YUCCA2 (YUC2)-Mediated 3-Indoleacetic Acid (IAA) Biosynthesis Regulates Chloroplast RNA Editing by Relieving the Auxin Response Factor 1 (ARF1)-Dependent Inhibition of Editing Factors in Arabidopsis thaliana
Abstract
:1. Introduction
2. Results
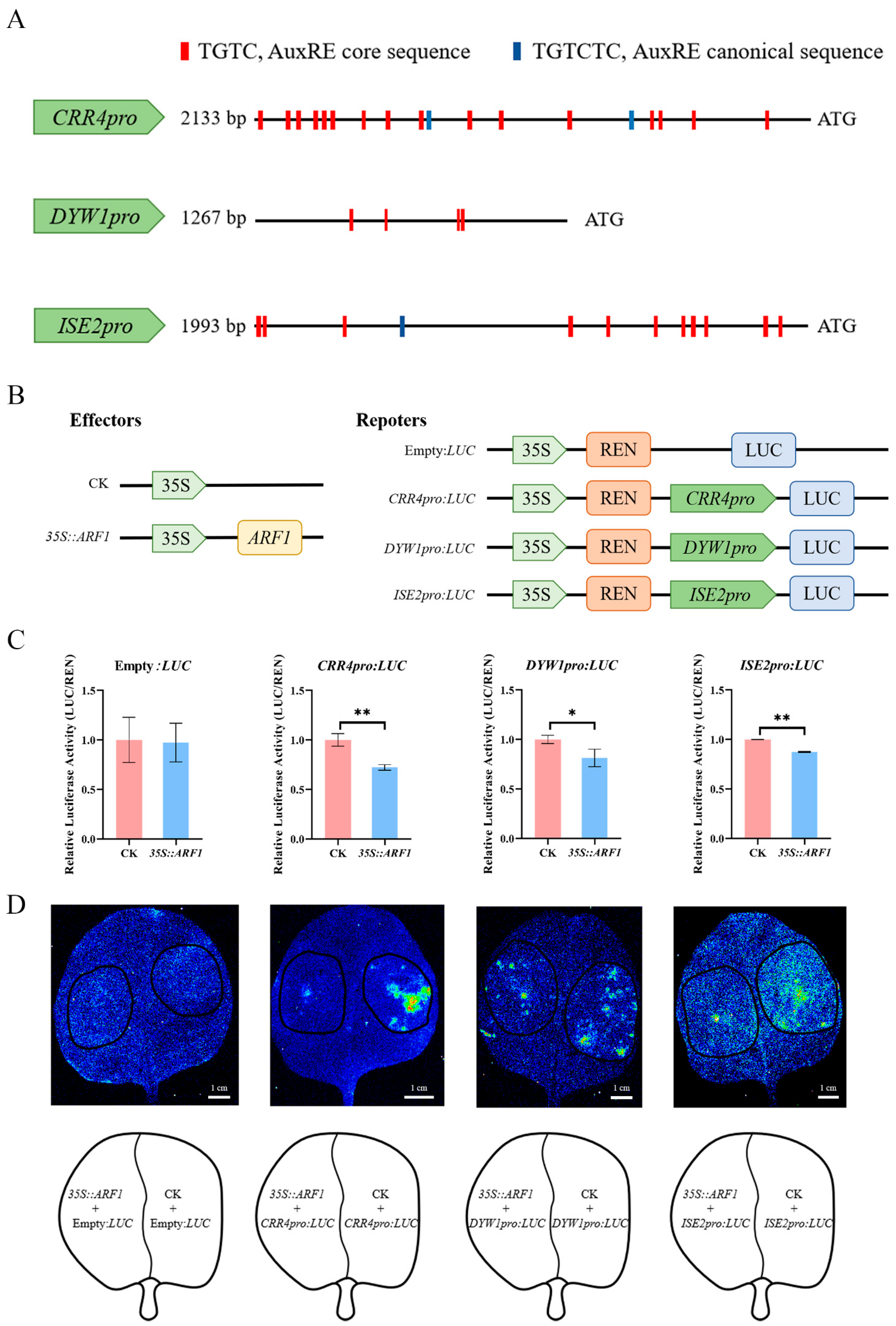
2.1. AuxREs in the Promoters of Chloroplast Editing Factors
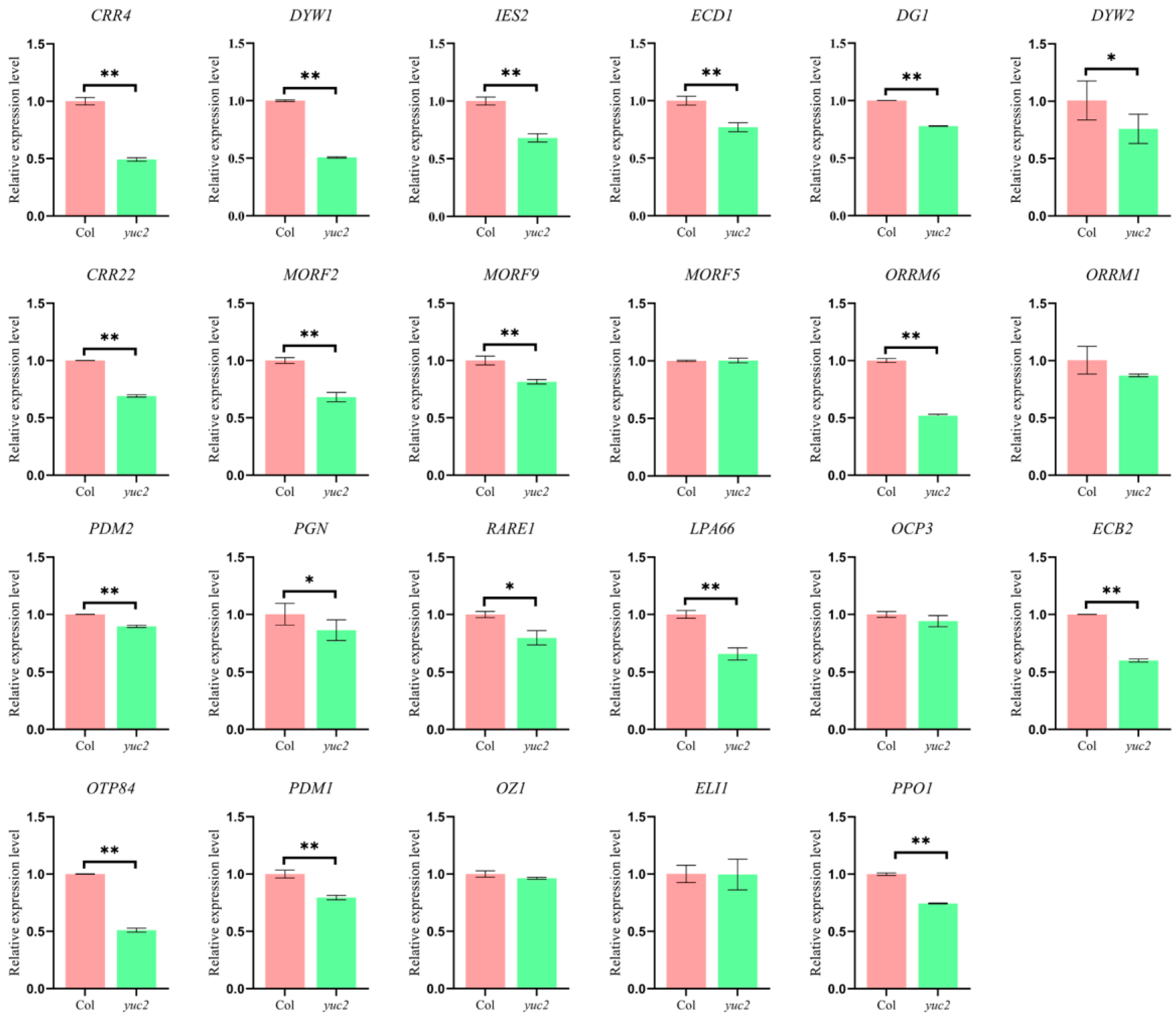
2.2. Loss of YUC2 Affects the Expression of Chloroplast Editing Factors
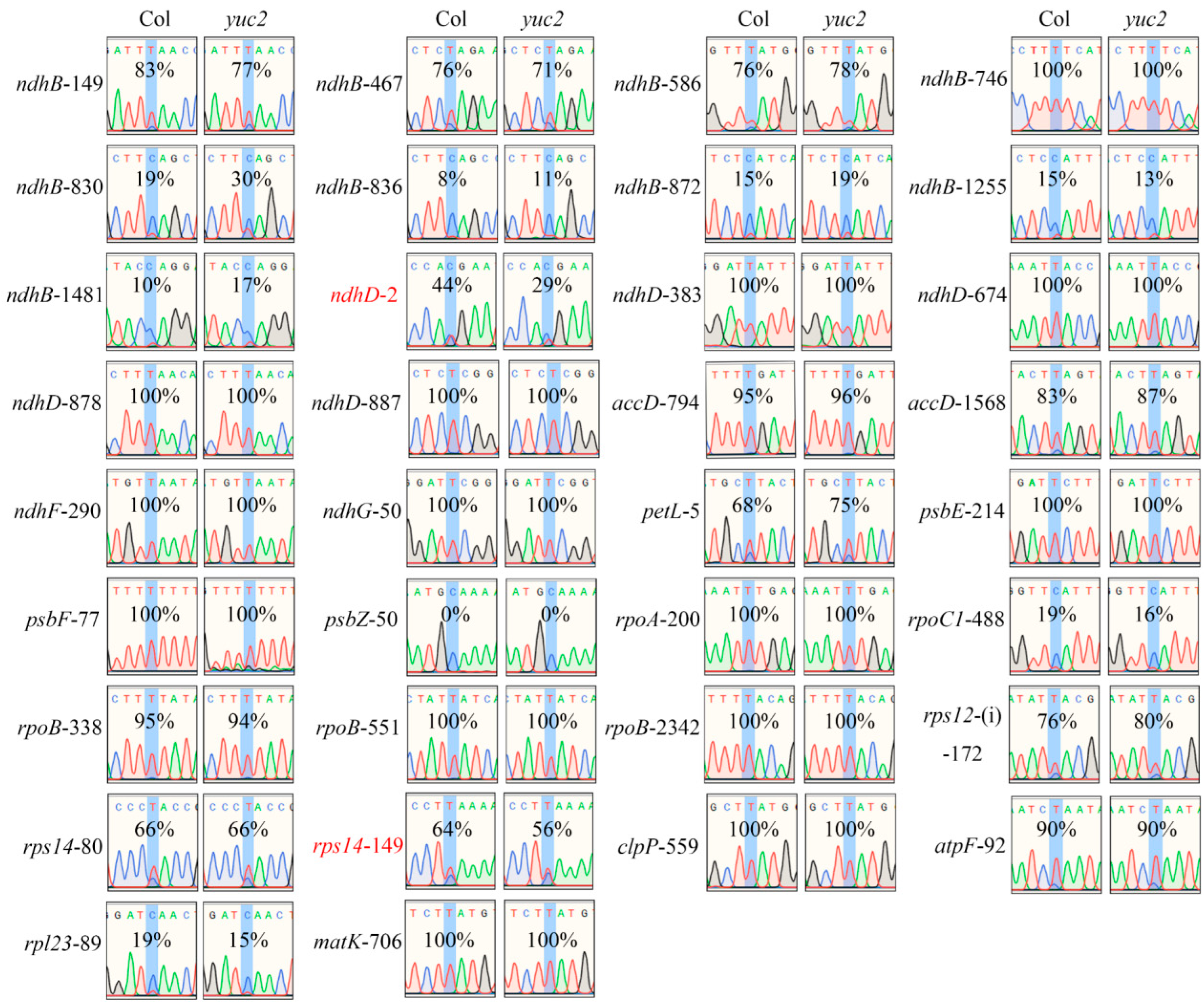
2.3. RNA-Editing Efficiency of the rps14-149 and ndhD-2 Sites Are Decreased in yuc2
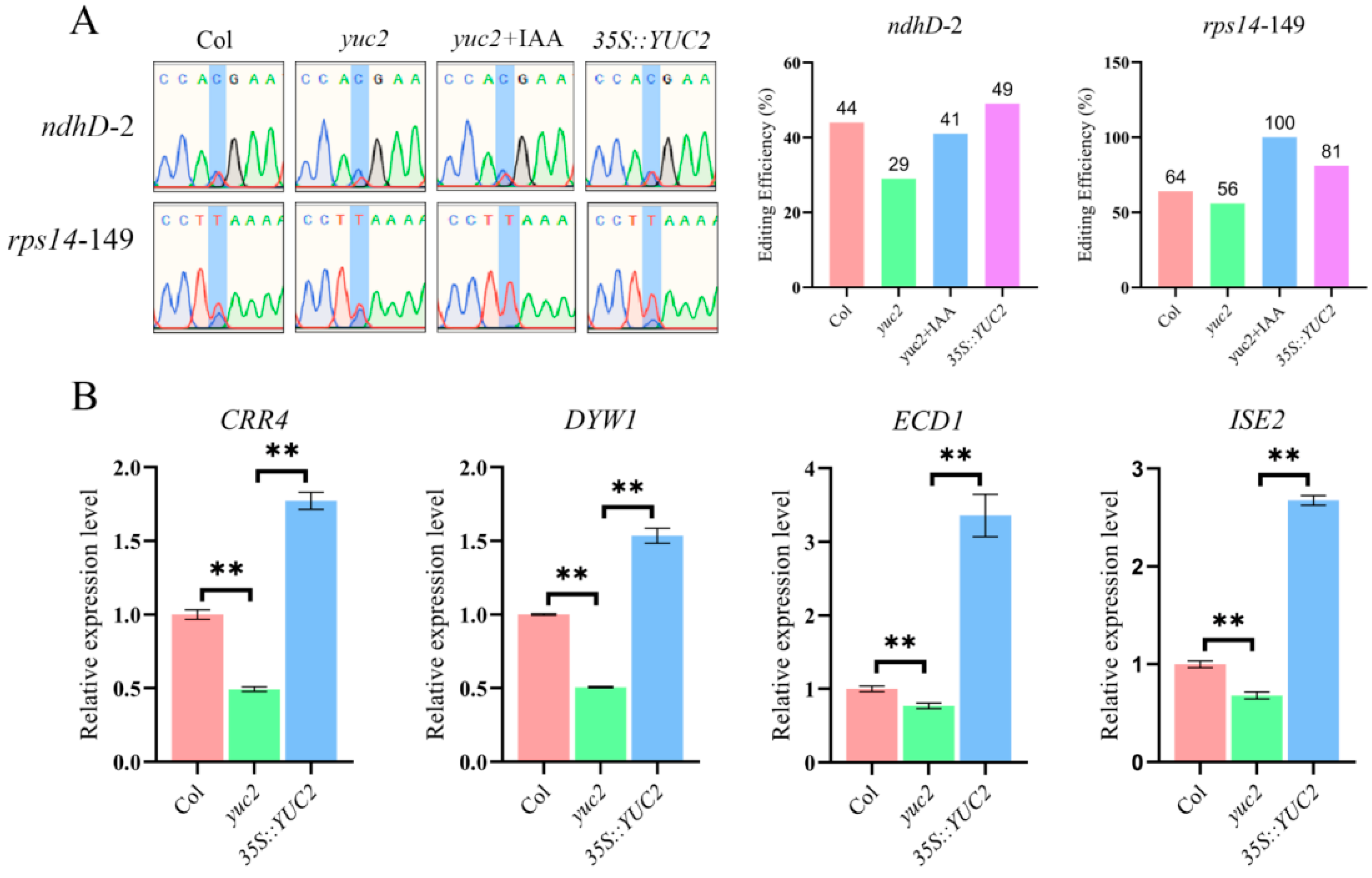
2.4. Auxin Positively Regulates the Editing of the rps14-149 and ndhD-2 Sites and Their Corresponding Editing Factors
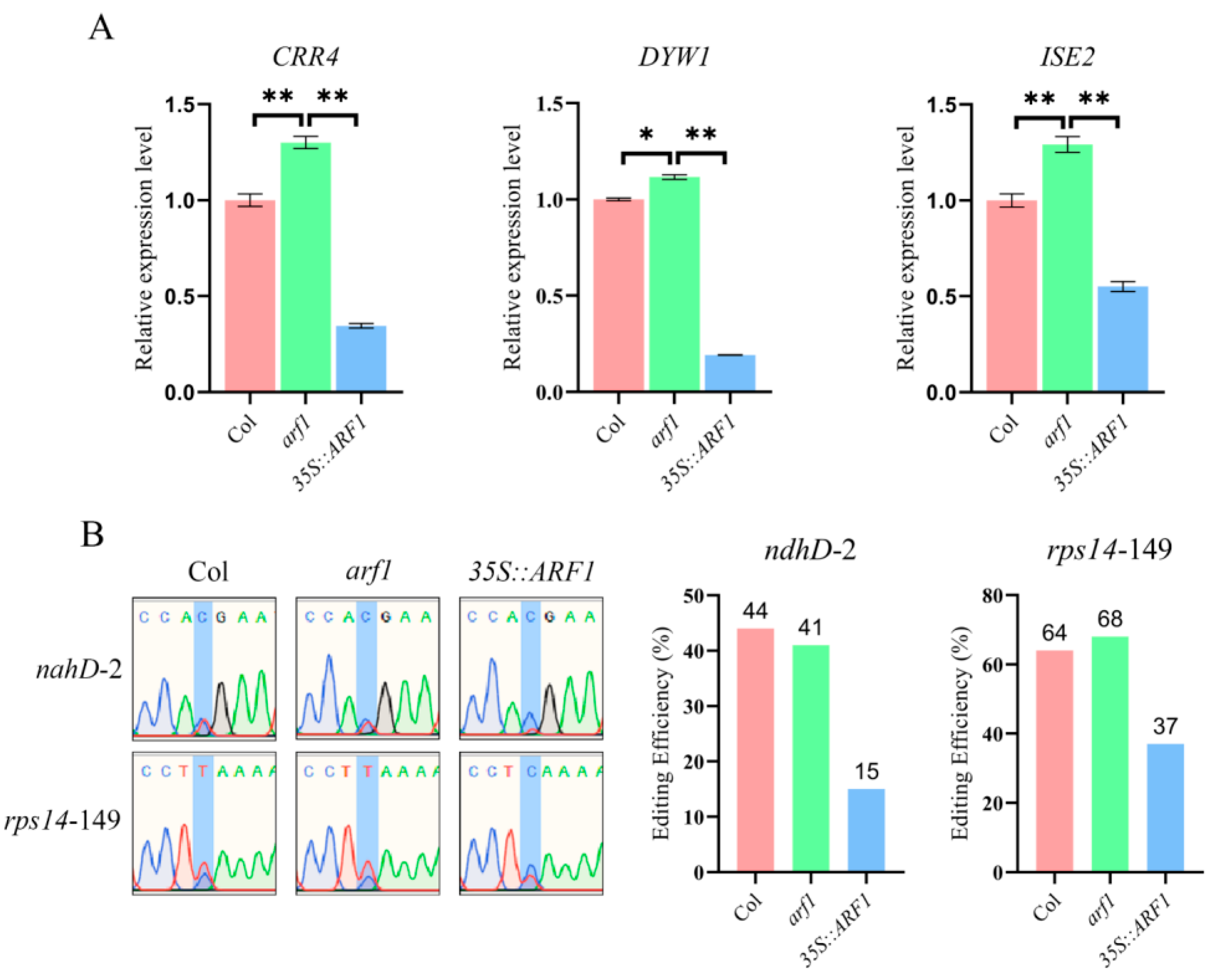
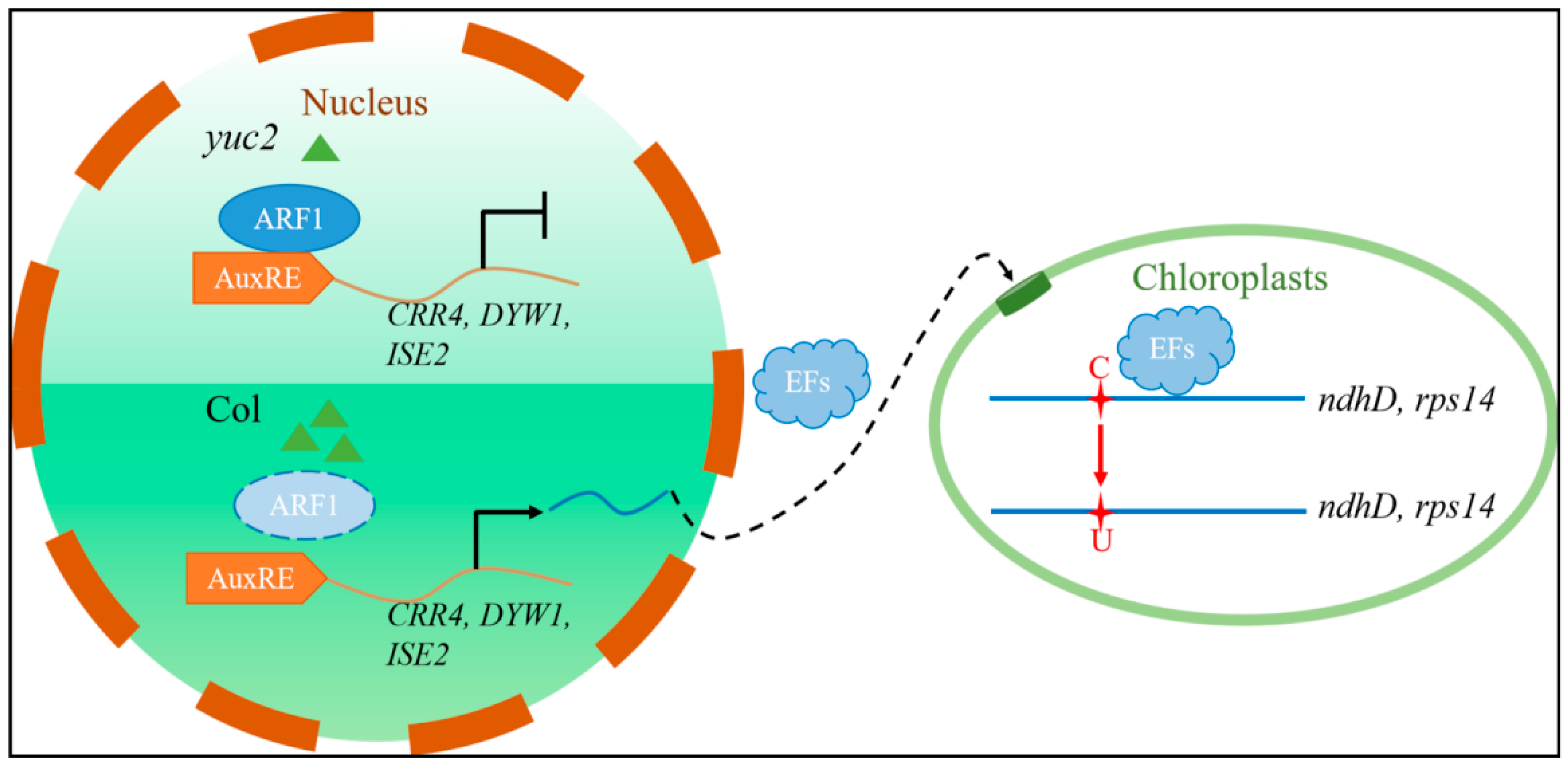
2.5. ARF1 Directly Regulates the Transcription of CRR4, DYW1, and ISE2
2.6. The Deletion of ARF1 Affects the Editing of the ndhD-2 and rps14-149 Sites
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Constructs and Plant Transformation
4.3. RNA Extraction and RT-qPCR
4.4. RNA Editing Efficiency Analysis
4.5. Luciferase Reporter Assay
4.6. Transient Expression in N. benthamiana
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kirchhoff, H. Chloroplast ultrastructure in plants. New Phytol. 2019, 223, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, Z.; Zhang, M.; Ai, P. A chloroplast-localized pentatricopeptide repeat protein involved in RNA editing and splicing and its effects on chloroplast development in rice. BMC Plant Biol. 2022, 22, 437. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, A.; Li, X.; Lu, C. The Role of Chloroplast Gene Expression in Plant Responses to Environmental Stress. Int. J. Mol. Sci. 2020, 21, 6082. [Google Scholar] [CrossRef] [PubMed]
- Hess, W.R.; Borner, T. Organellar RNA polymerases of higher plants. Int. Rev. Cytol. 1999, 190, 1–59. [Google Scholar]
- Zhelyazkova, P.; Sharma, C.M.; Forstner, K.U.; Liere, K.; Vogel, J.; Borner, T. The primary transcriptome of barley chloroplasts: Numerous noncoding RNAs and the dominating role of the plastid-encoded RNA polymerase. Plant Cell 2012, 24, 123–136. [Google Scholar] [CrossRef]
