Oomycete Soil Diversity Associated with Betula and Alnus in Forests and Urban Settings in the Nordic–Baltic Region
Abstract
:1. Introduction
2. Materials and Methods
2.1. Site Selection and Soil Collection
2.2. DNA Extraction, PCR Amplification and Sequencing
2.3. Data Analysis
2.4. Statistical Analysis
3. Results
3.1. The Taxonomic Diversity of Soil Oomycetes
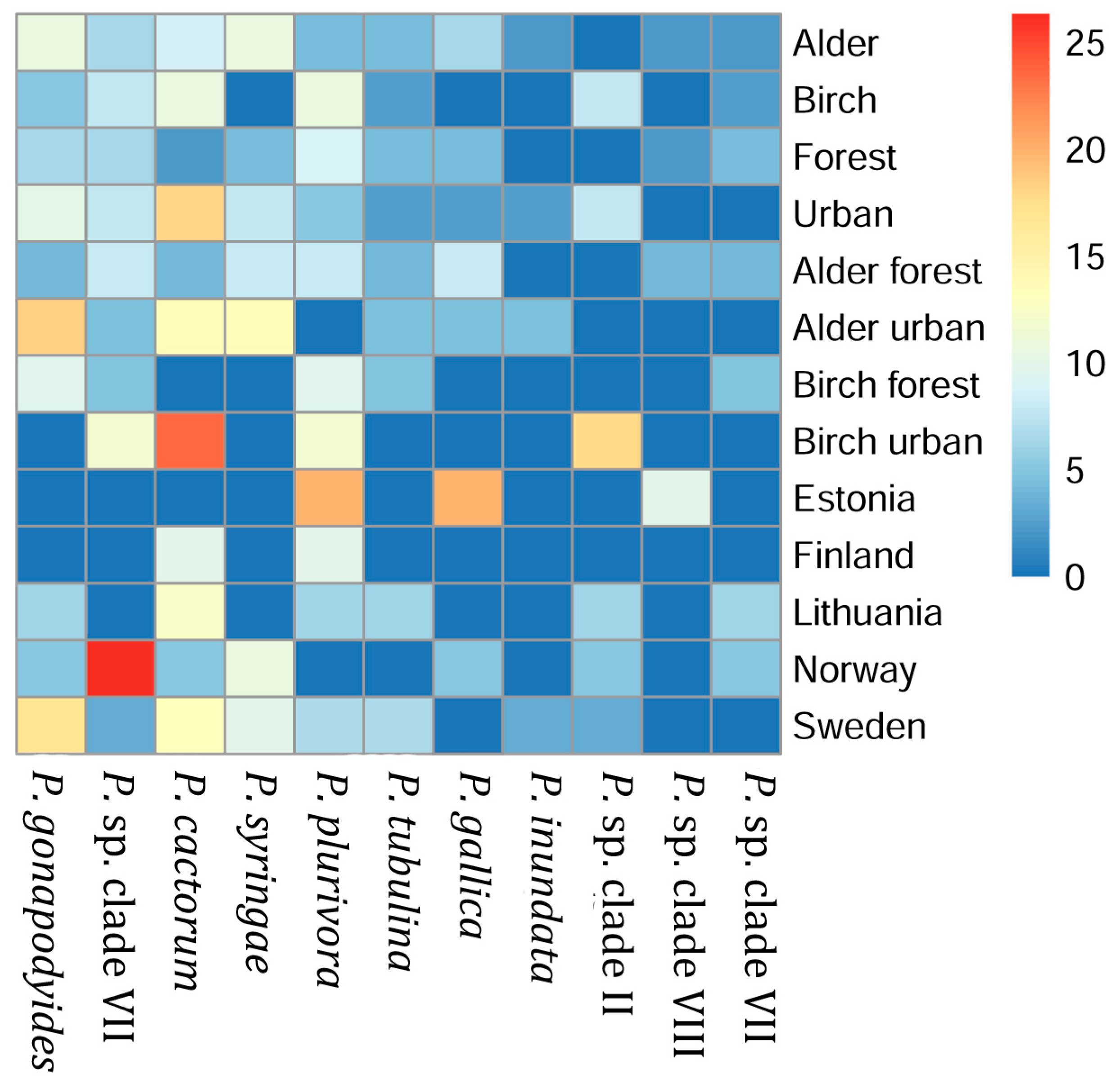
3.2. Phytophthora Species Prevalence in Different Sample Groups
3.3. Oomycete Richness and Composition Differences among Countries
3.4. Oomycete Richness and Composition Differences among Site Types
3.5. Effect of Climatic and Spatial Variables on Oomycete Community Composition and Species Richness
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Benavent-Celma, C.; López-García, N.; Ruba, T.; Ściślak, M.E.; Street-Jones, D.; van West, P.; Woodward, S.; Witzell, J. Current practices and emerging possibilities for reducing the spread of oomycete pathogens in terrestrial and aquatic production systems in the European Union. Fungal Biol. Rev. 2022, 40, 19–36. [Google Scholar] [CrossRef]
- Větrovský, T.; Kohout, P.; Kopecký, M.; Machac, A.; Man, M.; Bahnmann, B.D.; Brabcová, V.; Choi, J.; Meszárošová, L.; Human, Z.R.; et al. A meta-analysis of global fungal distribution reveals climate-driven patterns. Nat. Commun. 2019, 10, 5142. [Google Scholar] [CrossRef]
- Delavaux, C.S.; Schemanski, J.L.; House, G.L.; Tipton, A.G.; Sikes, B.; Bever, J.D. Root pathogen diversity and composition varies with climate in undisturbed grasslands, but less so in anthropogenically disturbed grasslands. ISME J. 2021, 15, 304–317. [Google Scholar] [CrossRef]
- Scales, N.C.; Chase, A.B.; Finks, S.S.; Malik, A.A.; Weihe, C.; Allison, S.D.; Martiny, A.C.; Martiny, J.B.H. Differential Response of Bacterial Microdiversity to Simulated Global Change. Appl. Environ. Microbiol. 2022, 88, e02429-21. [Google Scholar] [CrossRef] [PubMed]
- Derevnina, L.; Petre, B.; Kellner, R.; Dagdas, Y.F.; Sarowar, M.N.; Giannakopoulou, A.; De la Concepcion, J.C.; Chaparro-Garcia, A.; Pennington, H.G.; Van West, P.; et al. Emerging oomycete threats to plants and animals. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2016, 371, 20150459. [Google Scholar] [CrossRef] [PubMed]
