Revealing the Roles of the JAZ Family in Defense Signaling and the Agarwood Formation Process in Aquilaria sinensis
Abstract
:1. Introduction
2. Results
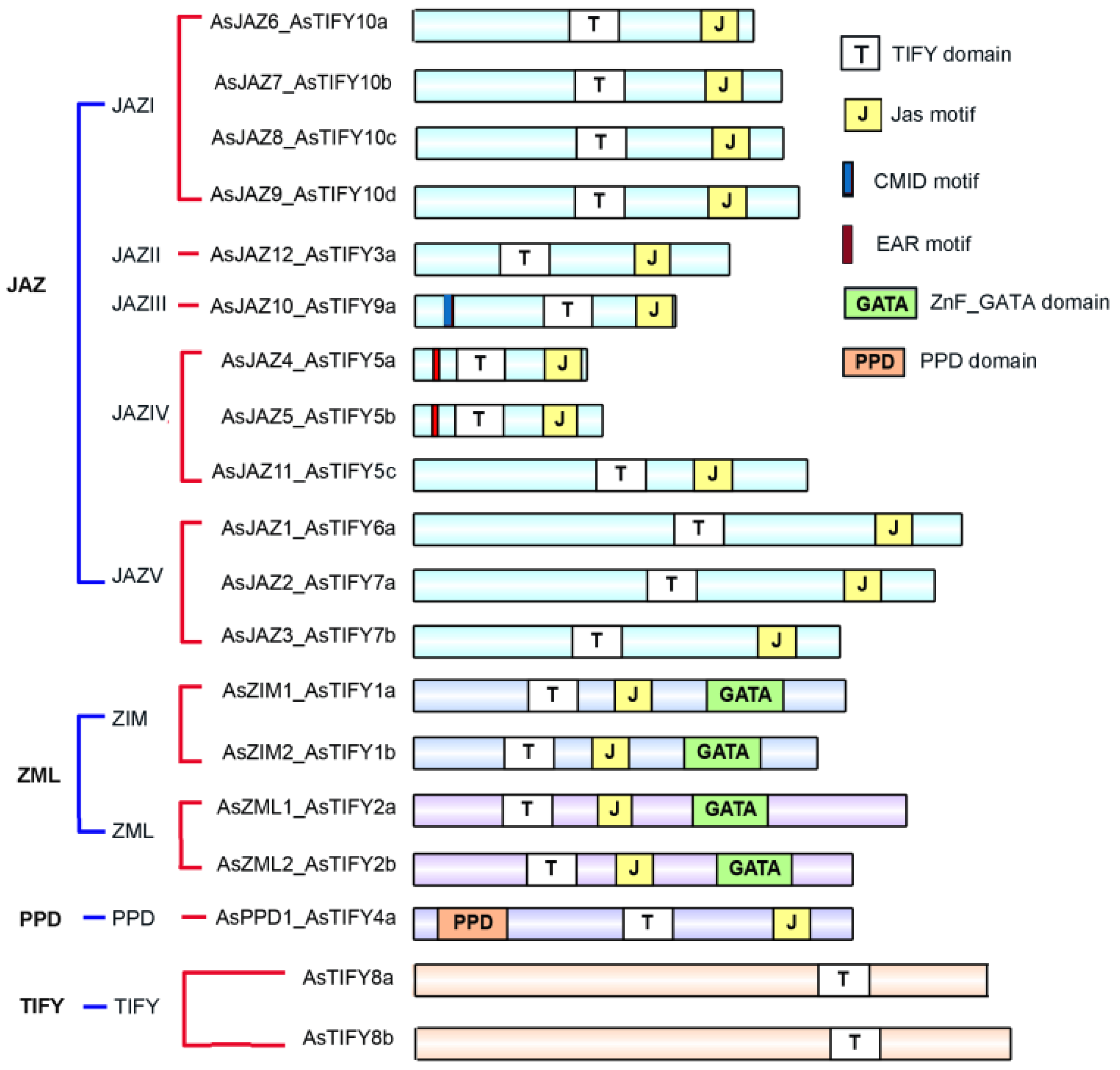
2.1. Phylogenetic Relationships and Major Domains of A. sinensis TIFY Family Members
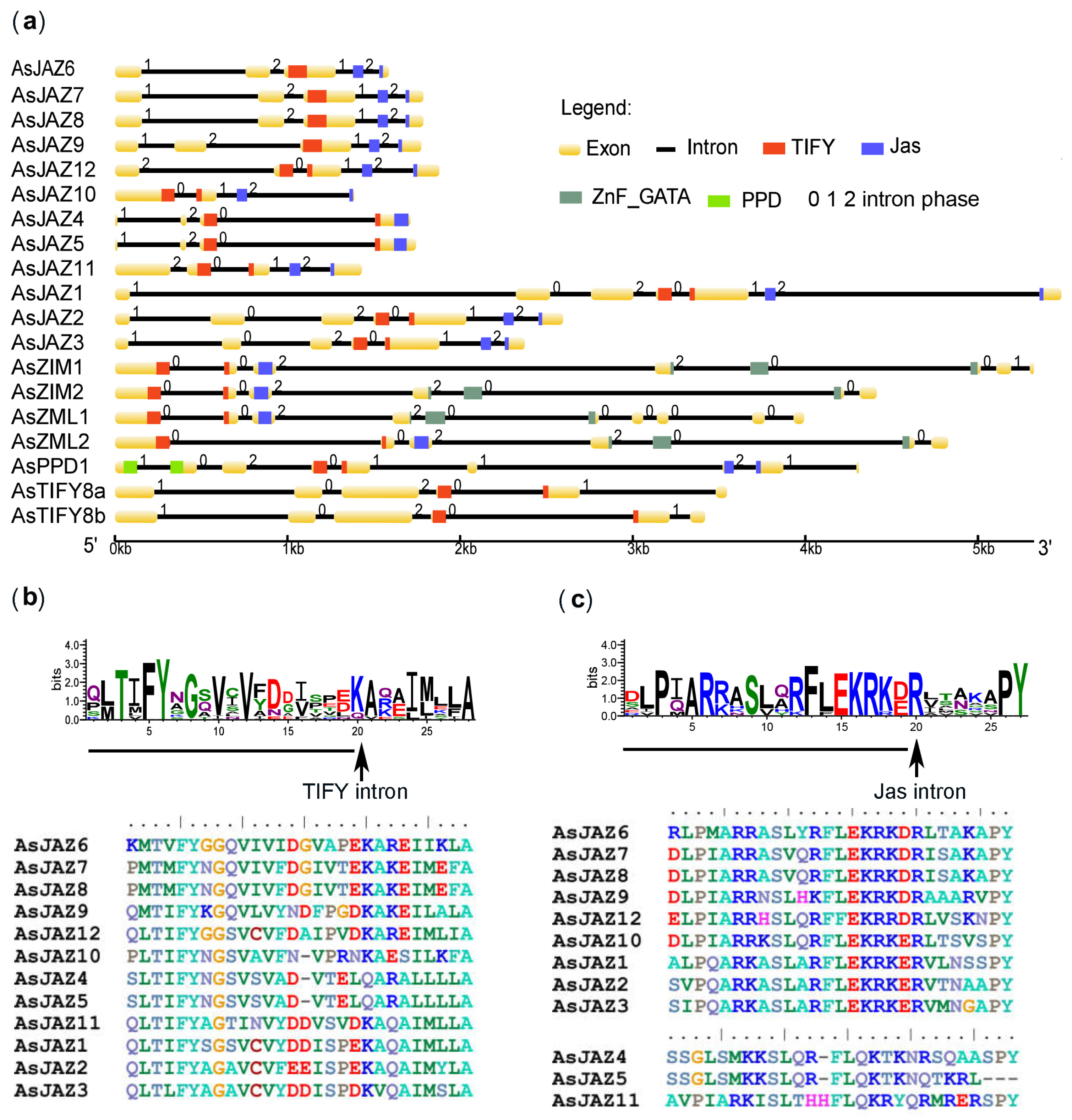
2.2. Locations and Structures of the AsTIFY Genes and Promoter Elements of the AsJAZ Genes
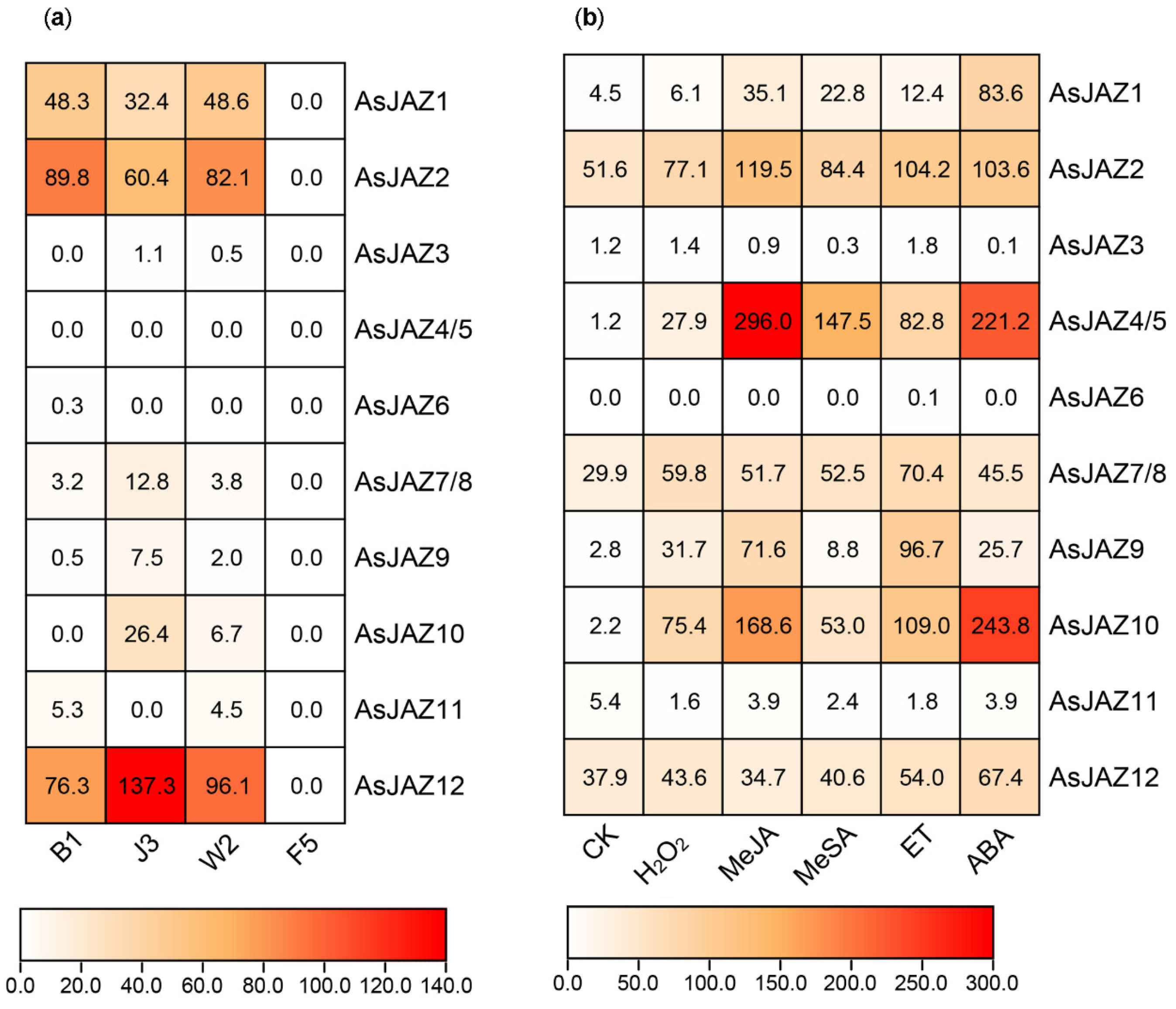
2.3. Expression Patterns of the AsJAZ Genes
2.4. Chromosome Location of AsWRKY Genes and Phylogenetic Analysis of AsWRKY Proteins
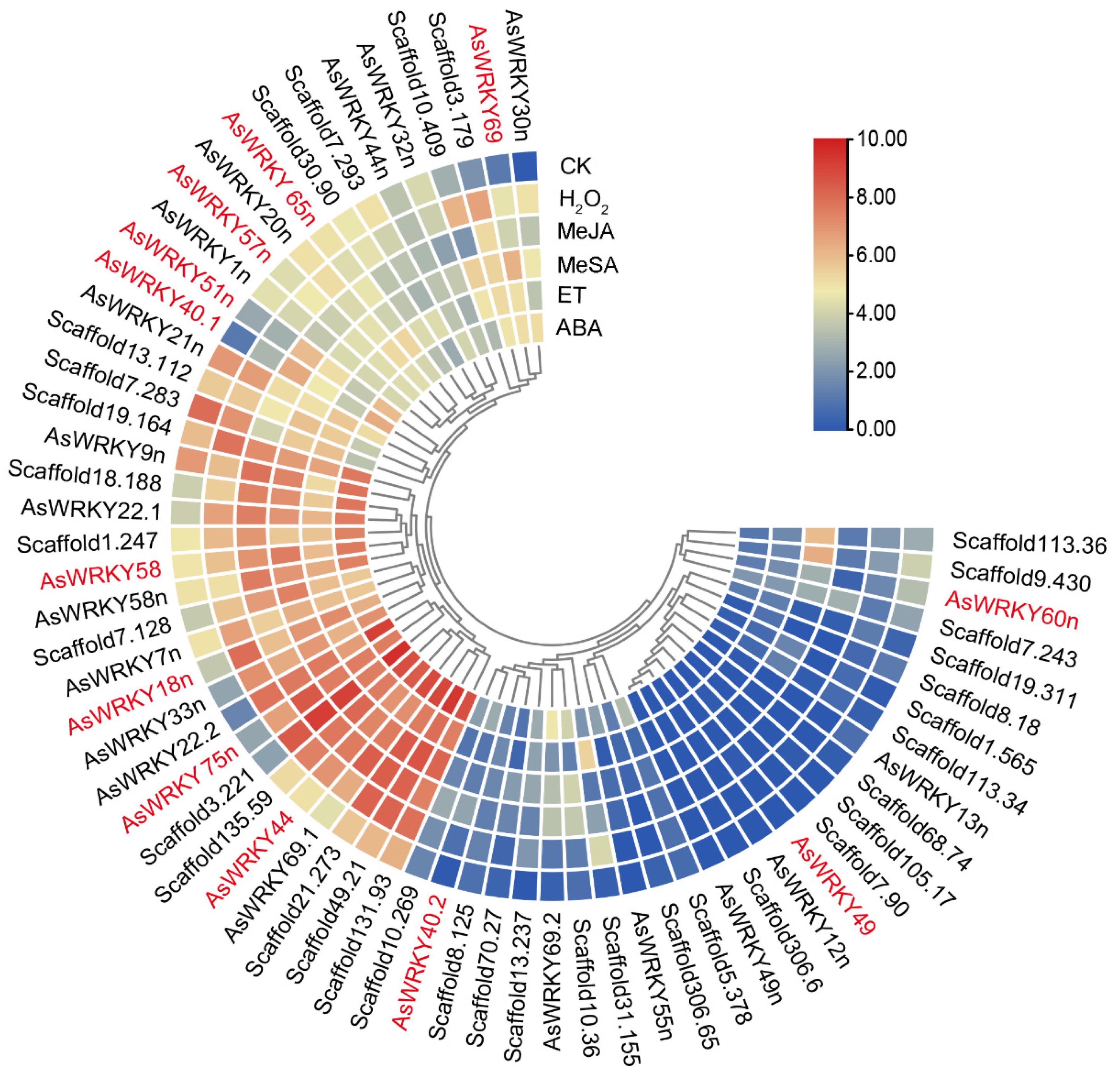
2.5. Expression Patterns of AsWRKY Genes
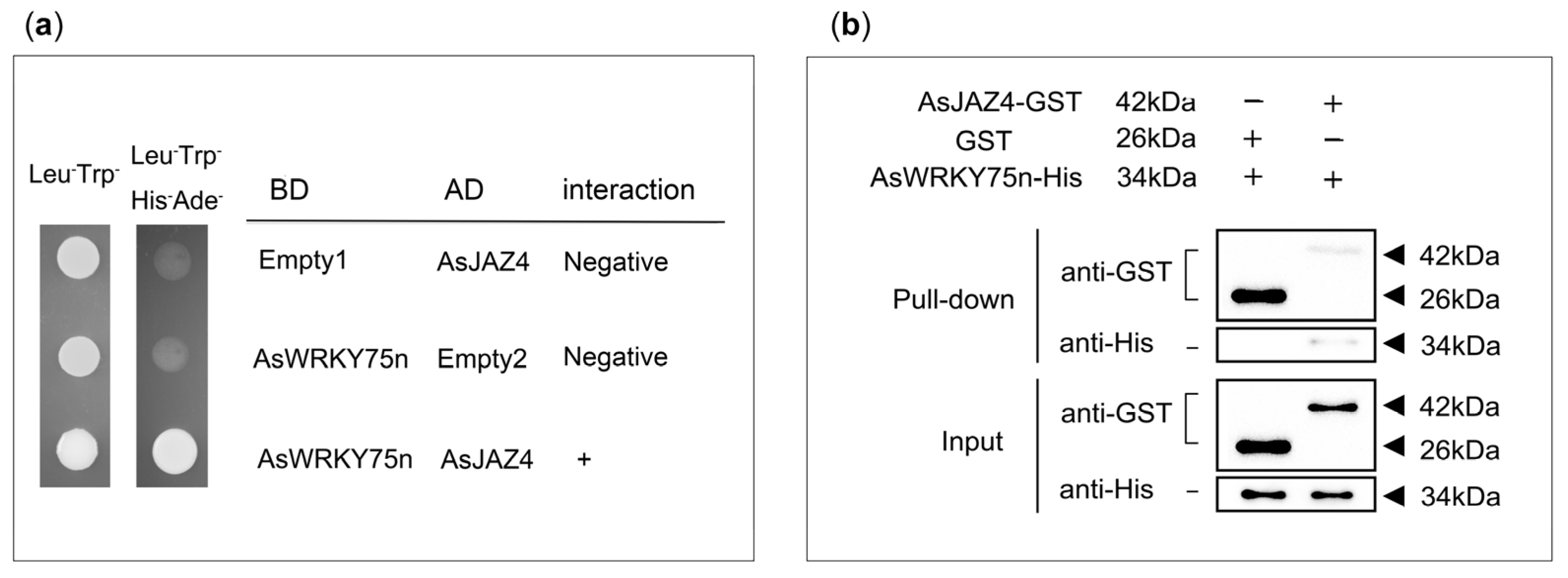
2.6. Experimental Validation of the Interaction between AsJAZ4 and AsWRKY75n
3. Discussion
3.1. Classification of the AsJAZ Family Members
3.2. Expression Patterns of the AsJAZ Family Members
3.3. Classification and Expression Patterns of the AsWRKY Genes
3.4. Possible Roles of the AsJAZ4/WRKY75n Complex
4. Materials and Methods
4.1. Bioinformatic Analysis of TIFY Family Members in A. sinensis
4.2. Bioinformatic Analysis of WRKY Family Members
4.3. Sequence Alignment and Phylogenetic Analysis
4.4. Plant Sample Preparation
4.5. Real-Time Quantitative PCR
4.6. Statistical Analysis
4.7. Transcriptomic Analysis and Heatmap Illustrating
4.8. Protein Expression and Pull-Down Assay
4.9. Yeast Two-Hybrid
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wasternack, C.; Strnad, M. Jasmonates: News on occurrence, biosynthesis, metabolism and action of an ancient group of signaling compounds. Int. J. Mol. Sci. 2018, 19, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vanholme, B.; Grunewald, W.; Bateman, A.; Kohchi, T.; Gheysen, G. The tify family previously known as ZIM. Trends Plant Sci. 2007, 12, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Howe, G.A.; Major, I.T.; Koo, A.J. Modularity in jasmonate signaling for multistress resilience. Annu. Rev. Plant Biol. 2018, 69, 387–415. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kazan, K.; Manners, J.M. JAZ repressors and the orchestration of phytohormone crosstalk. Trends Plant Sci. 2012, 17, 22–31. [Google Scholar] [CrossRef]
- Qi, T.C.; Huang, H.; Wu, D.W.; Yan, J.B.; Qi, Y.J.; Song, S.S.; Xie, D.X. Arabidopsis DELLA and JAZ proteins bind the WD-repeat/bHLH/MYB complex to modulate gibberellin and jasmonate signaling synergy. Plant Cell 2014, 26, 1118–1133. [Google Scholar] [CrossRef] [Green Version]
- Xie, Y.; Tan, H.J.; Ma, Z.X.; Huang, J.R. DELLA proteins promote anthocyanin biosynthesis via sequestering MYBL2 and JAZ suppressors of the MYB/bHLH/WD40 complex in Arabidopsis thaliana. Mol. Plant 2016, 9, 711–721. [Google Scholar] [CrossRef] [Green Version]
- Li, T.; Xu, Y.X.; Zhang, L.Z.; Ji, Y.L.; Tan, D.M.; Yuan, H.; Wang, A.D. The Jasmonate-activated transcription factor MdMYC2 regulates ETHYLENE RESPONSE FACTOR and ethylene biosynthetic genes to promote ethylene biosynthesis during apple fruit ripening. Plant Cell 2017, 29, 1316–1334. [Google Scholar] [CrossRef] [Green Version]
