Orchestrating Cellular Balance: ncRNAs and RNA Interactions at the Dominant of Autophagy Regulation in Cancer
Abstract
:1. Introduction
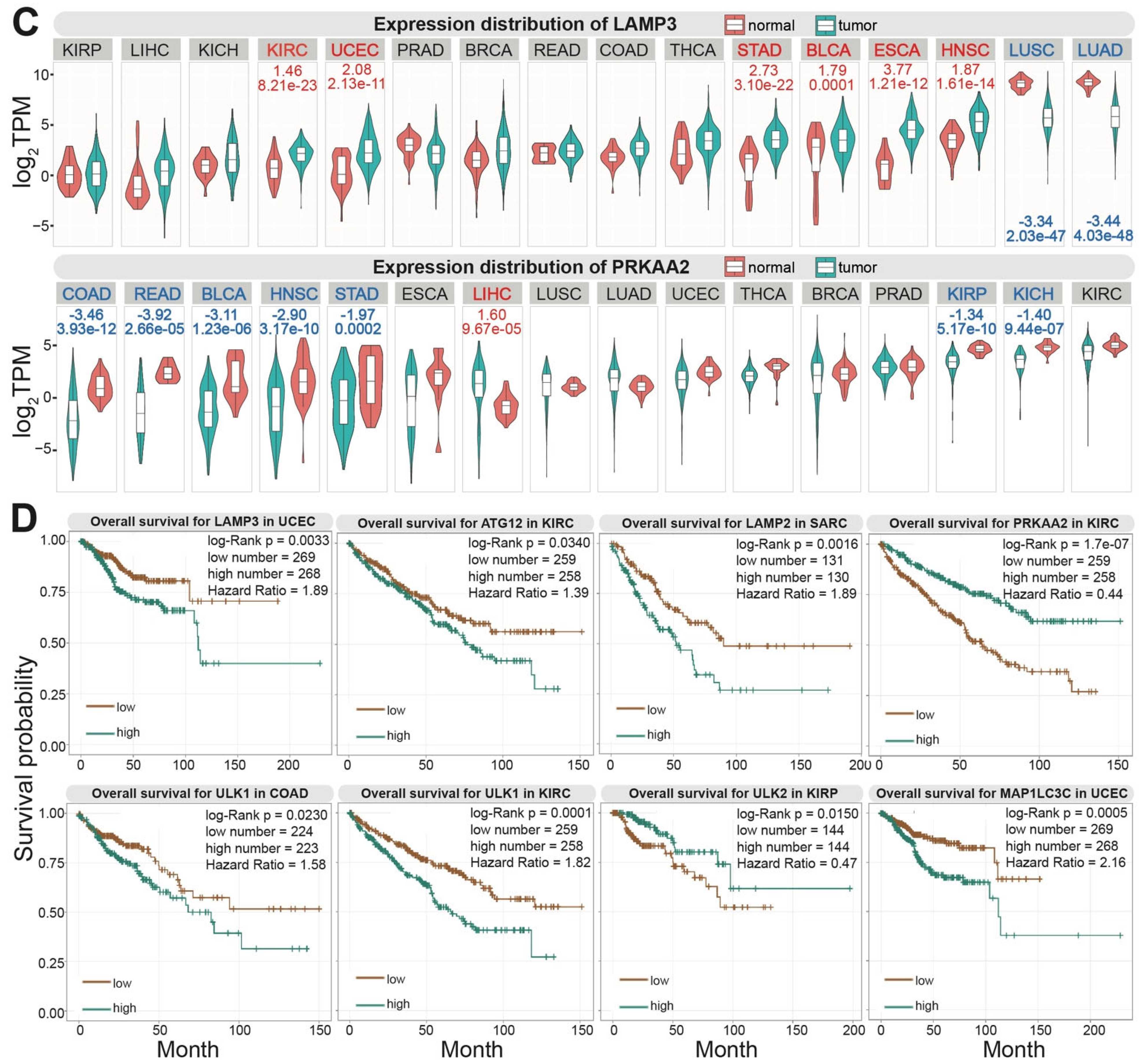
2. Autophagy-Related Genes and Cancer
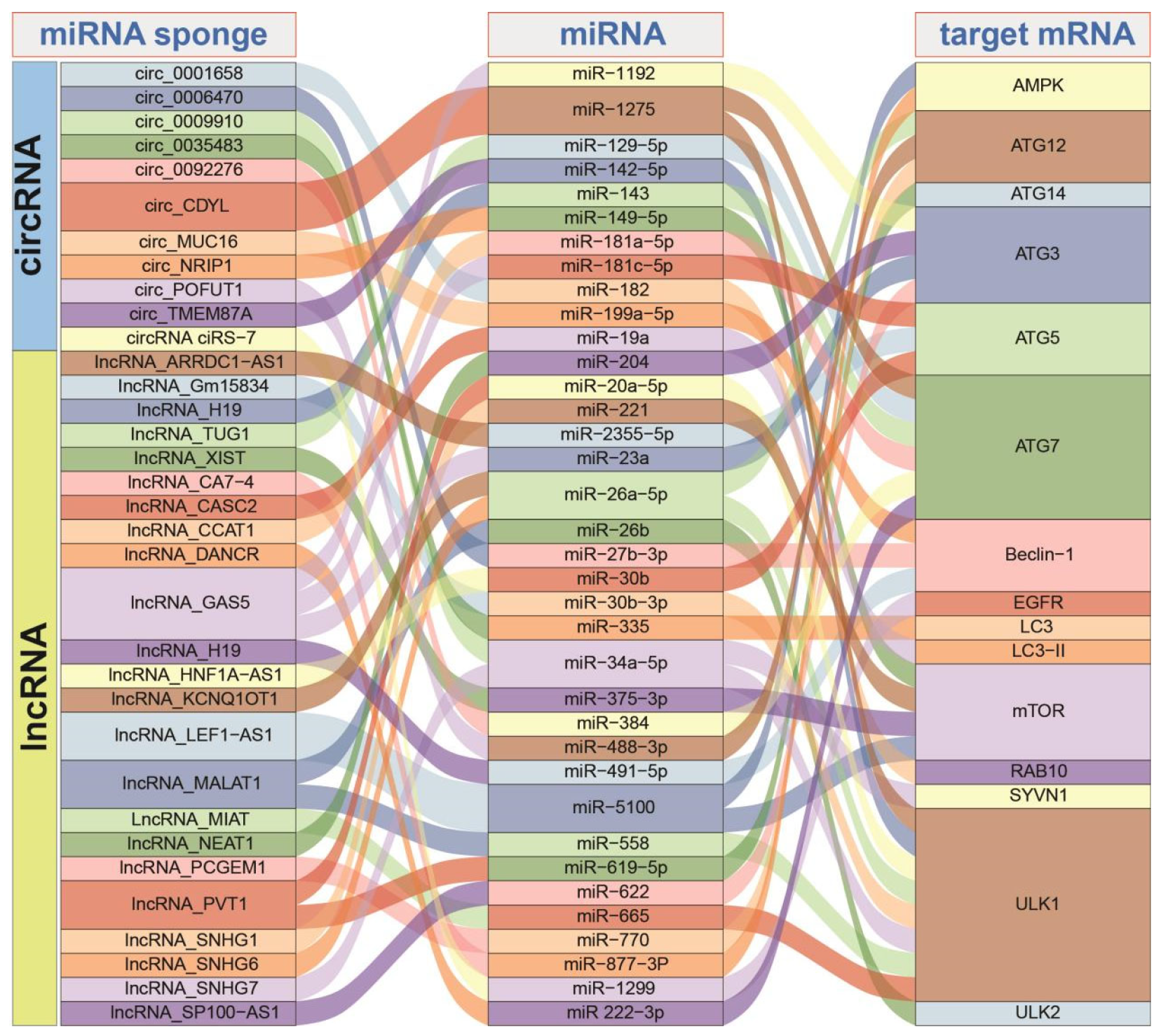
3. Regulation Roles of ncRNAs in Autophagy
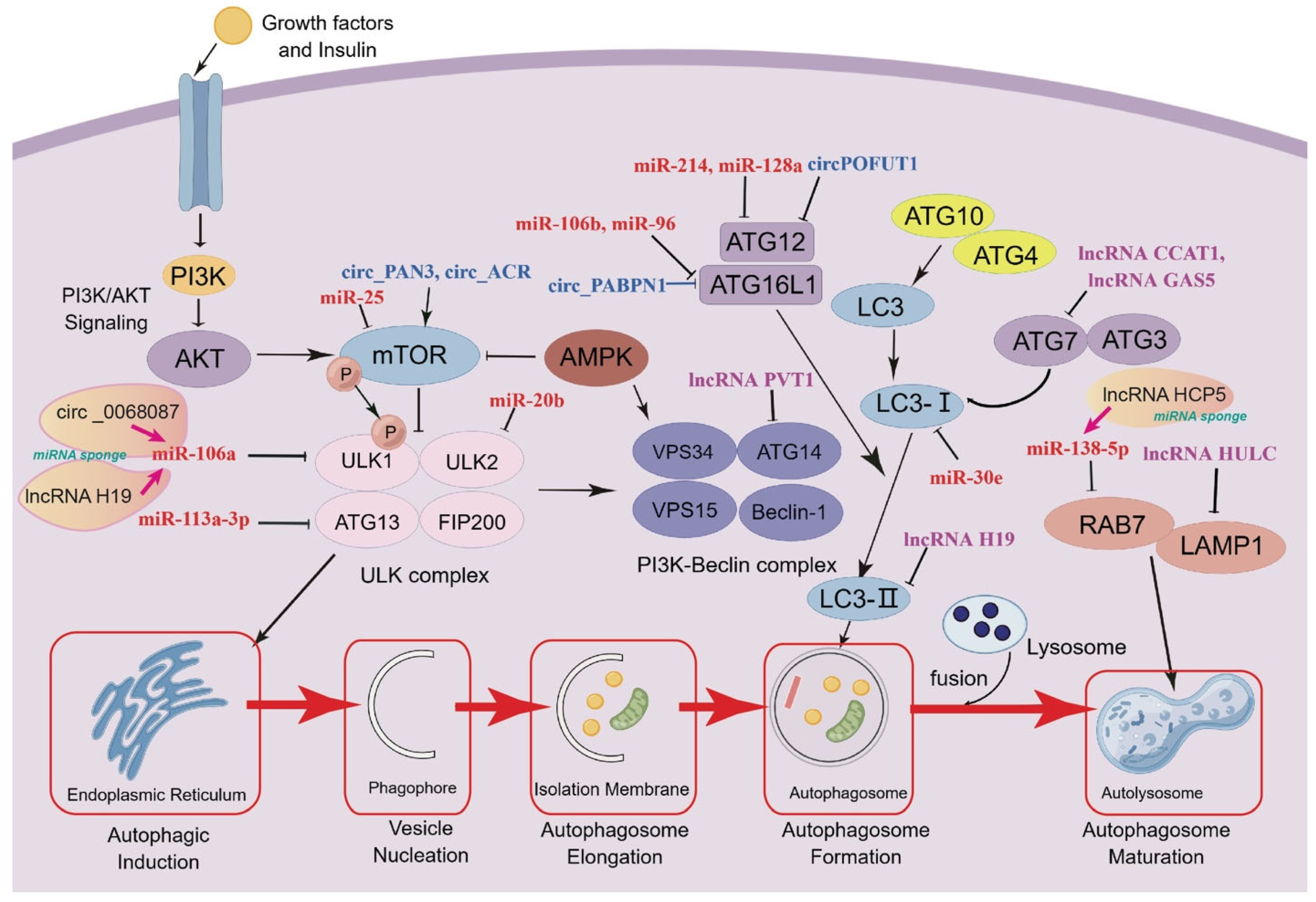
3.1. Regulation of ncRNAs in Autophagy Initiation
3.2. Regulation of ncRNAs in Vesicle Nucleation
3.3. Regulation of ncRNAs in Autophagic Vesicle Elongation
3.4. Regulation of ncRNAs in Autophagosome Formation and Maturation
4. Conclusions and Perspective
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yamamoto, K.; Iwadate, D.; Kato, H.; Nakai, Y.; Tateishi, K.; Fujishiro, M. Targeting autophagy as a therapeutic strategy against pancreatic cancer. J. Gastroenterol. 2022, 57, 603–618. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhang, X.; Yang, H.; Liang, T.; Bai, X. The “Self-eating” of cancer-associated fibroblast: A potential target for cancer. Biomed. Pharmacother. 2023, 163, 114762. [Google Scholar] [CrossRef] [PubMed]
- White, E. The role for autophagy in cancer. J. Clin. Investig. 2015, 125, 42–46. [Google Scholar] [CrossRef] [PubMed]
- White, E.; Mehnert, J.M.; Chan, C.S. Autophagy, Metabolism, and Cancer. Clin. Cancer Res. 2015, 21, 5037–5046. [Google Scholar] [CrossRef] [PubMed]
- Manent, J.; Banerjee, S.; de Matos Simoes, R.; Zoranovic, T.; Mitsiades, C.; Penninger, J.M.; Simpson, K.J.; Humbert, P.O.; Richardson, H.E. Autophagy suppresses Ras-driven epithelial tumourigenesis by limiting the accumulation of reactive oxygen species. Oncogene 2017, 36, 5576–5592. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Xu, L.; Wang, X.; Zhang, D.; Sun, G.; Wang, M.; Wang, M.; Han, Y.; Chai, R.; Wang, H.P. RDX1 activates autophagy via the PTEN-AKT signaling pathway to protect against cisplatin-induced spiral ganglion neuron damage. Autophagy 2021, 17, 4159–4181. [Google Scholar] [CrossRef]
- Zada, S.; Hwang, J.S.; Ahmed, M.; Lai, T.H.; Pham, T.M.; Elashkar, O.; Kim, D.R. Cross talk between autophagy and oncogenic signaling pathways and implications for cancer therapy. Biochim. Biophys. Acta Rev. Cancer 2021, 1876, 188565. [Google Scholar] [CrossRef]
- Amaravadi, R.; Kimmelman, A.C.; White, E. Recent insights into the function of autophagy in cancer. Genes Dev. 2016, 30, 1913–1930. [Google Scholar] [CrossRef]
- Li, X.; He, S.; Ma, B. Autophagy and autophagy-related proteins in cancer. Mol. Cancer 2020, 19, 12. [Google Scholar] [CrossRef]
- Slack, F.J.; Chinnaiyan, A.M. The role of non-coding RNAs in oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef]
- Kang, M.R.; Kim, M.S.; Oh, J.E.; Kim, Y.R.; Song, S.Y.; Kim, S.S.; Ahn, C.H.; Yoo, N.J.; Lee, S.H. Frameshift mutations of autophagy-related genes ATG2B, ATG5, ATG9B and ATG12 in gastric and colorectal cancers with microsatellite instability. J. Pathol. 2009, 217, 702–706. [Google Scholar] [CrossRef]
- Frangež, Ž.; Gérard, D.; He, Z.; Gavriil, M.; Fernández-Marrero, Y.; Seyed Jafari, S.M.; Hunger, R.E.; Lucarelli, P.; Yousefi, S.; Sauter, T.; et al. ATG5 and ATG7 Expression Levels Are Reduced in Cutaneous Melanoma and Regulated by NRF1. Front. Oncol. 2021, 11, 721624. [Google Scholar] [CrossRef]
- Wang, K. Autophagy and apoptosis in liver injury. Cell Cycle 2015, 14, 1631–1642. [Google Scholar] [CrossRef]
- Ren, Y.; Chen, Y.; Liang, X.; Lu, Y.; Pan, W.; Yang, M. MiRNA-638 promotes autophagy and malignant phenotypes of cancer cells via directly suppressing DACT3. Cancer Lett. 2017, 390, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, X.; Zhang, T.; Su, L.; Liu, B.; Zhu, Z.; Li, C. LncRNA MALAT1 promotes gastric cancer progression via inhibiting autophagic flux and inducing fibroblast activation. Cell Death Dis. 2021, 12, 368. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, H.; Zou, J.; Cao, G.; Li, Y.; Xing, C.; Wu, J. Exosomal circ_0091741 promotes gastric cancer cell autophagy and chemoresistance via the miR-330-3p/TRIM14/Dvl2/Wnt/beta-catenin axis. Hum. Cell 2023, 36, 258–275. [Google Scholar] [CrossRef]
- Liang, J.; Zhang, L.; Cheng, W. Non-coding RNA-mediated autophagy in cancer: A protumor or antitumor factor? Biochim. Biophys. Acta Rev. Cancer 2021, 1876, 188642. [Google Scholar] [CrossRef]
- Barnwal, S.K.; Bendale, H.; Banerjee, S. Non-coding RNAs associated with autophagy and their regulatory role in cancer therapeutics. Mol. Biol. Rep. 2022, 49, 7025–7037. [Google Scholar] [CrossRef] [PubMed]
- Levy, J.M.M.; Towers, C.G.; Thorburn, A. Targeting autophagy in cancer. Nat. Rev. Cancer 2017, 17, 528–542. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N.; Yoshimori, T.; Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell Dev. Biol. 2011, 27, 107–132. [Google Scholar] [CrossRef]
- Dowdle, W.E.; Nyfeler, B.; Nagel, J.; Elling, R.A.; Liu, S.; Triantafellow, E.; Menon, S.; Wang, Z.; Honda, A.; Pardee, G.; et al. Selective VPS34 inhibitor blocks autophagy and uncovers a role for NCOA4 in ferritin degradation and iron homeostasis in vivo. Nat. Cell Biol. 2014, 16, 1069–1079. [Google Scholar] [CrossRef]
- Rabinowitz, J.D.; White, E. Autophagy and metabolism. Science 2010, 330, 1344–1348. [Google Scholar] [CrossRef]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of cells and tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef] [PubMed]
