Osteoarthritis and microRNAs: Do They Provide Novel Insights into the Pathophysiology of This Degenerative Disorder?
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Strategy and Study Selection
2.2. Inclusion and Exclusion Criteria
2.3. Data Extraction
3. Results
3.1. MiRs Expression in OA Patients
3.2. MiRs Involvement in the Pathogenesis of OA
3.3. Effective Roles of miRs in OA Pathophysiology
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Trachana, V.; Ntoumou, E.; Anastasopoulou, L.; Tsezou, A. Studying miRs in OA: Critical overview of different analytical approaches. Mech. Ageing Dev. 2018, 171, 15–23. [Google Scholar] [CrossRef]
- Zhang, Y.; Jia, J.; Yang, S.; Liu, X.; Ye, S.; Tian, H. MiR-21 controls the development of oa by targeting GDF-5 in chondrocytes. Exp. Mol. Med. 2014, 46, e79. [Google Scholar] [CrossRef] [Green Version]
- Carossino, A.M.; Recenti, R.; Carossino, R.; Piscitelli, E.; Gozzini, A.; Martineti, V.; Mavilia, C.; Franchi, A.; Danielli, D.; Aglietti, P.; et al. Methodological models for in vitro amplification and maintenance of human articular chondrocytes from elderly patients. Biogerontology 2007, 8, 483–498. [Google Scholar] [CrossRef]
- Waarsing, J.H.; Kloppenburg, M.; Slagboom, P.; Kroon, H.M.; Houwing-Duistermaat, J.J.; Weinans, H.; Meulenbelt, I. OA susceptibility genes influence the association between hip morphology and OA. Arthritis Rheum. 2011, 63, 1349–1354. [Google Scholar] [CrossRef]
- Chapman, K.; Valdes, A.M. Genetic factors in OA pathogenesis. Bone 2012, 51, 258–264. [Google Scholar] [CrossRef]
- Borgonio Cuadra, V.M.; González-Huerta, N.C.; Romero-Cordoba, S.L.; Hidalgo-Miranda, A.; Miranda-Duarte, A. Altered Expression of Circulating miR in Plasma of Patients with Primary OA and In Silico Analysis of Their Pathways. PLoS ONE 2014, 9, e97690. [Google Scholar] [CrossRef]
- Dupont, C.; Armant, D.R.; Brenner, C.A. Epigenetics: Definition, Mechanisms and Clinical Perspective. Semin. Reprod. Med. 2009, 27, 351–357. [Google Scholar] [CrossRef] [Green Version]
- Macfarlane, L.-A.; Murphy, P.R. MiR: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef] [Green Version]
- Lu, T.X.; Rothenberg, M.E. MiR. J. Allergy Clin. Immunol. 2018, 141, 1202–1207. [Google Scholar] [CrossRef] [Green Version]
- Bhaskaran, M.; Mohan, M. MiRs: History, biogenesis, and their evolving role in animal development and disease. Veter Pathol. 2014, 51, 759–774. [Google Scholar] [CrossRef]
- Hartig, S.M.; Hamilton, M.P.; Bader, D.A.; McGuire, S.E. The miRNA Interactome in Metabolic Homeostasis. Trends Endocrinol. Metab. 2015, 26, 733–745. [Google Scholar] [CrossRef] [Green Version]
- Hamilton, M.P.; Rajapakshe, K.; Hartig, S.M.; Reva, B.; McLellan, M.D.; Kandoth, C.; Ding, L.; Zack, T.I.; Gunaratne, P.H.; Wheeler, D.A.; et al. Identification of a pan-cancer oncogenic miR superfamily anchored by a central core seed motif. Nat. Commun. 2013, 4, 2730. [Google Scholar] [CrossRef]
- Specjalski, K.; Jassem, E. MiRs: Potential Biomarkers and Targets of Therapy in Allergic Diseases? Arch. Immunol. Ther. Exp. 2019, 67, 213–223. [Google Scholar] [CrossRef] [Green Version]
- Ramanathan, K.; Padmanabhan, G. MiRNAs as potential biomarker of kidney diseases: A review. Cell Biochem. Funct. 2020, 38, 990–1005. [Google Scholar] [CrossRef]
- Timis, T.L.; Orasan, R.I. Understanding psoriasis: Role of miRNAs. Biomed. Rep. 2018, 9, 367–374. [Google Scholar] [CrossRef] [Green Version]
