Epigenetic Mechanisms of Tree Responses to Climatic Changes
Abstract
1. Introduction
2. Plant Epigenetic Regulation Mechanism
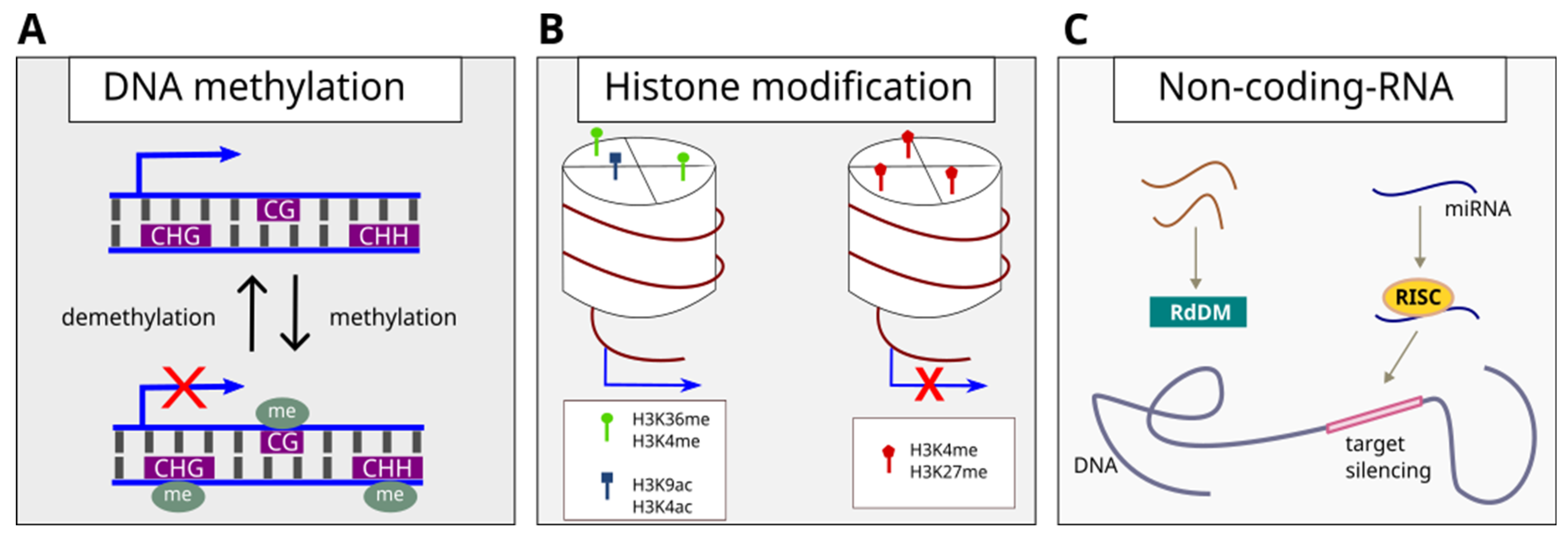
3. Epigenetics in Tree Development
| Environmental Stress | Plant Species | Epigenetic Regulation | Plant Probes | Reference |
|---|---|---|---|---|
| temperature | Picea abies | DNA methylation; small non-coding RNAs (miRNA and ta-siRNA) | grafts; embryogenetic cultures | [13,95] |
| Populus tremula×alba | DNA methylation; DML-dependent demethylation | adult trees | [96] | |
| Castanea sativa | gDNA methylation | 3-year-old trees; shoots from trunk base (juvenile phase) | [97,98] | |
| Malus × domestica | DNA methylation | adult trees | [99] | |
| Quercus suber L. | DNA methylation/histone H3 acetylation | 8-month-old plants | [100] | |
| Populus tremula × tremuloides | histone H3 acetylation | Tissue culture-grown plants transferred to soil and grown for six weeks | [101] | |
| Picea abies (L.) Karst. | epigenetic memory (DHNs) | adult trees | [102] | |
| temperature, drought | Populus deltoides × P. nigra | DNA methylation | cuttings | [103,104,105] |
| temperature, photoperiod | Picea abies | DNA methylation | 5-year-old plants | [106] |
| drought/water stress | Quercus ilex | DNA methylation | leaves from the upper part of the canopy | [107] |
| Populus tremula × P. alba | RNAi suppression of DNA methylation | leaves/shoot apices | [108] | |
| Fraxinus mandshurica and Fraxinus americana hybrids | DNA methylation | 2-year-old seedlings | [109] | |
| Populus trichocarpa | histone acetylation (H3K9ac) | 3-month-old plants | [110] | |
| salinity and nutrient variations | Laguncularia racemosa | DNA methylation | adult trees | [111] |
| the length of the day | Picea abies (L.) Karst. | miRNA | seedlings | [112] |
| radiation | Pinus sylvestris | genomic methylation, DNA methylation | plants from seeds | [113] |
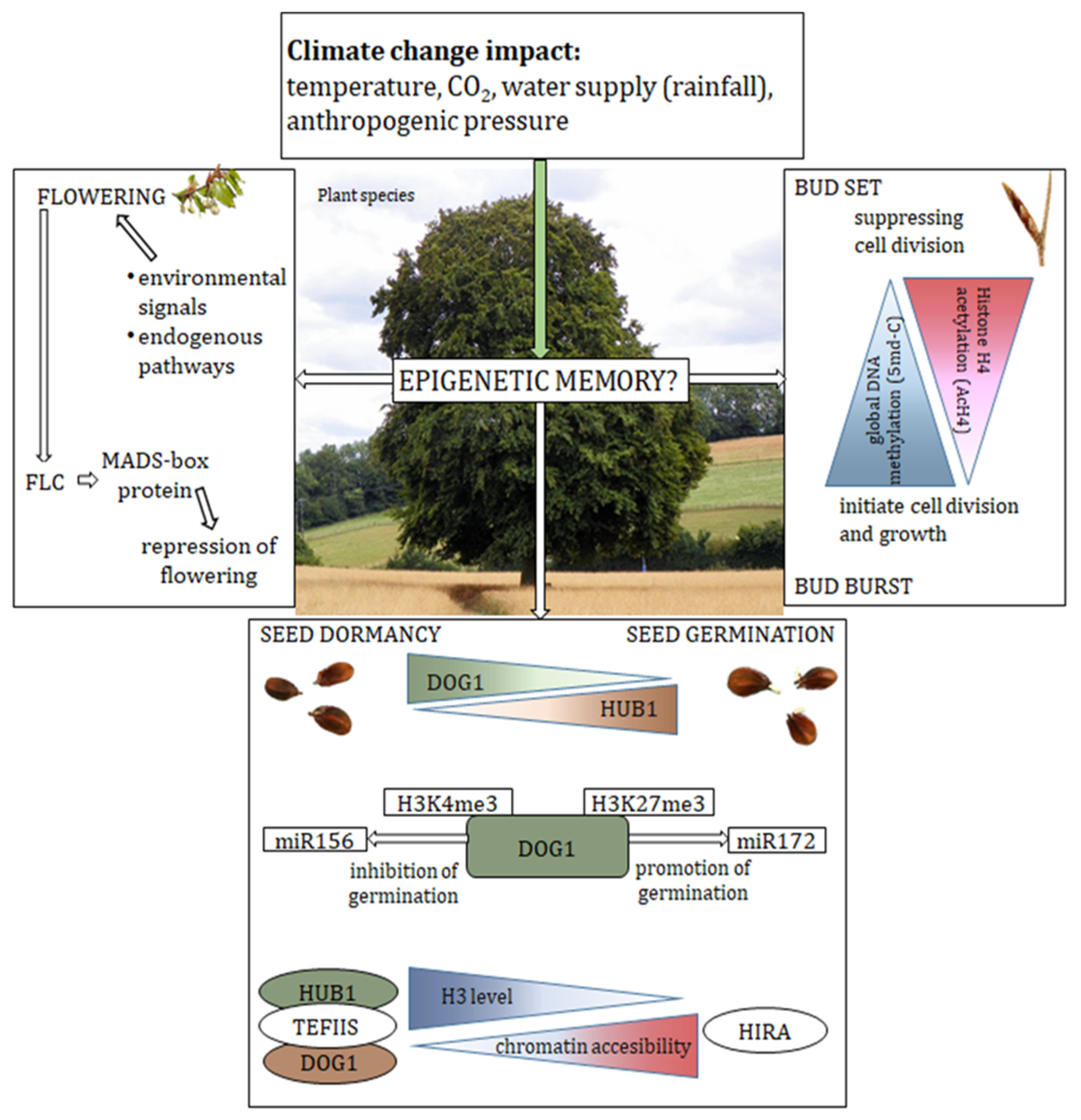
4. Dormancy and Germination of the Seeds
5. Bud Dormancy
6. Flowering
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Fladung, M.; Hoenicka, H. Fifteen Years of Forest Tree Biosafety Research in Germany. iForest 2012, 5, 126–130. [Google Scholar] [CrossRef]
- Cleland, E.; Chuine, I.; Menzel, A.; Mooney, H.; Schwartz, M. Shifting Plant Phenology in Response to Global Change. Trends Ecol. Evol. 2007, 22, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Partanen, J.; Koski, V.; Hanninen, H. Effects of Photoperiod and Temperature on the Timing of Bud Burst in Norway Spruce (Picea abies). Tree Physiol. 1998, 18, 811–816. [Google Scholar] [CrossRef]
- Primack, R.B.; Higuchi, H.; Miller-Rushing, A.J. The Impact of Climate Change on Cherry Trees and Other Species in Japan. Biol. Conserv. 2009, 142, 1943–1949. [Google Scholar] [CrossRef]
- García-García, I.; Méndez-Cea, B.; Martín-Gálvez, D.; Seco, J.I.; Gallego, F.J.; Linares, J.C. Challenges and Perspectives in the Epigenetics of Climate Change-Induced Forests Decline. Front. Plant Sci. 2022, 12, 797958. [Google Scholar] [CrossRef]
- Nicotra, A.B.; Atkin, O.K.; Bonser, S.P.; Davidson, A.M.; Finnegan, E.J.; Mathesius, U.; Poot, P.; Purugganan, M.D.; Richards, C.L.; Valladares, F.; et al. Plant Phenotypic Plasticity in a Changing Climate. Trends Plant Sci. 2010, 15, 684–692. [Google Scholar] [CrossRef]
- Rey, O.; Danchin, E.; Mirouze, M.; Loot, C.; Blanchet, S. Adaptation to Global Change: A Transposable Element–Epigenetics Perspective. Trends Ecol. Evol. 2016, 31, 514–526. [Google Scholar] [CrossRef]
- Staszak, A.M.; Pawlowski, T.A. Forest Tree Research in Post Genomic Era. Introduction to Systems Biology of Broadleaves. Dendrobiology 2012, 68, 113–123. [Google Scholar]
- Klupczyńska, E.A.; Pawłowski, T.A. Regulation of Seed Dormancy and Germination Mechanisms in a Changing Environment. Int. J. Mol. Sci. 2021, 22, 1357. [Google Scholar] [CrossRef]
- Niederhuth, C.E.; Schmitz, R.J. Putting DNA Methylation in Context: From Genomes to Gene Expression in Plants. Biochim. Et Biophys. Acta (BBA)—Gene Regul. Mech. 2017, 1860, 149–156. [Google Scholar] [CrossRef]
- Jablonka, E.; Oborny, B.; Molnár, I.; Kisdi, É.; Hofbauer, J.; Czárán, T. The Adaptive Advantage of Phenotypic Memory in Changing Environments. Phil. Trans. R. Soc. Lond. B 1995, 350, 133–141. [Google Scholar] [CrossRef]
- Richards, C.L.; Alonso, C.; Becker, C.; Bossdorf, O.; Bucher, E.; Colomé-Tatché, M.; Durka, W.; Engelhardt, J.; Gaspar, B.; Gogol-Döring, A.; et al. Ecological Plant Epigenetics: Evidence from Model and Non-model Species, and the Way Forward. Ecol. Lett. 2017, 20, 1576–1590. [Google Scholar] [CrossRef] [PubMed]
- Yakovlev, I.A.; Carneros, E.; Lee, Y.; Olsen, J.E.; Fossdal, C.G. Transcriptional Profiling of Epigenetic Regulators in Somatic Embryos during Temperature Induced Formation of an Epigenetic Memory in Norway Spruce. Planta 2016, 243, 1237–1249. [Google Scholar] [CrossRef] [PubMed]