- Karsli-Uzunbas, G.; Guo, J.Y.; Price, S.; Teng, X.; Laddha, S.V.; Khor, S.; Kalaany, N.Y.; Jacks, T.; Chan, C.S.; Rabinowitz, J.D.; et al. Autophagy is required for glucose homeostasis and lung tumor maintenance. Cancer Discov. 2014, 4, 914–927. [Google Scholar] [CrossRef]
- Kuma, A.; Hatano, M.; Matsui, M.; Yamamoto, A.; Nakaya, H.; Yoshimori, T.; Ohsumi, Y.; Tokuhisa, T.; Mizushima, N. The role of autophagy during the early neonatal starvation period. Nature 2004, 432, 1032–1036. [Google Scholar] [CrossRef] [PubMed]
- Takamura, A.; Komatsu, M.; Hara, T.; Sakamoto, A.; Kishi, C.; Waguri, S.; Eishi, Y.; Hino, O.; Tanaka, K.; Mizushima, N. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 2011, 25, 795–800. [Google Scholar] [CrossRef]
- Mathew, R.; Karp, C.M.; Beaudoin, B.; Vuong, N.; Chen, G.; Chen, H.Y.; Bray, K.; Reddy, A.; Bhanot, G.; Gelinas, C.; et al. Autophagy suppresses tumorigenesis through elimination of p62. Cell 2009, 137, 1062–1075. [Google Scholar] [CrossRef]
- Rosenfeldt, M.T.; O’Prey, J.; Morton, J.P.; Nixon, C.; MacKay, G.; Mrowinska, A.; Au, A.; Rai, T.S.; Zheng, L.; Ridgway, R.; et al. p53 status determines the role of autophagy in pancreatic tumour development. Nature 2013, 504, 296–300. [Google Scholar] [CrossRef]
- Jing, Z.; He, X.; Jia, Z.; Sa, Y.; Yang, B.; Liu, P.N. CAPD2 inhibits autophagy by regulating Ca(2+)/CAMKK2/AMPK/mTORC1 pathway and PARP-1/SIRT1 axis to promote colorectal cancer. Cancer Lett. 2021, 520, 26–37. [Google Scholar] [CrossRef] [PubMed]
- You, B.; Xia, T.; Gu, M.; Zhang, Z.; Zhang, Q.; Shen, J.; Fan, Y.; Yao, H.; Pan, S.; Lu, Y.; et al. AMPK-mTOR-Mediated Activation of Autophagy Promotes Formation of Dormant Polyploid Giant Cancer Cells. Cancer Res. 2022, 82, 846–858. [Google Scholar] [CrossRef]
- Cicchini, M.; Chakrabarti, R.; Kongara, S.; Price, S.; Nahar, R.; Lozy, F.; Zhong, H.; Vazquez, A.; Kang, Y.; Karantza, V. Autophagy regulator BECN1 suppresses mammary tumorigenesis driven by WNT1 activation and following parity. Autophagy 2014, 10, 2036–2052. [Google Scholar] [CrossRef] [PubMed]
- Al-Bari, M.A.A.; Xu, P. Molecular regulation of autophagy machinery by mTOR-dependent and -independent pathways. Ann. N. Y. Acad. Sci. 2020, 1467, 3–20. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.; Li, J.; Yang, K.; Cao, D. An overview of autophagy: Mechanism, regulation and research progress. Bull. Cancer 2021, 108, 304–322. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Chen, Y.; Tooze, S.A. Autophagy pathway: Cellular and molecular mechanisms. Autophagy 2018, 14, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kundu, M.; Viollet, B.; Guan, K.L.A. MPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 2011, 13, 132–141. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, H. Regulation of Autophagy by mTOR Signaling Pathway. Adv. Exp. Med. Biol. 2019, 1206, 67–83. [Google Scholar]
- Kocaturk, N.M.; Akkoc, Y.; Kig, C.; Bayraktar, O.; Gozuacik, D.; Kutlu, O. Autophagy as a molecular target for cancer treatment. Eur. J. Pharm. Sci. 2019, 134, 116–137. [Google Scholar] [CrossRef]
- Murugan, A.K. mTOR: Role in cancer, metastasis and drug resistance. Semin. Cancer Biol. 2019, 59, 92–111. [Google Scholar] [CrossRef]
- Gulia, S.; Chandra, P.; Das, A. The Prognosis of Cancer Depends on the Interplay of Autophagy, Apoptosis, and Anoikis within the Tumor Microenvironment. Cell Biochem. Biophys. 2023, 81, 621–658. [Google Scholar] [CrossRef]
- Maeser, D.; Gruener, R.F.; Huang, R.S. oncoPredict: An R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief Bioinform. 2021, 22, bbab260. [Google Scholar] [CrossRef]
- Yang, W.; Soares, J.; Greninger, P.; Edelman, E.J.; Lightfoot, H.; Forbes, S.; Bindal, N.; Beare, D.; Smith, J.A.; Thompson, I.R.; et al. Genomics of Drug Sensitivity in Cancer (GDSC): A resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013, 41, D955–D961. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef]
- Li, S.; Qiang, Q.; Shan, H.; Shi, M.; Gan, G.; Ma, F.; Chen, B. MiR-20a and miR-20b negatively regulate autophagy by targeting RB1CC1/FIP200 in breast cancer cells. Life Sci. 2016, 147, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, S.I.; Gautschi, O.; Batliner, J.; Gugger, M.; Fey, M.F.; Tschan, M.P. MicroRNA-106a targets autophagy and enhances sensitivity of lung cancer cells to Src inhibitors. Lung Cancer 2017, 107, 73–83. [Google Scholar] [CrossRef]
- Soni, M.; Patel, Y.; Markoutsa, E.; Jie, C.; Liu, S.; Xu, P.; Chen, H. Autophagy, cell viability, and chemoresistance are regulated by miR-489 in breast cancer. Mol. Cancer Res. 2018, 16, 1348–1360. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, N.; Liu, P.; Chen, Q.; Situ, H.; Xie, T.; Zhang, J.; Peng, C.; Lin, Y.; Chen, J. MicroRNA-25 regulates chemoresistance-associated autophagy in breast cancer cells, a process modulated by the natural autophagy inducer isoliquiritigenin. Oncotarget 2014, 5, 7013–7026. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, G.; Xu, P.; Lv, J.; Zhang, X.; Zhang, L.; Wang, S.; Cao, J.; Xia, Y.; Xuan, Z.; et al. Circular RNA TMEM87A promotes cell proliferation and metastasis of gastric cancer by elevating ULK1 via sponging miR-142-5p. J. Gastroenterol. 2021, 56, 125–138. [Google Scholar] [CrossRef]
- Duan, X.; Zhang, T.; Ding, S.; Wei, J.; Su, C.; Liu, H.; Xu, G. microRNA-17-5p modulates bacille calmette-guerin growth in RAW264. 7 cells by targeting ULK1. PLoS ONE 2015, 10, e0138011. [Google Scholar] [CrossRef]
