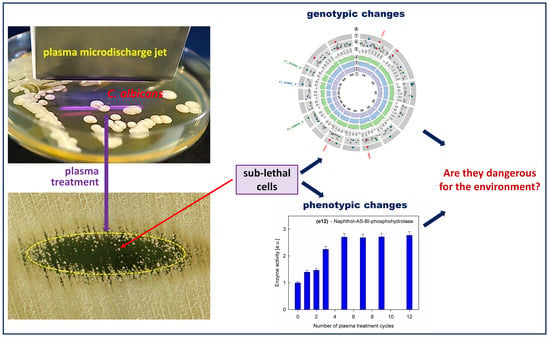
Genotypic and Phenotypic Changes in Candida albicans as a Result of Cold Plasma Treatment
Abstract
:1. Introduction
2. Results and Discussion
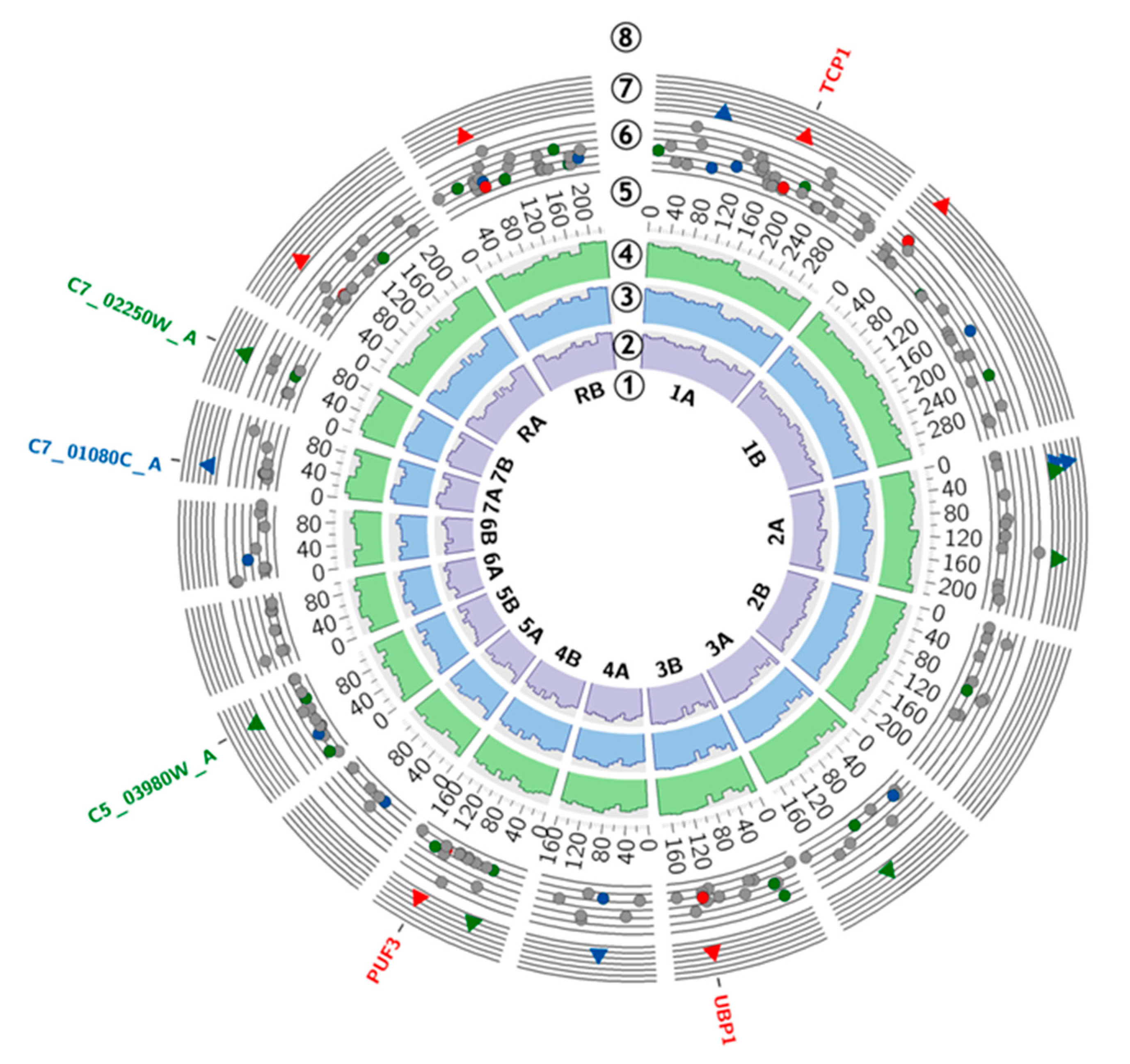
2.1. Genomic Alterations
2.2. Phenotypic Changes
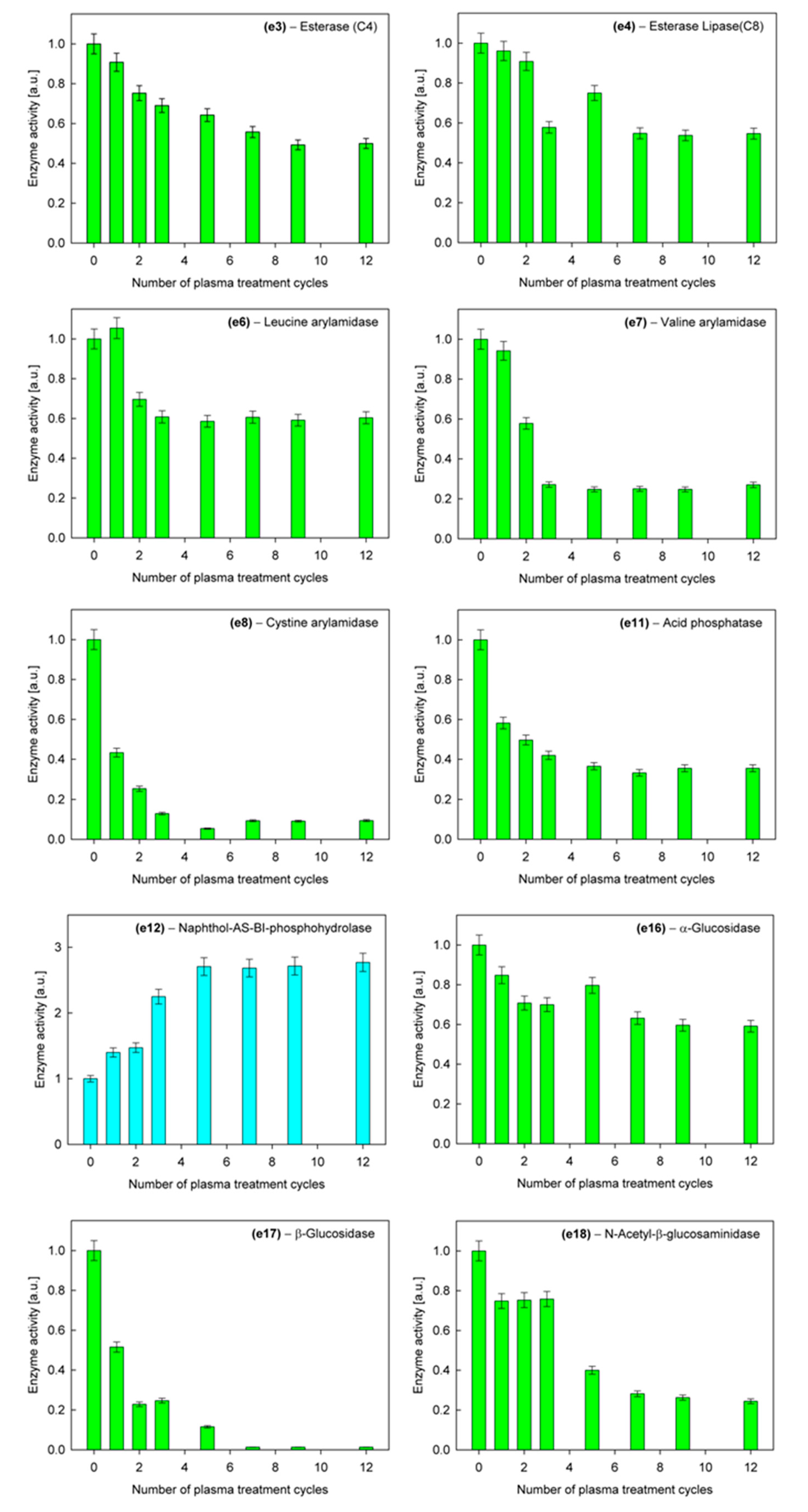
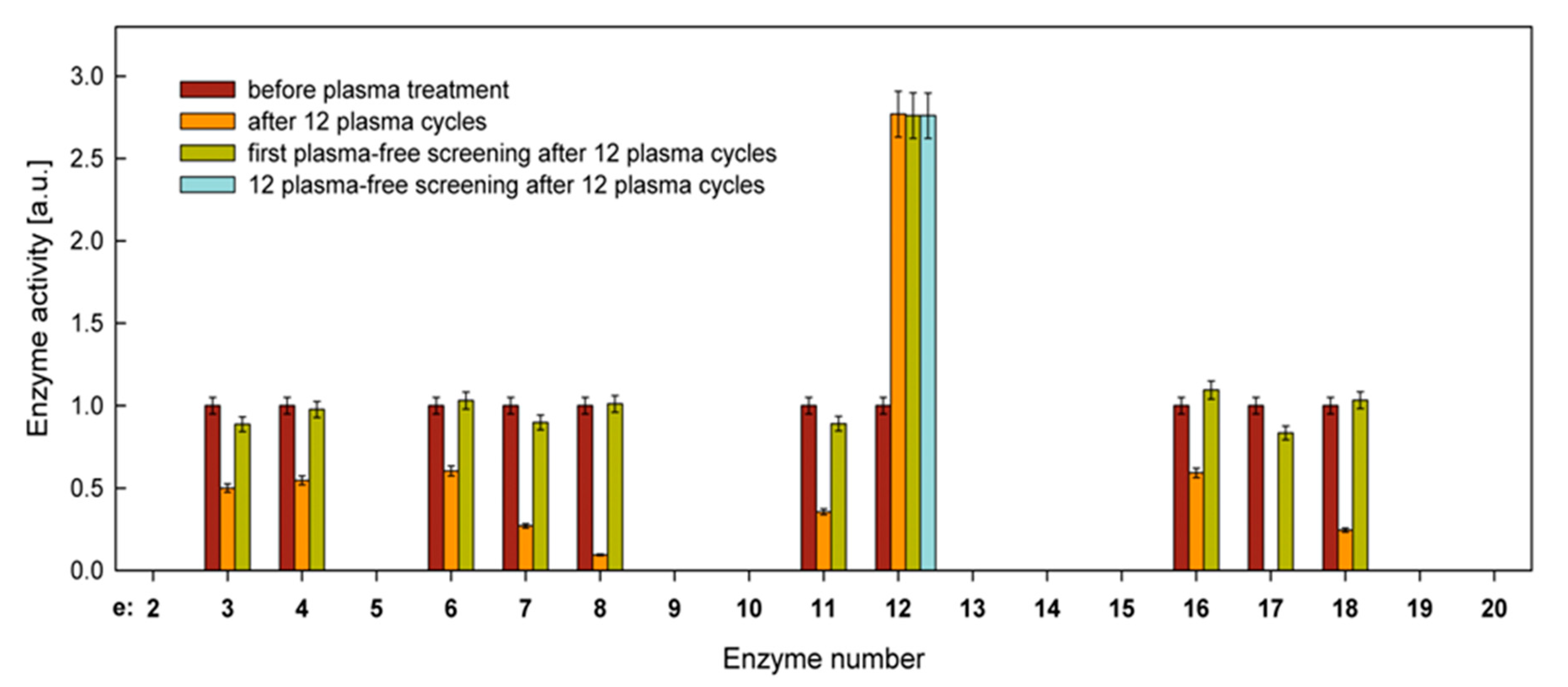
2.2.1. Carbon Assimilation and Hydrolytic Enzyme Activity
2.2.2. Susceptibility to Antifungal Drugs
3. Materials and Methods
3.1. Plasma Source
3.2. Microbiological and Plasma Treatment Procedures
3.3. DNA Isolation and Genome Bioinformatics Analysis
3.4. Carbohydrate Assimilation Test
3.5. Hydrolytic Enzyme Activity
3.6. Susceptibility Testing
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| C. albicans | Candida albicans |
| CFU/mL | Colony-forming unit per milliliter |
| CUG | Ttrinucleotide codon composed of cytosine, uracil and guanine |
| L/min | Liter per minute |
| MHz | Megahertz (frequency unit) |
| MIC | Minimum inhibitory concentration |
| SNVs | Single nucleotide variants |
| W | Watt (power unit) |
References
- Laroussi, M.; Kong, M.G.; Morfill, G.; Stolz, W. (Eds.) Plasma Medicine—Applications of Low-Temperature Gas. Plasmas in Medicine and Biology, 1st ed.; Cambridge University Press: Cambridge, UK, 2012. [Google Scholar]
- Fridman, A.; Friedman, G. Plasma Medicine, 1st ed.; Wiley: Chichester, UK, 2013. [Google Scholar]
- Shintani, H.; Sakudo, A. (Eds.) Gas Plasma Sterilization in Microbiology—Theory, Applications, Pitfalls and New Perspectives, 1st ed.; Caister Academic Press: Norfolk, UK, 2016. [Google Scholar]
- Laroussi, M. Cold plasma in medicine and healthcare: The new frontier in low temperature plasma applications. Front. Phys. 2020, 8, 74. [Google Scholar] [CrossRef]
- Dai, X.; Zhang, Z.; Zhang, J.; Ostrikov, K. Dosing: The key to precision plasma oncology. Plasma Process. Polym. 2020, e1900178. [Google Scholar] [CrossRef]
- Daeschlein, G.; Scholz, S.; Emmet, S.; von Podewils, S.; Haase, H.; von Woedtke, T.; Jüngen, M. Plasma medicine in dermatology: Basic antimicrobial efficacy testing as prerequisite to clinical plasma therapy. Plasma Med. 2012, 2, 33–69. [Google Scholar] [CrossRef] [Green Version]
- Emmert, S.; Brehmer, F.; Hänßle, H.; Helmke, A.; Mertens, N.; Ahmed, R.; Simon, D.; Wandke, D.; Maus-Friedrichs, W.; Däschlein, G.; et al. Atmospheric pressure plasma in dermatology: Ulcus treatment and much more. Clin. Plasma Med. 2013, 1, 24–29. [Google Scholar] [CrossRef]