- Ju, L.; Jing, Y.X.; Shi, P.T.; Liu, J.; Chen, J.S.; Yan, J.J.; Chu, J.F.; Chen, K.M.; Sun, J.Q. JAZ proteins modulate seed germination through interaction with ABI5 in bread wheat and Arabidopsis. New Phytol. 2019, 223, 246–260. [Google Scholar] [CrossRef]
- Liu, H.; Timko, M.P. Jasmonic acid signaling and molecular crosstalk with other phytohormones. Int. J. Mol. Sci. 2021, 22, 2914. [Google Scholar] [CrossRef]
- Chini, A.; Fonseca, S.; Fernandez, G.; Adie, B.; Chico, J.M.; Lorenzo, O.; Garcia-Casado, G.; Lopez-Vidriero, I.; Lozano, F.M.; Ponce, M.R.; et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 2007, 448, 666–671. [Google Scholar] [CrossRef]
- Thines, B.; Katsir, L.; Melotto, M.; Niu, Y.J.; Mandaokar, A.; Liu, G.H.; Nomura, K.; He, S.Y.; Howe, G.A.; Browse, J. JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signalling. Nature 2007, 448, 661–665. [Google Scholar] [CrossRef] [PubMed]
- Thireault, C.; Shyu, C.; Yoshida, Y.; St Aubin, B.; Campos, M.L.; Howe, G.A. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. Plant J. 2015, 82, 669–679. [Google Scholar] [CrossRef]
- Ye, H.Y.; Du, H.; Tang, N.; Li, X.H.; Xiong, L.Z. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol. Biol. 2009, 71, 291–305. [Google Scholar] [CrossRef]
- Wang, Y.K.; Qiao, L.Y.; Bai, J.F.; Wang, P.; Duan, W.J.; Yuan, S.H.; Yuan, G.L.; Zhang, F.T.; Zhang, L.P.; Zhao, C.P. Genome-wide characterization of JASMONATE-ZIM DOMAIN transcription repressors in wheat (Triticum aestivum L.). BMC Genom. 2017, 18, 152. [Google Scholar] [CrossRef] [Green Version]
- Ebel, C.; BenFeki, A.; Hanin, M.; Solano, R.; Chini, A. Characterization of wheat (Triticum aestivum) TIFY family and role of Triticum durum TdTIFY11a in salt stress tolerance. PLoS ONE 2018, 13, e0200566. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chini, A.; Ben-Romdhane, W.; Hassairi, A.; Aboul-Soud, M.A.M. Identification of TIFY/JAZ family genes in Solanum lycopersicum and their regulation in response to abiotic stresses. PLoS ONE 2017, 12, e0177381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, J.Z.; Zou, Z.W.; Xing, H.Q.; Duan, Y.; Zhu, X.J.; Ma, Y.C.; Wang, Y.H.; Fang, W.P. Genome-wide analysis reveals stress and hormone responsive patterns of JAZ family genes in Camellia sinensis. Int. J. Mol. Sci. 2020, 21, 2433. [Google Scholar] [CrossRef] [Green Version]
- Zheng, Y.C.; Chen, X.J.; Wang, P.J.; Sun, Y.; Yue, C.; Ye, N.X. Genome-wide and expression pattern analysis of JAZ family involved in stress responses and postharvest processing treatments in Camellia sinensis. Sci. Rep. 2020, 10, 2792. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.C.; Gao, M.; Singer, S.D.; Fei, Z.J.; Wang, H.; Wang, X.P. Genome-wide identification and analysis of the TIFY gene family in grape. PLoS ONE 2012, 7, e44465. [Google Scholar] [CrossRef] [Green Version]
- Chini, A.; Gimenez-Ibanez, S.; Goossens, A.; Solano, R. Redundancy and specificity in jasmonate signalling. Curr. Opin. Plant Biol. 2016, 33, 147–156. [Google Scholar] [CrossRef]
- Fonseca, S.; Chico, J.M.; Solano, R. The jasmonate pathway: The ligand, the receptor and the core signalling module. Curr. Opin. Plant Biol. 2009, 12, 539–547. [Google Scholar] [CrossRef] [PubMed]