- Vaillant, I.; Paszkowski, J. Role of Histone and DNA Methylation in Gene Regulation. Curr. Opin. Plant Biol. 2007, 10, 528–533. [Google Scholar] [CrossRef]
- Tessadori, F.; van Driel, R.; Fransz, P. Cytogenetics as a Tool to Study Gene Regulation. Trends Plant Sci. 2004, 9, 147–153. [Google Scholar] [CrossRef]
- Bräutigam, K.; Vining, K.J.; Lafon-Placette, C.; Fossdal, C.G.; Mirouze, M.; Marcos, J.G.; Fluch, S.; Fraga, M.F.; Guevara, M.Á.; Abarca, D.; et al. Epigenetic Regulation of Adaptive Responses of Forest Tree Species to the Environment. Ecol. Evol. 2013, 3, 399–415. [Google Scholar] [CrossRef]
- Boyko, A.; Kovalchuk, I. Epigenetic Regulation of Genome Stability in Plants in Response to Stress. In Epigenetic Memory and Control in Plants; Grafi, G., Ohad, N., Eds.; Signaling and Communication in Plants; Springer: Berlin/Heidelberg, Germany, 2013; Volume 18, pp. 41–56. ISBN 978-3-642-35226-3. [Google Scholar]
- Pien, S.; Grossniklaus, U. Polycomb Group and Trithorax Group Proteins in Arabidopsis. Biochim. Et Biophys. Acta (BBA)—Gene Struct. Expr. 2007, 1769, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Pascual, J.; Cañal, M.J.; Correia, B.; Escandon, M.; Hasbún, R.; Meijón, M.; Pinto, G.; Valledor, L. Can Epigenetics Help Forest Plants to Adapt to Climate Change? In Epigenetics in Plants of Agronomic Importance: Fundamentals and Applications; Springer International Publishing: Cham, Switzerland, 2014; pp. 125–146. ISBN 978-3-319-07970-7. [Google Scholar]
- Lämke, J.; Bäurle, I. Epigenetic and Chromatin-Based Mechanisms in Environmental Stress Adaptation and Stress Memory in Plants. Genome Biol. 2017, 18, 124. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Li, Z. Epigenetic Environmental Memories in Plants: Establishment, Maintenance, and Reprogramming. Trends Genet. 2018, 34, 856–866. [Google Scholar] [CrossRef]
- Tollefsbol, T.O. Transgenerational Epigenetics. In Transgenerational Epigenetics; Elsevier: Amsterdam, The Netherlands, 2014; pp. 1–8. ISBN 978-0-12-405944-3. [Google Scholar]
- Jablonka, E.; Lamb, M.J. Soft Inheritance: Challenging the Modern Synthesis. Genet. Mol. Biol. 2008, 31, 389–395. [Google Scholar] [CrossRef]
- Richards, E.J. Inherited Epigenetic Variation—Revisiting Soft Inheritance. Nat. Rev. Genet. 2006, 7, 395–401. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. The Significance of Responses of the Genome to Challenge. Science 1984, 226, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Hilker, M.; Schwachtje, J.; Baier, M.; Balazadeh, S.; Bäurle, I.; Geiselhardt, S.; Hincha, D.K.; Kunze, R.; Mueller-Roeber, B.; Rillig, M.C.; et al. Priming and Memory of Stress Responses in Organisms Lacking a Nervous System: Priming and Memory of Stress Responses. Biol. Rev. 2016, 91, 1118–1133. [Google Scholar] [CrossRef] [PubMed]
- Mauch-Mani, B.; Baccelli, I.; Luna, E.; Flors, V. Defense Priming: An Adaptive Part of Induced Resistance. Annu. Rev. Plant Biol. 2017, 68, 485–512. [Google Scholar] [CrossRef]
- Martinez-Medina, A.; Flors, V.; Heil, M.; Mauch-Mani, B.; Pieterse, C.M.J.; Pozo, M.J.; Ton, J.; van Dam, N.M.; Conrath, U. Recognizing Plant Defense Priming. Trends Plant Sci. 2016, 21, 818–822. [Google Scholar] [CrossRef]
- Walter, J.; Jentsch, A.; Beierkuhnlein, C.; Kreyling, J. Ecological Stress Memory and Cross Stress Tolerance in Plants in the Face of Climate Extremes. Environ. Exp. Bot. 2013, 94, 3–8. [Google Scholar] [CrossRef]
- Tronick, E.; Hunter, R.G. Waddington, Dynamic Systems, and Epigenetics. Front. Behav. Neurosci. 2016, 10, 107. [Google Scholar] [CrossRef]
- Bruce, T.J.A.; Matthes, M.C.; Napier, J.A.; Pickett, J.A. Stressful “Memories” of Plants: Evidence and Possible Mechanisms. Plant Sci. 2007, 173, 603–608. [Google Scholar] [CrossRef]
- Crisp, P.A.; Ganguly, D.; Eichten, S.R.; Borevitz, J.O.; Pogson, B.J. Reconsidering Plant Memory: Intersections between Stress Recovery, RNA Turnover, and Epigenetics. Sci. Adv. 2016, 2, e1501340. [Google Scholar] [CrossRef]
- Waddington, C.H. Canalization of development and the inheritance of acquired characters. Nature 1942, 150, 563–565. [Google Scholar] [CrossRef]
- Bird, A. Perceptions of Epigenetics. Nature 2007, 447, 396–398. [Google Scholar] [CrossRef] [PubMed]
- Grant-Downton, R.T.; Dickinson, H.G. Epigenetics and Its Implications for Plant Biology. 1. The Epigenetic Network in Plants. Ann. Bot. 2005, 96, 1143–1164. [Google Scholar] [CrossRef]
- Berger, S.L. The Complex Language of Chromatin Regulation during Transcription. Nature 2007, 447, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lang, Z.; Zhu, J.-K. Dynamics and Function of DNA Methylation in Plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, Maintaining and Modifying DNA Methylation Patterns in Plants and Animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- O’Brown, Z.K.; Greer, E.L. N6-Methyladenine: A Conserved and Dynamic DNA Mark. In DNA Methyltransferases—Role and Function; Advances in Experimental Medicine and Biology; Jeltsch, A., Jurkowska, R.Z., Eds.; Springer International Publishing: Cham, Switzerland, 2016; Volume 945, pp. 213–246. ISBN 978-3-319-43622-7. [Google Scholar]