- Fones, H.N.; Bebber, D.P.; Chaloner, T.M.; Kay, W.T.; Steinberg, G.; Gurr, S.J. Threats to global food security from emerging fungal and oomycete crop pathogens. Nat. Food 2020, 1, 332–342. [Google Scholar] [CrossRef]
- Scott, P.; Bader, M.K.F.; Burgess, T.; Hardy, G.; Williams, N. Global biogeography and invasion risk of the plant pathogen genus Phytophthora. Environ. Sci. Policy 2019, 101, 175–182. [Google Scholar] [CrossRef]
- Cobb, R.C.; Haas, S.E.; Kruskamp, N.; Dillon, W.W.; Swiecki, T.J.; Rizzo, D.M.; Frankel, S.J.; Meentemeyer, R.K. The magnitude of regional-scale tree mortality caused by the invasive pathogen Phytophthora ramorum. Earth’s Future 2020, 8, e2020EF001500. [Google Scholar] [CrossRef]
- Hassett, B.T.; Thines, M.; Buaya, A.; Ploch, S.; Gradinger, R. A glimpse into the biogeography, seasonality, and ecological functions of arctic marine Oomycota. IMA Fungus 2019, 10, 1–10. [Google Scholar] [CrossRef]
- Corredor-Moreno, P.; Saunders, D.G. Expecting the unexpected: Factors influencing the emergence of fungal and oomycete plant pathogens. New Phytol. 2020, 225, 118–125. [Google Scholar] [CrossRef]
- Bjelke, U.; Boberg, J.; Oliva, J.; Tattersdill, K.; McKie, B.-G. Dieback of riparian alder caused by the Phytophthora alni complex: Projected consequences for stream ecosystems. Freshw. Biol. 2016, 61, 565–579. [Google Scholar] [CrossRef]
- Fahlvik, N.; Hannerz, M.; Högbom, L.; Jacobson, S.; Liziniewicz, M.; Palm, J.; Rytter, L.; Sonesson, J.; Wallgren, M.; Weslien, J.-O. Björkens Möjligheter i ett Framtida Klimatanpassat Brukande av skog. Sammanställning av Nuläget och Förslag på Insatser för Framtiden; Skogforsk Rapport; Gävle Offset: Uppsala, Sweden, 2021; Volume 80. [Google Scholar]
- Sirkas, F.; Valgepea, M. Yearbook Forest 2020; Estonian Environment Agency: Tallinn, Estonia, 2022; p. 322. Available online: https://keskkonnaportaal.ee/sites/default/files/Teemad/Mets/Mets2020.pdf (accessed on 10 February 2023).