- Mohammed, T.; Firoz, A.; Ramadan, A.M. RNA Editing in Chloroplast: Advancements and Opportunities. Curr. Issues Mol. Biol. 2022, 44, 5593–5604. [Google Scholar] [CrossRef] [PubMed]
- del Campo, E.M. Post-transcriptional control of chloroplast gene expression. Gene Regul. Syst. Biol. 2009, 3, 31–47. [Google Scholar] [CrossRef] [PubMed]
- Takenaka, M.; Zehrmann, A.; Verbitskiy, D.; Hartel, B.; Brennicke, A. RNA editing in plants and its evolution. Annu. Rev. Genet. 2013, 47, 335–352. [Google Scholar] [CrossRef]
- Small, I.D.; Schallenberg-Rudinger, M.; Takenaka, M.; Mireau, H.; Ostersetzer-Biran, O. Plant organellar RNA editing: What 30 years of research has revealed. Plant J. 2020, 101, 1040–1056. [Google Scholar] [CrossRef]
- Yagi, Y.; Hayashi, S.; Kobayashi, K.; Hirayama, T.; Nakamura, T. Elucidation of the RNA recognition code for pentatricopeptide repeat proteins involved in organelle RNA editing in plants. PLoS ONE 2013, 8, e57286. [Google Scholar] [CrossRef]
- Cheng, S.; Gutmann, B.; Zhong, X.; Ye, Y.; Fisher, M.F.; Bai, F.; Castleden, I.; Song, Y.; Song, B.; Huang, J.; et al. Redefining the structural motifs that determine RNA binding and RNA editing by pentatricopeptide repeat proteins in land plants. Plant J. 2016, 85, 532–547. [Google Scholar] [CrossRef]
- Kotera, E.; Tasaka, M.; Shikanai, T. A pentatricopeptide repeat protein is essential for RNA editing in chloroplasts. Nature 2005, 433, 326–330. [Google Scholar] [CrossRef] [PubMed]
- Boussardon, C.; Salone, V.; Avon, A.; Berthome, R.; Hammani, K.; Okuda, K.; Shikanai, T.; Small, I.; Lurin, C. Two interacting proteins are necessary for the editing of the NdhD-1 site in Arabidopsis plastids. Plant Cell 2012, 24, 3684–3694. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Germain, A.; Giloteaux, L.; Hammani, K.; Barkan, A.; Hanson, M.R.; Bentolila, S. An RNA recognition motif-containing protein is required for plastid RNA editing in Arabidopsis and maize. Proc. Natl. Acad. Sci. USA 2013, 110, E1169–E1178. [Google Scholar] [CrossRef]
