Increased H3K9me3 and F-Actin Reorganization in the Rapid Adaptive Response to Hypergravity in Human T Lymphocytes
Abstract
:1. Introduction
2. Results
2.1. Hypergravity Did Not Compromise Cell Viability
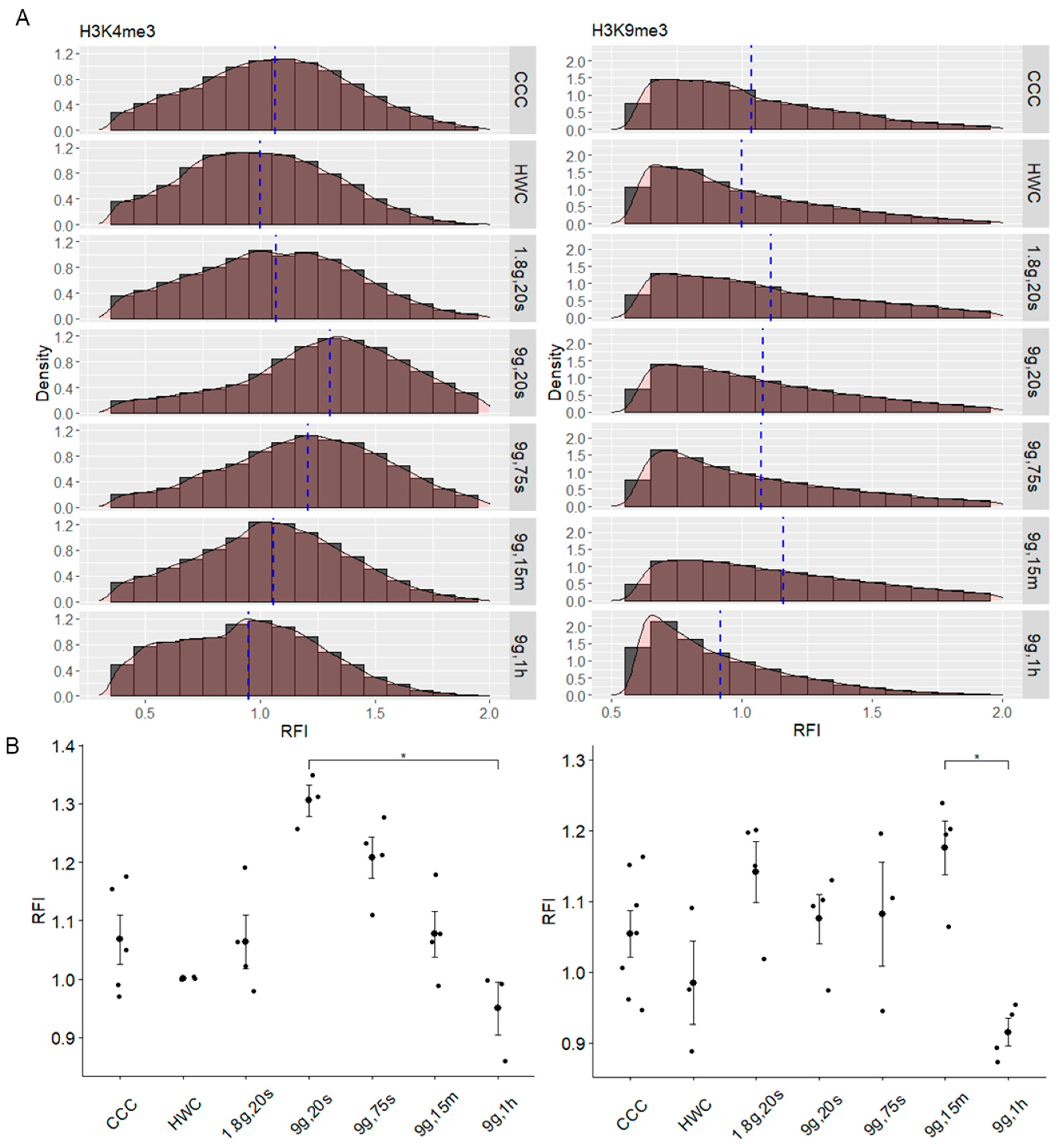
2.2. Hypergravity Only Minimally Affected H3K4me3 Histone Modification
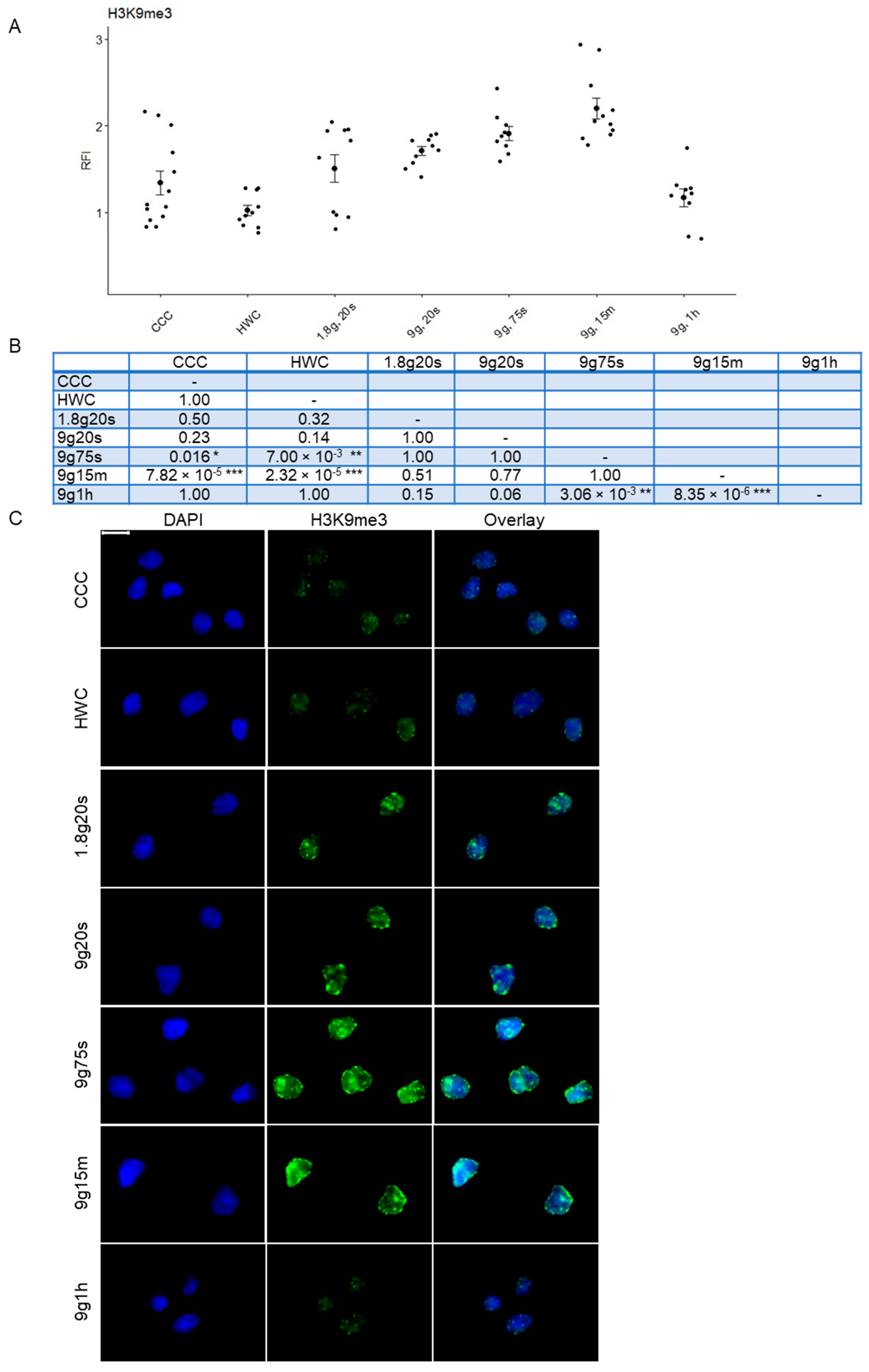
2.3. Dynamic Response of H3K9me3 Histone Modification in Hypergravity Conditions
2.4. Quantitative Analysis of Cytoskeletal Changes in Hypergravity
2.5. Dynamic Changes of Nuclear Morphology in Hypergravity
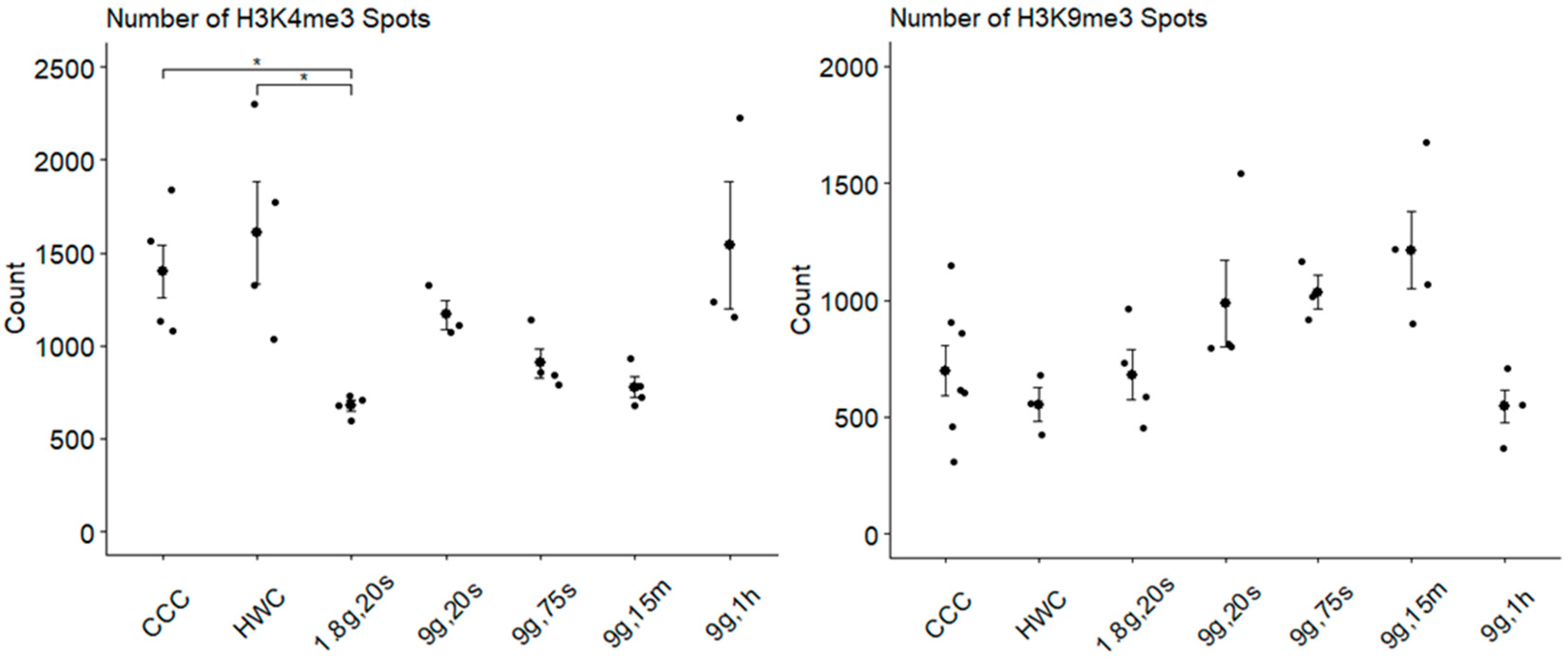
2.6. Characterization of Histone Modification Spots in Hypergravity
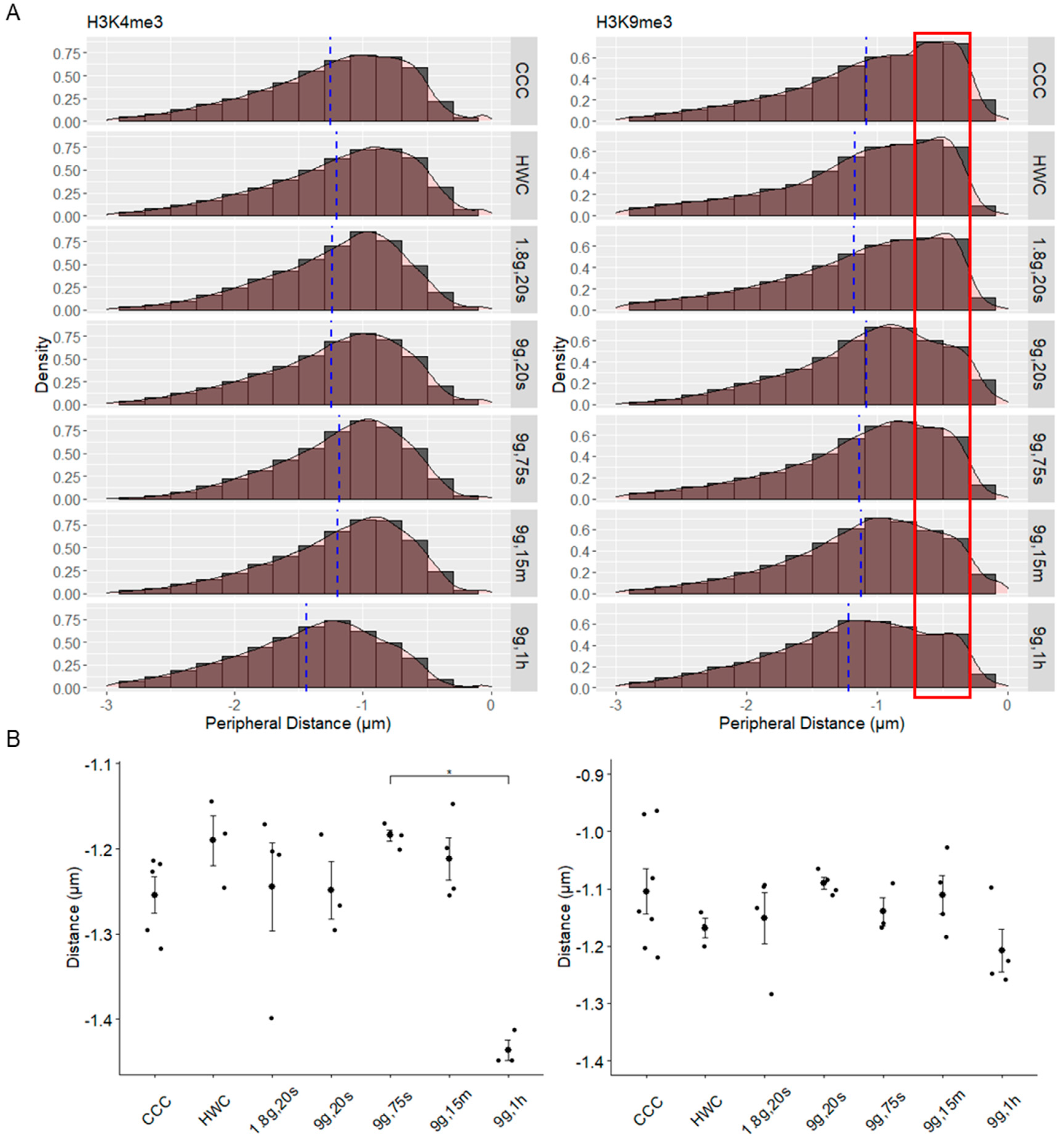
2.7. Spatial Distribution of Histone Modification Spots in Relation to the Nuclear Periphery in Hypergravity
2.8. Dynamic Response of H3K9me3 Histone Modification Spots to Hypergravity
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Centrifugation Experiment Design
4.3. Immunocytochemical Staining
4.4. Widefield Microscopy
4.5. Confocal Microscopy
4.6. Image Analysis
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| LINC | Linker of nucleoskeleton to cytoskeleton |
| RFI | Relative fluorescence intensity |
| SEM | Standard error of mean |
| CCC | Cell culture control |
| HWC | Hardware control |
| NC | Negative control |
| PC | Positive control |
| H3K4me3 | Histone 3 lysine 4 trimethylation |
| H3K9me3 | Histone 3 lysine 9 trimethylation |
| H3K4ac | Histone 3 lysine 4 acetylation |
| H3K27me3 | Histone 3 lysine 27 trimethylation |
| HM | Histone modification |
| TUNEL | Terminal deoxynucleotidyl transferase dUTP nick end labeling |
| COMPASS | Complex Proteins Associated with Set1 |
| HMT | Histone methyltransferase |
| LAD | Lamin associated domain |
References
- Pollock, R.D.; Hodkinson, P.D.; Smith, T.G. Oh G: The x, y and z of human physiological responses to acceleration. Exp. Physiol. 2021, 106, 2367–2384. [Google Scholar] [CrossRef] [PubMed]
- Isasi, E.; Isasi, M.E.; van Loon, J.J. The application of artificial gravity in medicine and space. Front. Physiol. 2022, 13, 1627. [Google Scholar] [CrossRef] [PubMed]
- Ford, W. Lymphocytes. 3. Distribution. Distribution of lymphocytes in health. J. Clin. Pathology. Suppl. (R. Coll. Pathol.) 1979, 13, 63. [Google Scholar] [CrossRef] [PubMed]
- Cucchini, F.; Di Mario, C.; Iavernaro, A.; Zeppellini, R.; Barilli, A.; Bolognesi, R. Peak aortic blood acceleration: A possible indicator of initial left ventricular impairment in patients with coronary artery disease. Eur. Heart J. 1991, 12, 860–868. [Google Scholar] [PubMed]
- Chemla, D.; Demolis, P.; Thyrault, M.; Annane, D.; Lecarpentier, Y.; Giudicelli, J. Blood flow acceleration in the carotid and brachial arteries of healthy volunteers: Respective contributions of cardiac performance and local resistance. Fundam. Clin. Pharmacol. 1996, 10, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Bennett, E.; Else, W.; Miller, G.; Sutton, G. Maximum accele ration of blood from the left ventricle in patients with is cha emic. Clin. Sci. Mol. Med. 1974, 46, 49–59. [Google Scholar] [PubMed]