- Hynynen, J.; Niemistö, P.; Viherä-Aarnio, A.; Brunner, A.; Hein, S.; and Velling, P. Silviculture of birch (Betula pendula Roth and Betula pubescens Ehrh.) in northern Europe. Forestry 2010, 83, 103–119. [Google Scholar] [CrossRef]
- Felton, A.; Felton, A.M.; Wam, H.K.; Witzell, J.; Wallgren, M.; Löf, M.; Sonesson, J.; Lindbladh, M.; Björkman, C.; Blennow, K.; et al. Forest biodiversity and ecosystem services from spruce-birch mixtures: The potential importance of tree spatial arrangement. Environ. Chall. 2022, 6, 100407. [Google Scholar] [CrossRef]
- Hanso, M.; Drenkhan, R. Lophodermium needle cast, insect defoliation and growth responses of young Scots pines in Estonia. For. Pathol. 2012, 42, 124–135. [Google Scholar] [CrossRef]
- Müller, M.M.; Hantula, J.; Wingfield, M.; Drenkhan, R. Diplodia sapinea found on Scots pine in Finland. For. Pathol. 2019, 49, e12483. [Google Scholar] [CrossRef]
- Cleary, M.; Laas, M.; Oskay, F.; Drenkhan, R. First report of Lecanosticta acicola on non-native Pinus mugo in southern Sweden. For. Pathol. 2019, 49, e12507. [Google Scholar] [CrossRef]
- Adamson, K.; Laas, M.; Blumenstein, K.; Busskamp, J.; Langer, G.J.; Klavina, D.; Kaur, A.; Maaten, T.; Mullett, M.S.; Müller, M.M.; et al. Highly Clonal Structure and Abundance of One Haplotype Characterise the Diplodia sapinea Populations in Europe and Western Asia. J. Fungi 2021, 7, 634. [Google Scholar] [CrossRef]
- Laas, M.; Adamson, K.; Barnes, I.; Janoušek, J.; Mullett, M.S.; Adamčíková, K.; Akiba, M.; Beenken, L.; Braganca, H.; Bulgakov, T.S.; et al. Diversity, migration routes, and worldwide population genetic structure of Lecanosticta acicola, the causal agent of brown spot needle blight. Mol. Plant Pathol. 2022, 23, 1620–1639. [Google Scholar] [CrossRef]
- Mozgeris, G.; Brukas, V.; Pivoriūnas, N.; Činga, G.; Makrickienė, E.; Byčenkienė, S.; Marozas, V.; Mikalajūnas, M.; Dudoitis, V.; Ulevičius, V.; et al. Spatial pattern of climate change effects on lithuanian forestry. Forests 2019, 10, 809. [Google Scholar] [CrossRef]
- Pliura, A.; Jankauskiene, J.; Lygis, V.; Suchockas, V.; Bajerkevičiene, G.; Verbylaite, R. Response of juvenile progeny of seven forest tree species and their populations to simulated climate change-related stressors, heat, elevated humidity and drought. iForest 2018, 11, 374–388. [Google Scholar] [CrossRef]
- Pliūra, A.; Jankauskienė, J.; Bajerkevičienė, G.; Lygis, V.; Suchockas, V.; Labokas, J.; Verbylaitė, R. Response of juveniles of seven forest tree species and their populations to different combinations of simulated climate change-related stressors: Spring-frost, heat, drought, increased UV radiation and ozone concentration under elevated CO2 level. J. Plant Res. 2019, 132, 789–811. [Google Scholar] [CrossRef] [PubMed]
- Pliūra, A.; Bajerkevičienė, G.; Labokas, J.; Lygis, V.; Jankauskienė, J.; Suchockas, V.; Verbylaitė, R. The effects of different combinations of simulated climate change-related stressors on juveniles of seven forest tree species grown as mono-species and mixed cultures. Balt. For. 2020, 26, 326. [Google Scholar] [CrossRef]
