A Deep-Learning Approach to ECG Classification Based on Adversarial Domain Adaptation
Abstract
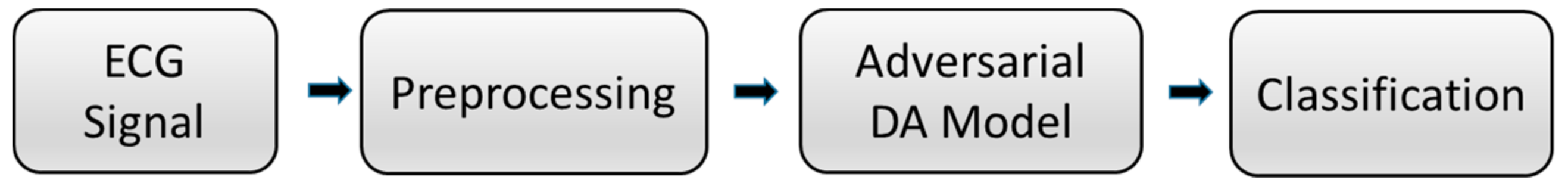
:1. Introduction
- (1)
- A new ECG signal classification method based on adversarial domain adaptive learning is proposed to solve the problem of insufficient labeled training samples and improve the phenomenon of different data distribution caused by individual differences.
- (2)
- Each component of this method, the F, D, and C modules, is optimized, respectively, to improve the feature diversity and feature abstraction.
- (3)
- Six crucial time features are proposed and concatenated with deep-learning features to increase the richness of the features.
2. Materials and Data Preprocessing
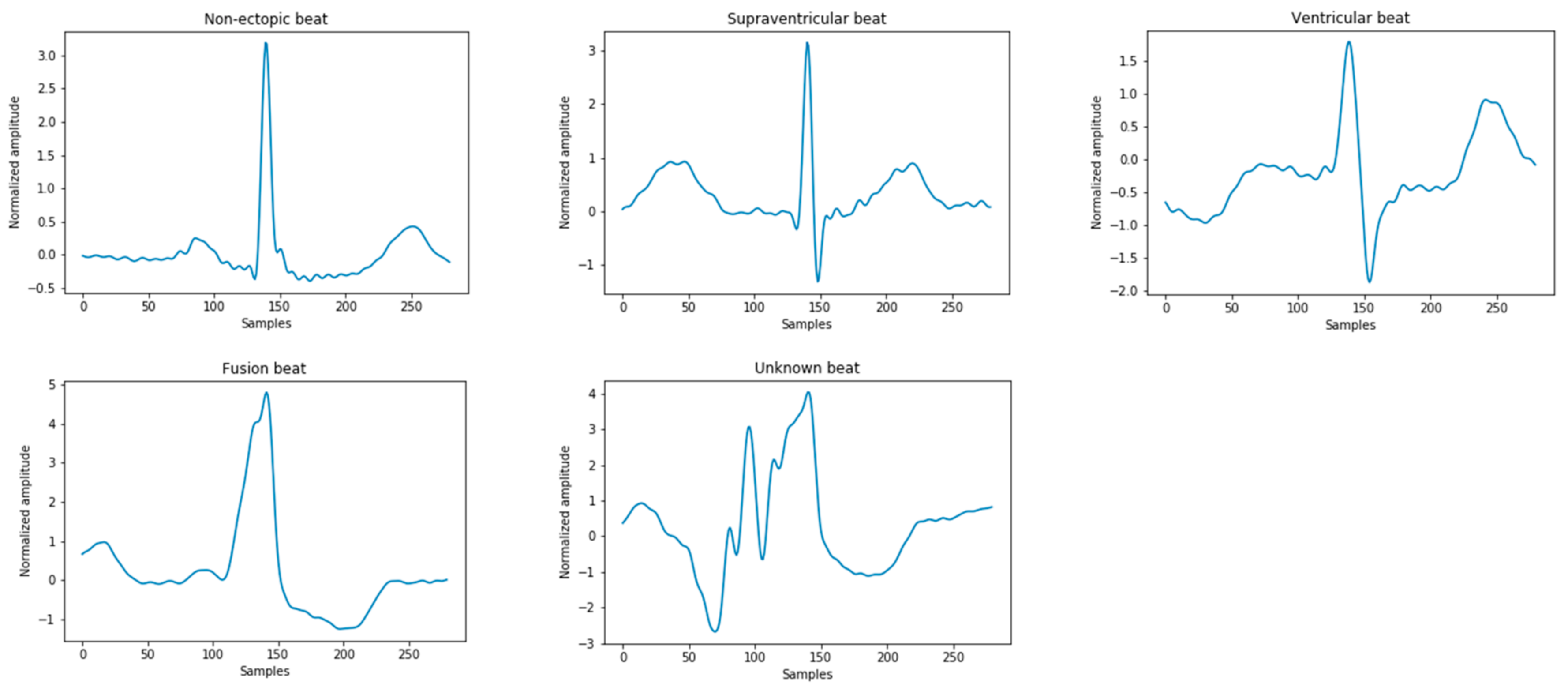
2.1. ECG Dataset
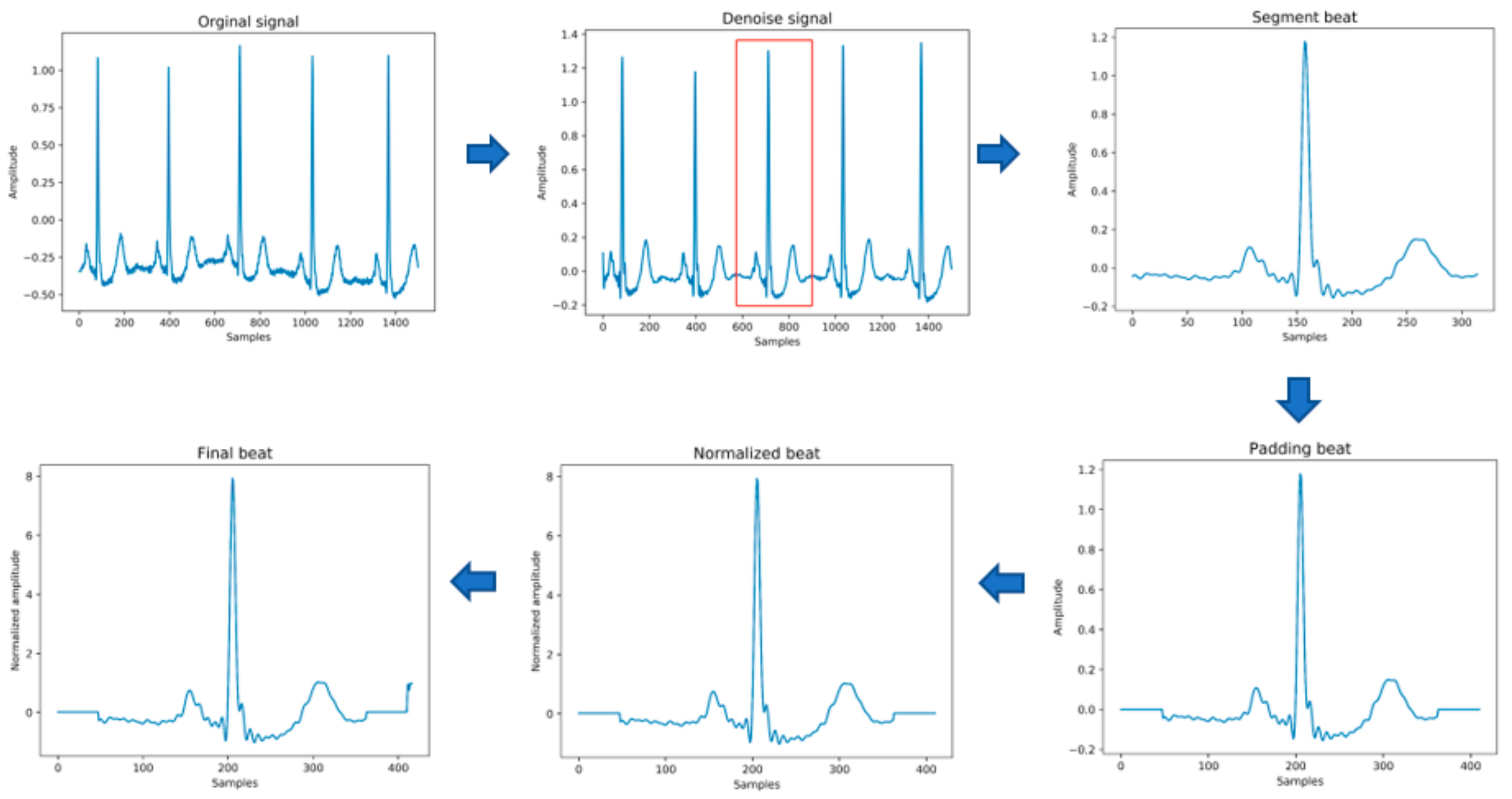
2.2. Data Preprocessing
- (1)
- Data denoising: Use the band pass filter F_{band} with a cutoff frequency of (0.5,40) and the discrete wavelet transform (DWT) with a basis function of db6 to eliminate the influence of electrode artifact noise (EMG), muscle artifact noise (MA) and baseline wander noise (BW) on the ECG signal.
- (2)
- Heartbeat segmentation: First, read the R peak position provided in the heartbeat label. Suppose is the position of the ith heartbeat R peak. The start position of the heartbeat is , and the end position of the heartbeat is , where represents rounding down to n. Then, the current number of heartbeat sampling points is .
- (3)
- Heartbeat alignment: After the heartbeats are divided, the number of sampling points for each heartbeat becomes different. In order to pass into the deep-learning model, the heartbeats must be aligned. Suppose the number of sampling points after unification is D; if is less than D, then fill with zero to D, and if is greater than D, then crop to D [28], and the heartbeat after final processing is . In the experiment of this paper, D is 411.
- (4)
- Data standardization: The aligned heartbeats are standardized by the formula for the Z-score to eliminate offset and amplitude scaling problems in the signal.
- (5)
- Extraction of time features: Manually extract six time features, including the previous RR intervals of the current heartbeats, the post-RR intervals, the local RR intervals lasting 10s, the average RR intervals of the entire record, and using the formula , to generate the normalized previous RR and normalized post-RR after normalization operation.
