Role of CpxR in Biofilm Development: Expression of Key Fimbrial, O-Antigen and Virulence Operons of Salmonella Enteritidis
Abstract
:1. Introduction
2. Results
2.1. Extracytoplasmic Stress-Response Regulators, RpoE and CpxR, Differentially Affect the Growth Rate of S. Enteritidis
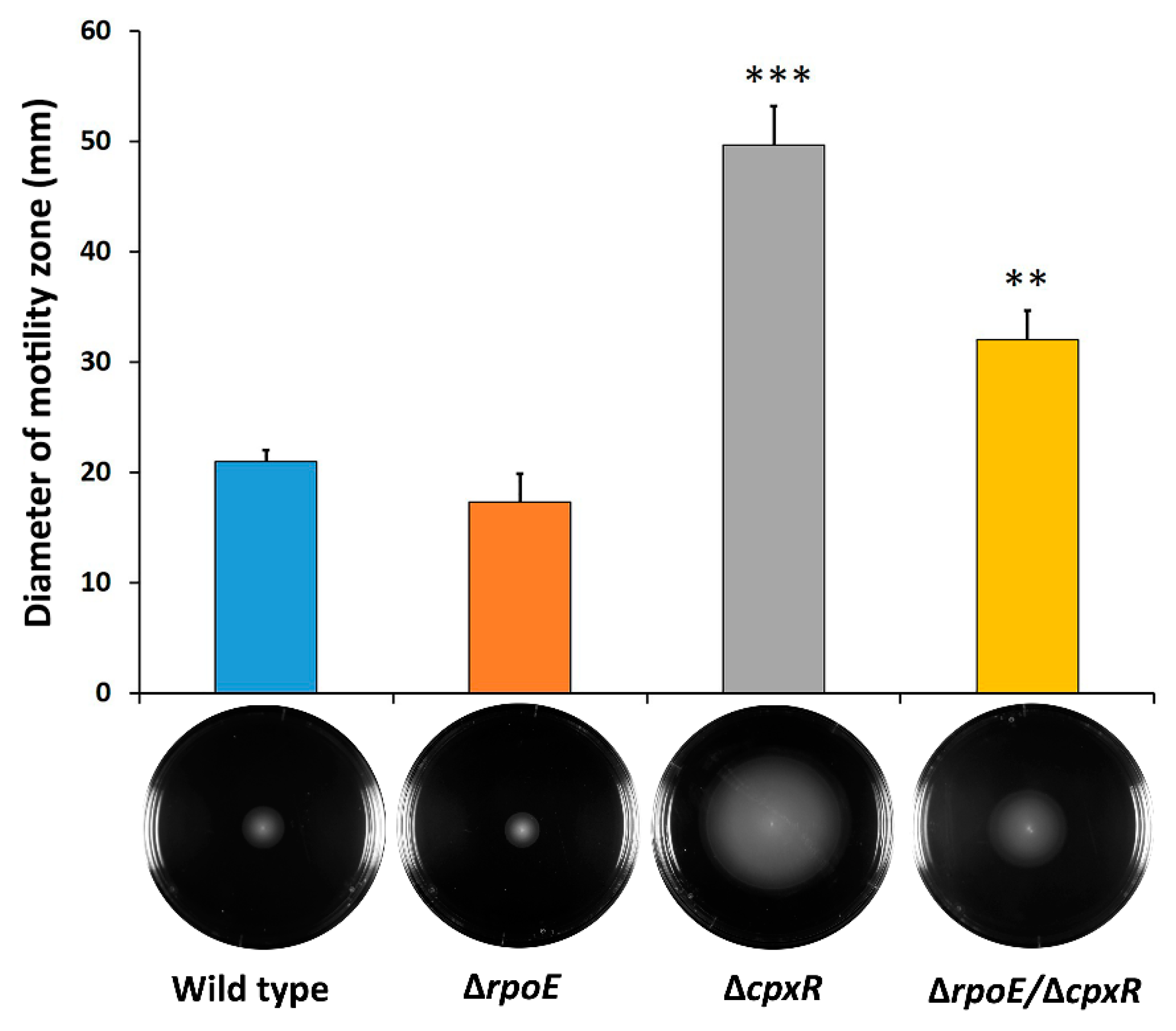
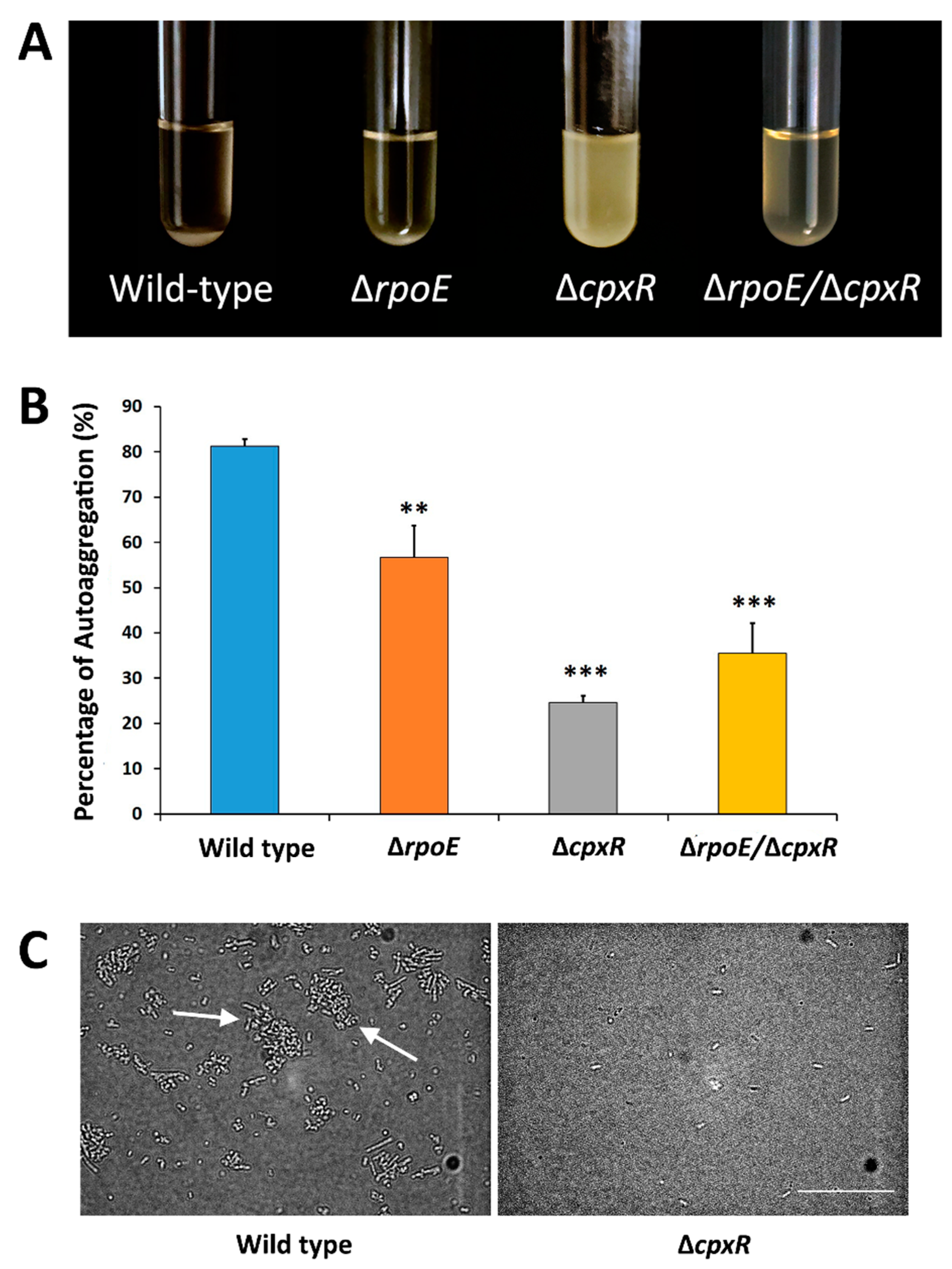
2.2. CpxR Impacts Motility and Auto-Aggregation of S. Enteritidis
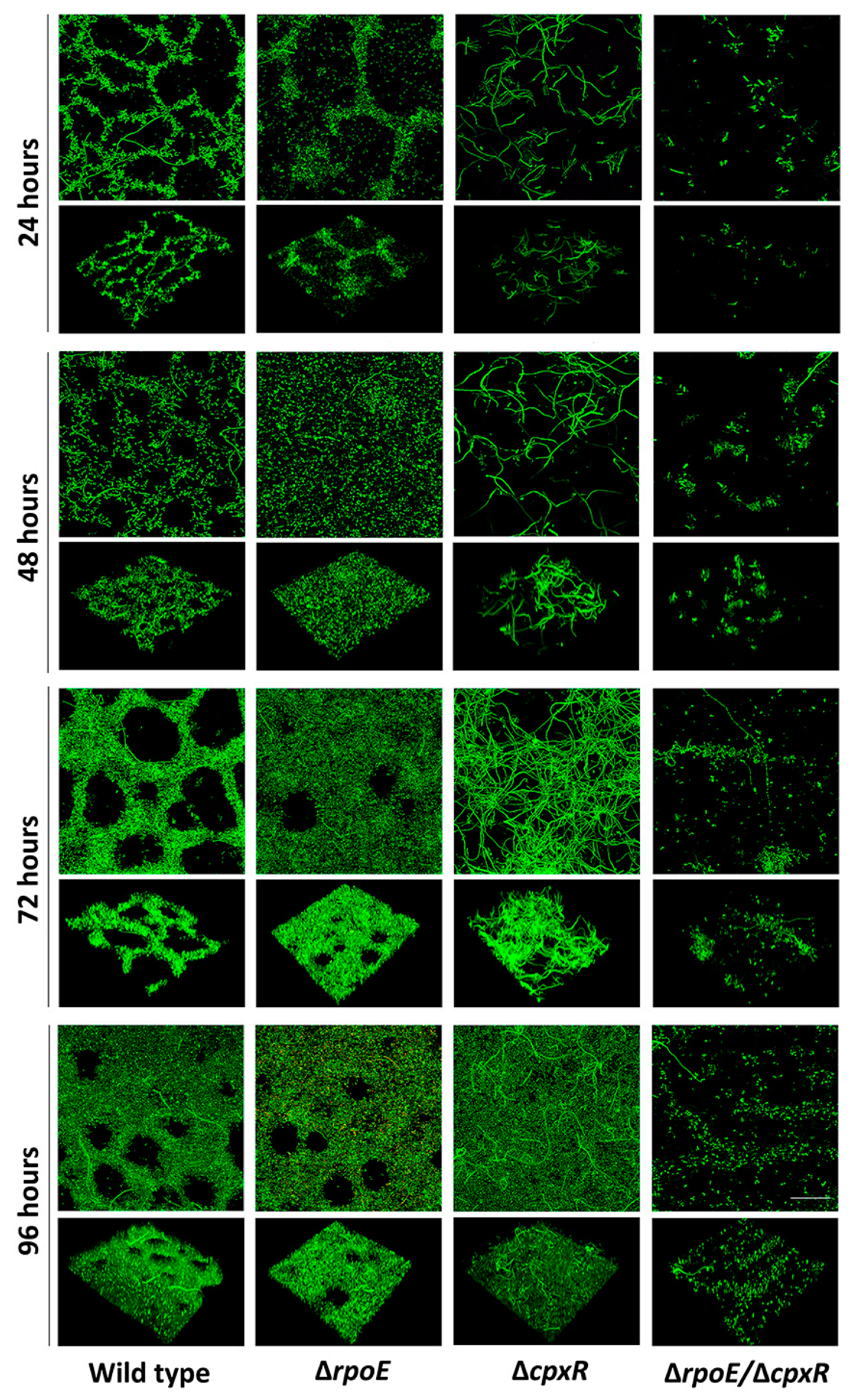
2.3. CpxR and CpxR/RpoE Have Profound Effects on Biofilm Formation
2.4. Effect of cpxR Deletion on the Biofilm Transcriptome
2.5. Effect of cpxR Deletion on the Transcriptome of Planktonic Cells
3. Discussion
4. Experimental Procedures