- Cogoli, A. The effect of hypogravity and hypergravity on cells of the immune system. J. Leukoc. Biol. 1993, 54, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Lorenzi, G.; Fuchs-Bislin, P.; Cogoli, A. Effects of hypergravity on “whole-blood” cultures of human lymphocytes. Aviat. Space Environ. Med. 1986, 57, 1131–1135. [Google Scholar]
- Genchi, G.G.; Rocca, A.; Marino, A.; Grillone, A.; Mattoli, V.; Ciofani, G. Hypergravity as a tool for cell stimulation: Implications in biomedicine. Front. Astron. Space Sci. 2016, 3, 26. [Google Scholar] [CrossRef]
- Lang, K.; Strell, C.; Niggemann, B.; Zänker, K.S.; Hilliger, A.; Engelmann, F.; Ullrich, O. Real-time video-microscopy of migrating immune cells in altered gravity during parabolic flights. Microgravity Sci. Technol. 2010, 22, 63–69. [Google Scholar] [CrossRef]
- Ivanova, K.; Block, I.; Das, P.K.; Gerzer, R. Role of cyclic GMP signaling in the melanocyte response to hypergravity. Signal Transduct. 2006, 6, 406–413. [Google Scholar] [CrossRef]
- Gaubin, Y.; Pianezzi, B.; Soleilhavoup, J.-P.; Croute, F. Modulation by hypergravity of extracellular matrix macromolecules in in vitro human dermal fibroblasts. Biochim. Biophys. Acta (BBA) Gen. Subj. 1995, 1245, 173–180. [Google Scholar] [CrossRef]
- Gebken, J.; Liiders, B.; Notbohm, H.; Klein, H.H.; Brinckmann, J.; Müller, P.K.; Bátge, B. Hypergravity stimulates collagen synthesis in human osteoblast-like cells: Evidence for the involvement of p44/42 MAP-kinases (ERK 1/2). J. Biochem. 1999, 126, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Saito, M.; Soshi, S.; Fujii, K. Effect of hyper-and microgravity on collagen post-translational controls of MC3T3-E1 osteoblasts. J. Bone Miner. Res. 2003, 18, 1695–1705. [Google Scholar] [CrossRef] [PubMed]
- Basile, V.; Romano, G.; Fusi, F.; Monici, M. Comparison between the effects of hypergravity and photomechanical stress on cells producing ECM. Microgravity Sci. Technol. 2009, 21, 151–157. [Google Scholar] [CrossRef]
- Oguro, A.; Sakurai, T.; Fujita, Y.; Lee, S.; Kubota, H.; Nagata, K.; Atomi, Y. The molecular chaperone HSP47 rapidly senses gravitational changes in myoblasts. Genes Cells 2006, 11, 1253–1265. [Google Scholar] [CrossRef] [PubMed]
- Ciofani, G.; Ricotti, L.; Rigosa, J.; Menciassi, A.; Mattoli, V.; Monici, M. Hypergravity effects on myoblast proliferation and differentiation. J. Biosci. Bioeng. 2012, 113, 258–261. [Google Scholar] [CrossRef]
- Ogneva, I.V.; Gnyubkin, V.; Laroche, N.; Maximova, M.V.; Larina, I.M.; Vico, L. Structure of the cortical cytoskeleton in fibers of postural muscles and cardiomyocytes of mice after 30-day 2-g centrifugation. J. Appl. Physiol. 2015, 118, 613–623. [Google Scholar] [CrossRef]
- Rocca, A.; Marino, A.; Rocca, V.; Moscato, S.; de Vito, G.; Piazza, V.; Mazzolai, B.; Mattoli, V.; Ngo-Anh, T.J.; Ciofani, G. Barium titanate nanoparticles and hypergravity stimulation improve differentiation of mesenchymal stem cells into osteoblasts. Int. J. Nanomed. 2015, 10, 433–445. [Google Scholar]
- Thiel, C.S.; Hauschild, S.; Huge, A.; Tauber, S.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B. Dynamic gene expression response to altered gravity in human T cells. Sci. Rep. 2017, 7, 5204. [Google Scholar] [CrossRef]