- Ozolinčius, R.; Lekevičius, E.; Stakėnas, V.; Galvonaitė, A.; Samas, A.; Valiukas, D. Lithuanian forests and climate change: Possible effects on tree species composition. Eur. J. Forest Res. 2014, 133, 51–60. [Google Scholar] [CrossRef]
- Stradaioli, M. Studies on the Pythiaceous Fungi Assemblage of Italian Beech Forest Soils along Altitudinal and Latitudinal Gradients under the Climate Change Scenario. Ph.D. Thesis, University of Tuscia, Viterbo, Italy, 26 June 2013. [Google Scholar]
- Vannini, A.; Bruni, N.; Tomassini, A.; Franceschini, S.; Vettraino, A.M. Pyrosequencing of environmental soil samples reveals biodiversity of the Phytophthora resident community in chestnut forests. FEMS Microbiol. 2013, 85, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Català, S.; Perez-Sierra, A.; Abad-Campos, P. The use of genus-specific amplicon pyrosequencing to assess Phytophthora species diversity using eDNA from soil and water in northern Spain. PLoS ONE 2015, 10, e0119311. [Google Scholar] [CrossRef] [PubMed]
- Ruiz Gómez, F.J.; Navarro-Cerrillo, R.M.; Pérez-de-Luque, A.; Oβwald, W.; Vannini, A.; Morales-Rodríguez, C. Assessment of functional and structural changes of soil fungal and oomycete communities in holm oak declined dehesas through metabarcoding analysis. Sci. Rep. 2019, 9, 5315. [Google Scholar] [CrossRef]
- Mora-Sala, B.; Gramaje, D.; Abad-Campos, P.; Berbegal, M. Diversity of Phytophthora species associated with Quercus ilex L. in three Spanish regions evaluated by NGS. Forests 2019, 10, 979. [Google Scholar] [CrossRef]
- Landa, B.B.; Arias-Giraldo, L.F.; Henricot, B.; Montes-Borrego, M.; Shuttleworth, L.A.; Pérez-Sierra, A. Diversity of Phytophthora species detected in disturbed and undisturbed British soils using high-throughput sequencing targeting ITS rRNA and COI mtDNA regions. Forests 2021, 12, 229. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Põlme, S.; Kõljalg, U.; Yorou, N.S.; Wijesundera, R.; Villarreal Ruiz, L.; Vasco-Palacios, A.M.; Thu, P.Q.; Suija, A.; et al. Global diversity and geography of soil fungi. Science 2014, 346, 1078. [Google Scholar] [CrossRef]
- Riit, T.; Tedersoo, L.; Drenkhan, R.; Runno-Paurson, E.; Kokko, H.; Anslan, S. Oomycete-specific ITS primers for identification and metabarcoding. MycoKeys 2016, 14, 17–30, Erratum in MycoKeys 2018, 41, 119. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2023; Available online: https://www.R-project.org/ (accessed on 10 February 2023).
- Cooke, D.E.L.; Drenth, A.; Duncan, J.M.; Wagels, G.; Brasier, C.M. A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genet. Biol. 2000, 30, 17–32. [Google Scholar] [CrossRef]
- Bahram, M.; Netherway, T.; Hildebrand, F.; Pritsch, K.; Drenkhan, R.; Loit, K.; Anslan, S.; Bork, P.; Tedersoo, L. Plant nutrient-acquisition strategies drive topsoil microbiome structure and function. New Phytol. 2020, 227, 1189–1199. [Google Scholar] [CrossRef]
- Tedersoo, L.; Anslan, S.; Bahram, M.; Drenkhan, R.; Pritsch, K.; Buegger, F.; Padari, A.; Hagh-Doust, N.; Mikryukov, V.; Gohar, D.; et al. Regional-scale in-depth analysis of soil fungal diversity reveals strong pH and plant species effects in Northern Europe. Front. Microbiol. 2020, 11, 1953. [Google Scholar] [CrossRef] [PubMed]