- Liu, D.; Zhang, Y.; Xu, M.; Chen, H.; Lu, X.; Ostrikov, K. Cold atmospheric pressure plasmas in dermatology: Sources, reactive agents, and therapeutic effects. Plasma Process. Polym. 2020, 17, e1900218. [Google Scholar] [CrossRef]
- Lloyd, G.; Friedman, G.; Jafri, S.; Schultz, G.; Fridman, A.; Harding, K. Gas plasma: Medical uses and developments in wound care. Plasma Process. Polym. 2010, 7, 194–211. [Google Scholar] [CrossRef]
- Scholtz, V.; Pazlarova, J.; Souskova, H.; Khun, J.; Julak, J. Nonthermal plasma—A tool for decontamination and disinfection. Biotechnol. Adv. 2015, 33, 1108–1119. [Google Scholar] [CrossRef]
- Sakudo, A.; Yagyu, Y.; Onodera, T. Disinfection and sterilization using plasma technology: Fundamentals and future perspectives for biological applications. Int. J. Mol. Sci. 2019, 20, 5216. [Google Scholar] [CrossRef] [Green Version]
- Isbary, G.; Heinlin, J.; Shimizu, T.; Zimmermann, J.L.; Morfill, G.; Schmidt, H.U.; Monetti, R.; Steffes, B.; Bunk, W.; Li, Y.; et al. Successful and safe use of 2 min cold atmospheric argon plasma in chronic wounds: Results of a randomized controlled trial. Br. J. Dermatol. 2012, 167, 404–410. [Google Scholar] [CrossRef]
- Park, G.Y.; Park, S.J.; Choi, M.Y.; Koo, I.G.; Byun, J.H.; Hong, J.W.; Sim, J.Y.; Collins, G.J.; Lee, J.K. Atmospheric-pressure plasma sources for biomedical applications. Plasma Sources Sci. Technol. 2012, 21, 043001. [Google Scholar] [CrossRef]
- Isbary, G.; Zimmermann, J.L.; Shimizu, T.; Li, Y.F.; Morfill, G.E.; Thomas, H.M.; Steffes, B.; Heinlin, J.; Karrer, S.; Stolz, W. Non-thermal plasma—More than five years of clinical experience. Clin. Plasma Med. 2013, 1, 19–23. [Google Scholar] [CrossRef]
- Ulrich, C.; Kluschke, F.; Patzelt, A.; Vandersee, S.; Czaika, V.A.; Richter, H.; Bob, A.; von Hutten, J.; Painsi, C.; Hügel, R.; et al. Clinical use of cold atmospheric pressure argon plasma in chronic leg ulcers: A pilot study. J. Wound Care 2015, 24, 196–203. [Google Scholar] [CrossRef]
- He, M.; Duan, J.; Xu, J.; Ma, M.; Chai, B.; He, G.; Gan, L.; Zhang, S.; Duan, X.; Lu, X.; et al. Candida albicans biofilm inactivated by cold plasma treatment in vitro and in vivo. Plasma Process. Polym. 2020, 17, e1900068. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, X.F.; Li, H.P.; Wang, L.Y.; Zhang, C.; Xing, X.H.; Bao, C.Y. Atmospheric and room temperature plasma (ARTP) as a new powerful mutagenesis tool. App. Microbiol. Biotechnol. 2014, 98, 5387–5396. [Google Scholar] [CrossRef] [PubMed]