- (6)
- Data augmentation: In order to overcome the problem of imbalance in the number of samples in the different categories, the SMOTE algorithm is used to generate different categories of data to balance the dataset.
3. Methodology
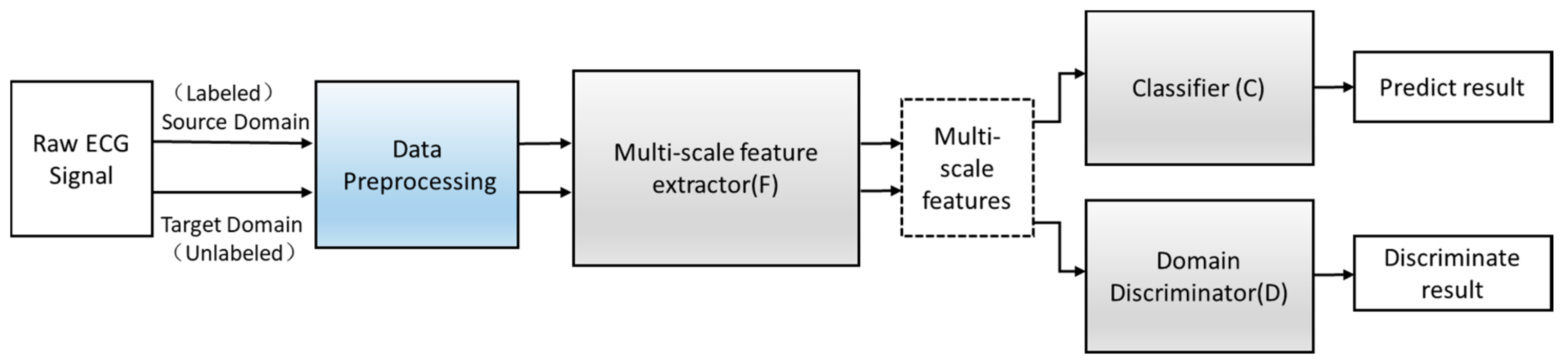
3.1. Model Architecture
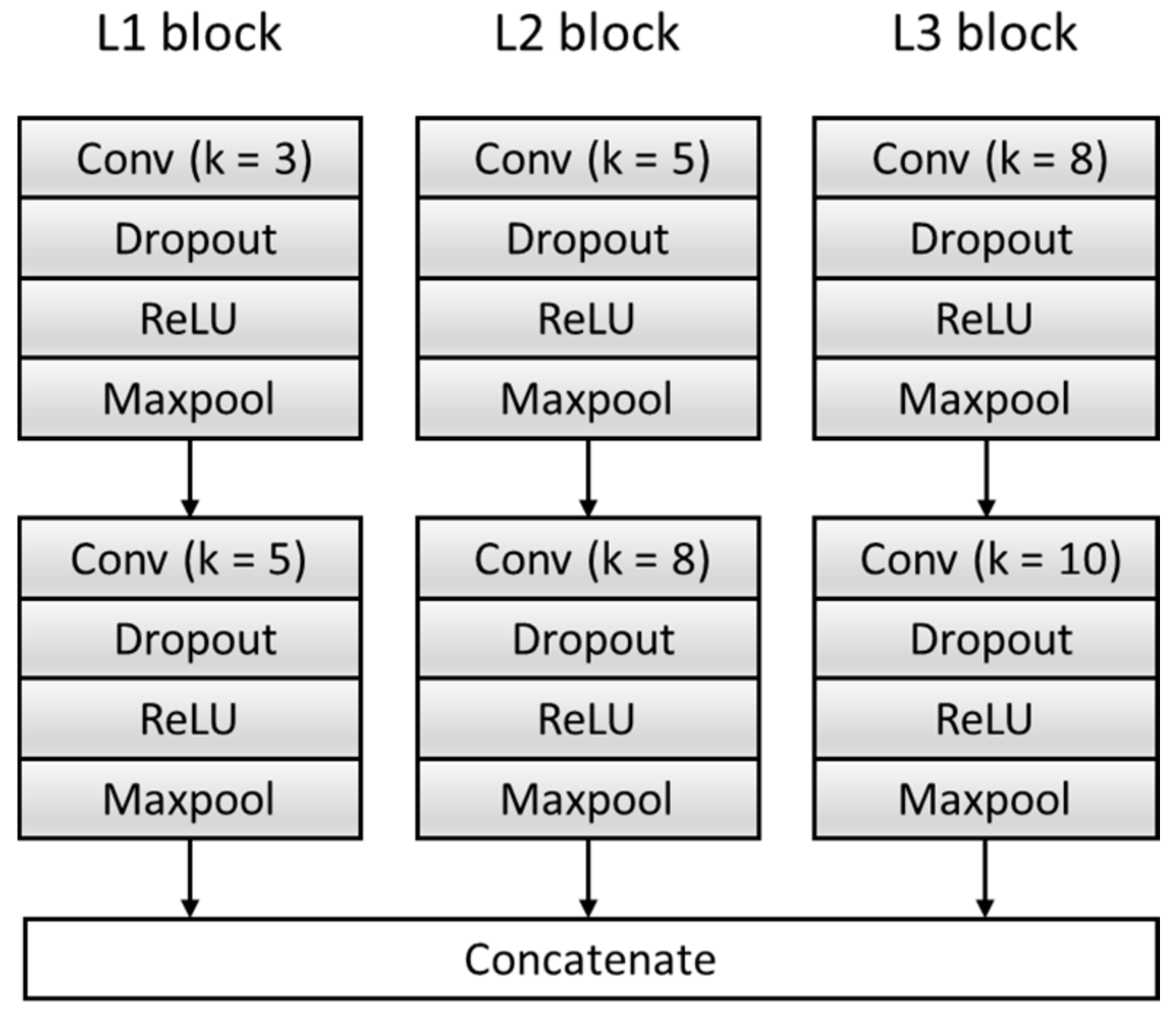
3.1.1. Multi-Scale Feature Extraction F
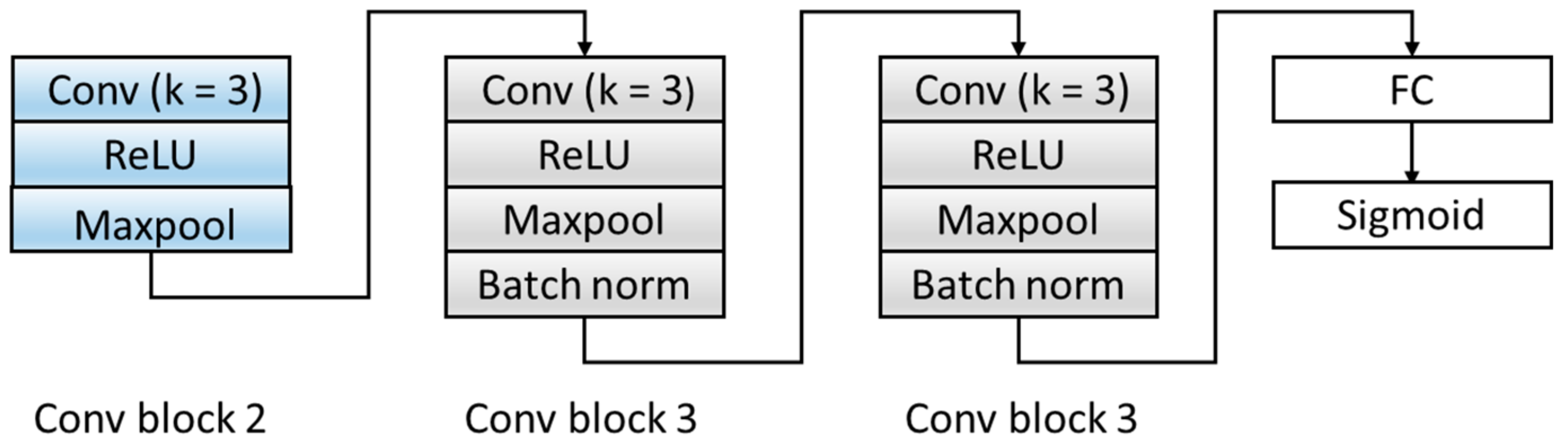
3.1.2. Domain Discrimination D
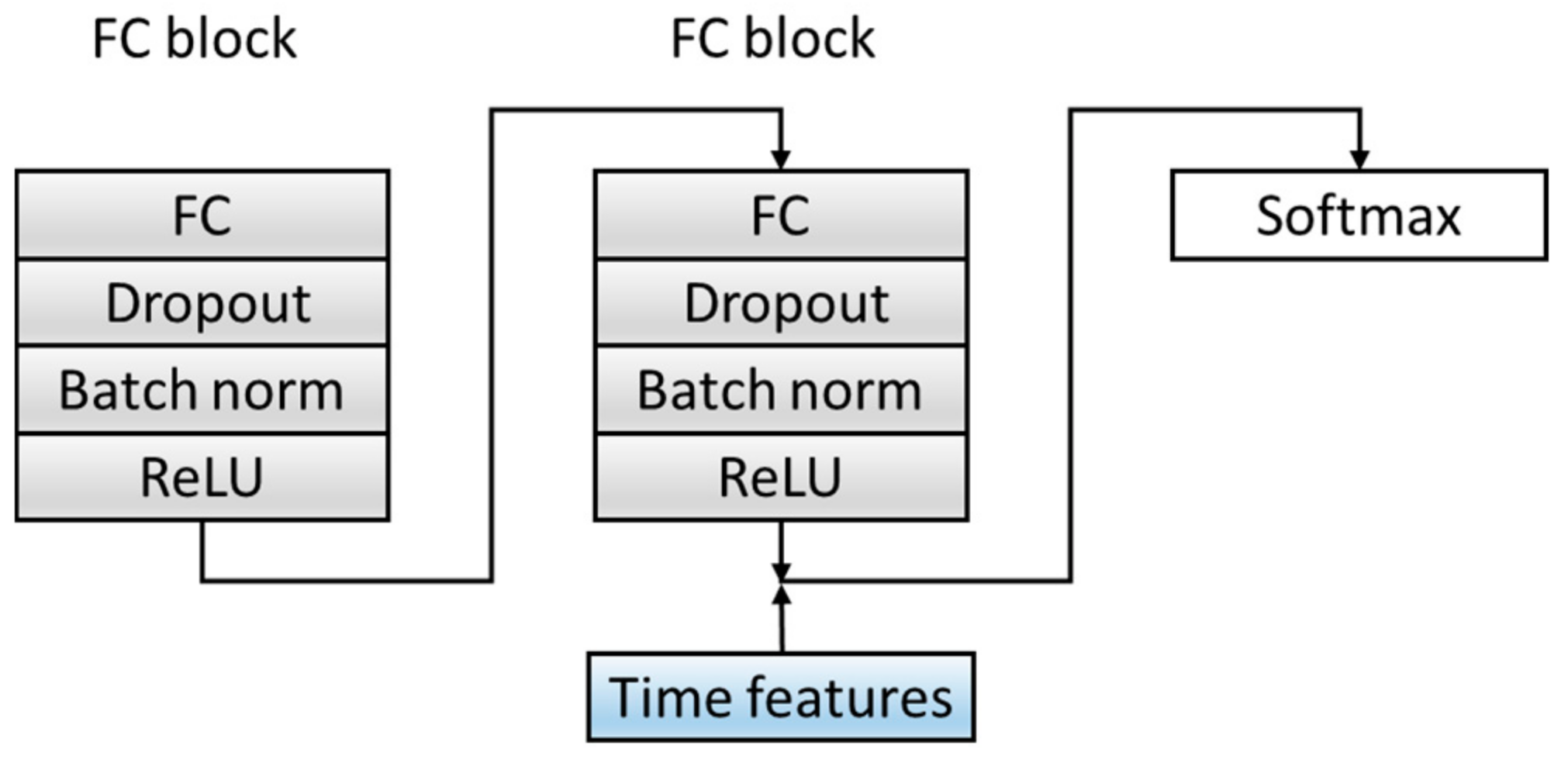
3.1.3. Classification C