4.1. Bacterial Strains, Plasmids and Growth Conditions
4.2. Construction of ΔcpxR, ΔrpoE and ΔcpxR/ΔrpoE Salmonella Enteritidis Mutant Strains
4.3. Growth Assay
4.4. Swarming-Motility Assay
4.5. Auto-Aggregation Assay
4.6. Microtiter-Plate Biofilm Formation Assay
4.7. Flow-Cell Biofilm Formation Assay
4.8. Confocal Laser Scanning Microscopy (CLSM) and Biofilm Quantification
4.9. Preparation of Planktonic and Biofilm Samples for RNA Extraction
4.10. RNA-Seq Analysis
4.11. Validation of RNA-Seq Data Using Real-Time Polymerase Chain Reaction (PCR)
4.12. Experimental Replications and Bioinformatics
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Majowicz, S.E.; Musto, J.; Scallan, E.; Angulo, F.J.; Kirk, M.; O’Brien, S.J.; Jones, T.F.; Fazil, A.; Hoekstra, R.M.; for the international Collaboration on Enteric Disease “Burden of Illness” Studies. The global burden of nontyphoidal Salmonella gastroenteritis. Clin. Infect. Dis. 2010, 50, 882–889. [Google Scholar] [CrossRef] [PubMed]
- Langridge, G.C.; Fookes, M.; Connor, T.R.; Feltwell, T.; Feasey, N.; Persons, B.N.; Seth-Smith, H.M.B.; Barquist, L.; Stedman, A.; Humphrey, T.; et al. Patterns of genome evolution that have accompanied host adaptation in Salmonella. Proc. Natl. Acad. Sci. USA 2015, 112, 863–868. [Google Scholar] [CrossRef]
- World Health Organization. Available online: https://www.who.int/news-room/fact-sheets/detail/salmonella-(non-typhoidal) (accessed on 2 October 2019).