- Meng, C.; Liu, Y.; Shen, Y.; Liu, S.; Wang, Z.; Ye, Q.; Liu, H.; Liu, X.; Jia, L. MicroRNA-26b suppresses autophagy in breast cancer cells by targeting DRAM1 mRNA, and is downregulated by irradiation. Oncol. Lett. 2018, 15, 1435–1440. [Google Scholar]
- Huang, Y.; Chuang, A.Y.; Ratovitski, E.A. Phospho-ΔNp63α/miR-885-3p axis in tumor cell life and cell death upon cisplatin exposure. Cell Cycle 2011, 10, 3938–3947. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, Z.; Xuan, Z.; Xu, P.; Wang, W.; Chen, Z.; Wang, S.; Sun, G.; Xu, J.; Xu, Z. Novel role of miR-133a-3p in repressing gastric cancer growth and metastasis via blocking autophagy-mediated glutaminolysis. J. Exp. Clin. Cancer Res. 2018, 37, 320. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Wang, F.; Hu, S.; Yin, C.; Li, X.; Zhao, S.; Wang, J.; Yan, X. MiR-20a and miR-106b negatively regulate autophagy induced by leucine deprivation via suppression of ULK1 expression in C2C12 myoblasts. Cell Signal. 2012, 24, 2179–2186. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhong, L.; Li, P.; Zhao, P. MicroRNA-100 Enhances Autophagy and Suppresses Migration and Invasion of Renal Cell Carcinoma Cells via Disruption of NOX4-Dependent mTOR Pathway. Clin. Transl. Sci. 2022, 15, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yu, Y.; Li, S.; Liu, Y.; Zhou, S.; Cao, S.; Yin, J.; Li, G. Micro RNA-30a ameliorates hepatic fibrosis by inhibiting Beclin1-mediated autophagy. J. Cell. Mol. Med. 2017, 21, 3679–3692. [Google Scholar] [CrossRef]
- Huang, T.; Wan, X.; Alvarez, A.A.; James, C.D.; Song, X.; Yang, Y.; Sastry, N.; Nakano, I.; Sulman, E.P.; Hu, B.M. IR93 (microRNA-93) regulates tumorigenicity and therapy response of glioblastoma by targeting autophagy. Autophagy 2019, 15, 1100–1111. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, B.; Long, H.; Yu, J.; Li, F.; Hou, H.; Yang, Q. Decreased miR-124-3p Expression Prompted Breast Cancer Cell Progression Mainly by Targeting Beclin-1. Clin. Lab. 2016, 62, 1139–1145. [Google Scholar] [CrossRef]
- Pan, W.; Zhong, Y.; Cheng, C.; Liu, B.; Wang, L.; Li, A.; Xiong, L.; Liu, S. MiR-30-Regulated Autophagy Mediates Angiotensin II-Induced Myocardial Hypertrophy. PLoS ONE 2013, 8, e53950. [Google Scholar] [CrossRef]
- Singh, S.V.; Dakhole, A.N.; Deogharkar, A.; Kazi, S.; Kshirsagar, R.; Goel, A.; Moiyadi, A.; Jalali, R.; Sridhar, E.; Gupta, T.; et al. Restoration of miR-30a expression inhibits growth, tumorigenicity of medulloblastoma cells accompanied by autophagy inhibition. Biochem. Biophys. Res. Commun. 2017, 491, 946–952. [Google Scholar] [CrossRef]
- Lai, L.; Chen, J.; Wang, N.; Zhu, G.; Duan, X.; Ling, F. MiRNA-30e mediated cardioprotection of ACE2 in rats with Doxorubicin-induced heart failure through inhibiting cardiomyocytes autophagy. Life Sci. 2017, 169, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Xu, J.; Zhao, J.; Bai, J. Mir-30d suppresses cell proliferation of colon cancer cells by inhibiting cell autophagy and promoting cell apoptosis. Tumor Biol. 2017, 39, 1010428317703984. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Jin, B.; Chen, L.; Zhuo, D.; Zhang, Z.; Gong, K.; Mao, Z. MiR-30d induces apoptosis and is regulated by the Akt/FOXO pathway in renal cell carcinoma. Cell Signal. 2013, 25, 1212–1221. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Han, X.; Hu, Z.; Chen, L. The PVT1/miR-216b/Beclin-1 regulates cisplatin sensitivity of NSCLC cells via modulating autophagy and apoptosis. Cancer Chemother. Pharmacol. 2019, 83, 921–931. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, A.; Chattopadhyay, D.; Chakrabarti, G. miR-17-5p downregulation contributes to paclitaxel resistance of lung cancer cells through altering beclin1 expression. PLoS ONE 2014, 9, e95716. [Google Scholar] [CrossRef] [PubMed]
- Moradi Marjaneh, R.; Khazaei, M.; Ferns, G.A.; Avan, A.; Aghaee-Bakhtiari, S.H. MicroRNAs as potential therapeutic targets to predict responses to oxaliplatin in colorectal cancer: From basic evidence to therapeutic implication. IUBMB Life 2019, 71, 1428–1441. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.-A.; Zhou, B.-R.; Xu, Y.; Chen, X.; Liu, J.; Gozali, M.; Wu, D.; Yin, Z.-Q.; Luo, D. MiR-23a-depressed autophagy is a participant in PUVA-and UVB-induced premature senescence. Oncotarget 2016, 7, 37420. [Google Scholar] [CrossRef]
- Ganesan, S.; Palani, H.K.; Lakshmanan, V.; Balasundaram, N.; Alex, A.A.; David, S.; Venkatraman, A.; Korula, A.; George, B.; Balasubramanian, P.; et al. Stromal cells downregulate miR-23a-5p to activate protective autophagy in acute myeloid leukemia. Cell Death Dis. 2019, 10, 736. [Google Scholar] [CrossRef]
- Capizzi, M.; Strappazzon, F.; Cianfanelli, V.; Papaleo, E.; Cecconi, F.M. IR7–3HG, a MYC-dependent modulator of cell proliferation, inhibits autophagy by a regulatory loop involving AMBRA1. Autophagy 2017, 13, 554–566. [Google Scholar] [CrossRef]
- Lian, W.-S.; Ko, J.-Y.; Wu, R.-W.; Sun, Y.-C.; Chen, Y.-S.; Wu, S.-L.; Weng, L.-H.; Jahr, H.; Wang, F.-S. MicroRNA-128a represses chondrocyte autophagy and exacerbates knee osteoarthritis by disrupting Atg12. Cell Death Dis. 2018, 9, 919. [Google Scholar] [CrossRef]
- Sun, L.; Liu, A.; Zhang, J.; Ji, W.; Li, Y.; Yang, X.; Wu, Z.; Guo, J. miR-23b improves cognitive impairments in traumatic brain injury by targeting ATG12-mediated neuronal autophagy. Behav. Brain Res. 2018, 340, 126–136. [Google Scholar] [CrossRef]