- Bewick, A.J.; Niederhuth, C.E.; Ji, L.; Rohr, N.A.; Griffin, P.T.; Leebens-Mack, J.; Schmitz, R.J. The Evolution of CHROMOMETHYLASES and Gene Body DNA Methylation in Plants. Genome Biol. 2017, 18, 65. [Google Scholar] [CrossRef]
- Amaral, J.; Ribeyre, Z.; Vigneaud, J.; Sow, M.D.; Fichot, R.; Messier, C.; Pinto, G.; Nolet, P.; Maury, S. Advances and Promises of Epigenetics for Forest Trees. Forests 2020, 11, 976. [Google Scholar] [CrossRef]
- Bewick, A.J.; Ji, L.; Niederhuth, C.E.; Willing, E.-M.; Hofmeister, B.T.; Shi, X.; Wang, L.; Lu, Z.; Rohr, N.A.; Hartwig, B.; et al. On the Origin and Evolutionary Consequences of Gene Body DNA Methylation. Proc. Natl. Acad. Sci. USA 2016, 113, 9111–9116. [Google Scholar] [CrossRef]
- Du, J.; Johnson, L.M.; Jacobsen, S.E.; Patel, D.J. DNA Methylation Pathways and Their Crosstalk with Histone Methylation. Nat. Rev. Mol. Cell Biol. 2015, 16, 519–532. [Google Scholar] [CrossRef]
- Biswas, S.; Rao, C.M. Epigenetic Tools (The Writers, The Readers and The Erasers) and Their Implications in Cancer Therapy. Eur. J. Pharmacol. 2018, 837, 8–24. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of Chromatin by Histone Modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef]
- Varga-Weisz, P. Chromatin Remodeling Factors and DNA Replication. In Epigenetics and Chromatin; Progress in Molecular and Subcellular Biology; Jeanteur, P., Ed.; Springer: Berlin/Heidelberg, Germany, 2005; Volume 38, pp. 1–30. ISBN 978-3-540-23372-5. [Google Scholar]
- Pu, L.; Sung, Z.R. PcG and TrxG in Plants—Friends or Foes. Trends Genet. 2015, 31, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Lu, F.; Cui, X.; Cao, X. Histone Methylation in Higher Plants. Annu. Rev. Plant Biol. 2010, 61, 395–420. [Google Scholar] [CrossRef]
- Schuettengruber, B.; Martinez, A.-M.; Iovino, N.; Cavalli, G. Trithorax Group Proteins: Switching Genes on and Keeping Them Active. Nat. Rev. Mol. Cell Biol. 2011, 12, 799–814. [Google Scholar] [CrossRef] [PubMed]
- Audergon, P.N.C.B.; Catania, S.; Kagansky, A.; Tong, P.; Shukla, M.; Pidoux, A.L.; Allshire, R.C. Restricted Epigenetic Inheritance of H3K9 Methylation. Science 2015, 348, 132–135. [Google Scholar] [CrossRef]
- Köhler, C.; Hennig, L. Regulation of Cell Identity by Plant Polycomb and Trithorax Group Proteins. Curr. Opin. Genet. Dev. 2010, 20, 541–547. [Google Scholar] [CrossRef] [PubMed]
- Metzger, E.; Wissmann, M.; Yin, N.; Müller, J.M.; Schneider, R.; Peters, A.H.F.M.; Günther, T.; Buettner, R.; Schüle, R. LSD1 Demethylates Repressive Histone Marks to Promote Androgen-Receptor-Dependent Transcription. Nature 2005, 437, 436–439. [Google Scholar] [CrossRef]
- Marmorstein, R.; Trievel, R.C. Histone Modifying Enzymes: Structures, Mechanisms, and Specificities. Biochim. Et Biophys. Acta (BBA)—Gene Regul. Mech. 2009, 1789, 58–68. [Google Scholar] [CrossRef]
- Tsukada, Y.; Fang, J.; Erdjument-Bromage, H.; Warren, M.E.; Borchers, C.H.; Tempst, P.; Zhang, Y. Histone Demethylation by a Family of JmjC Domain-Containing Proteins. Nature 2006, 439, 811–816. [Google Scholar] [CrossRef]
- Allfrey, V.G.; Faulkner, R.; Mirsky, A.E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl. Acad. Sci. USA 1964, 51, 786–794. [Google Scholar] [CrossRef]
- Luo, M.; Liu, X.; Singh, P.; Cui, Y.; Zimmerli, L.; Wu, K. Chromatin Modifications and Remodeling in Plant Abiotic Stress Responses. Biochim. Et Biophys. Acta (BBA)—Gene Regul. Mech. 2012, 1819, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.J.; Tian, L. Roles of Dynamic and Reversible Histone Acetylation in Plant Development and Polyploidy. Biochim. Et Biophys. Acta (BBA)—Gene Struct. Expr. 2007, 1769, 295–307. [Google Scholar] [CrossRef] [PubMed]
- Loury, R.; Sassone-Corsi, P. Analysis of Histone Phosphorylation: Coupling Intracellular Signaling to Chromatin Remodeling. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2003; Volume 377, pp. 197–212. ISBN 978-0-12-182781-6. [Google Scholar]
- Prigent, C.; Dimitrov, S. Phosphorylation of Serine 10 in Histone H3, What for? J. Cell Sci. 2003, 116, 3677–3685. [Google Scholar] [CrossRef] [PubMed]
- Pfluger, J.; Wagner, D. Histone Modifications and Dynamic Regulation of Genome Accessibility in Plants. Curr. Opin. Plant Biol. 2007, 10, 645–652. [Google Scholar] [CrossRef] [PubMed]
- Hicke, L. Protein Regulation by Monoubiquitin. Nat. Rev. Mol. Cell Biol. 2001, 2, 195–201. [Google Scholar] [CrossRef]
- Zhang, Y. Transcriptional Regulation by Histone Ubiquitination and Deubiquitination. Genes Dev. 2003, 17, 2733–2740. [Google Scholar] [CrossRef]
- Hanania, U.; Furman-Matarasso, N.; Ron, M.; Avni, A. Isolation of a Novel SUMO Protein from Tomato that Suppresses EIX-Induced Cell Death. Plant J. 1999, 19, 533–541. [Google Scholar] [CrossRef]