- Thiel, C.S.; Tauber, S.; Christoffel, S.; Huge, A.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B. Rapid coupling between gravitational forces and the transcriptome in human myelomonocytic U937 cells. Sci. Rep. 2018, 8, 13267. [Google Scholar] [CrossRef]
- Van Loon, J.J. Centrifuges for microgravity simulation. The reduced gravity paradigm. Front. Astron. Space Sci. 2016, 3, 21. [Google Scholar] [CrossRef]
- Thiel, C.S.; Christoffel, S.; Tauber, S.; Vahlensieck, C.; Zelicourt, D.; Layer, L.E.; Lauber, B.; Polzer, J.; Ullrich, O. Rapid Cellular Perception of Gravitational Forces in Human Jurkat T Cells and Transduction into Gene Expression Regulation. Int. J. Mol. Sci. 2020, 21, 514. [Google Scholar] [CrossRef]
- Vahlensieck, C.; Thiel, C.S.; Mosimann, M.; Bradley, T.; Caldana, F.; Polzer, J.; Lauber, B.A.; Ullrich, O. Transcriptional Response in Human Jurkat T Lymphocytes to a near Physiological Hypergravity Environment and to One Common in Routine Cell Culture Protocols. Int. J. Mol. Sci. 2023, 24, 1351. [Google Scholar] [CrossRef]
- Thiel, C.S.; Tauber, S.; Lauber, B.; Polzer, J.; Seebacher, C.; Uhl, R.; Neelam, S.; Zhang, Y.; Levine, H.; Ullrich, O. Rapid Morphological and Cytoskeletal Response to Microgravity in Human Primary Macrophages. Int. J. Mol. Sci. 2019, 20, 2402. [Google Scholar] [CrossRef]
- Vahlensieck, C.; Thiel, C.S.; Zhang, Y.; Huge, A.; Ullrich, O. Gravitational force—Induced 3D chromosomal conformational changes are associated with rapid transcriptional response in human T cells. Int. J. Mol. Sci. 2021, 22, 9426. [Google Scholar] [CrossRef]
- Iyer, K.V.; Pulford, S.; Mogilner, A.; Shivashankar, G. Mechanical activation of cells induces chromatin remodeling preceding MKL nuclear transport. Biophys. J. 2012, 103, 1416–1428. [Google Scholar] [CrossRef]
- Hsia, C.-R.; Melters, D.P.; Dalal, Y. The force is strong with this epigenome: Chromatin structure and mechanobiology. J. Mol. Biol. 2023, 435, 168019. [Google Scholar] [CrossRef]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Liang, G.; Lin, J.C.; Wei, V.; Yoo, C.; Cheng, J.C.; Nguyen, C.T.; Weisenberger, D.J.; Egger, G.; Takai, D.; Gonzales, F.A. Distinct localization of histone H3 acetylation and H3-K4 methylation to the transcription start sites in the human genome. Proc. Natl. Acad. Sci. USA 2004, 101, 7357–7362. [Google Scholar] [CrossRef]
- Lachner, M.; O’Sullivan, R.J.; Jenuwein, T. An epigenetic road map for histone lysine methylation. J. Cell Sci. 2003, 116, 2117–2124. [Google Scholar] [CrossRef]
- Poleshko, A.; Mansfield, K.M.; Burlingame, C.C.; Andrake, M.D.; Shah, N.R.; Katz, R.A. The human protein PRR14 tethers heterochromatin to the nuclear lamina during interphase and mitotic exit. Cell Rep. 2013, 5, 292–301. [Google Scholar] [CrossRef]
- Nava, M.M.; Miroshnikova, Y.A.; Biggs, L.C.; Whitefield, D.B.; Metge, F.; Boucas, J.; Vihinen, H.; Jokitalo, E.; Li, X.; Arcos, J.M.G. Heterochromatin-driven nuclear softening protects the genome against mechanical stress-induced damage. Cell 2020, 181, 800–817.e822. [Google Scholar] [CrossRef]
- Hsia, C.-R.; McAllister, J.; Hasan, O.; Judd, J.; Lee, S.; Agrawal, R.; Chang, C.-Y.; Soloway, P.; Lammerding, J. Confined migration induces heterochromatin formation and alters chromatin accessibility. Iscience 2022, 25, 104978. [Google Scholar] [CrossRef]