- Wager, A.; Browse, J. Social network: JAZ protein interactions expand our knowledge of Jasmonate signaling. Front. Plant Sci. 2012, 3, 41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takeuchi, J.; Fukui, K.; Seto, Y.; Takaoka, Y.; Okamoto, M. Ligand-receptor interactions in plant hormone signaling. Plant J. 2021, 105, 290–306. [Google Scholar] [CrossRef] [PubMed]
- Chini, A.; Fonseca, S.; Chico, J.M.; Fernandez-Calvo, P.; Solano, R. The ZIM domain mediates homo- and heteromeric interactions between Arabidopsis JAZ proteins. Plant J. 2009, 59, 77–87. [Google Scholar] [CrossRef]
- Chung, H.S.; Howe, G.A. A critical role for the TIFY motif in repression of jasmonate signaling by a stabilized splice variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis. Plant Cell 2009, 21, 131–145. [Google Scholar] [CrossRef] [Green Version]
- Pauwels, L.; Barbero, G.F.; Geerinck, J.; Tilleman, S.; Grunewald, W.; Perez, A.C.; Chico, J.M.; Bossche, R.V.; Sewell, J.; Gil, E.; et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 2010, 464, 788–791. [Google Scholar] [CrossRef] [Green Version]
- Pauwels, L.; Goossens, A. The JAZ proteins: A crucial interface in the jasmonate signaling cascade. Plant Cell 2011, 23, 3089–3100. [Google Scholar] [CrossRef] [Green Version]
- Melotto, M.; Mecey, C.; Niu, Y.; Chung, H.S.; Katsir, L.; Yao, J.; Zeng, W.P.; Thines, B.; Staswick, P.; Broese, J.; et al. A critical role of two positively charged amino acids in the Jas motif of Arabidopsis JAZ proteins in mediating coronatine- and jasmonoyl isoleucine-dependent interactions with the COI1 F-box protein. Plant J. 2008, 55, 979–988. [Google Scholar] [CrossRef] [Green Version]
- Zhang, F.; Yao, J.; Ke, J.; Zhang, L.; Lam, V.Q.; Xin, X.-F.; Zhou, X.E.; Chen, J.; Brunzelle, J.; Griffin, P.R.; et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 2015, 525, 269–273. [Google Scholar] [CrossRef] [Green Version]
- Dombrecht, B.; Xue, G.P.; Sprague, S.J.; Kirkegaard, J.A.; Ross, J.J.; Reid, J.B.; Fitt, G.P.; Sewelam, N.; Schenk, P.M.; Manners, J.M.; et al. MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 2007, 19, 2225–2245. [Google Scholar] [CrossRef] [Green Version]
- Fernández-Calvo, P.; Chini, A.; Fernández-Barbero, G.; Chico, J.-M.; Gimenez-Ibanez, S.; Geerinck, J.; Eeckhout, D.; Schweizer, F.; Godoy, M.; Franco-Zorrilla, J.M.; et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 2011, 23, 701–715. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.P.; Li, Y.; Pan, J.J.; Lou, D.J.; Hu, Y.R.; Yu, D.Q. The bHLH transcription factors MYC2, MYC3, and MYC4 are required for jasmonate-mediated inhibition of flowering in Arabidopsis. Mol. Plant 2017, 10, 1461–1464. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, Y.J.; Liang, G.; Yang, S.Z.; Yu, D.Q. Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence. Plant Cell 2014, 26, 230–245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, C.; Fan, M.; Yang, M.; Zhao, J.P.; Zhang, W.H.; Su, Y.; Xiao, L.T.; Deng, H.T.; Xie, D.X. Injury activates Ca2+/calmodulin-dependent phosphorylation of JAV1-JAZ8-WRKY51 complex for jasmonate biosynthesis. Mol. Cell 2018, 70, 136–149.e137. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.G.; Zhang, L.P.; Xiang, S.Y.; Chen, Y.L.; Zhang, H.Y.; Yu, D.Q. The transcription factor WRKY75 positively regulates jasmonate-mediated plant defense to necrotrophic fungal pathogens. J. Exp. Bot. 2021, 72, 1473–1489. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.X.; Yan, Q.; Gan, S.P.; Xue, D.; Wang, H.T.; Xing, H.; Zhao, J.M. GmWRKY40, a member of the WRKY transcription factor genes identified from Glycine max L., enhanced the resistance to Phytophthora sojae. BMC Plant Biol. 2019, 19, 598. [Google Scholar] [CrossRef] [Green Version]
- Pandey, S.P.; Roccaro, M.; Schon, M.; Logemann, E.; Somssich, I.E. Transcriptional reprogramming regulated by WRKY18 and WRKY40 facilitates powdery mildew infection of Arabidopsis. Plant J. 2010, 64, 912–923. [Google Scholar] [CrossRef]
- Jiang, Y.J.; Yu, D.Q. The WRKY57 transcription factor affects the expression of Jasmonate ZIM-Domain genes transcriptionally to compromise Botrytis cinerea resistance. Plant Physiol. 2016, 171, 2771–2782. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.; Mei, W.; Lin, Q.; Wang, H.; Wang, J.; Peng, S.; Li, H.; Zhu, J.; Li, W.; Wang, P.; et al. Genome sequence of the agarwood tree Aquilaria sinensis (Lour.) Spreng: The first chromosome-level draft genome in the Thymelaeceae family. Gigascience 2020, 9, giaa013. [Google Scholar] [CrossRef] [Green Version]
- Tan, C.S.; Isa, N.M.; Ismail, I.; Zainal, Z. Agarwood induction: Current developments and future perspectives. Front. Plant Sci. 2019, 10, 122. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Yang, Y.; Wei, J.H.; Meng, H.; Sui, C.; Chen, H.Q. Advances in studies on mechanism of agarwood formation in Aquilaria sinensis and its hypothesis of agarwood formation induced by defense response. Chin. Tradit. Herb. Drugs 2010, 41, 156–159. [Google Scholar]
- Schweizer, F.; Fernandez-Calvo, P.; Zander, M.; Diez-Diaz, M.; Fonseca, S.; Glauser, G.; Lewsey, M.G.; Ecker, J.R.; Solano, R.; Reymond, P. Arabidopsis basic helix-loop-helix transcription factors MYC2, MYC3, and MYC4 regulate glucosinolate biosynthesis, insect performance, and feeding behavior. Plant Cell 2013, 25, 3117–3132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, M.L.; Memelink, J. Jasmonate-responsive transcription factors regulating plant secondary metabolism. Biotechnol. Adv. 2016, 34, 441–449. [Google Scholar] [CrossRef]
- Mao, Y.B.; Liu, Y.Q.; Chen, D.Y.; Chen, F.Y.; Fang, X.; Hong, G.J.; Wang, J.W.; Chen, X.Y. Jasmonate response decay and defense metabolite accumulation contributes to age-regulated dynamics of plant insect resistance. Nat. Commun. 2017, 8, 13925. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schuman, M.C.; Meldau, S.; Gaquerel, E.; Diezel, C.; McGale, E.; Greenfield, S.; Baldwin, I.T. The active jasmonate JA-Ile regulates a specific subset of plant jasmonate-mediated resistance to herbivores in nature. Front. Plant Sci. 2018, 9, 787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.H.; Liao, Y.C.; Zhang, Z.; Liu, J.; Sun, P.W.; Gao, Z.H.; Sui, C.; Wei, J.H. Jasmonic acid is a crucial signal transducer in heat shock induced sesquiterpene formation in Aquilaria sinensis. Sci. Rep. 2016, 6, 21843. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.H.; Zhang, Z.; Wang, M.X.; Wei, J.H.; Chen, H.J.; Gao, Z.H.; Sui, C.; Luo, H.M.; Zhang, X.L.; Yang, Y.; et al. Identification of genes related to agarwood formation: Transcriptome analysis of healthy and wounded tissues of Aquilaria sinensis. BMC Genom. 2013, 14, 227. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.H.; Liao, Y.C.; Lv, F.F.; Zhang, Z.; Sun, P.W.; Gao, Z.H.; Hu, K.P.; Sui, C.; Jin, Y.; Wei, J.H. Transcription factor AsMYC2 controls the Jasmonate-responsive expression of ASS1 regulating sesquiterpene biosynthesis in Aquilaria sinensis (Lour.) Gilg. Plant Cell Physiol. 2017, 58, 2257. [Google Scholar] [CrossRef] [Green Version]
- Sun, P.W.; Xu, Y.H.; Yu, C.C.; Lv, F.F.; Tang, X.L.; Gao, Z.H.; Zhang, Z.; Wang, H.; Liu, Y.; Wei, J.H. WRKY44 represses expression of the wound-induced sesquiterpene biosynthetic gene ASS1 in Aquilaria sinensis. J. Exp. Bot. 2020, 71, 1128–1138. [Google Scholar] [CrossRef]
- Ye, W.; Wu, H.Q.; He, X.; Wang, L.; Zhang, W.M.; Li, H.H.; Fan, Y.F.; Tan, G.H.; Liu, T.M.; Gao, X.X. Transcriptome sequencing of chemically induced Aquilaria sinensis to identify genes related to agarwood formation. PLoS ONE 2016, 11, e0155505. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.H.; Sun, P.W.; Tang, X.L.; Gao, Z.H.; Zhang, Z.; Wei, J.H. Genome-wide analysis of WRKY transcription factors in Aquilaria sinensis (Lour.) Gilg. Sci. Rep. 2020, 10, 3018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, H.S.; Cooke, T.F.; Depew, C.L.; Patel, L.C.; Ogawa, N.; Kobayashi, Y.; Howe, G.A. Alternative splicing expands the repertoire of dominant JAZ repressors of jasmonate signaling. Plant J. 2010, 63, 613–622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef] [Green Version]
- Shyu, C.; Figueroa, P.; Depew, C.L.; Cooke, T.F.; Sheard, L.B.; Moreno, J.E.; Katsir, L.; Zheng, N.; Browse, J.; Howe, G.A. JAZ8 lacks a canonical degron and has an EAR motif that mediates transcriptional repression of jasmonate responses in Arabidopsis. Plant Cell 2012, 24, 536–550. [Google Scholar] [CrossRef] [Green Version]
- Ke, J.Y.; Ma, H.L.; Gu, X.; Thelen, A.; Brunzelle, J.S.; Li, J.Y.; Xu, H.E.; Melcher, K. Structural basis for recognition of diverse transcriptional repressors by the TOPLESS family of corepressors. Sci. Adv. 2015, 1, e1500107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moreno, J.E.; Shyu, C.; Campos, M.L.; Patel, L.C.; Chung, H.S.; Yao, J.; He, S.Y.; Howe, G.A. Negative feedback control of jasmonate signaling by an alternative splice variant of JAZ10. Plant Physiol. 2013, 162, 1006–1017. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, F.; Ke, J.Y.; Zhang, L.; Chen, R.Z.; Sugimoto, K.; Howe, G.A.; Xu, H.E.; Zhou, M.G.; He, S.Y.; Melcher, K. Structural insights into alternative splicing-mediated desensitization of jasmonate signaling. Proc. Natl. Acad. Sci. USA 2017, 114, 1720–1725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, Y.H.; Meng, Y.J.; Huang, D.L.; Qi, Y.H.; Chen, M. Origin and evolutionary analysis of the plant-specific TIFY transcription factor family. Genomics 2011, 98, 128–136. [Google Scholar] [CrossRef] [Green Version]