- Zhang, F.; Tang, W.; Hedtke, B.; Zhong, L.; Liu, L.; Peng, L.; Lu, C.; Grimm, B.; Lin, R. Tetrapyrrole biosynthetic enzyme protoporphyrinogen IX oxidase 1 is required for plastid RNA editing. Proc. Natl. Acad. Sci. USA 2014, 111, 2023–2028. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Shi, X.; Friso, G.; Van Wijk, K.; Bentolila, S.; Hanson, M.R. A zinc finger motif-containing protein is essential for chloroplast RNA editing. PLoS Genet. 2015, 11, e1005028. [Google Scholar] [CrossRef]
- Huang, C.; Yu, Q.B.; Li, Z.R.; Ye, L.S.; Xu, L.; Yang, Z.N. Porphobilinogen deaminase HEMC interacts with the PPR-protein AtECB2 for chloroplast RNA editing. Plant J. 2017, 92, 546–556. [Google Scholar] [CrossRef]
- Zhao, X.; Huang, J.; Chory, J. GUN1 interacts with MORF2 to regulate plastid RNA editing during retrograde signaling. Proc. Natl. Acad. Sci. USA 2019, 116, 10162–10167. [Google Scholar] [CrossRef]
- Sun, T.; Bentolila, S.; Hanson, M.R. The Unexpected Diversity of Plant Organelle RNA Editosomes. Trends Plant Sci. 2016, 21, 962–973. [Google Scholar] [CrossRef]
- Sun, Y.K.; Gutmann, B.; Yap, A.; Kindgren, P.; Small, I. Editing of Chloroplast rps14 by PPR Editing Factor EMB2261 Is Essential for Arabidopsis Development. Front. Plant Sci. 2018, 9, 841. [Google Scholar] [CrossRef]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 2007, 19, 2430–2439. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Wang, J.; Gao, Z.; Dong, J.; He, H.; Terzaghi, W.; Wei, N.; Deng, X.W.; Chen, H. Arabidopsis SAURs are critical for differential light regulation of the development of various organs. Proc. Natl. Acad. Sci. USA 2016, 113, 6071–6076. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Sun, N.; Yang, J.; Deng, Z.; Lan, J.; Qin, G.; He, H.; Deng, X.W.; Irish, V.F.; Chen, H.; et al. The Transcription Factors TCP4 and PIF3 Antagonistically Regulate Organ-Specific Light Induction of SAUR Genes to Modulate Cotyledon Opening during De-Etiolation in Arabidopsis. Plant Cell 2019, 31, 1155–1170. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.; Chang, Y.; Liu, G.; Shang, S.; Tian, L.; Shi, H. Molecular functional analysis of auxin/indole-3-acetic acid proteins (Aux/IAAs) in plant disease resistance in cassava. Physiol. Plant 2020, 168, 88–97. [Google Scholar] [CrossRef]