- Crisp, M.; Liu, Q.; Roux, K.; Rattner, J.; Shanahan, C.; Burke, B.; Stahl, P.D.; Hodzic, D. Coupling of the nucleus and cytoplasm: Role of the LINC complex. J. Cell Biol. 2006, 172, 41–53. [Google Scholar] [CrossRef]
- Thiel, C.S.; Huge, A.; Hauschild, S.; Tauber, S.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B. Stability of gene expression in human T cells in different gravity environments is clustered in chromosomal region 11p15.4. NPJ Microgravity 2017, 3, 22. [Google Scholar] [CrossRef]
- Wang, P.; Li, J.; Wei, Q. Understanding the Interplay between Cell Force and Cell Adhesion Processes. Eng. Regen. 2023, 4, 277–288. [Google Scholar] [CrossRef]
- Roca-Cusachs, P.; Gauthier, N.C.; Del Rio, A.; Sheetz, M.P. Clustering of α5β1 integrins determines adhesion strength whereas αvβ3 and talin enable mechanotransduction. Proc. Natl. Acad. Sci. USA 2009, 106, 16245–16250. [Google Scholar] [CrossRef]
- Saika, S.; Veldhuis, N.; Križaj, D.; Rahaman, S.O. New Insights into Mechanotransduction by Immune Cells in Physiological and Pathological Conditions. Front. Immunol. 2022, 13, 930362. [Google Scholar] [CrossRef]
- Alon, R.; Feigelson, S.W. Chemokine-triggered leukocyte arrest: Force-regulated bi-directional integrin activation in quantal adhesive contacts. Curr. Opin. Cell Biol. 2012, 24, 670–676. [Google Scholar] [CrossRef]
- Sun, Z.; Guo, S.S.; Fässler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 2016, 215, 445–456. [Google Scholar] [CrossRef]
- Santoni, G.; Amantini, C.; Santoni, M.; Maggi, F.; Morelli, M.B.; Santoni, A. Mechanosensation and mechanotransduction in natural killer cells. Front. Immunol. 2021, 12, 688918. [Google Scholar] [CrossRef]
- Tabdanov, E.; Gondarenko, S.; Kumari, S.; Liapis, A.; Dustin, M.L.; Sheetz, M.P.; Kam, L.C.; Iskratsch, T. Micropatterning of TCR and LFA-1 ligands reveals complementary effects on cytoskeleton mechanics in T cells. Integr. Biol. 2015, 7, 1272–1284. [Google Scholar] [CrossRef]
- Kim, S.T.; Takeuchi, K.; Sun, Z.-Y.J.; Touma, M.; Castro, C.E.; Fahmy, A.; Lang, M.J.; Wagner, G.; Reinherz, E.L. The αβ T cell receptor is an anisotropic mechanosensor. J. Biol. Chem. 2009, 284, 31028–31037. [Google Scholar] [CrossRef]
- Gunasinghe, S.D.; Peres, N.G.; Goyette, J.; Gaus, K. Biomechanics of T cell dysfunctions in chronic diseases. Front. Immunol. 2021, 12, 600829. [Google Scholar] [CrossRef]
- Liu, B.; Chen, W.; Evavold, B.D.; Zhu, C. Antigen-specific TCR–pMHC catch bonds trigger signaling by fast accumulation of force-prolonged bond lifetimes. Cell 2014, 157, 357. [Google Scholar] [CrossRef]
- Hong, J.; Persaud, S.P.; Horvath, S.; Allen, P.M.; Evavold, B.D.; Zhu, C. Force-regulated in situ TCR–peptide-bound MHC class II kinetics determine functions of CD4+ T cells. J. Immunol. 2015, 195, 3557–3564. [Google Scholar] [CrossRef]
- Plotnikov, S.V.; Waterman, C.M. Guiding cell migration by tugging. Curr. Opin. Cell Biol. 2013, 25, 619–626. [Google Scholar] [CrossRef]
- Kumar, S. Stiffness does matter. Nat. Mater. 2014, 13, 918–920. [Google Scholar] [CrossRef]