- Sheard, L.B.; Tan, X.; Mao, H.; Withers, J.; Ben-Nissan, G.; Hinds, T.R.; Kobayashi, Y.; Hsu, F.-F.; Sharon, M.; Browse, J.; et al. Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor. Nature 2010, 468, 400–405. [Google Scholar] [CrossRef] [Green Version]
- Li, H.L.; Guo, D.; Yang, Z.P.; Tang, X.; Peng, S.Q. Genome-wide identification and characterization of WRKY gene family in Hevea brasiliensis. Genomics 2014, 104, 14–23. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, W.; Fang, L.C.; Sun, X.M.; Su, L.Y.; Liang, Z.C.; Wang, N.; Londo, J.P.; Li, S.H.; Xin, H.P. Genome-wide identification of WRKY family genes and their response to cold stress in Vitis vinifera. BMC Plant Biol. 2014, 14, 103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, C.M.; Teixeira da Silva, J.A.; Tan, J.W.; Zhang, J.X.; Pan, X.P.; Li, M.Z.; Luo, J.P.; Duan, J. A genome-wide identification of the WRKY family genes and a survey of potential WRKY target genes in Dendrobium officinale. Sci. Rep. 2017, 7, 9200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qu, R.J.; Cao, Y.W.; Tang, X.Q.; Sun, L.Q.; Wei, L.; Wang, K.C. Identification and expression analysis of the WRKY gene family in Isatis indigotica. Gene 2021, 783, 145561. [Google Scholar] [CrossRef]
- Ling, J.; Jiang, W.J.; Zhang, Y.; Yu, H.J.; Mao, Z.C.; Gu, X.F.; Huang, S.W.; Xie, B.Y. Genome-wide analysis of WRKY gene family in Cucumis sativus. BMC Genom. 2011, 12, 471. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Brader, G.; Palva, E.T. The WRKY70 transcription factor: A node of convergence for jasmonate-mediated and salicylate-mediated signals in plant defense. Plant Cell 2004, 16, 319–331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, X.P.; Chen, C.H.; Fan, B.F.; Chen, Z.X. Physical and functional interactions between pathogen-induced Arabidopsis WRKY18, WRKY40, and WRKY60 transcription factors. Plant Cell 2006, 18, 1310–1326. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Pan, R.; Wei, J.; Lv, F.F. AsJAZ1 represses the expression of the sesquiterpene synthase gene based on the JA signaling pathway in Aquilaria sinensis (Lour.) Gilg. Plant Biotechnol. Rep. 2023, 17, 101–109. [Google Scholar] [CrossRef]
- Xu, Y.H.; Wang, J.W.; Wang, S.; Wang, J.Y.; Chen, X.Y. Characterization of GaWRKY1, a cotton transcription factor that regulates the sesquiterpene synthase gene (+)-δ-cadinene synthase-A. Plant Physiol. 2004, 135, 507–515. [Google Scholar] [CrossRef] [Green Version]
- Ma, D.M.; Pu, G.B.; Lei, C.Y.; Ma, L.Q.; Wang, H.H.; Guo, Y.W.; Chen, J.L.; Du, Z.G.; Wang, H.; Li, G.F.; et al. Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the amorpha-4,11-diene synthase gene, a key gene of artemisinin biosynthesis. Plant Cell Physiol. 2009, 50, 2146–2161. [Google Scholar] [CrossRef] [Green Version]