- Korber, D.R.; Choi, A.; Wolfaardt, G.M.; Ingham, S.C.; Caldwell, D.E. Substratum topography influences susceptibility of Salmonella enteritidis biofilms to trisodium phosphate. Appl. Env. Microbiol. 1997, 63, 3352–3358. [Google Scholar]
- Dantas, S.T.A.; Rossi, B.F.; Bonsaglia, E.C.R.; Castilho, I.G.; Hernandes, R.T.; Fernandes, A.; Rall, V.L.M. Cross-contamination and biofilm formation by Salmonella enterica serovar Enteritidis on various cutting boards. Foodborne Pathog Dis. 2018, 15, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Grabowicz, M.; Silhavy, T.J. Envelope stress responses: An interconnected safety net. Trends Biochem Sci. 2017, 42, 232–242. [Google Scholar] [CrossRef] [PubMed]
- Rhodius, V.A.; Suh, W.C.; Nonaka, G.; West, J.; Gross, C.A. Conserved and variable functions of the sigma E stress response in related genomes. PLoS Biol. 2006, 4, e2. [Google Scholar]
- Ravio, T.L. Everything old is new again: An update on current research on the Cpx envelope stress response. Biochim. Biophys Acta. 2014, 1843, 1529–1541. [Google Scholar] [CrossRef] [Green Version]
- Hung, D.L.; Raivio, T.L.; Jones, C.H.; Silhavy, T.J.; Hultgren, S.J. Cpx signalling pathway monitors biogenesis and affects assembly and expression of P pili. EMBO J. 2001, 20, 1508–1518. [Google Scholar] [CrossRef]
- Vidovic, S.; Elder, J.; Medihala, P.; Lawrence, J.R.; Predicala, B.; Zhang, H.; Korber, D.R. ZnO nanoparticles impose a panmetabolic toxic effect along with strong necrosis, inducing activation of the envelope stress response in Salmonella enterica serovar Enteritidis. Antimicrob Agents Chemother 2015, 59, 3317–3328. [Google Scholar] [CrossRef]
- Vidovic, S.; Medihala, P.; Dynes, J.J.; Daida, P.; Vujanovic, V.; Hitchcock, A.P.; Shetty, D.; Zhang, H.; Brown, D.R.; Lawrence, J.R.; et al. Importance of the RpoE regulon in maintaining the lipid bilayer during antimicrobial treatment with the polycationic agent, chlorhexidine. Proteomics 2018, 18. [Google Scholar] [CrossRef]
- Rattanama, P.; Thompson, J.R.; Kongkerd, M.; Srinitiwarawong, K.; Vuddhakul, V.; Mekalanos, J.J. Sigma E regulators control haemolytic activity and virulence in a shrimp pathogenic Vibrio harveyi. PLoS ONE 2012, 7, e32523. [Google Scholar] [CrossRef]
- Matter, L.B.; Ares, M.A.; Abundes-Gallegos, J.; Cedillo, M.L.; Yáñez, J.A.; Martinez-Laguna, Y.; De la Cruz, M.A.; Girón, J.A. The CpxRA stress response system regulates virulence features of avian pathogenic Escherichia coli. Env. Microbiol. 2018, 20, 3363–3377. [Google Scholar] [CrossRef] [PubMed]
- Barchinger, S.E.; Ades, S.E. Regulated proteolysis: Control of the Escherichia coli σE-dependent cell envelope stress response. Subcell Biochem. 2013, 66, 129–160. [Google Scholar] [PubMed]
- DiGiuseppe, P.A.; Silhavy, T.J. Signal detection and target gene induction by the CpxRA two-component system. J. Bacteriol. 2003, 185, 2432–2440. [Google Scholar] [CrossRef] [PubMed]
- Raivio, T.L.; Silhavy, T.J. Transduction of envelope stress in Escherichia coli by the Cpx two-component system. J. Bacteriol. 1997, 17, 7724–7733. [Google Scholar] [CrossRef]
- Batchelor, E.; Walthers, D.; Kenney, L.J.; Goulian, M. The Escherichia coli CpxA-CpxR envelope stress response system regulates expression of the porins OmpF and OmpC. J. Bacteriol. 2005, 187, 5723–5731. [Google Scholar] [CrossRef]