- Privat-Maldonado, A.; O’Connell, D.; Welch, E.; Vann, R.; van der Woude, M.W. Spatial dependence of DNA damage in bacteria due to low-temperature plasma application as assessed at the single cell level. Sci. Rep. 2016, 6, 35646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.Y.; Huang, Z.L.; Li, G.; Zhao, H.X.; Xing, X.H.; Sun, W.T.; Li, H.P.; Gou, Z.X.; Bao, C.Y. Novel mutation breeding method for Streptomyces avermitilis using an atmospheric pressure glow discharge plasma. J. Appl. Microbiol. 2010, 108, 851–858. [Google Scholar] [CrossRef]
- Mai-Prochnow, A.; Murphy, A.B.; McLean, K.M.; Kong, M.G.; Ostrikov, K. Atmospheric pressure plasmas: Infection control and bacterial responses. Int. J. Antimicrob. Agents 2014, 43, 508–517. [Google Scholar] [CrossRef]
- Li, G.; Li, H.P.; Wang, L.Y.; Wang, S.; Zhao, H.X.; Sun, W.T.; Xing, X.H.; Bao, C.Y. Genetic effects of radio-frequency, atmospheric-pressure glow discharges with helium. Appl. Phys. Lett. 2008, 92, 221504. [Google Scholar] [CrossRef] [Green Version]
- Alkawareek, M.Y.; Alshraiedeh, N.H.; Higginbotham., S.; Flynn, P.B.; Algwari, Q.T.; Gorman, S.P.; Graham, W.G.; Gilmore, B.F. Plasmid DNA damage following exposure to atmospheric pressure nonthermal plasma: Kinetics and influence of oxygen admixture. Plasma Med. 2014, 4, 211–219. [Google Scholar] [CrossRef]
- Edengeiser, E.; Lackmann, J.W.; Bründermann, E.; Schneider, S.; Benedikt, J.; Bandow, J.E.; Havenith, M. Synergistic effects of atmospheric pressure plasma-emitted components on DNA oligomers: A Raman spectroscopic study. J. Biophotonics 2015, 8, 918–924. [Google Scholar] [CrossRef]
- Li, H.P.; Wang, L.Y.; Li, G.; Jin, L.H.; Le, P.S.; Zhao, H.X.; Xing, X.H.; Bao, C.Y. Manipulation of lipase activity by the helium radio-frequency, atmospheric-pressure glow discharge plasma jet. Plasma Process. Polym. 2011, 8, 224–229. [Google Scholar] [CrossRef]
- Guo, L.; Zhao, Y.; Liu, D.; Liu, Z.; Chen, C.; Xu, R.; Tian, M.; Wang, X.; Chen, H.; Kong, M.G. Cold atmospheric-pressure plasma induces DNA−protein crosslinks through protein oxidation. Free Radic. Res. 2018, 52, 783–798. [Google Scholar] [CrossRef] [PubMed]
- Merz, W.G. Candida albicans strain delineation. Clin. Microbiol. Rev. 1990, 3, 321–334. [Google Scholar] [CrossRef] [PubMed]