- Han, J.L.; Wang, H.Z.; Lundgren, A.; Brodelius, P.E. Effects of overexpression of AaWRKY1 on artemisinin biosynthesis in transgenic Artemisia annua plants. Phytochemistry 2014, 102, 89–96. [Google Scholar] [CrossRef]
- Zheng, Z.Y.; Qamar, S.A.; Chen, Z.X.; Mengiste, T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 2006, 48, 592–605. [Google Scholar] [CrossRef] [PubMed]
- Lai, Z.B.; Li, Y.; Wang, F.; Cheng, Y.; Fan, B.F.; Yu, J.Q.; Chen, Z.X. Arabidopsis sigma factor binding proteins are activators of the WRKY33 transcription factor in plant defense. Plant Cell 2011, 23, 3824–3841. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, P.R.; Li, Z.H.; Huang, P.X.; Li, B.S.; Fang, S.; Chu, J.F.; Guo, H.W. A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence. Plant Cell 2017, 29, 2854–2870. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Zhang, L.; Ji, Y.; Jing, Y.; Li, L.; Chen, Y.; Wang, R.; Zhang, H.; Yu, D.; Chen, L. Arabidopsis SIGMA FACTOR BINDING PROTEIN1 (SIB1) and SIB2 inhibit WRKY75 function in abscisic acid-mediated leaf senescence and seed germination. J. Exp. Bot. 2022, 73, 182–196. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Chen, L.; Yu, D. Transcription factor WRKY75 interacts with DELLA proteins to affect flowering. Plant Physiol. 2018, 176, 790–803. [Google Scholar] [CrossRef] [Green Version]
- Lu, K.K.; Song, R.F.; Guo, J.X.; Zhang, Y.; Zuo, J.X.; Chen, H.H.; Liao, C.Y.; Hu, X.Y.; Ren, F.; Lu, Y.T.; et al. CycC1; 1-WRKY75 complex-mediated transcriptional regulation of SOS1 controls salt stress tolerance in Arabidopsis. Plant Cell 2023. ahead of print. [Google Scholar] [CrossRef]
- Xu, H.; Luo, D.; Zhang, F. DcWRKY75 promotes ethylene induced petal senescence in carnation (Dianthus caryophyllus L.). Plant J. 2021, 108, 1473–1492. [Google Scholar] [CrossRef]
- Xu, H.; Wang, S.; Larkin, R.M.; Zhang, F. The transcription factors DcHB30 and DcWRKY75 antagonistically regulate ethylene-induced petal senescence in carnation (Dianthus caryophyllus). J. Exp. Bot. 2022, 73, 7326–7343. [Google Scholar] [CrossRef]
- Meng, X.D.; Ji, Y.Q. Modern computational techniques for the HMMER sequence analysis. ISRN Bioinform. 2013, 2013, 252183. [Google Scholar] [CrossRef]
- Liu, W.Z.; Xie, Y.B.; Ma, J.Y.; Luo, X.T.; Nie, P.; Zuo, Z.X.; Lahrmann, U.; Zhao, Q.; Zheng, Y.Y.; Zhao, Y.; et al. IBS: An illustrator for the presentation and visualization of biological sequences. Bioinformatics 2015, 31, 3359–3361. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.; Jin, J.P.; Guo, A.Y.; Zhang, H.; Luo, J.C.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rombauts, S.; Dehais, P.; Van Montagu, M.; Rouze, P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999, 27, 295–296. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rushton, P.J.; Somssich, I.E.; Ringler, P.; Shen, Q.J. WRKY transcription factors. Trends Plant Sci. 2010, 15, 247–258. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef]
- Liu, J.; Xu, Y.H.; Zhang, Z.; Wei, J.H. Hydrogen peroxide promotes programmed cell death and salicylic acid accumulation during the induced production of sesquiterpenes in cultured cell suspensions of Aquilaria sinensis. Funct. Plant Biol. 2015, 42, 337–346. [Google Scholar] [CrossRef]