- Hu, J.L.; He, G.Y.; Lan, X.L.; Zeng, Z.C.; Guan, J.; Ding, Y.; Qian, X.L.; Liao, W.T.; Ding, Y.Q.; Liang, L. Inhibition of ATG12-mediated autophagy by miR-214 enhances radiosensitivity in colorectal cancer. Oncogenesis 2018, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Zhai, Z.; Wu, F.; Chuang, A.Y.; Kwon, J.H. miR-106b fine tunes ATG16L1 expression and autophagic activity in intestinal epithelial HCT116 cells. Inflamm. Bowel. Dis. 2013, 19, 2295–2301. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Guo, X.; Yu, C.; Sun, C.; Jiang, J. miR-138-5p suppresses autophagy in pancreatic cancer by targeting SIRT1. Oncotarget 2017, 8, 11071–11082. [Google Scholar] [CrossRef] [PubMed]
- Bao, L.; Lv, L.; Feng, J.; Chen, Y.; Wang, X.; Han, S.; Zhao, H. miR-487b-5p regulates temozolomide resistance of lung cancer cells through LAMP2-medicated autophagy. DNA Cell Biol. 2016, 35, 385–392. [Google Scholar] [CrossRef]
- Pennati, M.; Lopergolo, A.; Profumo, V.; De Cesare, M.; Sbarra, S.; Valdagni, R.; Zaffaroni, N.; Gandellini, P.; Folini, M. miR-205 impairs the autophagic flux and enhances cisplatin cytotoxicity in castration-resistant prostate cancer cells. Biochem. Pharmacol. 2014, 87, 579–597. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Jiang, J.; Liu, W.; Wang, H.; Zhao, L.; Liu, S.; Li, P.; Zhang, S.; Sun, C.; Wu, Y. microRNA-378 promotes autophagy and inhibits apoptosis in skeletal muscle. Proc. Natl. Acad. Sci. USA 2018, 115, E10849–E10858. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Xie, Y.; Zhou, Z.; Wu, Z.; Dai, X.; Xu, B. Curcumin regulates the progression of colorectal cancer via LncRNA NBR2/AMPK pathway. Technol. Cancer Res. Treat. 2019, 18, 1533033819870781. [Google Scholar] [CrossRef]
- Tang, S.; Tan, G.; Jiang, X.; Han, P.; Zhai, B.; Dong, X.; Qiao, H.; Jiang, H.; Sun, X. An artificial lncRNA targeting multiple miRNAs overcomes sorafenib resistance in hepatocellular carcinoma cells. Oncotarget 2016, 7, 73257. [Google Scholar] [CrossRef]
- Wang, X.; Lan, Z.; He, J.; Lai, Q.; Yao, X.; Li, Q.; Liu, Y.; Lai, H.; Gu, C.; Yan, Q.; et al. LncRNA SNHG6 promotes chemoresistance through ULK1-induced autophagy by sponging miR-26a-5p in colorectal cancer cells. Cancer Cell Int. 2019, 19, 234. [Google Scholar] [CrossRef]
- Li, Z.; Li, J.; Tang, N. Long noncoding RNA Malat1 is a potent autophagy inducer protecting brain microvascular endothelial cells against oxygen-glucose deprivation/reoxygenation-induced injury by sponging miR-26b and upregulating ULK2 expression. Neuroscience 2017, 354, 1–10. [Google Scholar] [CrossRef]
- Zhuo, C.; Jiang, R.; Lin, X.; Shao, M. LncRNA H19 inhibits autophagy by epigenetically silencing of DIRAS3 in diabetic cardiomyopathy. Oncotarget 2017, 8, 1429. [Google Scholar] [CrossRef]
- Qian, C.; Ye, Y.; Mao, H.; Yao, L.; Sun, X.; Wang, B.; Zhang, H.; Xie, L.; Zhang, H.; Zhang, Y.; et al. Downregulated lncRNA-SNHG1 enhances autophagy and prevents cell death through the miR-221/222/p27/mTOR pathway in Parkinson’s disease. Exp. Cell Res. 2019, 384, 111614. [Google Scholar] [CrossRef]
- Chen, Y.-N.; Cai, M.-Y.; Xu, S.; Meng, M.; Ren, X.; Yang, J.W.; Dong, Y.-Q.; Liu, X.; Yang, J.-M.; Xiong, X.-D. Identification of the lncRNA, AK156230, as a novel regulator of cellular senescence in mouse embryonic fibroblasts. Oncotarget 2016, 7, 52673. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-L.; Tseng, Y.-W.; Wu, J.-C.; Chen, G.-Y.; Lin, K.-C.; Hwang, S.-M.; Hu, Y.-C. Suppression of hepatocellular carcinoma by baculovirus-mediated expression of long non-coding RNA PTENP1 and MicroRNA regulation. Biomaterials 2015, 44, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Kats, L.; Salmena, L.; Weiss, D.; Tan, S.M.; Ala, U.; Karreth, F.; Poliseno, L.; Provero, P.; Di Cunto, F. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011, 147, 344–357. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Yao, R.; Huang, F.; Yi, J. LncRNA SNHG12 as a potent autophagy inducer exerts neuroprotective effects against cerebral ischemia/reperfusion injury. Biochem. Biophys. Res. Commun. 2019, 514, 490–496. [Google Scholar] [CrossRef]
- Ma, B.; Yuan, Z.; Zhang, L.; Lv, P.; Yang, T.; Gao, J.; Pan, N.; Wu, Q.; Lou, J.; Han, C. Long non-coding RNA AC023115. 3 suppresses chemoresistance of glioblastoma by reducing autophagy. Biochim. Biophys. Acta Mol. Cell Res. 2017, 1864, 1393–1404. [Google Scholar] [CrossRef]
- Zhou, C.; Yi, C.; Yi, Y.; Qin, W.; Yan, Y.; Dong, X.; Zhang, X.; Huang, Y.; Zhang, R.; Wei, J. LncRNA PVT1 promotes gemcitabine resistance of pancreatic cancer via activating Wnt/β-catenin and autophagy pathway through modulating the miR-619-5p/Pygo2 and miR-619-5p/ATG14 axes. Mol. Cancer 2020, 19, 118. [Google Scholar] [CrossRef]
- Luo, Y.; Zheng, S.; Wu, Q.; Wu, J.; Zhou, R.; Wang, C.; Wu, Z.; Rong, X.; Huang, N.; Sun, L.; et al. Long noncoding RNA (lncRNA) EIF3J-DT induces chemoresistance of gastric cancer via autophagy activation. Autophagy 2021, 17, 4083–4101. [Google Scholar] [CrossRef]
- Yan, W.; Chen, Z.-Y.; Chen, J.-Q.; Chen, H.-M. LncRNA NEAT1 promotes autophagy in MPTP-induced Parkinson’s disease through stabilizing PINK1 protein. Biochem. Biophys. Res. Commun. 2018, 496, 1019–1024. [Google Scholar] [CrossRef]
- Guo, J.; Ma, Y.; Peng, X.; Jin, H.; Liu, J. LncRNA CCAT1 promotes autophagy via regulating ATG7 by sponging miR-181 in hepatocellular carcinoma. J. Cell. Biochem. 2019, 120, 17975–17983. [Google Scholar] [CrossRef]
- Song, J.; Ahn, C.; Chun, C.H.; Jin, E.J. A long non-coding RNA, GAS5, plays a critical role in the regulation of miR-21 during osteoarthritis. J. Orthop. Res. 2014, 32, 1628–1635. [Google Scholar] [CrossRef]