- Novatchkova, M.; Budhiraja, R.; Coupland, G.; Eisenhaber, F.; Bachmair, A. SUMO Conjugation in Plants. Planta 2004, 220, 1–8. [Google Scholar] [CrossRef]
- Jenuwein, T.; Allis, C.D. Translating the Histone Code. Science 2001, 293, 1074–1080. [Google Scholar] [CrossRef]
- Margueron, R.; Trojer, P.; Reinberg, D. The Key to Development: Interpreting the Histone Code? Curr. Opin. Genet. Dev. 2005, 15, 163–176. [Google Scholar] [CrossRef]
- Zhang, X.; Clarenz, O.; Cokus, S.; Bernatavichute, Y.V.; Pellegrini, M.; Goodrich, J.; Jacobsen, S.E. Whole-Genome Analysis of Histone H3 Lysine 27 Trimethylation in Arabidopsis. PLoS Biol. 2007, 5, e129. [Google Scholar] [CrossRef] [PubMed]
- Gentry, M.; Hennig, L. Remodelling Chromatin to Shape Development of Plants. Exp. Cell Res. 2014, 321, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.A.; Mosher, R.A. RNA-Directed DNA Methylation: An Epigenetic Pathway of Increasing Complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef] [PubMed]
- Erdmann, R.M.; Picard, C.L. RNA-Directed DNA Methylation. PLoS Genet. 2020, 16, e1009034. [Google Scholar] [CrossRef] [PubMed]
- Haag, J.R.; Pikaard, C.S. Multisubunit RNA Polymerases IV and V: Purveyors of Non-Coding RNA for Plant Gene Silencing. Nat. Rev. Mol. Cell Biol. 2011, 12, 483–492. [Google Scholar] [CrossRef]
- Matzke, M.A.; Kanno, T.; Matzke, A.J.M. RNA-Directed DNA Methylation: The Evolution of a Complex Epigenetic Pathway in Flowering Plants. Annu. Rev. Plant Biol. 2015, 66, 243–267. [Google Scholar] [CrossRef]
- Xing, Y.; Xie, Z.; Sun, W.; Sun, Y.; Han, Z.; Zhang, S.; Tian, J.; Zhang, J.; Yao, Y. The RNA Directed DNA Methylation (RdDM) Pathway Regulates Anthocyanin Biosynthesis in Crabapple (Malus Cv. Spp.) Leaves by Methylating the McCOP1 Promoter. Plants 2021, 10, 2466. [Google Scholar] [CrossRef]
- Zhang, L.; Xiang, Y.; Chen, S.; Shi, M.; Jiang, X.; He, Z.; Gao, S. Mechanisms of MicroRNA Biogenesis and Stability Control in Plants. Front. Plant Sci. 2022, 13, 844149. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Iwakawa, H.; Tomari, Y. Molecular Insights into MicroRNA-Mediated Translational Repression in Plants. Mol. Cell 2013, 52, 591–601. [Google Scholar] [CrossRef]
- Wang, J.; Mei, J.; Ren, G. Plant MicroRNAs: Biogenesis, Homeostasis, and Degradation. Front. Plant Sci. 2019, 10, 360. [Google Scholar] [CrossRef] [PubMed]
- Villagómez-Aranda, A.L.; Feregrino-Pérez, A.A.; García-Ortega, L.F.; González-Chavira, M.M.; Torres-Pacheco, I.; Guevara-González, R.G. Activating Stress Memory: Eustressors as Potential Tools for Plant Breeding. Plant Cell Rep. 2022, 41, 1481–1498. [Google Scholar] [CrossRef] [PubMed]
- Thiebaut, F.; Hemerly, A.S.; Ferreira, P.C.G. A Role for Epigenetic Regulation in the Adaptation and Stress Responses of Non-Model Plants. Front. Plant Sci. 2019, 10, 246. [Google Scholar] [CrossRef] [PubMed]
- Remeš, J.; Pulkrab, K.; Bílek, L.; Podrázský, V. Economic and Production Effect of Tree Species Change as a Result of Adaptation to Climate Change. Forests 2020, 11, 431. [Google Scholar] [CrossRef]
- Hanel, M.; Rakovec, O.; Markonis, Y.; Máca, P.; Samaniego, L.; Kyselý, J.; Kumar, R. Revisiting the Recent European Droughts from a Long-Term Perspective. Sci. Rep. 2018, 8, 9499. [Google Scholar] [CrossRef]
- Brèteau-Amores, S.; Brunette, M.; Davi, H. An Economic Comparison of Adaptation Strategies Towards a Drought-Induced Risk of Forest Decline. Ecol. Econ. 2019, 164, 106294. [Google Scholar] [CrossRef]
- Bréda, N.; Badeau, V. Forest Tree Responses to Extreme Drought and Some Biotic Events: Towards a Selection According to Hazard Tolerance? Comptes Rendus Geosci. 2008, 340, 651–662. [Google Scholar] [CrossRef]
- Spiecker, H.; Lindner, M.; Schuler, J. What Science Can Tell Us. In Douglas-Fir—An Option for Europe; European Forest Institute: Joensuu, Finland, 2003; p. 121. [Google Scholar]
- Fox, R.J.; Donelson, J.M.; Schunter, C.; Ravasi, T.; Gaitán-Espitia, J.D. Beyond Buying Time: The Role of Plasticity in Phenotypic Adaptation to Rapid Environmental Change. Phil. Trans. R. Soc. B 2019, 374, 20180174. [Google Scholar] [CrossRef]
- Pigliucci, M. Evolution of Phenotypic Plasticity: Where Are We Going Now? Trends Ecol. Evol. 2005, 20, 481–486. [Google Scholar] [CrossRef]
- Sahu, P.P.; Pandey, G.; Sharma, N.; Puranik, S.; Muthamilarasan, M.; Prasad, M. Epigenetic Mechanisms of Plant Stress Responses and Adaptation. Plant Cell Rep. 2013, 32, 1151–1159. [Google Scholar] [CrossRef]
- Kinoshita, T.; Seki, M. Epigenetic Memory for Stress Response and Adaptation in Plants. Plant Cell Physiol. 2014, 55, 1859–1863. [Google Scholar] [CrossRef]
- Avramova, Z. Transcriptional ‘Memory’ of a Stress: Transient Chromatin and Memory (Epigenetic) Marks at Stress-Response Genes. Plant J. 2015, 83, 149–159. [Google Scholar] [CrossRef]
- Grativol, C.; Hemerly, A.S.; Ferreira, P.C.G. Genetic and Epigenetic Regulation of Stress Responses in Natural Plant Populations. Biochim. Et Biophys. Acta (BBA)—Gene Regul. Mech. 2012, 1819, 176–185. [Google Scholar] [CrossRef]