- Gao, Z.H.; Wei, J.H.; Yang, Y.; Zhang, Z.; Zhao, W.T. Selection and validation of reference genes for studying stress-related agarwood formation of Aquilaria sinensis. Plant Cell Rep. 2012, 31, 1759–1768. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Chan, C.K. Analysis of RNA-Seq data using TopHat and Cufflinks. Methods Mol. Biol. 2016, 1374, 339–361. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| Name | Synonym | a Chr. | b CDS (bp) | Protein | f ID | ||
|---|---|---|---|---|---|---|---|
| c Length (aa.) | d pI | e MW (Da) | |||||
| AsJAZ1 | AsTIFY6a | 8 (−) | 1161 | 386 | 9.39 | 40,533.48 | Scaffold 331.1 |
| AsJAZ2 | AsTIFY7a | 4 (−) | 1104 | 367 | 8.92 | 38,335.54 | Scaffold 5.283 |
| AsJAZ3 | AsTIFY7b | 7 (−) | 903 | 300 | 9.28 | 31,788.02 | Scaffold 134.44 |
| AsJAZ4 | AsTIFY5a | 3 (−) | 375 | 124 | 9.75 | 13,913.83 | Scaffold 262.13 g |
| AsJAZ5 | AsTIFY5b | 3 (+) | 408 | 135 | 10.01 | 15,241.81 | Scaffold 166.1 h |
| AsJAZ6 | AsTIFY10a | 8 (−) | 717 | 238 | 9.58 | 25,777.5 | Scaffold 46.130 |
| AsJAZ7 | AsTIFY10b | 4 (+) | 777 | 258 | 9.02 | 27,571.31 | Scaffold 43.66 |
| AsJAZ8 | AsTIFY10c | 4 (−) | 777 | 258 | 8.82 | 27,518.26 | Scaffold 5.466 |
| AsJAZ9 | AsTIFY11a | 6 (−) | 813 | 270 | 7.73 | 29,085.46 | Scaffold 12.166 |
| AsJAZ10 | AsTIFY9a | 1 (−) | 552 | 183 | 8.46 | 20,269.01 | Scaffold 9.133 |
| AsJAZ11 | AsTIFY5c | 4 (−) | 834 | 277 | 9.68 | 30,599.78 | Scaffold 5.102 |
| AsJAZ12 | AsTIFY3a | 8 (−) | 666 | 221 | 6.61 | 23,444.4 | Scaffold 46.99 |
| AsPPD1 | AsTIFY4a | 3 (+) | 987 | 328 | 6.46 | 35,656.83 | Scaffold 22.90 |
| AsZIM1 | AsTIFY1a | 2 (+) | 915 | 304 | 6.17 | 32,875.36 | Scaffold 60.49 |
| AsZIM2 | AsTIFY1b | 2 (+) | 855 | 284 | 6.65 | 30,628.17 | Scaffold 2.660 |
| AsZML1 | AsTIFY2a | 2 (+) | 1041 | 346 | 4.94 | 38,095.15 | Scaffold 60.50 |
| AsZML2 | AsTIFY2b | 2 (−) | 930 | 309 | 5.73 | 32,843.19 | Scaffold 2.738 |
| AsTIFY8a | 8 (−) | 1212 | 403 | 9.66 | 42,728.3 | Scaffold 21.46 | |
| AsTIFY8b | 4 (−) | 1254 | 417 | 8.53 | 44,097.7 | Scaffold 23.90 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Y.; Ran, J.; Li, G.; Wang, M.; Yang, C.; Wen, X.; Geng, X.; Zhang, L.; Li, Y.; Zhang, Z. Revealing the Roles of the JAZ Family in Defense Signaling and the Agarwood Formation Process in Aquilaria sinensis. Int. J. Mol. Sci. 2023, 24, 9872. https://doi.org/10.3390/ijms24129872
Ma Y, Ran J, Li G, Wang M, Yang C, Wen X, Geng X, Zhang L, Li Y, Zhang Z. Revealing the Roles of the JAZ Family in Defense Signaling and the Agarwood Formation Process in Aquilaria sinensis. International Journal of Molecular Sciences. 2023; 24(12):9872. https://doi.org/10.3390/ijms24129872
Chicago/Turabian StyleMa, Yimian, Jiadong Ran, Guoqiong Li, Mengchen Wang, Chengmin Yang, Xin Wen, Xin Geng, Liping Zhang, Yuan Li, and Zheng Zhang. 2023. "Revealing the Roles of the JAZ Family in Defense Signaling and the Agarwood Formation Process in Aquilaria sinensis" International Journal of Molecular Sciences 24, no. 12: 9872. https://doi.org/10.3390/ijms24129872