- Zhang, H.; Labouesse, M. Signalling through mechanical inputs—A coordinated process. J. Cell Sci. 2012, 125, 3039–3049. [Google Scholar] [CrossRef]
- Dutta, B.; Arya, R.K.; Goswami, R.; Alharbi, M.O.; Sharma, S.; Rahaman, S.O. Role of macrophage TRPV4 in inflammation. Lab. Investig. 2020, 100, 178–185. [Google Scholar] [CrossRef]
- Meli, V.S.; Veerasubramanian, P.K.; Atcha, H.; Reitz, Z.; Downing, T.L.; Liu, W.F. Biophysical regulation of macrophages in health and disease. J. Leukoc. Biol. 2019, 106, 283–299. [Google Scholar] [CrossRef]
- Koaykul, C.; Kim, M.-H.; Kawahara, Y.; Yuge, L.; Kino-Oka, M. Maintenance of neurogenic differentiation potential in passaged bone marrow-derived human mesenchymal stem cells under simulated microgravity conditions. Stem Cells Dev. 2019, 28, 1552–1561. [Google Scholar] [CrossRef]
- Schätzle, P.; Wuttke, R.; Ziegler, U.; Sonderegger, P. Automated quantification of synapses by fluorescence microscopy. J. Neurosci. Methods 2012, 204, 144–149. [Google Scholar] [CrossRef]
- Singh, K.P.; Kumari, R.; DuMond, J.W. Simulated microgravity-induced epigenetic changes in human lymphocytes. J. Cell. Biochem. 2010, 111, 123–129. [Google Scholar] [CrossRef]
- Tauber, S.; Hauschild, S.; Paulsen, K.; Gutewort, A.; Raig, C.; Hürlimann, E.; Biskup, J.; Philpot, C.; Lier, H.; Engelmann, F. Signal transduction in primary human T lymphocytes in altered gravity during parabolic flight and clinostat experiments. Cell. Physiol. Biochem. 2015, 35, 1034–1051. [Google Scholar] [CrossRef]
- Kamal, K.Y.; Herranz, R.; Van Loon, J.J.; Medina, F.J. Simulated microgravity, Mars gravity, and 2g hypergravity affect cell cycle regulation, ribosome biogenesis, and epigenetics in Arabidopsis cell cultures. Sci. Rep. 2018, 8, 6424. [Google Scholar] [CrossRef]
- Pokholok, D.K.; Harbison, C.T.; Levine, S.; Cole, M.; Hannett, N.M.; Lee, T.I.; Bell, G.W.; Walker, K.; Rolfe, P.A.; Herbolsheimer, E. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 2005, 122, 517–527. [Google Scholar] [CrossRef]
- Guillemette, B.; Drogaris, P.; Lin, H.-H.S.; Armstrong, H.; Hiragami-Hamada, K.; Imhof, A.; Bonneil, E.; Thibault, P.; Verreault, A.; Festenstein, R.J. H3 lysine 4 is acetylated at active gene promoters and is regulated by H3 lysine 4 methylation. PLoS Genet. 2011, 7, e1001354. [Google Scholar] [CrossRef]
- Tauber, S.; Hauschild, S.; Crescio, C.; Secchi, C.; Paulsen, K.; Pantaleo, A.; Saba, A.; Buttron, I.; Thiel, C.S.; Cogoli, A. Signal transduction in primary human T lymphocytes in altered gravity–results of the MASER-12 suborbital space flight mission. Cell Commun. Signal. 2013, 11, 32. [Google Scholar] [CrossRef]
- Pinskaya, M.; Gourvennec, S.; Morillon, A. H3 lysine 4 di-and tri-methylation deposited by cryptic transcription attenuates promoter activation. EMBO J. 2009, 28, 1697–1707. [Google Scholar] [CrossRef]
- Chong, S.Y.; Cutler, S.; Lin, J.-J.; Tsai, C.-H.; Tsai, H.-K.; Biggins, S.; Tsukiyama, T.; Lo, Y.-C.; Kao, C.-F. H3K4 methylation at active genes mitigates transcription-replication conflicts during replication stress. Nat. Commun. 2020, 11, 809. [Google Scholar] [CrossRef]
- Lee, M.; Kim, D.; Kwon, S. Hypergravity-induced changes in actin response of breast cancer cells to natural killer cells. Sci. Rep. 2021, 11, 7267. [Google Scholar] [CrossRef]