- Liu, Z.; Wei, X.; Zhang, A.; Li, C.; Bai, J.; Dong, J. Long non-coding RNA HNF1A-AS1 functioned as an oncogene and autophagy promoter in hepatocellular carcinoma through sponging hsa-miR-30b-5p. Biochem. Biophys. Res. Commun. 2016, 473, 1268–1275. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zhang, X.; Li, H.; Liu, J. The long noncoding RNA HOTAIR activates autophagy by upregulating ATG3 and ATG7 in hepatocellular carcinoma. Mol. Biosyst. 2016, 12, 2605–2612. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Wu, D.-D.; Sang, X.-B.; Wang, L.-L.; Zong, Z.-H.; Sun, K.-X.; Liu, B.-L.; Zhao, Y. The lncRNA HULC functions as an oncogene by targeting ATG7 and ITGB1 in epithelial ovarian carcinoma. Cell Death Dis. 2017, 8, e3118. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.-X.; Miao, C.-F.; Sang, L.-N.; Huang, Y.-M.; Zhang, R.; Sun, L.; Jiang, Z.-X. Circ_0009910 promotes imatinib resistance through ULK1-induced autophagy by sponging miR-34a-5p in chronic myeloid leukemia. Life Sci. 2020, 243, 117255. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Ling, Y.; Mehrpour, M.; Saw, P.E.; Liu, Z.; Tan, W.; Tian, Z.; Zhong, W.; Lin, W.; Luo, Q.; et al. Autophagy-associated circRNA circCDYL augments autophagy and promotes breast cancer progression. Mol. Cancer 2020, 19, 65. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Chen, W.-M.; Liu, S.; Wang, Z.-H.; Wei, T.-N.; Chen, Z.-Z.; Wu, W.-B. CircPAN3 contributes to drug resistance in acute myeloid leukemia through regulation of autophagy. Leuk. Res. 2019, 85, 106198. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chen, X.; Yao, J.; Kang, J. Circular RNA ACR relieves high glucose-aroused RSC96 cell apoptosis and autophagy via declining microRNA-145-3p. J. Cell. Biochem. 2019, 122, 1252. [Google Scholar]
- Meng, L.; Liu, S.; Ding, P.; Chang, S.; Sang, M. Circular RNA ciRS-7 inhibits autophagy of ESCC cells by functioning as miR-1299 sponge to target EGFR signaling. J. Cell. Biochem. 2020, 121, 1039–1049. [Google Scholar] [CrossRef]
- Gan, X.; Zhu, H.; Jiang, X.; Obiegbusi, S.C.; Yong, M.; Long, X.; Hu, J. CircMUC16 promotes autophagy of epithelial ovarian cancer via interaction with ATG13 and miR-199a. Mol. Cancer 2020, 19, 45. [Google Scholar] [CrossRef]
- Luo, M.; Deng, X.; Chen, Z.; Hu, Y.C. Circular RNA circPOFUT1 enhances malignant phenotypes and autophagy-associated chemoresistance via sequestrating miR-488-3p to activate the PLAG1-ATG12 axis in gastric cancer. Cell Death Dis. 2023, 14, 10. [Google Scholar] [CrossRef]
- Wang, Q.; Liang, D.; Shen, P.; Yu, Y.; Yan, Y.; You, W. Hsa_circ_0092276 promotes doxorubicin resistance in breast cancer cells by regulating autophagy via miR-348/ATG7 axis. Transl Oncol 2021, 14, 101045. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Liu, G.; Cao, H.; Zhang, H.; Shao, F. Hsa_circ_0035483 sponges hsa-miR-335 to promote the gemcitabine-resistance of human renal cancer cells by autophagy regulation. Biochem. Biophys. Res. Commun. 2019, 519, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-X.; Xiao, L.; Chung, H.K.; Ma, X.-X.; Liu, X.; Song, J.-L.; Jin, C.Z.; Rao, J.N.; Gorospe, M.; Wang, J.-Y. Interaction between HuR and circPABPN1 Modulates Autophagy in the Intestinal Epithelium by Altering ATG16L1 Translation. Mol. Cell. Biol. 2020, 40, e00492-19. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Li, X.; Wang, K.; Zhu, D.; Meng, B.; Liu, J.; Liang, X.; Jin, Y.; Liu, X.; Wen, Q.; et al. PURPL represses autophagic cell death to promote cutaneous melanoma by modulating ULK1 phosphorylation. Cell Death Dis. 2021, 12, 1070. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xie, S.; Yang, J.; Xiong, H.; Jia, Y.; Zhou, Y.; Chen, Y.; Ying, X.; Chen, C.; Ye, C. The long noncoding RNA H19 promotes tamoxifen resistance in breast cancer via autophagy. J. Hematol. Oncol. 2019, 12, 81. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Ma, J.; Yan, L.; Li, T.; Li, Z.; Han, X.; Shui, S. Down-regulation of Lncrna MALAT1 attenuates neuronal cell death through suppressing Beclin1-dependent autophagy by regulating Mir-30a in cerebral ischemic stroke. Cell. Physiol. Biochem. 2017, 43, 182–194. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Zeng, Z.; Zhang, Y.; Li, H.; Cheng, H.; Wang, J.; Wu, F. CircRNF144B/miR-342-3p/FBXL11 axis reduced autophagy and promoted the progression of ovarian cancer by increasing the ubiquitination of Beclin-1. Cell Death Dis. 2022, 13, 857. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Wang, P.; Xue, Y.; Qu, C.; Zheng, J.; Liu, X.; Ma, J.; Liu, Y.P. VT1 affects growth of glioma microvascular endothelial cells by negatively regulating miR-186. Tumor Biol. 2017, 39, 1010428317694326. [Google Scholar] [CrossRef]
- Yan, R.-L.; Luan, C.-L.; Liao, C.-C.; Liu, L.-H.; Chen, F.-Y.; Chen, H.-Y.; Chen, R.-H. Long noncoding RNA BCRP3 stimulates VPS34 and autophagy activities to promote protein homeostasis and cell survival. J. Biomed. Sci. 2022, 29, 30. [Google Scholar]
- Fujioka, Y.; Noda, N.N.; Nakatogawa, H.; Ohsumi, Y.; Inagaki, F. Dimeric coiled-coil structure of Saccharomyces cerevisiae Atg16 and its functional significance in autophagy. J. Biol. Chem. 2010, 285, 1508–1515. [Google Scholar] [CrossRef]