- Kabir, M.A.; Hussain, M.A.; Ahmed, Z. Candida albicans: A model organism for studying fungal pathogens. ISRN Microbiol. 2012, 538694. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yapar, N. Epidemiology and risk factors for invasive candidiasis. Ther. Clin. Risk Manag. 2014, 10, 95–105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pappas, P.G.; Kauffman, C.A.; Andes, D.R.; Clancy, C.J.; Marr, K.A.; Ostrosky-Zeichner, L.; Reboli, A.C.; Schuster, M.G.; Vazquez, J.A.; Walsh, T.J.; et al. Clinical practice guideline for the management of candidiasis: 2016 update by the infectious diseases society of America. Clin. Infect. Dis. 2016, 62, e1–e50. [Google Scholar] [CrossRef] [Green Version]
- Pfaller, M.A. Antifungal drug resistance: Mechanisms, epidemiology, and consequences for treatment. Am. J. Med. 2012, 125, S3–S13. [Google Scholar] [CrossRef]
- Pfaller, M.A.; Diekema, D.J.; Turnidge, J.D.; Castanheira, M.; Jones, R.N. Twenty years of the SENTRY antifungal surveillance program: Results for Candida species from 1997–2016. Open Forum Infect. Dis. 2019, 6, S79–S94. [Google Scholar] [CrossRef] [Green Version]
- Roemer, T.; Krysan, D.J. Antifungal drug development: Challenges, unmet clinical needs, and new approaches. Cold Spring Harb. Perspect. Med. 2014, 4, a019703. [Google Scholar] [CrossRef]
- Havlickova, B.; Czaika, V.A.; Friedrich, M. Epidemiological trends in skin mycoses worldwide. Mycoses 2008, 51, 2–15. [Google Scholar] [CrossRef]
- Raiesi, O.; Siavash, M.; Mohammadi, F.; Chabavizadeh, J.; Mahaki, B.; Maherolnaghsh, M.; Dehghan, P. Frequency of cutaneous fungal infections and azole resistance of the isolates in patients with diabetes mellitus. Adv. Biomed. Res. 2017, 6, 71. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.M.; Zhang, G.J.; Yuan, Y.K.; Ma, Y.; Xu, G.M.; Yang, Y. Research on the inactivation effect of low-temperature plasma on Candida albicans. IEEE Trans. Plasma Sci. 2008, 36, 498–503. [Google Scholar] [CrossRef]
- Song, Y.; Liu, D.; Ji, L.; Wang, W.; Zhao, P.; Quan, C.; Niu, J.; Zhang, X. The inactivation of resistant Candida albicans in a sealed package by cold atmospheric pressure plasmas. Plasma Process. Polym. 2012, 9, 17–21. [Google Scholar] [CrossRef]
- Song, Y.; Liu, D.; Ji, L.; Wang, W.; Niu, J.; Zhang, X. Plasma inactivation of Candida albicans by an atmospheric cold plasma brush composed of hollow fibers. IEEE Trans. Plasma Sci. 2012, 40, 1098–1102. [Google Scholar] [CrossRef]