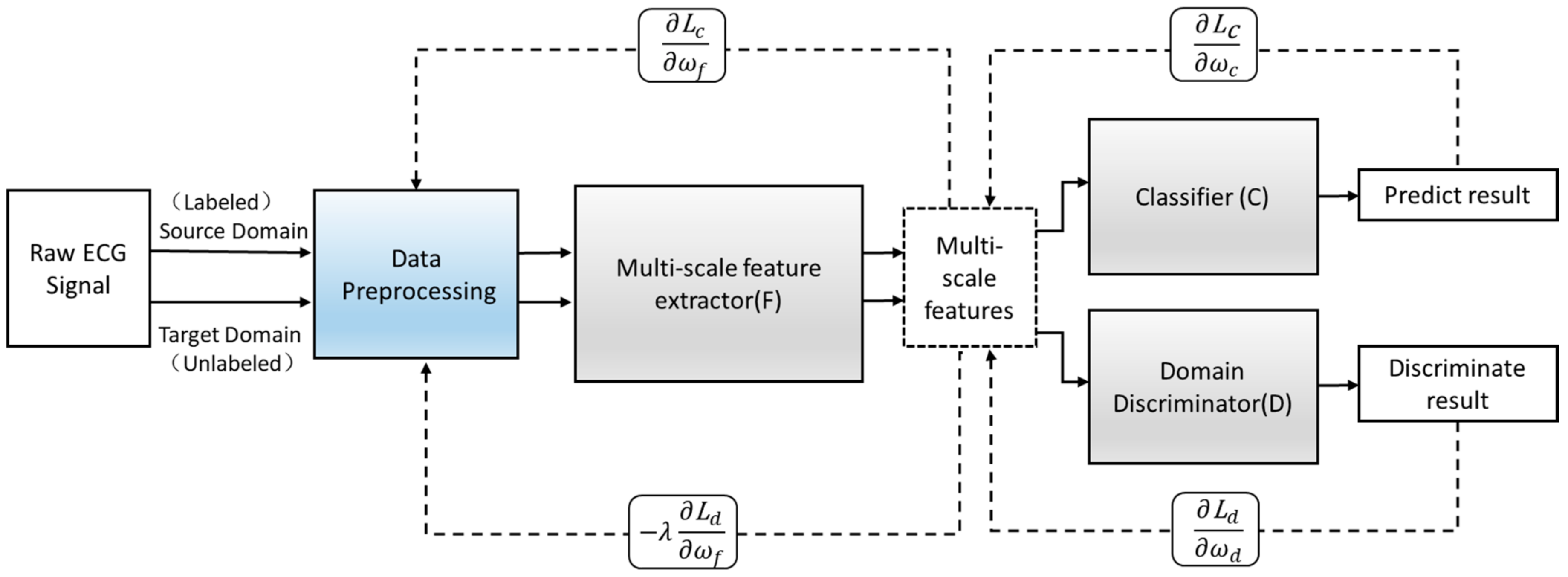
3.2. Training Process
- (1)
- The first step is to keep the parameters of the domain discrimination module unchanged and calculated by Formula (2) to maximize the loss of the domain discrimination module to update the parameters of the multi-scale feature extraction module to obtain the domain-invariant features. In this way, the invariant features can be fully obtained, which can summarize the source domain data and target domain data at the same time. Minimize the loss of the classification module to update the parameters of the classification module to obtain a classifier that accurately predicts the label. , ,and are the parameter values of the saddle point , , and , respectively.
- (2)
- The second step is to fix the parameters and , and keep them unchanged. Use Formula (3) to minimize the loss of the domain discrimination module and update the parameters of the domain discrimination module to obtain a strong discriminator that can distinguish the source of the feature.