- Homorogan, C.; Nitusca, D.; Seclaman, E.; Enatescu, V.; Marian, C. Uncovering the Roles of MiRs in Major Depressive Disorder: From Candidate Diagnostic Biomarkers to Treatment Response Indicators. Life 2021, 11, 1073. [Google Scholar] [CrossRef]
- Wang, J.; Cao, Y.; Lu, X.; Wang, T.; Li, S.; Kong, X.; Bo, C.; Li, J.; Wang, X.; Ma, H.; et al. MiRs and nervous system diseases: Network insights and computational challenges. Brief. Bioinform. 2020, 21, 863–875. [Google Scholar] [CrossRef]
- Kanwar, J.R.; Mahidhara, G.; Kanwar, R.K. MiR in human cancer and chronic inflammatory diseases. Front. Biosci. 2010, 2, 1113–1126. [Google Scholar] [CrossRef] [Green Version]
- Zhong, G.; Long, H.; Ma, S.; Shunhan, Y.; Li, J.; Yao, J. miRNA-335-5p relieves chondrocyte inflammation by activating autophagy in OA. Life Sci. 2019, 226, 164–172. [Google Scholar] [CrossRef]
- Skrzypa, M.; Szala, D.; Gablo, N.; Czech, J.; Pajak, J.; Kopańska, M.; Trzeciak, M.; Gargasz, K.; Snela, S.; Zawlik, I. miRNA-146a-5p is upregulated in serum and cartilage samples of patients with OA. Pol. J. Surg. 2018, 91, 1–5. [Google Scholar] [CrossRef]
- Wu, C.; Tian, B.; Qu, X.; Liu, F.; Tang, T.; Qin, A.; Zhu, Z.; Dai, K. MicroRNAs play a role in chondrogenesis and osteoarthritis (Review). Int. J. Mol. Med. 2014, 34, 13–23. [Google Scholar] [CrossRef] [Green Version]
- Mobasheri, A.; Rayman, M.P.; Gualillo, O.; Sellam, J.; van der Kraan, P.; Fearon, U. The role of metabolism in the pathogenesis of osteoarthritis. Nat. Rev. Rheumatol. 2017, 13, 302–311. [Google Scholar] [CrossRef]
- Shen, X.-F.; Cheng, Y.; Dong, Q.-R.; Zheng, M.-Q. MicroRNA-675-3p regulates IL-1β-stimulated human chondrocyte apoptosis and cartilage degradation by targeting GNG5. Biochem. Biophys. Res. Commun. 2020, 527, 458–465. [Google Scholar] [CrossRef]
- An, Y.; Wan, G.; Tao, J.; Cui, M.; Zhou, Q.; Hou, W. Down-regulation of microRNA-203a suppresses IL-1β-induced inflammation and cartilage degradation in human chondrocytes through Smad3 signaling. Biosci. Rep. 2020, 40, BSR20192723. [Google Scholar] [CrossRef] [Green Version]
- Lu, X.; Li, Y.; Chen, H.; Pan, Y.; Lin, R.; Chen, S. miR-335-5P contributes to human osteoarthritis by targeting HBP1. Exp. Ther. Med. 2021, 21, 109. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, C.; Zhao, J.; Xu, J.; Geng, Y.; Dai, L.; Huang, Y.; Fu, S.-C.; Dai, K.; Zhang, X. miR-146a facilitates osteoarthritis by regulating cartilage homeostasis via targeting Camk2d and Ppp3r2. Cell Death Dis. 2017, 8, e2734. [Google Scholar] [CrossRef] [Green Version]
- Gao, S.; Liu, L.; Zhu, S.; Wang, D.; Wu, Q.; Ning, G.; Feng, S. MicroRNA-197 regulates chondrocyte proliferation, migration, and inflammation in pathogenesis of osteoarthritis by targeting EIF4G2. Biosci. Rep. 2020, 40, BSR20192095. [Google Scholar] [CrossRef]
- He, W.; Cheng, Y. Inhibition of miR-20 promotes proliferation and autophagy in articular chondrocytes by PI3K/AKT/mTOR signaling pathway. Biomed. Pharmacother. 2018, 97, 607–615. [Google Scholar] [CrossRef]
- Duan, Z.-X.; Huang, P.; Tu, C.; Liu, Q.; Li, S.-Q.; Long, Z.-L.; Li, Z.-H. MicroRNA-15a-5p Regulates the Development of Osteoarthritis by Targeting PTHrP in Chondrocytes. BioMed Res. Int. 2019, 2019, 3904923. [Google Scholar] [CrossRef] [Green Version]
- Kapoor, M.; Martel-Pelletier, J.; Lajeunesse, D.; Pelletier, J.-P.; Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 2011, 7, 33–42. [Google Scholar] [CrossRef]
- Goldring, S.R.; Goldring, M. The Role of Cytokines in Cartilage Matrix Degeneration in Osteoarthritis. Clin. Orthop. Relat. Res. 2004, 427, S27–S36. [Google Scholar] [CrossRef]