- Qu, Z.L.; Santalahti, M.; Köster, K.; Berninger, F.; Pumpanen, J.; Heinonsalo, J.; Sun, H. Soil fungal community structure in boreal pine forests: From Southern to Subarctic areas of Finland. Front. Microbiol. 2021, 12, 653896. [Google Scholar] [CrossRef]
- Jarvis, S.; Woodward, S.; Alexander, I.J.; Taylor, A.F.S. Regional scale gradients of climate and nitrogen deposition drive variation in ectomycorrhizal fungal communities associated with native Scots pine. Glob. Chang. Biol. 2013, 19, 1688–1696. [Google Scholar] [CrossRef] [PubMed]
- Bahram, M.; Hildebrand, F.; Forslund, S.K.; Anderson, J.L.; Soudzilovskaia, N.A.; Bodegom, P.M.; Bengtsson-Palme, J.; Anslan, S.; Coelho, L.P.; Harend, H.; et al. Structure and function of the global topsoil microbiome. Nature 2018, 560, 233–237. [Google Scholar] [CrossRef]
- Bahram, M.; Espenberg, M.; Pärn, J.; Lehtovirta-Morley, L.; Anslan, S.; Kasak, K.; Kõljalg, U.; Liira, J.; Maddison, M.; Moora, M.; et al. Structure and function of the soil microbiome underlying N2O emissions from global wetlands. Nat. Commun. 2022, 13, 1430. [Google Scholar] [CrossRef] [PubMed]
- Barreiro, A.; Fox, A.; Jongen, M.; Melo, J.; Musyoki, M.; Vieira, A.; Zimmermann, J.; Carlsson, G.; Cruz, C.; Lüscher, A.; et al. Soil bacteria respond to regional edapho-climatic conditions while soil fungi respond to management intensity in grasslands along a European transect. Appl. Soil Ecol. 2022, 170, 104264. [Google Scholar] [CrossRef]
- Zhou, J.; Deng, Y.E.; Shen, L.; Wen, C.; Yan, Q.; Ning, D.; Qin, Y.; Xue, K.; Wu, L.; He, Z.; et al. Temperature mediates continental-scale diversity of microbes in forest soils. Nat. Commun. 2016, 7, 12083. [Google Scholar] [CrossRef]
- Willoughby, L.G.; Roberts, R.J. Towards strategic use of fungicides against Saprolegnia parasitica in salmonid fish hatcheries. J. Fish Dis. 1992, 15, 1–3. [Google Scholar] [CrossRef]
- Sformo, T.L.; De la Bastide, P.Y.; LeBlanc, J.; Givens, G.H.; Adams, B.; Seigle, J.C.; Kunaknana, S.C.; Moulton, L.L.; Hintz, W.E. Temperature response and salt tolerance of the opportunistic pathogen Saprolegnia parasitica: Implications for the broad whitefish subsistence fishery. Arct. Antarct. Alp. Res. 2021, 53, 271–285. [Google Scholar] [CrossRef]
- Herrero, M.L.; Brurberg, M.B.; Alayon, D.I.O.; Roleda, M. Occurrence and pathogenicity of Pythium (Oomycota) on Ulvaspecies (Chlorophyta) at different salinities. Algae 2020, 35, 79–89. [Google Scholar] [CrossRef]
- Costa, S.; Lopes, I. Saprolegniosis in amphibians: An integrated overview of a fluffy killer disease. J. Fungi 2022, 8, 537. [Google Scholar] [CrossRef] [PubMed]
- Rojas, J.A.; Jacobs, J.L.; Napieralski, S.; Karaj, B.; Bradley, C.A.; Chase, T.; Esker, P.D.; Giesler, L.J.; Jardine, D.J.; Malvick, D.K.; et al. Oomycete species associated with soybean seedlings in North America—Part II: Diversity and ecology in relation to environmental and edaphic factors. Phytopathology 2017, 107, 293–304. [Google Scholar] [CrossRef]