- (3)
- The third step is to repeat the operation of the first step, the parameters of the fixed domain discrimination module are unchanged, and the feature extraction module and the classification module are trained through Equation (2). Use this training process and update the parameters alternately.
- (4)
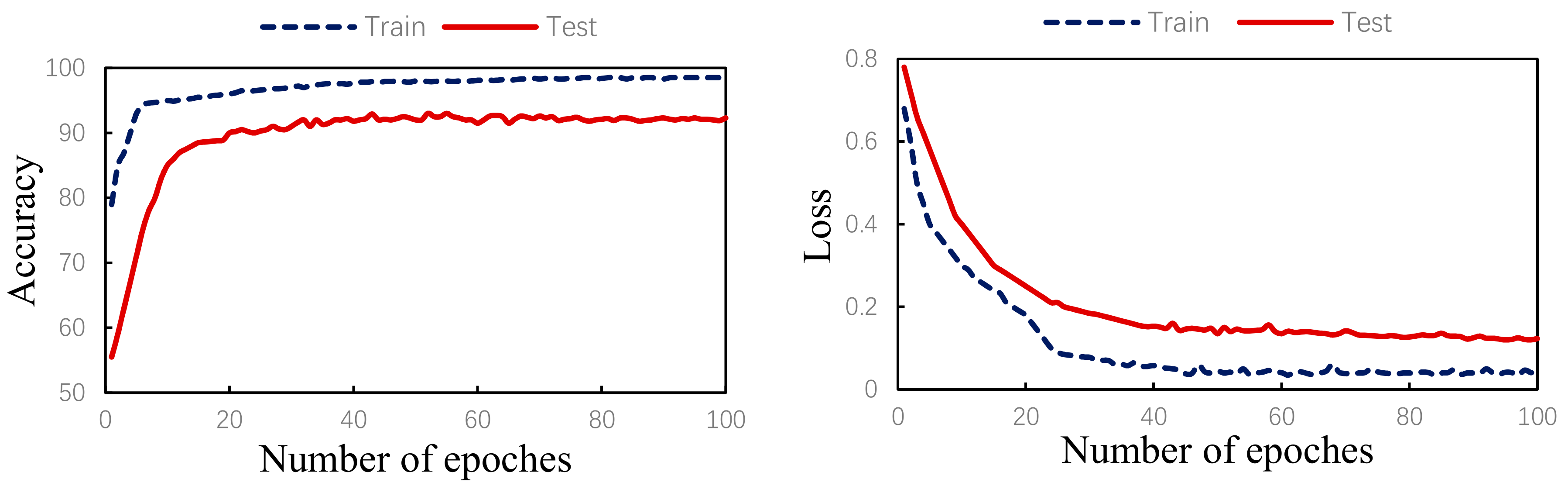
- In the end, the network maintains a dynamic balance. After reaching the preset number of iterations, the optimal value is obtained, and the optimal model is saved. Input the new heartbeat sample into the saved optimal model to obtain the final classification result. The training process is shown in Figure 8.
4. Results and Discussion
4.1. Experimental Setting
4.2. Performance Metrics
4.3. Experiments Results and Discussion
4.3.1. Experiment 1: Algorithm Validity
4.3.2. Experiment 2: Comparison of Different Models
- (1)
- Single-scale Model A. Among them, Model A uses three convolutional layers, three pooling layers, and two fully connected layers to build. The kernel size of the convolutional layer is 3.
- (2)
- Model A+RR. The six time features proposed in Section 2.2 are added to the last fully connected layer of Model A, and the stitched overall features are input into the Softmax classifier for classification.
- (3)
- The model proposed in the paper. During the experiment, with the use of the same input data, the comparison of the classification results is shown in Table 7 below.
4.3.3. Experiment 3: Comparison of Other Paper Methods
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- World Health Organization. Cardiovascular Diseases (CVDs). Available online: http://www.who.int/mediacentre/factsheets/fs317/en/ (accessed on 10 September 2017).