- Mashiguchi, K.; Tanaka, K.; Sakai, T.; Sugawara, S.; Kawaide, H.; Natsume, M.; Hanada, A.; Yaeno, T.; Shirasu, K.; Yao, H.; et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 18512–18517. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Yang, H.; Shang, C.; Ma, S.; Liu, L.; Cheng, J. The Roles of Auxin Biosynthesis YUCCA Gene Family in Plants. Int. J. Mol. Sci. 2019, 20, 6343. [Google Scholar] [CrossRef] [PubMed]
- Della Rovere, F.; Fattorini, L.; D’Angeli, S.; Veloccia, A.; Falasca, G.; Altamura, M.M. Auxin and cytokinin control formation of the quiescent centre in the adventitious root apex of Arabidopsis. Ann. Bot. 2013, 112, 1395–1407. [Google Scholar] [CrossRef]
- Veloccia, A.; Fattorini, L.; Della Rovere, F.; Sofo, A.; D’Angeli, S.; Betti, C.; Falasca, G.; Altamura, M.M. Ethylene and auxin interaction in the control of adventitious rooting in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 6445–6458. [Google Scholar] [CrossRef]
- Kasahara, H. Current aspects of auxin biosynthesis in plants. Biosci. Biotechnol. Biochem. 2016, 80, 34–42. [Google Scholar] [CrossRef]
- Kavai-Ool, U.N. Genetic regulation of polar auxin transport and its role in control of shoot morphogenesis. Biol. Bull. Rev. 2011, 1, 517–525. [Google Scholar] [CrossRef]
- Leyser, O. Auxin Signaling. Plant Physiol. 2018, 176, 465–479. [Google Scholar] [CrossRef]
- Chandler, J.W. Auxin response factors. Plant Cell Environ. 2016, 39, 1014–1028. [Google Scholar] [CrossRef] [PubMed]
- Li, S.B.; Xie, Z.Z.; Hu, C.G.; Zhang, J.Z. A Review of Auxin Response Factors (ARFs) in Plants. Front. Plant Sci. 2016, 7, 47. [Google Scholar] [CrossRef] [PubMed]