- Dai, Z.; Tan, Y.; Yang, F.; Qu, L.; Zhang, H.; Wan, Y.; Li, Y. Altered actin dynamics and functions of osteoblast-like cells in parabolic flight may involve ERK1/2. Microgravity Sci. Technol. 2011, 23, 19–27. [Google Scholar] [CrossRef]
- Kloc, M.; Chanana, P.; Vaughn, N.; Uosef, A.; Kubiak, J.Z.; Ghobrial, R.M. New insights into cellular functions of nuclear actin. Biology 2021, 10, 304. [Google Scholar] [CrossRef]
- Vitavska, O.; Merzendorfer, H.; Wieczorek, H. The V-ATPase subunit C binds to polymeric F-actin as well as to monomeric G-actin and induces cross-linking of actin filaments. J. Biol. Chem. 2005, 280, 1070–1076. [Google Scholar] [CrossRef] [PubMed]
- Maxson, M.E.; Grinstein, S. The vacuolar-type H(+)-ATPase at a glance—More than a proton pump. J. Cell Sci. 2014, 127, 4987–4993. [Google Scholar] [CrossRef]
- Kaminski, A.; Fedorchak, G.R.; Lammerding, J. The cellular mastermind (?)—Mechanotransduction and the nucleus. Prog. Mol. Biol. Transl. Sci. 2014, 126, 157–203. [Google Scholar]
- Geng, J.; Kang, Z.; Sun, Q.; Zhang, M.; Wang, P.; Li, Y.; Li, J.; Su, B.; Wei, Q. Microtubule Assists Actomyosin to Regulate Cell Nuclear Mechanics and Chromatin Accessibility. Research 2023, 6, 0054. [Google Scholar] [CrossRef]
- Puech, P.-H.; Bongrand, P. Mechanotransduction as a major driver of cell behaviour: Mechanisms, and relevance to cell organization and future research. Open Biol. 2021, 11, 210256. [Google Scholar] [CrossRef]
- Pawłowski, R.; Rajakylä, E.K.; Vartiainen, M.K.; Treisman, R. An actin-regulated importin α/β-dependent extended bipartite NLS directs nuclear import of MRTF-A. EMBO J. 2010, 29, 3448–3458. [Google Scholar] [CrossRef]
- Scheer, U.; Hinssen, H.; Franke, W.W.; Jockusch, B.M. Microinjection of actin-binding proteins and actin antibodies demonstrates involvement of nuclear actin in transcription of lampbrush chromosomes. Cell 1984, 39, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, P.; Chen, M.; Winkler, D.D.; Luger, K.; Shen, X. Evidence for monomeric actin function in INO80 chromatin remodeling. Nat. Struct. Mol. Biol. 2013, 20, 426–432. [Google Scholar] [CrossRef] [PubMed]
- Kelpsch, D.J.; Tootle, T.L. Nuclear actin: From discovery to function. Anat. Rec. 2018, 301, 1999–2013. [Google Scholar] [CrossRef] [PubMed]
- Heo, S.-J.; Han, W.M.; Szczesny, S.E.; Cosgrove, B.D.; Elliott, D.M.; Lee, D.A.; Duncan, R.L.; Mauck, R.L. Mechanically induced chromatin condensation requires cellular contractility in mesenchymal stem cells. Biophys. J. 2016, 111, 864–874. [Google Scholar] [CrossRef]
- Montavon, T.; Shukeir, N.; Erikson, G.; Engist, B.; Onishi-Seebacher, M.; Ryan, D.; Musa, Y.; Mittler, G.; Meyer, A.G.; Genoud, C. Complete loss of H3K9 methylation dissolves mouse heterochromatin organization. Nat. Commun. 2021, 12, 4359. [Google Scholar] [CrossRef]
- Ayrapetov, M.K.; Gursoy-Yuzugullu, O.; Xu, C.; Xu, Y.; Price, B.D. DNA double-strand breaks promote methylation of histone H3 on lysine 9 and transient formation of repressive chromatin. Proc. Natl. Acad. Sci. USA 2014, 111, 9169–9174. [Google Scholar] [CrossRef]