- Shi, H.; Wang, H.; Zhang, F.; Huang, Y.; Zhao, L.; Liu, C. Inter-patient heartbeat classification based on region feature extraction and ensemble classifier. Biomed. Signal Process. Control 2019, 51, 97–105. [Google Scholar] [CrossRef]
- Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-time patient-specific ECG classification by 1-D convolutional neural networks. IEEE Trans. Biomed. Eng. 2016, 63, 664–675. [Google Scholar] [CrossRef] [PubMed]
- Minami, K.; Nakajima, H.; Toyoshima, T. Real-time discrimination of ventricular tachyarrhythmia with Fourier-transform neural network. IEEE Trans. Biomed. Eng. 1999, 46, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Inan, O.T.; Giovangrandi, L.; Kovacs, G.T. Robust neural-network-based classification of premature ventricular contractions using wavelet transform and timing interval features. IEEE Trans. Biomed. Eng. 2006, 53, 2507–2515. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Sudarshan, V.K.; Koh, J.E.; Martis, R.J.; Tan, J.H.; Oh, S.L.; Muhammad, A.; Hagiwara, Y.; Mookiah, M.R.K.; Chua, K.P.; et al. Application of higher order spectra for the characterization of coronary artery disease using electrocardiogram signals. Biomed. Signal Process. Control 2017, 31, 31–43. [Google Scholar] [CrossRef]
- Melgani, F.; Bazi, Y. Classification of electrocardiogram signals with support vector machines and particle swarm optimization. IEEE Trans. Inf. Technol. Biomed. 2008, 12, 667–677. [Google Scholar] [CrossRef] [PubMed]
- Martis, R.J.; Acharya, U.R.; Prasad, H.; Chua, C.K.; Lim, C.M.; Suri, J.S. Application of higher order statistics for atrial arrhythmia classification. Biomed. Signal Process. Control 2013, 8, 888–900. [Google Scholar] [CrossRef]
- Acharya, U.R.; Fujita, H.; Adam, M.; Lih, O.S.; Hong, T.J.; Sudarshan, V.K.; Koh, J.E. Automated Characterization of Arrhythmias Using Nonlinear Features from Tachycardia ECG Beats. In Proceedings of the IEEE International Conference on Systems Man, and Cybernetics, Budapest, Hungary, 9–12 October 2016; IEEE: New York, NY, USA, 2016; pp. 533–538. [Google Scholar]
- Ye, C.; Kumar, B.V.K.V.; Coimbra, M.T. Heartbeat classification using morphological and dynamic features of ECG signals. IEEE Trans. Biomed. Eng. 2012, 59, 2930–2941. [Google Scholar] [CrossRef]
- Shi, H.; Wang, H.; Huang, Y.; Zhao, L.; Qin, C.; Liu, C. A hierarchical method based on weighted extreme gradient boosting in ECG heartbeat classification. Comput. Methods Programs Biomed. 2019, 171, 1–10. [Google Scholar] [CrossRef]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M.; Gertych, A.; Tan, R.S. A deep convolutional neural network model to classify heartbeats. Comput. Biol. Med. 2017, 89, 389–396. [Google Scholar] [CrossRef]
- Yildirim, Ö. A novel wavelet sequence based on deep bidirectional LSTM network model for ECG signal classification. Comput. Biol. Med. 2018, 96, 189–202. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.; Wang, H.; Lu, W. A noval two-lead arrhythmia classification system based on cnn and lstm. J. Mech. Med. Biol. 2019, 19, 1950004. [Google Scholar] [CrossRef]
- Mathews, S.M.; Kambhamettu, C.; Barner, K.E. A novel application of deep learning for single-lead ECG classification. Comput. Biol. Med. 2018, 99, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Sellami, A.; Hwang, H. A robust deep convolutional neural network with batch-weighted loss for heartbeat classification. Expert Syst. Appl. 2019, 122, 75–84. [Google Scholar] [CrossRef]
- Wang, H.; Shi, H.; Chen, X.; Zhao, L.; Huang, Y.; Liu, C. An Improved Convolutional Neural Network Based Approach for Automated Heartbeat Classification. J. Med. Syst. 2020, 44, 35. [Google Scholar] [CrossRef] [PubMed]
- Golrizkhatami, Z.; Acan, A. ECG classification using three-level fusion of different feature descriptors. Expert Syst. Appl. 2018, 114, 54–64. [Google Scholar] [CrossRef]
- Association for the Advancement of Medical Instrumentation; American National Standards Institute. Testing and Reporting Performance Results of Cardiac Rhythm and ST Segment Measurement Algorithms; AAMI: Arlington, VA, USA, 2012. [Google Scholar]
- Nurmaini, S.; Partan, R.U.; Caesarendra, W.; Dewi, T.; Rachmatullah, M.N.; Darmawahyuni, A.; Bhayyu, V.; Firdaus, F. An Automated ECG Beat Classification System Using Deep Neural Networks with an Unsupervised Feature Extraction Technique. Appl. Sci. 2019, 9, 2921. [Google Scholar] [CrossRef] [Green Version]
- Rajpurkar, P.; Hannun, A.Y.; Haghpanahi, M.; Bourn, C.; Ng, A.Y. Cardiologist-level arrhythmia detection with convolutional neural networks. arXiv 2017, arXiv:1707.01836. [Google Scholar]