- Cao, Y.; Pan, L.; Zhang, X.; Guo, W.; Huang, D. LncRNA SNHG3 promotes autophagy-induced neuronal cell apoptosis by acting as a ceRNA for miR-485 to up-regulate ATG7 expression. Metab. Brain Dis. 2020, 35, 1361–1369. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Lu, W.; Ge, D.; Meng, N.; Li, Y.; Su, L.; Zhang, S.; Zhang, Y.; Zhao, B.; Miao, J. Anew microRNA signal pathway regulated by long noncoding RNA TGFB2-OT1 in autophagy and inflammation of vascular endothelial cells. Autophagy 2015, 11, 2172–2183. [Google Scholar] [CrossRef] [PubMed]
- Tang, B.; Li, N.; Gu, J.; Zhuang, Y.; Li, Q.; Wang, H.-G.; Fang, Y.; Yu, B.; Zhang, J.-Y.; Xie, Q.-H. Compromised autophagy by MIR30B benefits the intracellular survival of Helicobacter pylori. Autophagy 2012, 8, 1045–1057. [Google Scholar] [CrossRef]
- Gan, J.; Cai, Q.; Qu, Y.; Zhao, F.; Wan, C.; Luo, R.; Mu, D. miR-96 attenuates status epilepticus-induced brain injury by directly targeting Atg7 and Atg16L1. Sci. Rep. 2017, 7, 10270. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, M.; Liu, H. LncRNA17A regulates autophagy and apoptosis of SH-SY5Y cell line as an in vitro model for Alzheimer’s disease. Biosci. Biotechnol. Biochem. 2019, 83, 609–621. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, H.; Shen, Q.; Feng, L.; Jin, H. Long non-coding RNAs involved in autophagy regulation. Cell Death Dis. 2017, 8, e3073. [Google Scholar] [CrossRef]
- Bento, C.F.; Puri, C.; Moreau, K.; Rubinsztein, D.C. The role of membrane-trafficking small GTPases in the regulation of autophagy. J. Cell Sci. 2013, 126, 1059–1069. [Google Scholar] [CrossRef]
- Ao, X.; Zou, L.; Wu, Y. Regulation of autophagy by the Rab GTPase network. Cell Death Differ. 2014, 21, 348–358. [Google Scholar] [CrossRef]
- Tao, J.; Liu, W.; Shang, G.; Zheng, Y.; Huang, J.; Lin, R.; Chen, L. MiR-207/352 regulate lysosomal-associated membrane proteins and enzymes following ischemic stroke. Neuroscience 2015, 305, 1–14. [Google Scholar] [CrossRef]
- Su, C.; Yang, X.; Lou, J. Geniposide reduces α-synuclein by blocking microRNA-21/lysosome-associated membrane protein 2A interaction in Parkinson disease models. Brain Res. 2016, 1644, 98–106. [Google Scholar] [CrossRef] [PubMed]
- Kato, M.; Abdollahi, M.; Tunduguru, R.; Tsark, W.; Chen, Z.; Wu, X.; Wang, J.; Chen, Z.B.; Lin, F.M.; Lanting, L.; et al. miR-379 deletion ameliorates features of diabetic kidney disease by enhancing adaptive mitophagy via FIS1. Commun. Biol. 2021, 4, 30. [Google Scholar]
- Pan, J.; Fang, S.; Tian, H.; Zhou, C.; Zhao, X.; Tian, H.; He, J.; Shen, W.; Meng, X.; Jin, X.; et al. lncRNA JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis of lung cancer by activating Wnt/β-catenin signaling. Mol. Cancer 2020, 19, 9. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Yu, X.; Li, Z. Circular RNA hsa_circ_0001658 regulates apoptosis and autophagy in gastric cancer through microRNA-182/Ras-related protein Rab-10 signaling axis. Bioengineered 2022, 13, 2387–2397. [Google Scholar] [CrossRef]

| Autophagy Stage | Name | Composition |
|---|---|---|
| Initiation | Energy depletion | AMPK |
| mTOR complex | mTORC1 | |
| mTORC2 | ||
| Class III PI3K complex | Vps34 | |
| Vps15 | ||
| ULK complexes | FIP200 | |
| ULK1 | ||
| ULK2 | ||
| ATG13 | ||
| ATG101 | ||
| Class II PI3K complex | Beclin-1 | |
| Vesicle nucleation | Class III PI3K complex | Vps34 |
| Vps15 | ||
| Class II PI3K complex | Beclin-1 | |
| Bcl-2 family | Bcl-2 | |
| - | ATG14 | |
| Vesicle elongation | Atg12-Atg5-Atg16 | ATG7 |
| ATG5 | ||
| ATG12 | ||
| ATG16 | ||
| LC3 | LC3-I | |
| LC3-II | ||
| ATG4B | ||
| Autophagosome formation (Vesicle fusion) | STX17 | |
| - | ATG10 | |
| LC3 | LC3-I | |
| LC3-II | ||
| - | Rab7 | |
| - | Rab5 | |
| - | Rab9 | |
| Maturation and degradation | - | p62 |
| LC3 | LC3II | |
| - | Rab7 | |
| - | Rab8B | |
| - | Rab24 | |
| LAMP | LAMP1 | |
| LAMP2 | ||
| LAMP3 |
| Autophagy Stage | miRNA | Disease/Cancer | Target | Function | Signaling Pathway/Axis | Ref. |
|---|---|---|---|---|---|---|
| initiation | miR-20a/20b | breast cancer | RB1CC1/FIP200 | Overexpression of miR-20a and miR-20b attenuates autophagy | - | [44] |
| initiation | miR-106a | lung adenocarcinoma | ULK1 | miRNA-106a targeted ULK1 results in death of different NSCLC cells | miR-106a-ULK1 | [45] |
| initiation | miR-489 | breast cancer | ULK1, LAPTM4B | miR-489 affects autophagy by targeting ULK1 | - | [46] |
| initiation | miR-25 | breast cancer | ULK1 | miR-25 functions as a regulator of autophagy by targeting ULK1 | - | [47] |
| initiation | miR-142-5p | gastric cancer | ULK1 | miR-142-5p can regulate ULK1 expression | - | [48] |
| initiation | miR-26 | non-small cell lung cancer | TGF-β1 | miR-26 reduces autophagy via targeting TGF-β1 | TGF-β1-JNK | |
| initiation | miR-17-5p | cells | ULK1 | miR-17-5p inhibits ULK1 expression in cellular autophagy | - | [49] |
| initiation | miR-26b | breast cancer | DRAM1 | miR-26b can suppress autophagy in breast cancer cells | TGF-β1-JNK | [50] |
| initiation | miR-885-3p | squamous cell carcinoma | ULK2 | miR-885-3p contributes to the regulation of squamous cell carcinoma cell autophagy | - | [51] |
| initiation | miR-133a-3p | gastric cancer | ATG13, GABARAPL1 | miR-133a-3p expression inhibits autophagy to hinder gastric cancer metastasis via blocking GABARAPL1 and ATG13 expression | - | [52] |
| initiation | miR-20a | C2C12 myoblasts | ULK1 | miR-20a inhibits the expression of ULK1, which leads to a reduction in autophagy induced by leucine deprivation | PI3K-AKT- MTOR | [53] |