- Yakovlev, I.; Fossdal, C.G.; Skrøppa, T.; Olsen, J.E.; Jahren, A.H.; Johnsen, Ø. An Adaptive Epigenetic Memory in Conifers with Important Implications for Seed Production. Seed Sci. Res. 2012, 22, 63–76. [Google Scholar] [CrossRef]
- Angers, B.; Castonguay, E.; Massicotte, R. Environmentally Induced Phenotypes and DNA Methylation: How to Deal with Unpredictable Conditions until the next Generation and After. Mol. Ecol. 2010, 19, 1283–1295. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. DNA Methylation Patterns and Epigenetic Memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Silva, H.G.; Sobral, R.S.; Magalhães, A.P.; Morais-Cecílio, L.; Costa, M.M.R. Genome-Wide Identification of Epigenetic Regulators in Quercus suber L. Int. J. Mol. Sci. 2020, 21, 3783. [Google Scholar] [CrossRef]
- Yakovlev, I.A.; Asante, D.K.A.; Fossdal, C.G.; Junttila, O.; Johnsen, Ø. Differential Gene Expression Related to an Epigenetic Memory Affecting Climatic Adaptation in Norway Spruce. Plant Sci. 2011, 180, 132–139. [Google Scholar] [CrossRef]
- Conde, D.; Le Gac, A.; Perales, M.; Dervinis, C.; Kirst, M.; Maury, S.; González-Melendi, P.; Allona, I. Chilling-responsive DEMETER-LIKE DNA Demethylase Mediates in Poplar Bud Break. Plant Cell Environ. 2017, 40, 2236–2249. [Google Scholar] [CrossRef]
- Santamaría, M.; Hasbún, R.; Valera, M.; Meijón, M.; Valledor, L.; Rodríguez, J.L.; Toorop, P.E.; Cañal, M.; Rodríguez, R. Acetylated H4 Histone and Genomic DNA Methylation Patterns during Bud Set and Bud Burst in Castanea Sativa. J. Plant Physiol. 2009, 166, 1360–1369. [Google Scholar] [CrossRef]
- Hasbún, R.; Valledor, L.; Santamaría, E.; Cañal, M.J.; Rodríguez, R. Dynamics of dna methylation in chestnut trees development. Acta Hortic. 2007, 760, 563–566. [Google Scholar] [CrossRef]
- Kumar, G.; Rattan, U.K.; Singh, A.K. Chilling-Mediated DNA Methylation Changes during Dormancy and Its Release Reveal the Importance of Epigenetic Regulation during Winter Dormancy in Apple (Malus × Domestica Borkh.). PLoS ONE 2016, 11, e0149934. [Google Scholar] [CrossRef] [PubMed]
- Correia, B.; Hancock, R.D.; Amaral, J.; Gomez-Cadenas, A.; Valledor, L.; Pinto, G. Combined Drought and Heat Activates Protective Responses in Eucalyptus Globulus That Are Not Activated When Subjected to Drought or Heat Stress Alone. Front. Plant Sci. 2018, 9, 819. [Google Scholar] [CrossRef]
- Karlberg, A.; Englund, M.; Petterle, A.; Molnar, G.; Sjödin, A.; Bako, L.; Bhalerao, R.P. Analysis of Global Changes in Gene Expression during Activity-Dormancy Cycle in Hybrid Aspen Apex. Plant Biotechnol. 2010, 27, 1–16. [Google Scholar] [CrossRef]
- Carneros, E.; Yakovlev, I.; Viejo, M.; Olsen, J.E.; Fossdal, C.G. The Epigenetic Memory of Temperature during Embryogenesis Modifies the Expression of Bud Burst-Related Genes in Norway Spruce Epitypes. Planta 2017, 246, 553–566. [Google Scholar] [CrossRef]
- Raj, S.; Bräutigam, K.; Hamanishi, E.T.; Wilkins, O.; Thomas, B.R.; Schroeder, W.; Mansfield, S.D.; Plant, A.L.; Campbell, M.M. Clone History Shapes Populus Drought Responses. Proc. Natl. Acad. Sci. USA 2011, 108, 12521–12526. [Google Scholar] [CrossRef]
- Gourcilleau, D.; Bogeat-Triboulot, M.-B.; Le Thiec, D.; Lafon-Placette, C.; Delaunay, A.; El-Soud, W.A.; Brignolas, F.; Maury, S. DNA Methylation and Histone Acetylation: Genotypic Variations in Hybrid Poplars, Impact of Water Deficit and Relationships with Productivity. Ann. For. Sci. 2010, 67, 208. [Google Scholar] [CrossRef]
- Thapliyal, G.; Vemanna, R.S.; Pawar, P.M.; Barthwal, S.; Meena, R.K.; Pandey, S.; Bhandari, M.S. First Record of Off-Season Flowering in Populus Deltoides from India: Paradigm of Climate Change Indicator. Int. J. Biometeorol. 2020, 64, 1629–1634. [Google Scholar] [CrossRef]
- Johnsen, O.; Fossdal, C.G.; Nagy, N.; Molmann, J.; DaeHLEN, O.G.; Skroppa, T. Climatic Adaptation in Picea Abies Progenies Is Affected by the Temperature during Zygotic Embryogenesis and Seed Maturation. Plant Cell Env. 2005, 28, 1090–1102. [Google Scholar] [CrossRef]
- Rico, L.; Ogaya, R.; Barbeta, A.; Peñuelas, J. Changes in DNA Methylation Fingerprint of Quercus Ilex Trees in Response to Experimental Field Drought Simulating Projected Climate Change. Plant Biol. J. 2014, 16, 419–427. [Google Scholar] [CrossRef]
- Sow, M.D.; Le Gac, A.; Fichot, R.; Lanciano, S.; Delaunay, A.; Le Jan, I.; Lesage-Descauses, M.; Citerne, S.; Caius, J.; Brunaud, V.; et al. RNAi Suppression of DNA Methylation Affects the Drought Stress Response and Genome Integrity in Transgenic Poplar. New Phytol. 2021, 232, 80–97. [Google Scholar] [CrossRef] [PubMed]
- Zeng, F.-S.; Zhou, S.; Zhan, Y.-G.; Dong, J. Drought Resistance and DNA Methylation of Interspecific Hybrids between Fraxinus Mandshurica and Fraxinus Americana. Trees 2014, 28, 1679–1692. [Google Scholar] [CrossRef]
