado, L.C.; de Paula, M.B.; da Silva Pessoa Vieira, C.J.; de Morais Bronzoni, R.V.; de Melo Santos, M.A.V.; Wallau, G.L. Culicidae Evolutionary History Focusing on the Culicinae Subfamily Based on Mitochondrial Phylogenomics. Sci. Rep. 2020, 10, 18823. [Google Scholar] [CrossRef]
- Tyson, W.H. Contributions to the Mosquito Fauna of Southeast Asia. VII. Genus Aedeomyia Theobald in South-East Asia. Contrib. Am. Entomol. Inst. 1970, 6, 1–27. [Google Scholar]
- Belkin, J.N. The Mosquitoes of the South Pacific (Diptera, Culicidae); Cambridge University Press: London, UK, 1962; Volume 2. [Google Scholar]
- Reinert, J.F. List of Abbreviations for Currently Valid Generic-Level Taxa in Family Culicidae (Diptera). J. Eur. Mosq. Control Assoc. 2009, 27, 68–76. [Google Scholar]
- Reinert, J.F. List of Species in Tribe Mansoniini with Published Illustrations and/or Descriptions of Eggs (Diptera: Culicidae). Eur. Mosq. Bull. 2010, 28, 143–147. [Google Scholar]
- Reinert, J.F.; Harbach, R.E.; Kitching, I.J. Phylogeny and Classification of Tribe aedini (Diptera: Culicidae). Zool. J. Linn. Soc. 2009, 157, 700–794. [Google Scholar] [CrossRef] [Green Version]
- Reinert, J.F.; Harbach, R.E.; Kitching, I.J. Phylogeny and Classification of Aedini (Diptera: Culicidae), Based on Morphological Characters of All Life Stages. Zool. J. Linn. Soc. 2004, 142, 289–368. [Google Scholar] [CrossRef] [Green Version]
- Reinert, J.F. Comparative Anatomy of the Female Genitalia of Generic-Level Taxa in Tribe aedini (Diptera: Culicidae). Part XV. Genus Georgecraigius Reinert, Harbach and Kitching. Contrib. Am. Entomol. Inst. 2008, 35, 1–10. [Google Scholar]
- Wilkerson, R.C.; Linton, Y.M.; Fonseca, D.M.; Schultz, T.R.; Price, D.C.; Strickman, D.A. Making Mosquito Taxonomy Useful: A Stable Classification of Tribe Aedini That Balances Utility with Current Knowledge of Evolutionary Relationships. PLoS ONE 2015, 10, e133602. [Google Scholar] [CrossRef]
- Soghigian, J.; Andreadis, T.G.; Livdahl, T.P. From ground pools to treeholes: Convergent evolution of habitat and phenotype in Aedes mosquitoes. BMC Evol. Biol. 2017, 17, 262. [Google Scholar] [CrossRef] [PubMed]
- Liria, J.; Navarro, J.-C. Phylogenetic Relationships in the Genus Psorophora Robineau-Desvoidy (Diptera: Culicidae), Based on Morphological Characters. Entomol. Appl. Sci. Lett. 2014, 4, 22–50. [Google Scholar]






| Species | ID-Code | Collected | Used | Sex | Collection Site | State | Geographical Coordinates | Year |
|---|---|---|---|---|---|---|---|---|
| Ad. squamipennis | AR_867100 | 6 | 5 | ♀ | Altamira | Pará | −3.203333, −52.206389 | 2017 |
| Ae. fluviatilis | AR_866516 | 13 | 5 | ♀ | Canaã dos Carajás | Pará | −6.496944, −49.878333 | 2019 |
| Cq. nigricans | AR_850512 | 45 | 5 | ♀ | Altamira | Pará | −3.203333, −52.206389 | 2017 |
| Ps. albipes | AR_859743 | 30 | 5 | ♀ | Belém | Pará | −1.495795, −48.454260 | 2017 |
| Ps. ferox | AR_849712 | 14 | 5 | ♀ | Santa Bárbara | Pará | −1.209000, −48.271722 | 2017 |
| Species Evaluated | Nucleotide Identity (%)—COI (Barcode DNA) | ||||
|---|---|---|---|---|---|
| Ad. squamipennis AF417729 | Oc. fluviatilis KF314736 | Cq. nigricans KM593033 | Ps. albipes MN997524 | Ps. ferox MN997516 | |
| Ad. squamipennis | 98.99 | - | - | - | - |