- Gerken, H.; Charlson, E.S.; Cicirelli, E.M.; Kenney, L.J.; Misra, R. MzrA: A novel modulator of the EnvZ/OmpR two-component regulon. Mol. Microbiol. 2009, 72, 1408–1422. [Google Scholar] [CrossRef]
- Grabowicz, M.; Koren, D.; Silhavy, T.J. The CpxQ sRNA negatively regulates Skp to prevent mistargeting of β-barrel outer membrane proteins into the cytoplasmic membrane. mBio 2016, 7, e00312-16. [Google Scholar] [CrossRef]
- Li, H.; Liu, F.; Peng, W.; Yan, K.; Zhao, H.; Liu, T.; Cheng, H.; Chang, P.; Yuan, F.; Chen, H.; et al. The CpxA/CpxR two-component system affects biofilm formation and virulence in Actinobacillus pleuropmeumoniae. Front Cell Infect Microbiol. 2018, 8, 72. [Google Scholar] [CrossRef]
- Vestby, L.K.; Møretrø, T.; Balance, S.; Langsrud, S.; Nesse, L.L. Survival potential of wild type cellulose deficient Salmonella from the feed industry. BMC Vet. Res. 2009, 5, 43. [Google Scholar] [CrossRef]
- Yaron, S.; Romling, U. Biofilm formation by enteric pathogens and its role in plant colonization and persistence. Microb Biotechnol. 2014, 7, 496–516. [Google Scholar] [CrossRef] [PubMed]
- Baloda, S.B.; Christensen, L.; Trajcevska, S. Persistence of a Salmonella enterica serovar Typhimurium DT12 clone in a piggery and in agriculture soil amended with Salmonella-contaminated slurry. Appl. Env. Microbiol. 2001, 67, 2859–2862. [Google Scholar] [CrossRef] [PubMed]
- Greene, S.K.; Daly, E.R.; Talbot, E.A.; Demma, L.J.; Holzbauer, S.; Patel, N.J.; Hill, T.A.; Walderhaug, M.O.; Hoekstra, R.M.; Lynch, M.F.; et al. Recurrent multistate outbreak of Salmonella Newport associated with tomatoes from contaminated fields, 2005. Epidemiol. Infect. 2008, 136, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Tremblay, Y.D.N.; Vogeleer, P.; Jacques, M.; Harel, J. High-throughput microfluidic method to study biofilm formation and host-pathogen interactions in pathogenic Escherichia coli. Appl. Env. Microbiol. 2015, 81, 2827–2840. [Google Scholar] [CrossRef] [PubMed]
- Otto, K.; Silhavy, T.J. Surface sensing and adhesion of Escherichia coli controlled by the Cpx-signalling pathway. Proc. Natl. Acad. Sci. USA 2002, 99, 2287–2292. [Google Scholar] [CrossRef] [PubMed]
- Raivio, T.L. Envelope stress response and Gram-negative bacterial pathogenesis. Mol. Microbiol. 2005, 56, 1119–1128. [Google Scholar] [CrossRef]
- Hernday, A.D.; Braaten, B.A.; Broitman-Maduro, G.; Engelberts, P.; Low, D.A. Regulation of the Pap epigenetic switch by CpxAR: Phosphorylated CpxR inhibits transition to the phase ON state by competition with Lrp. Mol. Cell 2004, 16, 537–547. [Google Scholar]
- Nevesinjac, A.Z.; Raivio, T.L. The Cpx envolope stress response affects expression of the type IV bundle-forming pili of enteropathogenic Escherichia coli. J. Bacteriol. 2005, 187, 672–686. [Google Scholar] [CrossRef]
- Vogt, S.L.; Nevesinjac, A.Z.; Humphries, R.M.; Donnenberg, M.S.; Armstrong, G.D.; Raivio, T.L. The Cpx envelope stress response both facilitates and inhibits elaboration of the enteropathogenic Escherichia coli bundle-forming pilus. Mol. Microbiol. 2010, 76, 1095–1110. [Google Scholar] [CrossRef]