- Rienstra, J.; Hernández-García, J.; Weijers, D. To bind or not to bind: How Auxin Response Factors select their target genes. J. Exp. Bot. 2023, erad259. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. ARF1, a transcription factor that binds to auxin response elements. Science 1997, 276, 1865–1868. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Dai, X.; De-Paoli, H.; Cheng, Y.; Takebayashi, Y.; Kasahara, H.; Kamiya, Y.; Zhao, Y. Auxin overproduction in shoots cannot rescue auxin deficiencies in Arabidopsis roots. Plant Cell Physiol. 2014, 55, 1072–1079. [Google Scholar] [CrossRef]
- Zhang, M.; Hu, X.; Zhu, M.; Xu, M.; Wang, L. Transcription factors NF-YA2 and NF-YA10 regulate leaf growth via auxin signaling in Arabidopsis. Sci. Rep. 2017, 7, 1395. [Google Scholar] [CrossRef]
- Salazar-Iribe, A.; De-la-Pena, C. Auxins, the hidden player in chloroplast development. Plant Cell Rep. 2020, 39, 1595–1608. [Google Scholar] [CrossRef]
- Sagar, M.; Chervin, C.; Mila, I.; Hao, Y.; Roustan, J.P.; Benichou, M.; Gibon, Y.; Biais, B.; Maury, P.; Latche, A.; et al. SlARF4, an auxin response factor involved in the control of sugar metabolism during tomato fruit development. Plant Physiol. 2013, 161, 1362–1374. [Google Scholar] [CrossRef]
- Yuan, Y.; Mei, L.; Wu, M.; Wei, W.; Shan, W.; Gong, Z.; Zhang, Q.; Yang, F.; Yan, F.; Zhang, Q.; et al. SlARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development. J. Exp. Bot. 2018, 69, 5507–5518. [Google Scholar] [CrossRef]
- Zubo, Y.O.; Yamburenko, M.V.; Kusnetsov, V.V.; Borner, T. Methyl jasmonate, gibberellic acid, and auxin affect transcription and transcript accumulation of chloroplast genes in barley. J. Plant Physiol. 2011, 168, 1335–1344. [Google Scholar] [CrossRef] [PubMed]
- Melo, N.K.; Bianchetti, R.E.; Lira, B.S.; Oliveira, P.M.; Zuccarelli, R.; Dias, D.L.; Demarco, D.; Peres, L.E.; Rossi, M.; Freschi, L. Nitric Oxide, Ethylene, and Auxin Cross Talk Mediates Greening and Plastid Development in Deetiolating Tomato Seedlings. Plant Physiol. 2016, 170, 2278–2294. [Google Scholar] [CrossRef]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 2006, 20, 1790–1799. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Baba, S.; Obayashi, T.; Sato, M.; Toyooka, K.; Keränen, M.; Aro, E.-M.; Fukaki, H.; Ohta, H.; Sugimoto, K.; et al. Regulation of Root Greening by Light and Auxin/Cytokinin Signaling in Arabidopsis. Plant Cell 2012, 24, 1081–1095. [Google Scholar] [CrossRef]