- Ren, W.; Fan, H.; Grimm, S.A.; Guo, Y.; Kim, J.J.; Yin, J.; Li, L.; Petell, C.J.; Tan, X.-F.; Zhang, Z.-M. Direct readout of heterochromatic H3K9me3 regulates DNMT1-mediated maintenance DNA methylation. Proc. Natl. Acad. Sci. USA 2020, 117, 18439–18447. [Google Scholar] [CrossRef]
- Wang, Y.-H.; Hariharan, A.; Bastianello, G.; Toyama, Y.; Shivashankar, G.; Foiani, M.; Sheetz, M.P. DNA damage causes rapid accumulation of phosphoinositides for ATR signaling. Nat. Commun. 2017, 8, 2118. [Google Scholar] [CrossRef]
- Caridi, C.P.; D’Agostino, C.; Ryu, T.; Zapotoczny, G.; Delabaere, L.; Li, X.; Khodaverdian, V.Y.; Amaral, N.; Lin, E.; Rau, A.R. Nuclear F-actin and myosins drive relocalization of heterochromatic breaks. Nature 2018, 559, 54–60. [Google Scholar] [CrossRef]
- Raurell-Vila, H.; Bosch-Presegue, L.; Gonzalez, J.; Kane-Goldsmith, N.; Casal, C.; Brown, J.P.; Marazuela-Duque, A.; Singh, P.B.; Serrano, L.; Vaquero, A. An HP1 isoform-specific feedback mechanism regulates Suv39h1 activity under stress conditions. Epigenetics 2017, 12, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Chaturvedi, P.; Parnaik, V.K. Lamin A rod domain mutants target heterochromatin protein 1α and β for proteasomal degradation by activation of F-box protein, FBXW10. PLoS ONE 2010, 5, e10620. [Google Scholar] [CrossRef] [PubMed]
- Mungamuri, S.K.; Qiao, R.F.; Yao, S.; Manfredi, J.J.; Gu, W.; Aaronson, S.A. USP7 enforces heterochromatinization of p53 target promoters by protecting SUV39H1 from MDM2-mediated degradation. Cell Rep. 2016, 14, 2528–2537. [Google Scholar] [CrossRef] [PubMed]
- Stephens, A.D.; Liu, P.Z.; Banigan, E.J.; Almassalha, L.M.; Backman, V.; Adam, S.A.; Goldman, R.D.; Marko, J.F. Chromatin histone modifications and rigidity affect nuclear morphology independent of lamins. Mol. Biol. Cell 2018, 29, 220–233. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Tajik, A.; Chen, J.; Jia, Q.; Chowdhury, F.; Wang, L.; Chen, J.; Zhang, S.; Hong, Y.; Yi, H. Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression. Nat. Commun. 2014, 5, 4619. [Google Scholar] [CrossRef] [PubMed]
- Le, H.Q.; Ghatak, S.; Yeung, C.-Y.C.; Tellkamp, F.; Günschmann, C.; Dieterich, C.; Yeroslaviz, A.; Habermann, B.; Pombo, A.; Niessen, C.M. Mechanical regulation of transcription controls Polycomb-mediated gene silencing during lineage commitment. Nat. Cell Biol. 2016, 18, 864–875. [Google Scholar] [CrossRef]
- Vahlensieck, C.; Thiel, C.S.; Adelmann, J.; Lauber, B.A.; Polzer, J.; Ullrich, O. Rapid transient transcriptional adaptation to hypergravity in jurkat t cells revealed by comparative analysis of microarray and RNA-Seq data. Int. J. Mol. Sci. 2021, 22, 8451. [Google Scholar] [CrossRef]
- Hiragami-Hamada, K.; Soeroes, S.; Nikolov, M.; Wilkins, B.; Kreuz, S.; Chen, C.; De La Rosa-Velazquez, I.A.; Zenn, H.M.; Kost, N.; Pohl, W. Dynamic and flexible H3K9me3 bridging via HP1β dimerization establishes a plastic state of condensed chromatin. Nat. Commun. 2016, 7, 11310. [Google Scholar] [CrossRef]
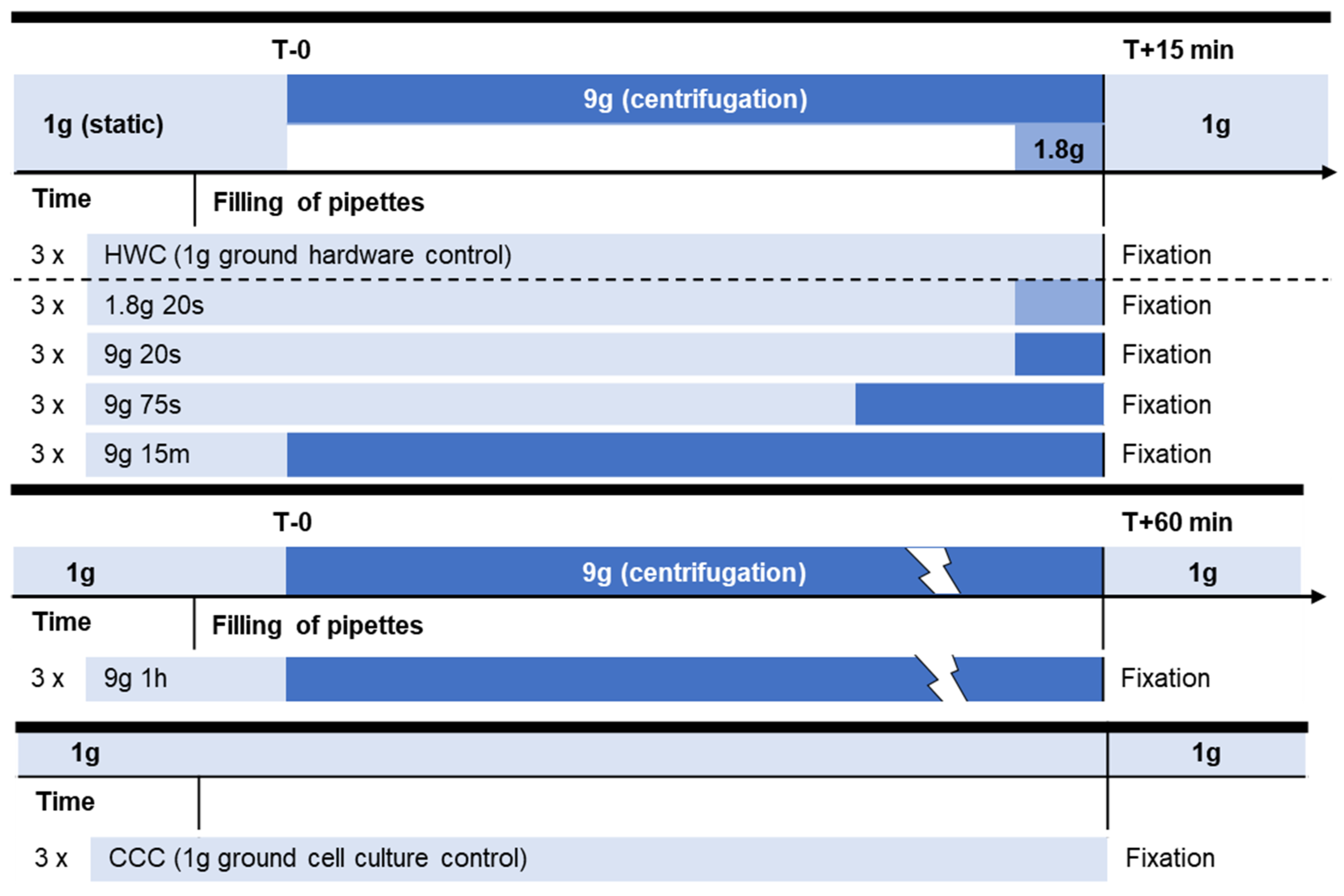



| ID | Group | Gravitational Force (× g) | Centrifuge Time (Tc) | Incubation Time (Ti) |
|---|---|---|---|---|
| CCC | Cell culture control (CCC) | 1 | 0 | 0 |