- De la Cruz, M.A.; Ruiz-Tagle, A.; Ares, M.A.; Pacheco, S.; Yanez, J.A.; Cedillo, L.; Torres, J.; Giron, J.A. The expression of Longus type 4 pilus of enterotoxigenic Escherichia coli is regulated by LngR and LngS and by H-NS, CpxR and CRP global regulators. Environ. Microbiol. 2017, 19, 1761–1775. [Google Scholar] [CrossRef]
- Dudin, O.; Geiselmann, J.; Ogasawara, H.; Ishihama, A.; Lacour, S. Repression of flagellar genes in exponential phase by CsgD and CpxR, two crucial modulators of Escherichia coli biofilm formation. J. Bacteriol. 2014, 196, 707–715. [Google Scholar] [CrossRef] [PubMed]
- Dorel, C.; Vidal, O.; Prigent-Combaret, C.; Vallet, I.; Lejeune, P. Involvement of the Cpx signal transduction pathway of E. coli in biofilm formation. FEMS Microbiol Lett. 1999, 178, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.; Wood, T.K. OmpA influences Escherichia coli biofilm formation by repressing cellulose production through the CpxRA two-component system. Env. Microbiol. 2009, 11, 2735–2746. [Google Scholar] [CrossRef] [PubMed]
- Haraga, A.; Ohlson, M.B.; Miller, S.I. Salmonellae interplay with host cells. Nat. Rev. Microbiol. 2008, 6, 53–66. [Google Scholar] [CrossRef] [PubMed]
- Shah, D.H.; Zhou, X.; Kim, H.Y.; Call, D.R.; Guard, J. Transposon mutagenesis of Salmonella enterica serovar Enteritidis identifies genes that contribute to invasiveness in human and chicken cells and survival in egg albumen. Infect. Immun. 2012, 80, 4203–4215. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Zhang, L.; Wang, P.; Meng, G. Structural basis of host recognition and biofilm formation by Salmonella Saf pili. eLife 2017, 6, e28619. [Google Scholar] [CrossRef]
- Partridge, J.D.; Harshey, R.M. More than motility: Salmonella flagella contribute to overriding friction and facilitating colony hydration during swarming. J. Bacteriol. 2013, 195, 919–929. [Google Scholar] [CrossRef]
- Ditmarsch, D.; Boyle, K.E.; Sakhtah, H.; Oyler, J.E.; Nadell, C.D.; Déziel, É.; Dietrich, L.E.P.; Xavier, J.B. Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria. Cell Rep. 2013, 4, 697–708. [Google Scholar] [CrossRef]
- Belas, P. Biofilms, flagella, and mechanosensing of surfaces by bacteria. Trends Microbiol. 2014, 22, 517–527. [Google Scholar] [CrossRef]
- Barnhart, M.M.; Chapman, M.R. Curli biogenesis and function. Annu Rev. Microbiol. 2006, 60, 131–147. [Google Scholar] [CrossRef]
- Weening, E.H.; Barker, J.D.; Laarakker, M.C.; Humphries, A.D.; Tsolis, R.M.; Bäumler, A.J. The Salmonella enterica serotype Typhimurium lpf, bcf, stb, stc, std, and sth fimbrial operons are required for intestinal persistence in mice. Infect. Immun. 2005, 73, 3358–3366. [Google Scholar] [CrossRef] [PubMed]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef] [PubMed]
- Widdel, F. Theory and measurement of bacterial growth. Di dalam Grundpraktikum Mikrobiologie 2017, 4, 1–11. [Google Scholar]
- Legendre, G.; Fay, F.; Linossier, I.; Vallee-Rehel, K. Evaluation of antibacterial activity against Salmonella Enteritidis. J. Microbiol. 2011, 49, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Shanks, R.M.Q.; Meehl, M.A.; Brothers, K.M.; Martinez, R.M.; Donegan, N.P.; Graber, M.; Cheung, A.L.; O’Toole, G.A. Genetic evidence for an alternative citrate-dependent biofilm formation pathway in Staphylococcus aureus that is dependent on fibronectin binding proteins and the GraRS two-component regulatory system. Infect. Immun. 2008, 76, 2469–2477. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A.; Kolter, R. The initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds with multiple, convergent signaling pathways: A genetic analysis. Mol. Microbiol. 1998, 28, 449–461. [Google Scholar] [CrossRef]