- Cackett, L.; Luginbuehl, L.H.; Schreier, T.B.; Lopez-Juez, E.; Hibberd, J.M. Chloroplast development in green plant tissues: The interplay between light, hormone, and transcriptional regulation. New Phytol. 2021, 233, 2000–2016. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Krieger, A.M.; Yekutieli, D. Adaptive linear step-up procedures that control the false discovery rate. Biometrika 2006, 93, 491–507. [Google Scholar] [CrossRef]
- Ellis, C.M.; Nagpal, P.; Young, J.C.; Hagen, G.; Guilfoyle, T.J.; Reed, J.W. AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 2005, 132, 4563–4574. [Google Scholar] [CrossRef] [PubMed]
- Tseng, C.C.; Lee, C.J.; Chung, Y.T.; Sung, T.Y.; Hsieh, M.H. Differential regulation of Arabidopsis plastid gene expression and RNA editing in non-photosynthetic tissues. Plant Mol. Biol. 2013, 82, 375–392. [Google Scholar] [CrossRef]
- Bobik, K.; McCray, T.N.; Ernest, B.; Fernandez, J.C.; Howell, K.A.; Lane, T.; Staton, M.; Burch-Smith, T.M. The chloroplast RNA helicase ISE2 is required for multiple chloroplast RNA processing steps in Arabidopsis thaliana. Plant J. 2017, 91, 114–131. [Google Scholar] [CrossRef]
- Jiang, T.; Zhang, J.; Rong, L.; Feng, Y.; Wang, Q.; Song, Q.; Zhang, L.; Ouyang, M. ECD1 functions as an RNA-editing trans-factor of rps14-149 in plastids and is required for early chloroplast development in seedlings. J. Exp. Bot. 2018, 69, 3037–3051. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Tian, F.; Yu, J.; Zhang, Y.; Xie, Y.; Wu, B.; Miao, Y. MORF9 Functions in Plastid RNA Editing with Tissue Specificity. Int. J. Mol. Sci. 2019, 20, 4635. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Bassham, D. Using Arabidopsis Mesophyll Protoplasts to Study Unfolded Protein Response Signaling. Bio-Protocol 2018, 8, e3101. [Google Scholar] [CrossRef] [PubMed]
| Gene | Promoter Length | TGTCNN | TGTCTC | TGTCGG |
|---|---|---|---|---|
| ATPF editing factor 1 (AEF1) | 2031 | 9 | 0 | 0 |
| chloroplast biogenesis 19 (CLB19) | 822 | 5 | 0 | 0 |