| Ae. fluviatilis | - | 98.78 | - | - | - |
| Cq. nigricans | - | - | 99.85 | - | - |
| Ps. albipes | - | - | - | 95.14 | - |
| Ps. ferox | - | - | - | - | 99.7 |
| Species | Size (bp) | A% | T% | C% | G% | AT% | AT-Skew | GC-Skew |
|---|---|---|---|---|---|---|---|---|
| Whole mitogenome | ||||||||
| Aedeomyia squamipennis | 15,087 | 40.3 | 37.9 | 13.0 | 8.8 | 78.2 | 0.0307 | −0.1927 |
| Aedes fluviatilis | 14,910 | 40.4 | 37.1 | 13.3 | 9.2 | 77.5 | 0.0426 | −0.1822 |
| Coquillettidia nigricans | 14,789 | 40.0 | 38.9 | 13.3 | 7.8 | 78.9 | 0.0139 | −0.2607 |
| Psorophora albipes | 15,137 | 39.3 | 38.6 | 13.1 | 9.0 | 77.9 | 0.0090 | −0.1855 |
| Psorophora ferox | 15,307 | 39.1 | 38.5 | 13.2 | 9.1 | 77.6 | 0.0068 | −0.1839 |
| PCGs | ||||||||
| Aedeomyia squamipennis | 11,170 | 32.6 | 44.1 | 11.3 | 12.0 | 76.7 | −0.1499 | 0.0300 |
| Aedes fluviatilis | 11,216 | 32.6 | 43.3 | 11.8 | 12.2 | 75.9 | −0.1410 | 0.0167 |
| Coquillettidia nigricans | 11,357 | 33.4 | 44.4 | 10.8 | 11.3 | 77.8 | −0.1414 | 0.0226 |
| Psorophora albipes | 11,269 | 32.5 | 44.1 | 11.4 | 12.0 | 76.6 | −0.1514 | 0.0256 |
| Psorophora ferox | 11,270 | 32.1 | 44.0 | 11.6 | 12.4 | 76.1 | −0.1564 | 0.0333 |
| tRNAs | ||||||||
| Aedeomyia squamipennis | 1470 | 40.8 | 38.9 | 8.9 | 11.4 | 79.7 | 0.0238 | 0.1232 |
| Aedes fluviatilis | 1486 | 40.4 | 39.4 | 8.7 | 11.4 | 79.8 | 0.0125 | 0.1343 |
| Coquillettidia nigricans | 1479 | 41.3 | 39.7 | 8.0 | 11.0 | 81.0 | 0.0198 | 0.1579 |
| Psorophora albipes | 1493 | 39.5 | 39.2 | 9.2 | 12.0 | 78.7 | 0.0038 | 0.1321 |
| Psorophora ferox | 1494 | 40.1 | 39.3 | 8.8 | 11.8 | 79.4 | 0.0101 | 0.1456 |
| rRNAs | ||||||||
| Aedeomyia squamipennis | 2160 | 38.9 | 43.8 | 6.0 | 11.3 | 82.7 | −0.0593 | 0.3064 |
| Aedes fluviatilis | 2151 | 39.1 | 43.6 | 6.1 | 11.2 | 82.7 | −0.0544 | 0.2948 |
| Coquillettidia nigricans | 2097 | 39.6 | 42.7 | 5.4 | 12.2 | 82.3 | −0.0377 | 0.3864 |
| Psorophora albipes | 2154 | 39.3 | 42.7 | 6.3 | 11.7 | 82.0 | −0.0415 | 0.3000 |
| Psorophora ferox | 2160 | 39.4 | 42.6 | 6.3 | 11.7 | 82.0 | −0.0390 | 0.3000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
do Nascimento, B.L.S.; da Silva, F.S.; Nunes-Neto, J.P.; de Almeida Medeiros, D.B.; Cruz, A.C.R.; da Silva, S.P.; da Silva e Silva, L.H.; de Oliveira Monteiro, H.A.; Dias, D.D.; Vieira, D.B.R.; et al. First Description of the Mitogenome and Phylogeny of Culicinae Species from the Amazon Region. Genes 2021, 12, 1983. https://doi.org/10.3390/genes12121983
do Nascimento BLS, da Silva FS, Nunes-Neto JP, de Almeida Medeiros DB, Cruz ACR, da Silva SP, da Silva e Silva LH, de Oliveira Monteiro HA, Dias DD, Vieira DBR, et al. First Description of the Mitogenome and Phylogeny of Culicinae Species from the Amazon Region. Genes. 2021; 12(12):1983. https://doi.org/10.3390/genes12121983
Chicago/Turabian Styledo Nascimento, Bruna Laís Sena, Fábio Silva da Silva, Joaquim Pinto Nunes-Neto, Daniele Barbosa de Almeida Medeiros, Ana Cecília Ribeiro Cruz, Sandro Patroca da Silva, Lucas Henrique da Silva e Silva, Hamilton Antônio de Oliveira Monteiro, Daniel Damous Dias, Durval Bertram Rodrigues Vieira, and et al. 2021. "First Description of the Mitogenome and Phylogeny of Culicinae Species from the Amazon Region" Genes 12, no. 12: 1983. https://doi.org/10.3390/genes12121983