- Li, S.; Lin, Y.-C.J.; Wang, P.; Zhang, B.; Li, M.; Chen, S.; Shi, R.; Tunlaya-Anukit, S.; Liu, X.; Wang, Z.; et al. The AREB1 Transcription Factor Influences Histone Acetylation to Regulate Drought Responses and Tolerance in Populus trichocarpa. Plant Cell 2019, 31, 663–686. [Google Scholar] [CrossRef] [PubMed]
- Lira-Medeiros, C.F.; Parisod, C.; Fernandes, R.A.; Mata, C.S.; Cardoso, M.A.; Ferreira, P.C.G. Epigenetic Variation in Mangrove Plants Occurring in Contrasting Natural Environment. PLoS ONE 2010, 5, e10326. [Google Scholar] [CrossRef]
- Yakovlev, I.A.; Fossdal, C.G.; Johnsen, Ø. MicroRNAs, the Epigenetic Memory and Climatic Adaptation in Norway Spruce. New Phytol. 2010, 187, 1154–1169. [Google Scholar] [CrossRef]
- Kovalchuk, O.; Burke, P.; Arkhipov, A.; Kuchma, N.; James, S.J.; Kovalchuk, I.; Pogribny, I. Genome Hypermethylation in Pinus Silvestris of Chernobyl—A Mechanism for Radiation Adaptation? Mutat. Res. Fundam. Mol. Mech. Mutagen. 2003, 529, 13–20. [Google Scholar] [CrossRef]
- Reich, P.B.; Oleksyn, J. Climate Warming Will Reduce Growth and Survival of Scots Pine except in the Far North: Scots Pine Growth and Survival Following Climate Transfer. Ecol. Lett. 2008, 11, 588–597. [Google Scholar] [CrossRef]
- Pawłowski, T.A.; Klupczyńska, E.A.; Staszak, A.M.; Suszka, J. Proteomic Analysis of Black Poplar (Populus nigra L.) Seed Storability. Ann. For. Sci. 2019, 76, 104. [Google Scholar] [CrossRef]
- Sano, N.; Rajjou, L.; North, H.M.; Debeaujon, I.; Marion-Poll, A.; Seo, M. Staying Alive: Molecular Aspects of Seed Longevity. Plant Cell Physiol. 2016, 57, 660–674. [Google Scholar] [CrossRef]
- Alakärppä, E.; Salo, H.M.; Valledor, L.; Cañal, M.J.; Häggman, H.; Vuosku, J. Natural Variation of DNA Methylation and Gene Expression May Determine Local Adaptations of Scots Pine Populations. J. Exp. Bot. 2018, 69, 5293–5305. [Google Scholar] [CrossRef]
- Walck, J.L.; Hidayati, S.N.; Dixon, K.W.; Thompson, K.; Poschlod, P. Climate Change and Plant Regeneration from Seed: Climate change and plant regeneration. Glob. Change Biol. 2011, 17, 2145–2161. [Google Scholar] [CrossRef]
- Cochrane, A. Are We Underestimating the Impact of Rising Summer Temperatures on Dormancy Loss in Hard-Seeded Species? Aust. J. Bot. 2017, 65, 248–256. [Google Scholar] [CrossRef]
- Davis, M.B.; Shaw, R.G. Range Shifts and Adaptive Responses to Quaternary Climate Change. Science 2001, 292, 673–679. [Google Scholar] [CrossRef]
- Pearson, R.G.; Dawson, T.P. Predicting the Impacts of Climate Change on the Distribution of Species: Are Bioclimate Envelope Models Useful? Eval. Bioclimate Envel. Models. Glob. Ecol. Biogeogr. 2003, 12, 361–371. [Google Scholar] [CrossRef]
- Footitt, S.; Müller, K.; Kermode, A.R.; Finch-Savage, W.E. Seed Dormancy Cycling in A Rabidopsis: Chromatin Remodelling and Regulation of DOG 1 in Response to Seasonal Environmental Signals. Plant J. 2015, 81, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Xuan, L.; Jiang, Y.; Yu, H. Regulation by Flowering locus T and terminal flower 1 in Flowering Time and Plant Architecture. Small Struct. 2021, 2, 2000125. [Google Scholar] [CrossRef]
- Tai, H.H.; Tai, G.C.C.; Beardmore, T. Dynamic Histone Acetylation of Late Embryonic Genes during Seed Germination. Plant Mol. Biol. 2005, 59, 909–925. [Google Scholar] [CrossRef]
- Tanaka, M.; Kikuchi, A.; Kamada, H. The Arabidopsis Histone Deacetylases HDA6 and HDA19 Contribute to the Repression of Embryonic Properties after Germination. Plant Physiol. 2008, 146, 149–161. [Google Scholar] [CrossRef]
- Yashiroda, H.; Tanaka, K. Hub1 Is an Essential Ubiquitin-like Protein without Functioning as a Typical Modifier in Fission Yeast: Role of Hub1 in S. pombe. Genes Cells 2004, 9, 1189–1197. [Google Scholar] [CrossRef]
- McNally, T.; Huang, Q.; Janis, R.S.; Liu, Z.; Olejniczak, E.T.; Reilly, R.M. Structural Analysis of UBL5, a Novel Ubiquitin-like Modifier. Protein Sci. 2003, 12, 1562–1566. [Google Scholar] [CrossRef]
- Du, H.-N. Transcription, DNA Damage and Beyond: The Roles of Histone Ubiquitination and Deubiquitination. CPPS 2012, 13, 447–466. [Google Scholar] [CrossRef] [PubMed]
- Layat, E.; Bourcy, M.; Cotterell, S.; Zdzieszyńska, J.; Desset, S.; Duc, C.; Tatout, C.; Bailly, C.; Probst, A.V. The Histone Chaperone HIRA Is a Positive Regulator of Seed Germination. Int. J. Mol. Sci. 2021, 22, 4031. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.B.; Kao, C.-F.; Hillyer, C.; Pikaart, M.; Osley, M.A. H2B Ubiquitylation Plays a Role in Nucleosome Dynamics during Transcription Elongation. Mol. Cell 2008, 31, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Long, R.L.; Gorecki, M.J.; Renton, M.; Scott, J.K.; Colville, L.; Goggin, D.E.; Commander, L.E.; Westcott, D.A.; Cherry, H.; Finch-Savage, W.E. The Ecophysiology of Seed Persistence: A Mechanistic View of the Journey to Germination or Demise: The Ecophysiology of Seed Persistence. Biol. Rev. 2015, 90, 31–59. [Google Scholar] [CrossRef]