- Kostov, K.G.; Borges, A.C.; Koga-Ito, Y.; Nishime, T.M.C.; Prysiazhnyi, V.; Honda, R.Y. Inactivation of Candida albicans by cold atmospheric pressure plasma jet. IEEE Trans. Plasma Sci. 2015, 43, 770–775. [Google Scholar] [CrossRef]
- Tyczkowska-Sieroń, E.; Kapica, R.; Markiewicz, J.; Tyczkowski, J. Linear microdischarge jet for microbiological applications. Plasma Med. 2018, 8, 57–71. [Google Scholar] [CrossRef]
- Rahimi-Verki, N.; Shapoorzadeh, A.; Razzaghi-Abyaneh, M.; Atyabi, S.M.; Shams-Ghahfarokhi, M.; Jahanshiri, Z.; Gholami-Shabani, M. Cold atmospheric plasma inhibits the growth of Candida albicans by affecting ergosterol biosynthesis and suppresses the fungal virulence factors in vitro. Photodiagnosis Photodyn. Ther. 2016, 13, 66–72. [Google Scholar] [CrossRef]
- Borges, A.C.; Nishime, T.M.C.; Kostov, K.G.; de Morais Gouvêa Lima, G.; Gontijo, A.V.L.; de Carvalho, J.N.M.M.; Honda, R.Y.; Koga-Ito, C.Y. Cold atmospheric pressure plasma jet modulates Candida albicans virulence traits. Clin. Plasma Med. 2017, 7/8, 9–15. [Google Scholar] [CrossRef] [Green Version]
- Yan, L.; Yang, C.; Tang, J. Disruption of the intestinal mucosal barrier in Candida albicans infections. Microbiol. Res. 2013, 168, 389–395. [Google Scholar] [CrossRef]
- Zhou, Y.; Liao, M.; Zhu, C.; Hu, Y.; Tong, T.; Peng, X.; Li, M.; Feng, M.; Cheng, L.; Ren, B.; et al. ERG3 and ERG11 genes are critical for the pathogenesis of Candida albicans during the oral mucosal infection. Int. J. Oral Sci. 2018, 10, 9. [Google Scholar] [CrossRef]
- Skrzypek, M.S.; Binkley, J.; Binkley, G.; Miyasato, S.R.; Simison, M.; Sherlock, G. The Candida Genome Database (CGD): Incorporation of Assembly 22, systematic identifiers and visualization of high throughput sequencing data. Nucleic Acids Res. 2017, 45, D592–D596. [Google Scholar] [CrossRef] [Green Version]
- Selmecki, A.; Forche, A.; Berman, J. Genomic plasticity of the human fungal pathogen Candida albicans. Eukaryot. Cell 2010, 9, 991–1008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schaller, M.; Borelli, C.; Korting, H.C.; Hube, B. Hydrolytic enzymes as virulence factors of Candida albicans. Mycoses 2005, 48, 365–377. [Google Scholar] [CrossRef]