| initiation | miR-100 | renal cell carcinoma | mTOR | miR-100 can inactivate mTOR and thus increase autophagy in renal cancer cells. | mTOR | [54] |
| vesicle nucleation | miR-30a | hepatic fibrosis | Beclin-1 | Overexpression of miR-30a inhibits Beclin1-mediated autophagy to prevent the occurrence of liver fibrosis | - | [55] |
| vesicle nucleation | miR-93 | glioblastoma | BECN1, Beclin-1, ATG5, ATG4B, SQSTM1/p62 | miR-93 inhibits autophagy functions by targeting multiple autophagy regulators | PI3K-AKT | [56] |
| vesicle nucleation | miR-124-3p | breast cancer | Beclin-1, LC3-I | miR-124-3p promotes the progression of breast cancer cells by enhancing the expression of Beclin-1 | - | [57] |
| vesicle nucleation | miR-30a | cardiomyocyte | Beclin-1 | Downregulation of miR-30a expression upregulates beclin-1 expression and enhances autophagy in cardiomyocytes | - | [58] |
| vesicle nucleation | miR-30a | medulloblastoma | Beclin-1, LC3B | miR-30a inhibits autophagy by downregulating the expression of Beclin-1and LC3B | - | [59] |
| vesicle nucleation | miR-30e | cardiomyopathy | Beclin-1, LC3-I, LC3-II | miR-30e can downregulate the expression of Beclin-1 | - | [60] |
| vesicle nucleation | miR-30d | colon cancer | Beclin-1 | Overexpression of miR-30d inhibits the proliferation of colon cancer cells | - | [61] |
| vesicle nucleation | miR-30d | renal cell carcinoma | MTDH | miR-30d targets MTDH and inhibits renal cancer cells | AKT/FOXO | [62] |
| vesicle nucleation | miR-124-3p | breast cancer | Beclin-1 | Decreased miR-124-3p expression prompts breast cancer cell progression | - | [57] |
| vesicle nucleation | miR-216b | non-small cell lung cancer | Beclin-1 | miR-216b can inhibit cisplatin sensitivity of NSCLC through regulating apoptosis and autophagy via miR-216b/Beclin-1 pathway | miR-216b/Beclin-1 axis | [63] |
| vesicle nucleation | miR-17-5p | non-small cell lung cancer | Beclin-1 | miR-17-5p facilitates the ability of cell proliferation, inhibits autophagy and apoptosis by modulating Beclin-1 | - | [64] |
| vesicle nucleation | miR-143 | colorectal cancer | Beclin-1 | miR-143 targets various cellular that are involved in the autophagy pathways pathogenesis of colorectal cancer | PI3K/AKT/Wnt | [65] |
| elongation | miR-23a | fibroblasts | AMBRA1 | miR-23a inhibits the autophagy of fibroblasts during UV-induced photoaging | - | [66] |
| elongation | miR-23a-5p | acute myeloid leukemia | TLR2 | Downregulation of miR-23a-5p in leukemic cells can lead to the upregulation of protective autophagy | - | [67] |
| elongation | miR-7 | lung cancer | AMBRA1 | AMBRA1 is targeted by miR-7, leading to the promotion of lung cancer cell proliferation | AKT | [68] |
| elongation | miR-128a | osteoarthritis | ATG12 | ATG12, induced by miR-128a, loss represses chondrocyte autophagy to aggravate OA progression | - | [69] |
| elongation | miR-23b | traumatic brain injury | ATG12 | miR-23b directly targets to the 3′UTR region of ATG12 to suppress the activation of neuronal autophagy | - | [70] |
| elongation | miR-214 | colorectal cancer | ATG12, LC3 | miR-214 inhibits autophagy and induction of apoptosis by targeting ATG12 | - | [71] |
| autophagosome formation | miR-106b | colorectal cancer | ATG16L1 | miR-106b inhibits starvation-induced autophagy by inhibiting the expression of ATG16L1 | - | [72] |
| autophagosome maturation | miR-138-5p | pancreatic cancer | SIRT1 | miR-138-5p specifically targets SIRT1, thereby inhibiting autophagy. | - | [73] |
| autophagosome maturation | miR-487b-5p | lung cancer | LAMP2 | miR-487b-5p directly targets LAMP2 to affect the latter stage of autophagy flux in lung cancer | - | [74] |
| autophagosome maturation | miR-205 | prostate cancer | RAB27A, LAMP3 | miR-205 inhibits autophagy in prostate cancer cells | - | [75] |
| autophagosome maturation | miR-378 | - | PDK1 | miR-378 promotes autophagy initiation through the mammalian target of rapamycin mTOR/ULK1 pathway and sustains autophagy by targeting phosphoinositide-dependent protein kinase 1 (PDK1) | mTOR/ULK1 | [76] |
| Autophagy Stage | lncRNA | Cancer/Disease | Target | Function | Signaling Pathways/Axis | Refs. |
|---|---|---|---|---|---|---|
| initiation | lncRNA NBR2 | colorectal cancer | AMPK | AMPK promotes the activation of autophagy by binding to lncRNA NBR2 | mTOR | [77] |
| initiation | lncRNA AD5-A lncRNA | hepatocellular carcinoma (HCC) | AKT, mTOR | Overexpression of AD5-A lncRNA can block the function of miRNAs to inhibit AKT/mTOR activity and promote autophagy activation | AKT/mTOR | [78] |
| initiation | lncRNA SNHG6 | colorectal cancer | ULK1 | lncRNA SNHG6 is able to promote colorectal cancer chemoresistance and enhance autophagy through regulation of ULK1 | - | [79] |
| initiation | lncRNA MALAT1 | brain microvascular endothelial cell injury | ULK2 | lncRNA MALAT1 can promote the expression of ULK2, suggesting that MALAT1 protects brain microvascular endothelial cells from ischemia-reperfusion injury by promoting autophagy | - | [80] |
| initiation | lncRNA H19 | cardiomyocytes | DIRAS3 | H19 could inhibit cardiomyocyte autophagy by epigenetically silencing DIRAS3 | mTOR | [81] |
| initiation | lncRNA SNHG1 | parkinson’s disease | LC3-II | Downregulated lncRNA SNHG1 inhibits the mTOR pathway and initiates autophagy | mTOR | [82] |
| initiation | lncRNA AK156230 | mouse embryonic fibroblasts | mTOR | AK156230 can inhibit replicative senescence (RS); meanwhile, the mTOR signaling pathway leads to autophagy deficiency, which may accelerate aging | mTOR | [83] |