- Makiola, A.; Holdaway, R.J.; Wood, J.R.; Orwin, K.H.; Glare, T.R.; Dickie, I.A. Environmental and plant community drivers of plant pathogen composition and richness. New Phytol. 2021, 233, 496–504. [Google Scholar] [CrossRef]
- Blakney, A.J.; Bainard, L.D.; St-Arnaud, M.; Hijri, M. Soil chemistry and soil history significantly structure oomycete communities in Brassicaceae crop rotations. Appl. Environ. Microbiol. 2023, 89, e01314-22. [Google Scholar] [CrossRef]
- Castaño, C.; Lindahl, B.D.; Alday, J.G.; Hagenbo, A.; Martínez de Aragón, J.; Parladé, J.; Pera, J.; Bonet, J.A. Soil microclimate changes affect soil fungal communities in a Mediterranean pine forest. New Phytol. 2018, 220, 1211–1221. [Google Scholar] [CrossRef]
- Ogris, N.; Drenkhan, R.; Vahalík, P.; Cech, T.; Mullett, M.; Tubby, K. The potential global distribution of an emerging forest pathogen, Lecanosticta acicola, under a changing climate. Front. For. Glob. Chang. 2023, 6, 1221339. [Google Scholar] [CrossRef]
- Tubby, K.; Adamčikova, K.; Adamson, K.; Akiba, M.; Barnes, I.; Boroń, P.; Bragança, H.; Bulgakov, T.; Burgdorf, N.; Capretti, P.; et al. The increasing threat to European forests from the invasive foliar pine pathogen, Lecanosticta acicola. For. Ecol. Manag. 2023, 536, 120847. [Google Scholar] [CrossRef]
- Rytkönen, A.; Lilja, A.; Vercauteren, A.; Sirkiä, S.; Parikka, P.; Soukainen, M.; Hantula, J. Identity and potential pathogenicity of Phytophthora species found on symptomatic Rhododendron plants in a Finnish nursery. Can. J. Plant Pathol. 2012, 34, 255–267. [Google Scholar] [CrossRef]
- Lilja, A.; Rytkönen, A.; Hantula, J. Introduced pathogens found on ornamentals, strawberry and trees in Finland over the past 20 years. Agric. Food Sci. 2011, 20, 74–85. [Google Scholar] [CrossRef]
- Talgø, V.; Herrero, M.-L.; Toppe, B.; Brurberg, M.B.; Thurston, R.; Stensvand, A. Phytophthora plurivora, ny skadegjerar på tre i Noreg [Phytophthora plurivora, a new pathogen on trees in Norway]. Bioforsk FOKUS 2009, 5, 238–239. [Google Scholar]
- Telfer, K.H.; Brurberg, M.B.; Haukeland, S.; Stensvand, A.; Talgø, V. Phytophthora survives the digestive system of the invasive slug Arion vulgaris. Eur. J. Plant Pathol. 2015, 142, 125–132. [Google Scholar] [CrossRef]
- Dale, A.L.; Feau, N.; Berube, J.A.; Ponchart, J.; Bilodeau, G.J.; Hamelin, R.C. Urban environments harbor greater oomycete and Phytophthora diversity, creating a bridgehead for potential new pathogens to natural ecosystems. Environ. DNA 2022, 4, 1039–1051. [Google Scholar] [CrossRef]
- Trzewik, A.; Orlikowski, L.B.; Oszako, T.; Nowakowska, J.A.; Orlikowska, T. The characterization of Phytophthora isolates obtained from diseased Alnus glutinosa in Poland. Balt. For. 2015, 21, 44–50. [Google Scholar]
- Lilja, A.; Rikala, R.; Hietala, A.; Heinonen, R. Stem lesions on Betula pendula seedlings in Finnish forest nurseries and the pathogenicity of Phytophthora cactorum. Eur. J. Plant Pathol. 1996, 26, 89–96. [Google Scholar] [CrossRef]
- Brasier, C.M.; Kirk, S.A.; Delcan, J.; Cooke, D.E.L.; Jung, T.; Man in’t Veld, W.A. Phytophthora alni sp. nov.; its variants: Designation of emerging heteroploid hybrid pathogens spreading on Alnus trees. Mycol. Res. 2004, 108, 1172–1184. [Google Scholar] [CrossRef]