| chloroplast RNA-binding protein 31a (CP31A) | 2096 | 7 | 2 | 0 |
| chloroplast RNA-binding protein 31b (CP31B) | 1935 | 11 | 0 | 0 |
| chloroplast RNA editing factor 3 (CREF3) | 1849 | 18 | 0 | 1 |
| chloroplast RNA editing factor 7 (CREF7) | 1678 | 19 | 1 | 0 |
| chlororespiratory reduction 2 (CRR2) | 1851 | 11 | 0 | 0 |
| chlororespiratory reduction 2 (CRR4) | 2143 | 18 | 2 | 0 |
| chlororespiratory reduction 21 (CRR21) | 1567 | 17 | 1 | 1 |
| chlororespiratory reduction 22 (CRR22) | 1817 | 19 | 2 | 0 |
| chlororespiratory reduction 28 (CRR28) | 2216 | 8 | 0 | 0 |
| delayed greening 1 (DG1) | 1550 | 16 | 1 | 0 |
| defectively organized tributaries 4 (DOT4) | 1149 | 8 | 1 | 1 |
| dyw domain protein 1 (DYW1) | 1267 | 4 | 0 | 0 |
| dyw domain protein 2 (DYW2) | 2106 | 16 | 2 | 0 |
| Arabidopsis early chloroplast biogenesis 2 (ECB2) | 1610 | 9 | 2 | 0 |
| early chloroplast development 1 (ECD1) | 2000 | 10 | 1 | 1 |
| ectopic lignification 1 (ELI1) | 2162 | 18 | 3 | 0 |
| hydroxymethylbilane synthase (HEMC) | 655 | 4 | 0 | 0 |
| increased size exclusion limit 2 (ISE2) | 1993 | 12 | 0 | 0 |
| low PSII accumulation 66 (LPA66) | 1291 | 6 | 1 | 0 |
| mitochondrial editing factor 37 (MEF37) | 1873 | 11 | 1 | 0 |
| multiple organellar rna editing factor 2 (MORF2) | 2073 | 13 | 2 | 0 |
| multiple organellar rna editing factor 5 (MORF5) | 1976 | 6 | 1 | 0 |
| multiple organellar rna editing factor 8 (MORF8) | 2109 | 18 | 2 | 0 |
| multiple organellar rna editing factor 9 (MORF9) | 1872 | 15 | 0 | 0 |
| NUWA (named after the well-known goddess in ancient Chinese mythology) | 1914 | 6 | 0 | 0 |
| overexpressor of cationic peroxidase 3 (OCP3) | 878 | 3 | 1 | 0 |
| organelle rna recognition motif protein 1 (ORRM1) | 1938 | 5 | 0 | 0 |
| organelle rna recognition motif protein 6 (ORMM6) | 1114 | 8 | 2 | 1 |
| organelle transcript processing 80 (OTP80) | 1873 | 9 | 0 | 0 |
| organelle transcript processing 81 (OTP81) | 1983 | 10 | 0 | 1 |
| organelle transcript processing 82 (OTP82) | 1801 | 2 | 0 | 0 |
| organelle transcript processing 84 (OTP84) | 1597 | 9 | 1 | 0 |