- Cohen, D. Optimizing Reproduction in a Randomly Varying Environment. J. Theor. Biol. 1966, 12, 119–129. [Google Scholar] [CrossRef]
- Graeber, K.; Linkies, A.; Steinbrecher, T.; Mummenhoff, K.; Tarkowská, D.; Turečková, V.; Ignatz, M.; Sperber, K.; Voegele, A.; de Jong, H.; et al. DELAY OF GERMINATION 1 Mediates a Conserved Coat-Dormancy Mechanism for the Temperature- and Gibberellin-Dependent Control of Seed Germination. Proc. Natl. Acad. Sci. USA 2014, 111, E3571–E3580. [Google Scholar] [CrossRef]
- Nakabayashi, K.; Bartsch, M.; Xiang, Y.; Miatton, E.; Pellengahr, S.; Yano, R.; Seo, M.; Soppe, W.J.J. The Time Required for Dormancy Release in Arabidopsis Is Determined by DELAY OF GERMINATION1 Protein Levels in Freshly Harvested Seeds. Plant Cell 2012, 24, 2826–2838. [Google Scholar] [CrossRef]
- Bentsink, L.; Jowett, J.; Hanhart, C.J.; Koornneef, M. Cloning of DOG1, a Quantitative Trait Locus Controlling Seed Dormancy in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 17042–17047. [Google Scholar] [CrossRef]
- Cyrek, M.; Fedak, H.; Ciesielski, A.; Guo, Y.; Sliwa, A.; Brzezniak, L.; Krzyczmonik, K.; Pietras, Z.; Kaczanowski, S.; Liu, F.; et al. Seed Dormancy in Arabidopsis Is Controlled by Alternative Polyadenylation of DOG1. Plant Physiol. 2016, 170, 947–955. [Google Scholar] [CrossRef]
- Müller, K.; Bouyer, D.; Schnittger, A.; Kermode, A.R. Evolutionarily Conserved Histone Methylation Dynamics during Seed Life-Cycle Transitions. PLoS ONE 2012, 7, e51532. [Google Scholar] [CrossRef]
- Zeng, Y.; Raimondi, N.; Kermode, A.R. Role of an ABI3 Homologue in Dormancy Maintenance of Yellow-Cedar Seeds and in the Activation of Storage Protein and Em Gene Promoters. Plant Mol. Biol. 2003, 51, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Kermode, A.R. A Gymnosperm ABI3 Gene Functions in a Severe Abscisic Acid-Insensitive Mutant of Arabidopsis (Abi3-6) to Restore the Wild-Type Phenotype and Demonstrates a Strong Synergistic Effect with Sugar in the Inhibition of Post-Germinative Growth. Plant Mol. Biol. 2004, 56, 731–746. [Google Scholar] [CrossRef] [PubMed]
- Huo, H.; Wei, S.; Bradford, K.J. DELAY OF GERMINATION1 (DOG1) Regulates Both Seed Dormancy and Flowering Time through MicroRNA Pathways. Proc. Natl. Acad. Sci. USA 2016, 113, E2199–E2206. [Google Scholar] [CrossRef] [PubMed]
- Nonogaki, H. Seed Germination and Dormancy: The Classic Story, New Puzzles, and Evolution. J. Integr. Plant Biol. 2019, 61, 541–563. [Google Scholar] [CrossRef]
- Martin, R.C.; Liu, P.-P.; Goloviznina, N.A.; Nonogaki, H. MicroRNA, Seeds, and Darwin?: Diverse Function of MiRNA in Seed Biology and Plant Responses to Stress. J. Exp. Bot. 2010, 61, 2229–2234. [Google Scholar] [CrossRef] [PubMed]
- Reyes, J.L.; Chua, N.-H. ABA Induction of MiR159 Controls Transcript Levels of Two MYB Factors during Arabidopsis Seed Germination: MiR159 Regulation of ABA Responses during Germination. Plant J. 2007, 49, 592–606. [Google Scholar] [CrossRef]
- Horvath, D.P.; Anderson, J.V.; Chao, W.S.; Foley, M.E. Knowing When to Grow: Signals Regulating Bud Dormancy. Trends Plant Sci. 2003, 8, 534–540. [Google Scholar] [CrossRef]
- Yamane, H.; Singh, A.K.; Cooke, J.E.K. Plant Dormancy Research: From Environmental Control to Molecular Regulatory Networks. Tree Physiol. 2021, 41, 523–528. [Google Scholar] [CrossRef]
- Rohde, A.; Bhalerao, R.P. Plant Dormancy in the Perennial Context. Trends Plant Sci. 2007, 12, 217–223. [Google Scholar] [CrossRef]
- Lang, G.A.; Early, J.D.; Martin, G.C.; Darnell, R.L. Endo-, Para-, and Ecodormancy: Physiological Terminology and Classification for Dormancy Research. HortScience 1987, 22, 371–377. [Google Scholar] [CrossRef]
- Martin, G.C. Bud Dormancy in Deciduous Fruit Trees. In Plant Physiology: A Treatise; Steward, F.C., Ed.; Academic Press: New York, NY, USA, 1991; pp. 183–225. [Google Scholar]
- Santamaría, M.E.; Rodríguez, R.; Cañal, M.J.; Toorop, P.E. Transcriptome Analysis of Chestnut (Castanea sativa) Tree Buds Suggests a Putative Role for Epigenetic Control of Bud Dormancy. Ann. Bot. 2011, 108, 485–498. [Google Scholar] [CrossRef] [PubMed]
- Fraga, M.F.; Rodriguez, R.; Canal, M.J. Genomic DNA Methylation-Demethylation during Aging and Reinvigoration of Pinus Radiata. Tree Physiol. 2002, 22, 813–816. [Google Scholar] [CrossRef] [PubMed]
- Valledor, L.; Jorrín, J.V.; Rodríguez, J.L.; Lenz, C.; Meijón, M.; Rodríguez, R.; Cañal, M.J. Combined Proteomic and Transcriptomic Analysis Identifies Differentially Expressed Pathways Associated to Pinus Radiata Needle Maturation. J. Proteome Res. 2010, 9, 3954–3979. [Google Scholar] [CrossRef] [PubMed]
- Monteuuis, O.; Baurens, F.C.; Goh, D.K.S.; Quimado, M.; Doulbeau, S.; Verdeil, J.L. DNA Methylation in Acacia Mangium In Vitro and Ex-Vitro Buds, in Relation to Their Within-Shoot Position, Age and Leaf Morphology of the Shoots. Silvae Genet. 2009, 58, 287–292. [Google Scholar] [CrossRef]