- Duan, L.; Liang, Y.; Xu, X.; Xiao, Y.; Wang, D. Recent progress on the role of miR-140 in cartilage matrix remodelling and its implications for osteoarthritis treatment. Arthritis Res. Ther. 2020, 22, 194. [Google Scholar] [CrossRef]
- Wang, W.-T.; Huang, Z.-P.; Sui, S.; Liu, J.-H.; Yu, D.-M.; Wang, W.-B. microRNA-1236 promotes chondrocyte apoptosis in osteoarthritis via direct suppression of PIK3R3. Life Sci. 2020, 253, 117694. [Google Scholar] [CrossRef]
- Beyer, C.; Zampetaki, A.; Lin, N.-Y.; Kleyer, A.; Perricone, C.; Iagnocco, A.; Distler, A.; Langley, S.R.; Gelse, K.; Sesselmann, S.M.; et al. Signature of circulating microRNAs in osteoarthritis. Ann. Rheum. Dis. 2015, 74, e18. [Google Scholar] [CrossRef] [Green Version]
- Ntoumou, E.; Tzetis, M.; Braoudaki, M.; Lambrou, G.; Poulou, M.; Malizos, K.; Stefanou, N.; Anastasopoulou, L.; Tsezou, A. Serum microRNA array analysis identifies miR-140-3p, miR-33b-3p and miR-671-3p as potential osteoarthritis biomarkers involved in metabolic processes. Clin. Epigenet. 2017, 9, 127. [Google Scholar] [CrossRef] [Green Version]
- Lian, W.-S.; Ko, J.-Y.; Wu, R.-W.; Sun, Y.-C.; Chen, Y.-S.; Wu, S.-L.; Weng, L.-H.; Jahr, H.; Wang, F.-S. MicroRNA-128a represses chondrocyte autophagy and exacerbates knee osteoarthritis by disrupting Atg12. Cell Death Dis. 2018, 9, 919. [Google Scholar] [CrossRef]
- Hu, J.; Wang, Z.; Pan, Y.; Ma, J.; Miao, X.; Qi, X.; Zhou, H.; Jia, L. MiR-26a and miR-26b mediate osteoarthritis progression by targeting FUT4 via NF-κB signaling pathway. Int. J. Biochem. Cell Biol. 2018, 94, 79–88. [Google Scholar] [CrossRef]
- Endisha, H.; Datta, P.; Sharma, A.; Nakamura, S.; Rossomacha, E.; Younan, C.; Ali, S.A.; Tavallaee, G.; Lively, S.; Potla, P.; et al. MicroRNA-34a-5p Promotes Joint Destruction during Osteoarthritis. Arthritis Rheumatol. 2021, 73, 426–439. [Google Scholar] [CrossRef]
- Mao, G.; Zhang, Z.; Huang, Z.; Chen, W.; Huang, G.; Meng, F.; Zhang, Z.; Kang, Y. MicroRNA-92a-3p regulates the expression of cartilage-specific genes by directly targeting histone deacetylase 2 in chondrogenesis and degradation. Osteoarthr. Cartil. 2017, 25, 521–532. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Wu, X. MicroRNA-103 contributes to osteoarthritis development by targeting Sox6. Biomed. Pharmacother. 2019, 118, 109186. [Google Scholar] [CrossRef]
- A Karlsen, T.; de Souza, G.A.; Ødegaard, B.; Engebretsen, L.; E Brinchmann, J. microRNA-140 Inhibits Inflammation and Stimulates Chondrogenesis in a Model of Interleukin 1β-induced Osteoarthritis. Mol. Ther. Nucleic Acids 2016, 5, e373. [Google Scholar] [CrossRef] [Green Version]
- Qiu, W.-J.; Xu, M.-Z.; Zhu, X.-D.; Ji, Y.-H. MicroRNA-27a alleviates IL-1β-induced inflammatory response and articular cartilage degradation via TLR4/NF-κB signaling pathway in articular chondrocytes. Int. Immunopharmacol. 2019, 76, 105839. [Google Scholar] [CrossRef]
- Ding, B.; Xu, S.; Sun, X.; Gao, J.; Nie, W.; Xu, H. miR-18a-3p Encourages Apoptosis of Chondrocyte in Osteoarthritis via HOXA1 Pathway. Curr. Mol. Pharmacol. 2020, 13, 328–341. [Google Scholar] [CrossRef]
- Hwang, H.S.; Kim, H.A. Chondrocyte Apoptosis in the Pathogenesis of Osteoarthritis. Int. J. Mol. Sci. 2015, 16, 26035–26054. [Google Scholar] [CrossRef]
- Kim, H.A.; Lee, Y.J.; Seong, S.C.; Choe, K.W.; Song, Y.W. Apoptotic chondrocyte death in human OA. J. Rheumatol. 2000, 27, 455–462. [Google Scholar]