| initiation | lncRNA PTENP1 | hepatocellular carcinoma cells | AKT | Overexpression of lncRNA PTENP1 indirectly inhibits the PI3K/AKT pathway and then induces pro-death autophagy, leading to the death of hepatocellular carcinoma cells | PI3K/AKT | [84,85] |
| vesicle nucleation | lncRNA SNHG12 | SH-SY5Y cells | LC3-II, Beclin-1 | The expression of lncRNA SNHG12 promotes LC3-II and Beclin-1 expression levels, thus inducing autophagy activation | - | [86] |
| vesicle nucleation | lncRNA AC023115.3 | human glioblastoma cells | Beclin-1 | AC023115.3 is induced by cisplatin, and elevated AC023115.3 promotes cisplatin-induced apoptosis by inhibiting autophagy | miR-26a-GSK3β-Mcl1 axis | [87] |
| vesicle nucleation | lncRNA PVT1 | - | ATG14 | PVT1 interacts with ATG14 in the cytoplasm, and PVT1 can upregulate the expression of both Pygo2 and ATG14, thus regulating autophagic activity | - | [88] |
| vesicle nucleation | lncRNA EIF3J-DT | gastric cancer | ATG14 | EIF3J-DT activates autophagy and induces drug resistance in gastric cancer cells by targeting ATG14, thus contributing to activation of autophagy | - | [89] |
| vesicle nucleation | lncRNA NEAT1 | Parkinson’s disease | LC3-II | lncRNA NEAT1 can induce abnormal autophagy by stabilizing PINK1, which is an LC3-II upstream regulatory factor and plays a role in the pathogenesis of PD | - | [90] |
| elongation | lncRNA CCAT1 | hepatocellular carcinoma cell | ATG7 | lncRNA CCAT1 facilitates hepatocellular carcinoma cell autophagy and cell proliferation, and then regulates ATG7 expression | - | [91] |
| elongation | lncRNA GAS5 | osteoarthritis | Beclin-1, ATG3, ATG5, ATG7, ATG12 | lncRNA GAS5, upregulating in osteoarthritis (OA), contributes to the pathogenesis of OA and thereby represses autophagy | - | [92] |
| elongation | lncRNA HNF1A-AS1 | hepatocellular carcinoma | ATG5, Beclin-1, ATG12 | lncRNA HNF1A-AS1, binding to its target Beclin-1, ATG5, and ATG12, can provoke autophagy in hepatocellular carcinoma | - | [93] |
| elongation | lncRNA HOTAIR | hepatocellular carcinoma | ATG3, ATG7 | lncRNA HOTAIR is upregulated to promote hepatocellular carcinoma cell proliferation, probably by enhancing ATG3 and ATG7 expression | - | [94] |
| elongation | lncRNA HULC | epithelial ovarian carcinoma | ATG7, LC3-II, LAMP1 | lncRNA HULC overexpression reduces ATG7, LC3-II, and LAMP1 expression, and then reduces apoptosis and inhibits autophagy | - | [95] |
| Autophagy Stage | circRNA | Cancer/Disease | Target | Function | Signaling Pathway/Axis | Ref. |
|---|---|---|---|---|---|---|
| initiation | circ_0009910 | chronic myeloid leukemia | ULK1 | circ_0009910 can regulate the expression of ULK1, thereby activating the level of autophagy | - | [96] |
| initiation | circ_CDYL | breast cancer | ATG7, ULK1 | circ_CDYL regulates the expression of autophagy-related genes ATG7 and ULK1, thus promoting autophagy | - | [97] |
| initiation | circ_PAN3 | acute myeloid leukemia | mTOR | circ-PAN3 regulates autophagy via the AMPK/mTOR signaling pathway in acute myeloid leukemia | AMPK/mTOR | [98] |
| initiation | circRNA ACR | RSC96 cells | mTOR | circRNA ACR in RSC96 cells promotes the activation of the PI3K/AKT/mTOR pathway to alleviate autophagy | PI3K/AKT/mTOR | [99] |
| initiation | circRNA ciRS-7 | esophageal squamous cell carcinoma | mTOR | circRNA ciRS-7 affects the AKT–mTOR signaling pathway, thus inhibiting autophagy of ESCC cells | AKT-mTOR | [100] |
| vesicle nucleation | circ_MUC16 | epithelial ovarian cancer | Beclin1, RUNX1, ATG13 | circ_MUC16 promotes autophagy in epithelial ovarian cancer by regulating Beclin1, RUNX1, and ATG13 | - | [101] |
| vesicle nucleation | circPOFUT1 | gastric cancer | ATG12 | circPOFUT1 promotes ATG12 expression to regulate autophagy-associated chemoresistance in gastric cancer | - | [102] |
| elongation | circ_0092276 | breast cancer | ATG7, LC3-II, LC3-I, Beclin-1 | circ_0092276 affects autophagy and proliferation, and represses apoptosis of breast cancer cells | - | [103] |
| elongation | circ_0035483 | renal clear cell carcinoma cells | LC3-II, LC3-I | when circ_0035483 expression is downregulated, the LC3II/LC3I ratio is significantly reduced, thus inhibiting autophagy | - | [104] |
| Autophagosome formation | circ_ PABPN1 | intestinal epithelial cells | ATG16L1 | circ_ PABPN1 inhibits ATG16L1 translation and thus regulates autophagy in intestinal epithelial cells | - | [105] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, X.; Xiong, S.; Zhao, X.; Jin, J.; Yang, X.; Du, Y.; Zhao, L.; He, Z.; Gong, C.; Guo, L.; et al. Orchestrating Cellular Balance: ncRNAs and RNA Interactions at the Dominant of Autophagy Regulation in Cancer. Int. J. Mol. Sci. 2024, 25, 1561. https://doi.org/10.3390/ijms25031561
Yang X, Xiong S, Zhao X, Jin J, Yang X, Du Y, Zhao L, He Z, Gong C, Guo L, et al. Orchestrating Cellular Balance: ncRNAs and RNA Interactions at the Dominant of Autophagy Regulation in Cancer. International Journal of Molecular Sciences. 2024; 25(3):1561. https://doi.org/10.3390/ijms25031561
Chicago/Turabian StyleYang, Xueni, Shizheng Xiong, Xinmiao Zhao, Jiaming Jin, Xinbing Yang, Yajing Du, Linjie Zhao, Zhiheng He, Chengjun Gong, Li Guo, and et al. 2024. "Orchestrating Cellular Balance: ncRNAs and RNA Interactions at the Dominant of Autophagy Regulation in Cancer" International Journal of Molecular Sciences 25, no. 3: 1561. https://doi.org/10.3390/ijms25031561