- Silva, S.; Negri, M.; Henriques, M.; Oliviera, R.; Williams, D.W.; Azeredo, J. Candida glabrata, Candida parapsilosis and Candida tropicalis: Biology, epidemiology, pathogenicity and antifungal resistance. FEMS Microbiol. Rev. 2012, 36, 288–305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Staniszewska, M.; Gizińska, M.; Kurządkowski, W. The role of leucine arylamidase in the virulence of Candida albicans. Postep. Microbiol. 2013, 52, 373–380. [Google Scholar]
- Majumdar, S.; Pal, S. Information transmission in microbial and fungal communication: From classical to quantum. J. Cell Commun. Signal. 2018, 12, 491–502. [Google Scholar] [CrossRef] [PubMed]
- Bloemendal, S.; Kück, U. Cell-to-cell communication in plants, animals, and fungi: A comparative review. Sci. Nat. 2013, 100, 3–19. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.P.; Budge, S.; Kaloriti, D.; Tillmann, A.; Jacobsen, M.D.; Yin, Z.; Ene, I.V.; Bohovych, I.; Sandai, D.; Kastora, S.; et al. Stress adaptation in a pathogenic fungus. J. Exp. Biol. 2014, 217, 144–155. [Google Scholar] [CrossRef] [Green Version]
- Mohammadi, F.; Ghasemi, Z.; Familsatarian, B.; Salehi, E.; Sharifynia, S.; Marikani, A.; Mirzadeh, M.; Hosseini, M.A. Relationship between antifungal susceptibility profile and virulence factors in Candida albicans isolated from nail specimens. Rev. Soc. Bras. Med. Trop. 2020, 53, e20190214. [Google Scholar] [CrossRef]
- Tyczkowski, J.; Kazimierski, P.; Zieliński, J. Microplasma Electrode Reactor for the Surface Treatment under Atmospheric Pressure. Patent PL 212569, 31 October 2012. [Google Scholar]
- Babraham Bioinformatics; FastQC A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 26 August 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Koboldt, D.C.; Chen, K.; Wylie, T.; Larson, D.E.; McLellan, M.D.; Mardis, E.R.; Weinstock, G.M.; Wilson, R.K.; Ding, L. VarScan: Variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 2009, 25, 2283–2285. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [Green Version]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [Green Version]
- Ramírez, F.; Ryan, D.P.; Grüning, B.; Bhardwaj, V.; Kilpert, F.; Richter, A.S.; Heyne, S.; Dündar, F.; Manke, T. DeepTools2: A next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016, 44, W160–W165. [Google Scholar] [CrossRef]
- Collin, R.; Starr, M.J. Comparative ontogenetic changes in enzyme activity during embryonic development of calyptraeid gastropods. Biol. Bull. 2013, 225, 8–17. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Standard Name | Systematic Name | Localization | Mutation | Sample 12_1 p-Value | Sample 12_2 p-Value | Description [44] |