| HWC | Hardware control (HWC) | 1 | 0 | 15 min |
| 1.8g, 20s | Hypergravity (parabolic flight simulation) | 1.8 | 20 s | 14 min 40 s |
| 9g, 20s | Hypergravity | 9 | 20 s | 14 min 40 s |
| 9g, 75s | Hypergravity (sounding rocket simulation) | 9 | 75 s | 13 min 45 s |
| 9g, 15m | Hypergravity | 9 | 15 min | 0 |
| 9g, 1h | Hypergravity | 9 | 60 min | 0 |
| Analyses | Epitope/Parameter | CCC | HWC | 1.8g20s | 9g20s | 9g75s | 9g15m | 9g1h | |
|---|---|---|---|---|---|---|---|---|---|
| Widefield Overall RFI | H3K4me3 | 0.80 | 1.00 | 0.74 | 0.92 | 0.86 | 0.97 | 0.89 | |
| H3K9me3 | 1.00 | 1.00 | 1.63 | 1.69 | 1.90 | 2.18 | 0.95 | ||
| F-Actin | 1.01 | 0.94 | 0.68 | 0.59 | 0.70 | 0.66 | 1.22 | ||
| Nuclear Shape | Width (µm) | 5.20 | 5.40 | 5.56 | 5.62 | 5.77 | 5.82 | 5.96 | |
| Height (µm) | 3.62 | 3.44 | 3.21 | 3.15 | 3.03 | 3.07 | 2.98 | ||
| Oblate | 0.36 | 0.39 | 0.43 | 0.46 | 0.47 | 0.45 | 0.46 | ||
| Sphericity | 0.92 | 0.92 | 0.91 | 0.90 | 0.89 | 0.90 | 0.90 | ||
| HM Spot Number | H3K4me3 | 1404 | 1617 | 681 | 1162 | 897 | 778 | 1454 | |
| H3K9me3 | 576 | 533 | 561 | 814 | 735 | 957 | 437 | ||
| HM Spot Peripheral Distance | H3K4me3 | Skew | −0.69 | −0.77 | −0.72 | −0.66 | −0.65 | −0.73 | −0.52 |
| Mean (µm) | −1.25 | −1.20 | −1.24 | −1.25 | −1.18 | −1.20 | −1.44 | ||
| H3K9me3 | Skew | −0.88 | −1.00 | −0.98 | −0.69 | −0.95 | −0.78 | −0.62 | |
| Mean (µm) | −1.09 | −1.17 | −1.18 | −1.08 | −1.14 | −1.12 | −1.22 | ||
| HM Spot RFI | H3K4me3 | Skew | 0.25 | 0.21 | 0.66 | 0.18 | 0.12 | 0.33 | 0.31 |
| Mean | 1.07 | 1.00 | 1.07 | 1.30 | 1.20 | 1.06 | 0.95 | ||
| H3K9me3 | Skew | 1.08 | 1.08 | 0.88 | 0.95 | 1.08 | 0.89 | 1.22 | |
| Mean | 1.04 | 1.00 | 1.11 | 1.08 | 1.07 | 1.16 | 0.92 | ||
| Epitope | Excitation (nm) | Emission (nm) | Exposure Time (ms) | Laser Power | Gain |
|---|---|---|---|---|---|
| DAPI | 390 | 460–480 | 100 | 60% | low noise & high well |
| H3K4me3/H3K9me3 (Alexa 488) | 475 | 535–570 | 200 | 60% | low noise & high well |
| Phalloidin (Alexa 647) | 635 | >695 | 300 | 60% | low noise & high well |
| TUNEL | 635 | >695 | 200 | 40% | low noise & high well |
| Epitope | Excitation (nm) | Emission (nm) | Exposure Time (ms) | Laser Power | Gain |
|---|---|---|---|---|---|
| DAPI | 405 | 425–506 | 100 | 30% | 2 |
| H3K4me3/H3K9me3 (Alexa 488) | 488 | 506–620 | 200 | 50% | 2 |
| Phalloidin (Alexa 647) | 647 | 650–776 | 300 | 50% | 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wernlé, K.; Thiel, C.S.; Ullrich, O. Increased H3K9me3 and F-Actin Reorganization in the Rapid Adaptive Response to Hypergravity in Human T Lymphocytes. Int. J. Mol. Sci. 2023, 24, 17232. https://doi.org/10.3390/ijms242417232
Wernlé K, Thiel CS, Ullrich O. Increased H3K9me3 and F-Actin Reorganization in the Rapid Adaptive Response to Hypergravity in Human T Lymphocytes. International Journal of Molecular Sciences. 2023; 24(24):17232. https://doi.org/10.3390/ijms242417232
Chicago/Turabian StyleWernlé, Kendra, Cora S. Thiel, and Oliver Ullrich. 2023. "Increased H3K9me3 and F-Actin Reorganization in the Rapid Adaptive Response to Hypergravity in Human T Lymphocytes" International Journal of Molecular Sciences 24, no. 24: 17232. https://doi.org/10.3390/ijms242417232