- Korber, D.R.; Greer, G.G.; Wolfaardt, G.M.; Kohlman, S. Efficacy enhancement of trisodium phosphate against spoilage and pathogenic bacteria in model biofilms and on adipose tissue. J. Food Prot. 2002, 65, 627–635. [Google Scholar] [CrossRef]
- Mangalappalli-Illathu, A.K.; Vidović, S.; Korber, D.R. Differential adaptive response and survival of Salmonella enterica serovar enteritidis planktonic and biofilm cells exposed to benzalkonium chloride. Antimicrob Agents Chemother. 2008, 52, 3669–3680. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liao, Y.; Smyth, G.K.; Shi, W. The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 2013, 41, e108. [Google Scholar] [CrossRef] [PubMed]
- Winer, J.; Jung, C.K.; Shackel, I.; Williams, P.M. Development and validation of real-time quantitative reverse transcriptase- polymerase chain reaction for monitoring gene expression in cardiac myocytes in vitro. Anal. Biochem. 1999, 270, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Sievert, C.; Parmer, C.; Hocking, T.; Chamberlain, S.; Ram, K.; Corvellec, M.; Despouy, P. Plotly for R: Create Interactive Web Graphics via ‘plotly.js’. R Package Version 4.1. 2017, p. 110. Available online: https://CRAN.R-project.org/package=plotly (accessed on 15 May 2019).




| Primer Name | Sequence (5′-3′) |
|---|---|
| Primers for rpoE and cpxR deletion | |
| rpoE - Forward | ATG AGC GAG CAG TTA ACG GAC CAG GTC CTG GTT GAA CGG TGT AGG CTG GAG CTG CTT CG |
| rpoE - Reverse | TCA ACG CCT GAT AAG CGG TTG AAC TTT ATT ATC AAT AGC CAT ATG AAT ATC CTC CTT AG |
| cpxR - Forward | ATT AGC GAC GCC TGA TGA CGT AAT TTC TGC CTC GGA GGT ACG TAA ACA TGT AGG CTG GAG CTG CTT CG |
| cpxR - Reverse | CCA GCG TCA ACC AGA AGA TGG CGA AGA TGC GCG CGG TTA AAC TTC CTA CAT ATG AAT ATC CTC CTT AG |
| Primers for confirmation of rpoE and cpxR gene deletions | |
| rpoE mutant F | GAC CTG TCT ACA ACA TGA CAA ACA |
| rpoE mutant R | CGG ATC AGG TGA TAA CTC TCC CAG |
| cpxR mutant F | CGC TTG CTC CCA AAA TCT TTT CTG |
| cpxR mutant R | GTT GCT CTA TCA TCA ATC CCT GGC |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shetty, D.; Abrahante, J.E.; Chekabab, S.M.; Wu, X.; Korber, D.R.; Vidovic, S. Role of CpxR in Biofilm Development: Expression of Key Fimbrial, O-Antigen and Virulence Operons of Salmonella Enteritidis. Int. J. Mol. Sci. 2019, 20, 5146. https://doi.org/10.3390/ijms20205146
Shetty D, Abrahante JE, Chekabab SM, Wu X, Korber DR, Vidovic S. Role of CpxR in Biofilm Development: Expression of Key Fimbrial, O-Antigen and Virulence Operons of Salmonella Enteritidis. International Journal of Molecular Sciences. 2019; 20(20):5146. https://doi.org/10.3390/ijms20205146
Chicago/Turabian StyleShetty, Deeksha, Juan E. Abrahante, Samuel M. Chekabab, Xuxiaochen Wu, Darren R. Korber, and Sinisa Vidovic. 2019. "Role of CpxR in Biofilm Development: Expression of Key Fimbrial, O-Antigen and Virulence Operons of Salmonella Enteritidis" International Journal of Molecular Sciences 20, no. 20: 5146. https://doi.org/10.3390/ijms20205146