|---|---|---|---|---|---|---|
| Single Nucleotide Variants | ||||||
| TCP1 | C1_08560W_A | Chr1A:1874103 | G>C (p. Gly>Arg) | 0.0369 | 0.0095 | Chaperonin- containing T-complex subunit, induced by alpha pheromone in SpiderM medium; stationary phase enriched protein |
| Chr1A:1874104 | G>A(p.Gly>Glu) | 0.0471 | 0.0112 | |||
| Uncharacterized region | Chr1B:275481 | T>A | 0.0033 | 0.0104 | – | |
| UBP1 | C3_05870C_A | Chr3B: 1315611 | C>T (p.Gln>STOP) | 0.0208 | 0.0119 | Ortholog(s) have thiol-dependent ubiquitin-specific protease activity, role in negative regulation of protein autoubiquitination, protein deubiquitination and cytoplasm, endoplasmic reticulum localization |
| PUF3 | C4_05370W_A | Chr4B:1172318 | T>C (p.Gly>Gly) | 0.0304 | 0.0304 | RNA-binding protein involved in the regulation of mitochondrial biogenesis |
| Uncharacterized region | ChrRA:655094 | A>G | 0.0405 | 0.0082 | – | |
| Uncharacterized region | ChrRA:655095 | C>G | 0.0336 | 0.0031 | – | |
| Insertions and Deletions | ||||||
| Uncharacterized region | Chr1A:837098 | -AAAAGAAAAG | 0.0064 | 0.0428 | – | |
| Uncharacterized region | Chr2A:254001 | -TAGAAGAAGAAT | 0.0027 | 0.0345 | – | |
| Uncharacterized region | Chr2A:342691 | +C | 0.0028 | 0.0257 | – | |
| Uncharacterized region | Chr2A:239531 | -T | 0.0105 | 0.0337 | – | |
| Uncharacterized region | Chr2A:1421683 | +GATATTTAAGG | 0.0153 | 0.0256 | – | |
| Uncharacterized region | Chr3A:980388 | +T | 0.0338 | 0.0291 | – | |
| Uncharacterized region | Chr4A:686524 | -T | 0.0373 | 0.0346 | – | |
| Uncharacterized region | Chr4B:456519 | +TAAAT | 0.0185 | 0.0285 | – | |
| − | C5_03980W_A | Chr5B: 878800 | +CAACAACAA | 0.0403 | 0.0364 | Protein of unknown function; Spider biofilm induced |
| − | C7_01080C_A | Chr7A:228708 | -TC | 0.0182 | 0.0231 | Major repeat sequence (MRS) on Chromosome 7; MRS units are composed of variable numbers of RPS units flanked by HOK and RB2 sequences; found on most chromosomes; may serve as recombination hot spot |
| – | C7_02250W_A | Chr7B:490089 | +TTCCAA | 0.0022 | 0.0208 | Ortholog of C. dubliniensis CD36: Cd36_72050, C. parapsilosis CDC317: CPAR2_301140, C. tenuis NRRL Y-1498: CANTEDRAFT_ 135055, and Debaryomyces hansenii CBS767: DEHA2E07678g |
| Carbohydrate | Carbohydrate Symbol | Before Plasma Treatment | After 12 × Plasma Treatment |
|---|---|---|---|
| D-glucose | GLU | + | + |