| organelle transcript processing 85 (OTP85) | 1967 | 11 | 2 | 0 |
| organelle transcript processing 86 (OTP86) | 913 | 5 | 1 | 0 |
| organelle transcript processing 90 (OTP90) | 2028 | 12 | 1 | 0 |
| organelle zinc finger 1 (OZ1) | 2129 | 10 | 2 | 0 |
| pentratricopeptide repeat protein pigment-defective mutant 1 (PDM1) | 1797 | 21 | 6 | 2 |
| pentratricopeptide repeat protein pigment-defective mutant 2 (PDM2) | 2028 | 16 | 1 | 1 |
| pentatricopeptide repeat protein for germination on NaCl (PGN) | 2202 | 15 | 1 | 1 |
| protoporphyrinogen ix oxidase 1 (PPO1) | 2045 | 19 | 1 | 0 |
| required for accD RNA editing 1 (RARE1) | 2188 | 12 | 1 | 0 |
| tRNA arginine adenosine deaminase (TADA) | 2075 | 10 | 0 | 0 |
| white to green 1 (WTG1) | 1623 | 11 | 0 | 1 |
| yellow seedling1 (YS1) | 1266 | 7 | 0 | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Z.-A.; Li, Y.; Liu, D.; Molloy, D.P.; Luo, Z.-F.; Li, H.-O.; Zhao, J.; Zhou, J.; Su, Y.; Wang, R.-Z.; et al. YUCCA2 (YUC2)-Mediated 3-Indoleacetic Acid (IAA) Biosynthesis Regulates Chloroplast RNA Editing by Relieving the Auxin Response Factor 1 (ARF1)-Dependent Inhibition of Editing Factors in Arabidopsis thaliana. Int. J. Mol. Sci. 2023, 24, 16988. https://doi.org/10.3390/ijms242316988
Li Z-A, Li Y, Liu D, Molloy DP, Luo Z-F, Li H-O, Zhao J, Zhou J, Su Y, Wang R-Z, et al. YUCCA2 (YUC2)-Mediated 3-Indoleacetic Acid (IAA) Biosynthesis Regulates Chloroplast RNA Editing by Relieving the Auxin Response Factor 1 (ARF1)-Dependent Inhibition of Editing Factors in Arabidopsis thaliana. International Journal of Molecular Sciences. 2023; 24(23):16988. https://doi.org/10.3390/ijms242316988
Chicago/Turabian StyleLi, Zi-Ang, Yi Li, Dan Liu, David P. Molloy, Zhou-Fei Luo, Hai-Ou Li, Jing Zhao, Jing Zhou, Yi Su, Ruo-Zhong Wang, and et al. 2023. "YUCCA2 (YUC2)-Mediated 3-Indoleacetic Acid (IAA) Biosynthesis Regulates Chloroplast RNA Editing by Relieving the Auxin Response Factor 1 (ARF1)-Dependent Inhibition of Editing Factors in Arabidopsis thaliana" International Journal of Molecular Sciences 24, no. 23: 16988. https://doi.org/10.3390/ijms242316988