- Bose, T.; Wingfield, M.J.; Roux, J.; Vivas, M.; Burgess, T.I. Phytophthora species associated with roots of native and non-native trees in natural and managed forests. Microb. Ecol. 2021, 81, 122–133. [Google Scholar] [CrossRef]
- Gonzalez, J.M.; Portillo, M.C.; Belda-Ferre, P.; Mira, A. Amplification by PCR artificially reduces the proportion of the rare biosphere in microbial communities. PLoS ONE 2012, 7, e29973. [Google Scholar] [CrossRef]
- Burgess, T.I.; White, D.; Sapsford, S.J. Comparison of Primers for the Detection of Phytophthora (and Other Oomycetes) from Environmental Samples. J. Fungi 2022, 8, 980. [Google Scholar] [CrossRef]
- Suzuki, K.; Takahashi, K.; Harada, N. Evaluation of primer pairs for studying arbuscular mycorrhizal fungal community compositions using a MiSeq platform. Biol. Fertil. Soils 2020, 56, 853–858. [Google Scholar] [CrossRef]
- Raza, M.M.; Bebber, D.P. Climate change and plant pathogens. Curr. Opin. Microbiol. 2022, 70, 102233. [Google Scholar] [CrossRef] [PubMed]
- Schoebel, C.N.; Stewart, J.; Gruenwald, N.J.; Rigling, D.; Prospero, S. Population history and pathways of spread of the plant pathogen Phytophthora plurivora. PLoS ONE 2014, 9, e85368. [Google Scholar] [CrossRef] [PubMed]






| Alder-Dominated Sites | Lit | Est | Fin | Swe | Nor | Significant Pairs | |
|---|---|---|---|---|---|---|---|
| Pythiales | Rel. abund. | 1.001 | 0.981 | 0.830 | 0.711 | 0.822 | - |
| OTU rich. | 23.1 | 24.9 | 21.2 | 19.1 | 20.2 | - | |
| Shannon div. | 1.803 | 1.195 | 1.562 | 0.992 | 0.931 | LN * | |
| Saprolegniales | Rel. abund. | −0.032 | 0.008 | 0.142 | 0.221 | 0.155 | - |
| OTU rich. | 2.78 | 5.74 | 7.42 | 6.33 | 11.72 | LN **, EN * | |
| Shannon div. | 0.068 | 0.383 | 0.384 | 0.667 | 1.196 | LN **, EN * | |
| Peronosporales | Rel. abund. | 0.018 | 0.006 | 0.012 | 0.042 | 0.018 | - |
| OTU rich. | 2.04 | 2.80 | 1.33 | 1.89 | 1.99 | - | |
| Shannon div. | 0.051 | 0.298 | 0.042 | 0.281 | 0.123 | - | |
| Birch-dominated sites | |||||||
| Pythiales | Rel. abund. | 0.983 | - | 0.989 | 0.733 | 0.726 | - |
| OTU rich. | 17.5 | - | 15.7 | 13.0 | 22.3 | - | |
| Shannon div. | 1.094 | - | 0.795 | 1.234 | 0.591 | - | |
| Saprolegniales | Rel. abund. | 0.005 | - | 0.011 | 0.256 | 0.264 | - |
| OTU rich. | 3.54 | - | 3.27 | 9.17 | 9.10 | LS **, FS **, LN **, FN * | |
| Shannon div. | 0.101 | - | 0.058 | 0.837 | 0.858 | LS **, FS **, LN **, FN * | |
| Peronosporales | Rel. abund. | 0.007 | - | 0.000 | 0.008 | 0.007 | - |
| OTU rich. | 2.553 | - | 0.965 | 2.632 | 1.327 | - | |
| Shannon div. | 0.183 | - | 0.030 | 0.103 | 0.019 | - | |
| Forest sites | |||||||
| Pythiales | Rel. abund. | 0.947 | 0.914 | 0.886 | 0.602 | 0.690 | LS * |
| OTU rich. | 19.8 | 21.1 | 19.0 | 19.0 | 14.7 | - | |
| Shannon div. | 1.310 | 0.904 | 1.054 | 1.008 | 0.609 | - | |
| Saprolegniales | Rel. abund. | 0.0367 | 0.089 | 0.095 | 0.383 | 0.292 | LS * |
| OTU rich. | 3.66 | 4.95 | 5.36 | 8.44 | 11.45 | LN ***, EN * | |
| Shannon div. | 0.171 | 0.344 | 0.296 | 0.956 | 1.250 | LN ***, EN **, LS **, FN * | |
| Peronosporales | Rel. abund. | 0.011 | -0.003 | 0.015 | 0.011 | 0.016 | - |