| glycerol | GLY | + | + |
| 2-keto-D-gluconate | 2KG | + | + |
| L-arabinose | ARA | − | − |
| D-xylose | XYL | + | + |
| adonitol | ADO | + | + |
| xylitol | XLT | + | + |
| D-galactose | GAL | + | + |
| inositol | INO | − | − |
| D-sorbitol | SOR | + | + |
| methyl-αD-glucopyranoside | MDG | + | + |
| N-acetyl-D-glucosamine | NAG | + | + |
| D-cellobiose | CEL | − | − |
| D-lactose | LAC | − | − |
| D-maltose | MAL | + | + |
| D-sucrose | SAC | + | + |
| D-trehalose | TRE | + | + |
| D-melesitose | MLZ | − | − |
| D-raffinose | RAF | − | − |
| No. | Enzyme | Wavelength [nm] |
|---|---|---|
| e2 | Alkaline phosphatase | 537 |
| e3 | Esterase (C4) | 537 |
| e4 | Esterase Lipase (C8) | 537 |
| e5 | Lipase (C14) | 537 |
| e6 | Leucine arylamidase | 494 |
| e7 | Valine arylamidase | 494 |
| e8 | Cystine arylamidase | 494 |
| e9 | Trypsin | 494 |
| e10 | α-Chymotrypsin | 494 |
| e11 | Acid phosphatase | 537 |
| e12 | Naphthol-AS-BI-phosphohydrolase | 582 |
| e13 | α-Galactosidase | 547 |
| e14 | β-Galactosidase | 547 |
| e15 | β-Glucuronidase | 582 |
| e16 | α-Glucosidase | 537 |
| e17 | β-Glucosidase | 537 |
| e18 | N-Acetyl-β-glucosaminidase | 454 |
| e19 | α-Mannosidase | 537 |
| e20 | β-Fucosidase | 537 |
| Antifungal Agent | Before Plasma Treatment [MIC Value] | After 12 × Plasma Treatment [MIC Value] |
|---|---|---|
| Voriconazole | 0.094 | 0.094 |
| Fluconazole | 2.0 | 2.0 |
| Caspofungin | 0.125 | 0.125 |
| Amphotericin B | 0.125 | 0.125 |
| Micafungin | 0.012 | 0.012 |
| Anidulafungin | 0.004 | 0.004 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tyczkowska-Sieroń, E.; Kałużewski, T.; Grabiec, M.; Kałużewski, B.; Tyczkowski, J. Genotypic and Phenotypic Changes in Candida albicans as a Result of Cold Plasma Treatment. Int. J. Mol. Sci. 2020, 21, 8100. https://doi.org/10.3390/ijms21218100
Tyczkowska-Sieroń E, Kałużewski T, Grabiec M, Kałużewski B, Tyczkowski J. Genotypic and Phenotypic Changes in Candida albicans as a Result of Cold Plasma Treatment. International Journal of Molecular Sciences. 2020; 21(21):8100. https://doi.org/10.3390/ijms21218100
Chicago/Turabian StyleTyczkowska-Sieroń, Ewa, Tadeusz Kałużewski, Magdalena Grabiec, Bogdan Kałużewski, and Jacek Tyczkowski. 2020. "Genotypic and Phenotypic Changes in Candida albicans as a Result of Cold Plasma Treatment" International Journal of Molecular Sciences 21, no. 21: 8100. https://doi.org/10.3390/ijms21218100