- Yıldırım, Ö.; Pławiak, P.; Tan, R.S.; Acharya, U.R. Arrhythmia detection using deep convolutional neural network with long duration ECG signals. Comput. Biol. Med. 2018, 102, 411–420. [Google Scholar] [CrossRef]
- De Chazal, P.; O’Dwyer, M.; Reilly, R.B. Automatic classification of heartbeats using ECG morphology and heartbeat interval features. IEEE Trans. Biomed. Eng. 2004, 51, 1196–1206. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Hua, W.; Li, Z.; Li, J.; Gao, X. Heartbeat classification using projected and dynamic features of ECG signal. Biomed. Signal Process. Control 2017, 31, 165–173. [Google Scholar] [CrossRef]
- Li, T.; Zhou, M. ECG classification using wavelet packet entropy and random forests. Entropy 2016, 18, 285. [Google Scholar] [CrossRef]
- Association for the Advancement of Medical Instrumentation; American National Standards Institute. Testing and Reporting Performance Results of Cardiac Rhythm and ST-Segment Measurement Algorithms; EC57; AAMI: Arlington, VA, USA, 1998. [Google Scholar]
- Mar, T.; Zaunseder, S.; Martínez, J.P.; Llamedo, M.; Poll, R. Optimization of ECG classification by means of feature selection. IEEE Trans. Biomed. Eng. 2011, 58, 2168–2177. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.S.; Mak, M.W.; Cheung, C.C. Towards End-to-End ECG Classification With Raw Signal Extraction and Deep Neural Networks. IEEE J. Biomed. Health Inform. 2019, 23, 1574–1584. [Google Scholar] [CrossRef]

| N | S | V | F | Q |
|---|---|---|---|---|
| Normal | Atrial premature | Premature ventricular contraction | Fusion of ventricular and normal | Paced |
| Left bundle branch block | Aberrant atrial premature | Ventricular escape | Fusion of paced and normal | |
| Right bundle branch block | Nodal(junctional) premature | Unclassifiable | ||
| Atrial escape | Supra-ventricular premature | |||
| Nodal (junctional) escape |
| Datasets | Number of Heartbeats | Total | |||
|---|---|---|---|---|---|
| N | S | V | F | ||
| DS1 | 45824 | 943 | 3787 | 414 | 50,968 |
| DS2 | 44213 | 1836 | 3219 | 388 | 49,656 |
| Total | 90,037 | 2779 | 7006 | 802 | 100,624 |
| Block | Layers | Output Size |
|---|---|---|
| Input (preprocessed ECG signal) | 411 × 1 | |
| L1 block | 204 × 16 | |
| L1 block | 100 × 32 | |
| L2 block | 203 × 16 | |
| L2 block | 98 × 32 | |
| L3 block | 202 × 16 | |
| L3 block | 96 × 32 | |
| Concatenate | 9408 × 1 |
| Block | Layers | Output Size |
|---|---|---|
| Input (the output of the F model) | 9408 × 1 | |
| Conv block 2 | 4691 × 6 | |
| Conv block 3 | 2336 × 6 | |
| Conv Block 3 | 1159 × 6 | |
| FC layer (3474,100) | 100 × 1 | |
| Sigmoid | 2 |
| Block | Layers | Output Size |
|---|---|---|
| Input (the output of the F model) | 9408 × 1 | |
| FC block | 100 × 1 | |
| FC block | 10 × 1 | |
| Add(10,6) | 16 × 1 | |
| Softmax | 4 |
| Type | Sen (%) | PPV (%) | Acc (%) |
|---|---|---|---|
| N | 94.0 | 97.4 | 91.8 |
| S | 76.6 | 73.2 | 98.0 |
| V | 85.2 | 57.8 | 93.5 |
| F | 38.4 | 44.9 | 99.0 |
| Average | 73.6 | 68.3 | 95.5 |
| Model | Acc | N (Sen, PPV) | S (Sen, PPV) | V (Sen, PPV) | F (Sen, PPV) |
|---|---|---|---|---|---|
| Model A | 89.0 | 90.1, 97.6 | 63.5, 35.7 | 90.4, 54.6 | 20.6, 28.4 |
| Model A+RR | 91.3 | 92.5, 97.8 | 81.9, 59.1 | 88.8, 56.1 | 23.7, 43.8 |
| Proposed | 92.3 | 93.9, 97.4 | 76.6, 73.2 | 85.1, 57.8 | 38.4, 44.9 |
| Method | Year | Acc (%) | N | S | V | F | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sen (%) | PPV (%) | Sen (%) | PPV (%) | Sen (%) | PPV (%) | Sen (%) | PPV (%) | |||
| Chazal | 2004 | 85.8 | 86.8 | 99.1 | 75.9 | 38.5 | 77.7 | 81.9 | 68.2 | 26.5 |
| Ye | 2012 | 86.4 | 88.6 | 97.5 | 60.8 | 52.3 | 81.5 | 63.1 | 19.5 | 2.5 |
| Mathews | 2018 | 74.81 | 73.89 | 99.66 | 88.39 | 33.63 | 77.24 | 69.20 | 73.7 | 4.67 |
| Sellami | 2019 | 88.34 | 88.51 | 98.80 | 82.04 | 30.44 | 92.05 | 72.13 | 89.4 | 8.6 |
| Shi | 2019 | 92.1 | 92.1 | 99.5 | 91.7 | 46.2 | 95.1 | 88.1 | 61.6 | 15.2 |
| Proposed | - | 92.3 | 93.9 | 97.4 | 76.6 | 73.2 | 85.1 | 57.8 | 38.4 | 44.9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niu, L.; Chen, C.; Liu, H.; Zhou, S.; Shu, M. A Deep-Learning Approach to ECG Classification Based on Adversarial Domain Adaptation. Healthcare 2020, 8, 437. https://doi.org/10.3390/healthcare8040437
Niu L, Chen C, Liu H, Zhou S, Shu M. A Deep-Learning Approach to ECG Classification Based on Adversarial Domain Adaptation. Healthcare. 2020; 8(4):437. https://doi.org/10.3390/healthcare8040437
Chicago/Turabian StyleNiu, Lisha, Chao Chen, Hui Liu, Shuwang Zhou, and Minglei Shu. 2020. "A Deep-Learning Approach to ECG Classification Based on Adversarial Domain Adaptation" Healthcare 8, no. 4: 437. https://doi.org/10.3390/healthcare8040437