- Charron, J.-B.F.; He, H.; Elling, A.A.; Deng, X.W. Dynamic Landscapes of Four Histone Modifications during Deetiolation in Arabidopsis. Plant Cell 2010, 21, 3732–3748. [Google Scholar] [CrossRef]
- Meijón, M.; Feito, I.; Valledor, L.; Rodríguez, R.; Cañal, M.J. Dynamics of DNA Methylation and Histone H4 Acetylation during Floral Bud Differentiation in Azalea. BMC Plant Biol. 2010, 10, 10. [Google Scholar] [CrossRef]
- Lloret, A.; Badenes, M.L.; Ríos, G. Modulation of Dormancy and Growth Responses in Reproductive Buds of Temperate Trees. Front. Plant Sci. 2018, 9, 1368. [Google Scholar] [CrossRef]
- De la Fuente, L.; Conesa, A.; Lloret, A.; Badenes, M.L.; Ríos, G. Genome-Wide Changes in Histone H3 Lysine 27 Trimethylation Associated with Bud Dormancy Release in Peach. Tree Genet. Genomes 2015, 11, 45. [Google Scholar] [CrossRef]
- Leida, C.; Conesa, A.; Llácer, G.; Badenes, M.L.; Ríos, G. Histone Modifications and Expression of DAM6 Gene in Peach Are Modulated during Bud Dormancy Release in a Cultivar-dependent Manner. New Phytol. 2012, 193, 67–80. [Google Scholar] [CrossRef]
- Lloret, A.; Martínez-Fuentes, A.; Agustí, M.; Badenes, M.L.; Ríos, G. Chromatin-Associated Regulation of Sorbitol Synthesis in Flower Buds of Peach. Plant Mol. Biol. 2017, 95, 507–517. [Google Scholar] [CrossRef]
- Amasino, R.M.; Michaels, S.D. The Timing of Flowering: Figure 1. Plant Physiol. 2010, 154, 516–520. [Google Scholar] [CrossRef]
- Bouché, F.; Woods, D.P.; Amasino, R.M. Winter Memory throughout the Plant Kingdom: Different Paths to Flowering. Plant Physiol. 2017, 173, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, C.C.; Conn, A.B.; Dennis, E.S.; Peacock, W.J. Different Regulatory Regions Are Required for the Vernalization-Induced Repression of FLOWERING LOCUS C and for the Epigenetic Maintenance of Repression. Plant Cell 2002, 14, 2527–2537. [Google Scholar] [CrossRef] [PubMed]
- Bastow, R.; Mylne, J.S.; Lister, C.; Lippman, Z.; Martienssen, R.A.; Dean, C. Vernalization Requires Epigenetic Silencing of FLC by Histone Methylation. Nature 2004, 427, 164–167. [Google Scholar] [CrossRef]
- Ringrose, L.; Paro, R. Polycomb/Trithorax Response Elements and Epigenetic Memory of Cell Identity. Development 2007, 134, 223–232. [Google Scholar] [CrossRef]
- Sheldon, C.C.; Rouse, D.T.; Finnegan, E.J.; Peacock, W.J.; Dennis, E.S. The Molecular Basis of Vernalization: The Central Role of FLOWERING LOCUS C (FLC). Proc. Natl. Acad. Sci. USA 2000, 97, 3753–3758. [Google Scholar] [CrossRef] [PubMed]
- Sung, S.; Amasino, R.M. Vernalization in Arabidopsis Thaliana Is Mediated by the PHD Finger Protein VIN3. Nature 2004, 427, 159–164. [Google Scholar] [CrossRef]
- Wood, C.C.; Robertson, M.; Tanner, G.; Peacock, W.J.; Dennis, E.S.; Helliwell, C.A. The Arabidopsis thaliana Vernalization Response Requires a Polycomb-like Protein Complex That Also Includes vernalization insensitive 3. Proc. Natl. Acad. Sci. USA 2006, 103, 14631–14636. [Google Scholar] [CrossRef]
- Wellensiek, S.J. Dividing Cells as the Prerequisite for Vernalization. Plant Physiol. 1964, 39, 832–835. [Google Scholar] [CrossRef]
- Schwabe, W.W. Factors Controlling Flowering in the Chrysanthemum: V. de-vernalization in relation to high temperature and low light intensity treatments. J. Exp. Bot. 1955, 6, 435–450. [Google Scholar] [CrossRef]
- Finnegan, E.J.; Dennis, E.S. Vernalization-Induced Trimethylation of Histone H3 Lysine 27 at FLC Is Not Maintained in Mitotically Quiescent Cells. Curr. Biol. 2007, 17, 1978–1983. [Google Scholar] [CrossRef] [PubMed]
- Böhlenius, H.; Huang, T.; Charbonnel-Campaa, L.; Brunner, A.M.; Jansson, S.; Strauss, S.H.; Nilsson, O. CO/FT Regulatory Module Controls Timing of Flowering and Seasonal Growth Cessation in Trees. Science 2006, 312, 1040–1043. [Google Scholar] [CrossRef] [PubMed]
- Orru, M.; Mattana, E.; Pritchard, H.W.; Bacchetta, G. Thermal Thresholds as Predictors of Seed Dormancy Release and Germination Timing: Altitude-Related Risks from Climate Warming for the Wild Grapevine Vitis vinifera Subsp. sylvestris. Ann. Bot. 2012, 110, 1651–1660. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kurpisz, B.; Pawłowski, T.A. Epigenetic Mechanisms of Tree Responses to Climatic Changes. Int. J. Mol. Sci. 2022, 23, 13412. https://doi.org/10.3390/ijms232113412
Kurpisz B, Pawłowski TA. Epigenetic Mechanisms of Tree Responses to Climatic Changes. International Journal of Molecular Sciences. 2022; 23(21):13412. https://doi.org/10.3390/ijms232113412
Chicago/Turabian StyleKurpisz, Barbara, and Tomasz Andrzej Pawłowski. 2022. "Epigenetic Mechanisms of Tree Responses to Climatic Changes" International Journal of Molecular Sciences 23, no. 21: 13412. https://doi.org/10.3390/ijms232113412
APA StyleKurpisz, B., & Pawłowski, T. A. (2022). Epigenetic Mechanisms of Tree Responses to Climatic Changes. International Journal of Molecular Sciences, 23(21), 13412. https://doi.org/10.3390/ijms232113412