| OTU rich. | 2.03 | 2.45 | 1.15 | 2.00 | 1.64 | - | |
| Shannon div. | 0.094 | 0.241 | 0.034 | 0.145 | 0.123 | - | |
| Urban sites | |||||||
| Pythiales | Rel. abund. | 0.948 | - | 0.967 | 0.827 | 0.846 | - |
| OTU rich. | 17.0 | - | 19.1 | 22.9 | 17.5 | - | |
| Shannon div. | 1.331 | - | 1.310 | 1.237 | 0.823 | - | |
| Saprolegniales | Rel. abund. | 0.018 | - | 0.025 | 0.117 | 0.138 | - |
| OTU rich. | 2.02 | - | 4.83 | 7.42 | 9.08 | LN * | |
| Shannon div. | 0.016 | - | 0.078 | 0.590 | 0.791 | - | |
| Peronosporales | Rel. abund. | 0.017 | - | -0.002 | 0.034 | 0.010 | - |
| OTU rich. | 2.77 | - | 1.13 | 2.54 | 1.55 | FS * | |
| Shannon div. | 0.286 | - | 0.028 | 0.226 | 0.029 | - | |
| Alder-Dominated Sites | Forest | Urban | Significance (p-Value) | |
|---|---|---|---|---|
| Pythiales | Rel. abund. | 0.823 | 0.915 | 0.2500 # |
| OTU rich. | 21.7 | 21.7 | 0.9912 # | |
| Shannon div. | 1.29 | 1.30 | 0.9579 # | |
| Saprolegniales | Rel. abund. | 0.156 | 0.042 | 0.1293 # |
| OTU rich. | 6.73 | 6.86 | 0.9349 # | |
| Shannon div. | 0.638 | 0.441 | 0.4057 # | |
| Peronosporales | Rel. abund. | 0.015 | 0.023 | 0.7553 # |
| OTU rich. | 1.83 | 2.19 | 0.2887 # | |
| Shannon div. | 0.114 | 0.203 | 0.5013 # | |
| Birch-dominated sites | ||||
| Pythiales | Rel. abund. | 0.741 | 0.975 | 0.0050 ** |
| OTU rich. | 14.6 | 19.6 | 0.1336 # | |
| Shannon div. | 0.681 | 1.176 | 0.0139 * | |
| Saprolegniales | Rel. abund. | 0.249 | 0.019 | 0.0062 ** |
| OTU rich. | 7.26 | 5.28 | 0.0812 # | |
| Shannon div. | 0.619 | 0.308 | 0.0478 * | |
| Peronosporales | Rel. abund. | 0.007 | 0.004 | 0.4565 # |
| OTU rich. | 1.58 | 2.16 | 0.2236 # | |
| Shannon div. | 0.100 | 0.067 | 0.7103 # | |
| Variable | Variance (%) | F Value | p Value |
|---|---|---|---|
| Temperature seasonality | 5.99 | 5.66 | 0.001 |
| Annual mean temperature | 5.75 | 5.43 | 0.001 |
| Mean temperature of warmest quarter | 5.38 | 5.08 | 0.001 |
| Precipitation of wettest quarter | 4.13 | 3.90 | 0.001 |
| Annual precipitation | 3.15 | 2.97 | 0.009 |
| Mean temperature of driest quarter | 2.29 | 2.16 | 0.037 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Riit, T.; Cleary, M.; Adamson, K.; Blomquist, M.; Burokienė, D.; Marčiulynienė, D.; Oliva, J.; Poimala, A.; Redondo, M.A.; Strømeng, G.M.; et al. Oomycete Soil Diversity Associated with Betula and Alnus in Forests and Urban Settings in the Nordic–Baltic Region. J. Fungi 2023, 9, 926. https://doi.org/10.3390/jof9090926
Riit T, Cleary M, Adamson K, Blomquist M, Burokienė D, Marčiulynienė D, Oliva J, Poimala A, Redondo MA, Strømeng GM, et al. Oomycete Soil Diversity Associated with Betula and Alnus in Forests and Urban Settings in the Nordic–Baltic Region. Journal of Fungi. 2023; 9(9):926. https://doi.org/10.3390/jof9090926
Chicago/Turabian StyleRiit, Taavi, Michelle Cleary, Kalev Adamson, Mimmi Blomquist, Daiva Burokienė, Diana Marčiulynienė, Jonàs Oliva, Anna Poimala, Miguel Angel Redondo, Gunn Mari Strømeng, and et al. 2023. "Oomycete Soil Diversity Associated with Betula and Alnus in Forests and Urban Settings in the Nordic–Baltic Region" Journal of Fungi 9, no. 9: 926. https://doi.org/10.3390/jof9090926