- Huang, Z.; Zhang, N.; Ma, W.; Dai, X.; Liu, J. MiR-337-3p promotes chondrocytes proliferation and inhibits apoptosis by regulating PTEN/AKT axis in osteoarthritis. Biomed. Pharmacother. 2017, 95, 1194–1200. [Google Scholar] [CrossRef]
- Cheng, F.; Hu, H.; Sun, K.; Yan, F.; Geng, Y. miR-455-3p enhances chondrocytes apoptosis and inflammation by targeting COL2A1 in the in vitro osteoarthritis model. Biosci. Biotechnol. Biochem. 2020, 84, 695–702. [Google Scholar] [CrossRef]
- Mathiessen, A.; Conaghan, P.G. Synovitis in osteoarthritis: Current understanding with therapeutic implications. Arthritis Res. Ther. 2017, 19, 18. [Google Scholar] [CrossRef] [Green Version]
- Jin, Z.; Ren, J.; Qi, S. Human bone mesenchymal stem cells-derived exosomes overexpressing microRNA-26a-5p alleviate osteoarthritis via down-regulation of PTGS2. Int. Immunopharmacol. 2020, 78, 105946. [Google Scholar] [CrossRef]
- Murata, K.; Yoshitomi, H.; Tanida, S.; Ishikawa, M.; Nishitani, K.; Ito, H.; Nakamura, T. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res. Ther. 2010, 12, R86. [Google Scholar] [CrossRef] [Green Version]
- Aae, T.F.; Karlsen, T.A.; Haugen, I.K.; Risberg, M.A.; Lian, B.; Brinchmann, J.E. Evaluating plasma extracellular vesicle microRNAs as possible biomarkers for osteoarthritis. Osteoarthr. Cartil. Open 2020, 1, 100018. [Google Scholar] [CrossRef]
- Soyocak, A.; Kurt, H.; Ozgen, M.; Cosan, D.T.; Colak, E.; Gunes, H.V. miRNA-146a, miRNA-155 and JNK expression levels in peripheral blood mononuclear cells according to grade of knee osteoarthritis. Gene 2017, 627, 207–211. [Google Scholar] [CrossRef]
- Zakaria, S.S.; Gaballah, H.H.; El Saadany, H.M. Micro RNA-146a expression, NF-κB/P65 activity and serum pentosidine levels as potential biomarkers for disease severity in primary knee osteoarthritis patients. Egypt. Rheumatol. 2016, 38, 319–325. [Google Scholar] [CrossRef] [Green Version]
- Papathanasiou, I.; Mourmoura, E.; Balis, C.; Tsezou, A. Impact of miR-SNP rs2910164 on miR-146a expression in osteoarthritic chondrocytes. Adv. Med. Sci. 2020, 65, 78–85. [Google Scholar] [CrossRef]
- Li, Y.-H.; Tavallaee, G.; Tokar, T.; Nakamura, A.; Sundararajan, K.; Weston, A.; Sharma, A.; Mahomed, N.; Gandhi, R.; Jurisica, I.; et al. Identification of synovial fluid microRNA signature in knee osteoarthritis: Differentiating early- and late-stage knee osteoarthritis. Osteoarthr. Cartil. 2016, 24, 1577–1586. [Google Scholar] [CrossRef] [Green Version]
- Fan, Z.; Liu, Y.; Shi, Z.; Deng, K.; Zhang, H.; Li, Q.; Cao, S.; Li, S.; Zhang, H. MiR-155 promotes interleukin-1β-induced chondrocyte apoptosis and catabolic activity by targeting PIK3R1-mediated PI3K/Akt pathway. J. Cell. Mol. Med. 2020, 24, 8441–8451. [Google Scholar] [CrossRef]
- Li, L.; Jia, J.; Liu, X.; Yang, S.; Ye, S.; Yang, W.; Zhang, Y. MicroRNA-16-5p Controls Development of Osteoarthritis by Targeting SMAD3 in Chondrocytes. Curr. Pharm. Des. 2015, 21, 5160–5167. [Google Scholar] [CrossRef]
- Condrat, C.E.; Thompson, D.C.; Barbu, M.G.; Bugnar, O.L.; Boboc, A.; Cretoiu, D.; Suciu, N.; Cretoiu, S.M.; Voinea, S.C. miRNAs as Biomarkers in Disease: Latest Findings Regarding Their Role in Diagnosis and Prognosis. Cells 2020, 9, 276. [Google Scholar] [CrossRef] [Green Version]
- Chang, W.-W.; Wen, L.-Y.; Zhang, L.; Tong, X.; Jin, Y.-L.; Chen, G.-M. Association of rs2910164 in miR-146a with type 2 diabetes mellitus: A case–control and meta-analysis study. Front. Endocrinol. 2022, 13, 961635. [Google Scholar] [CrossRef]
- Rousseau, J.-C.; Millet, M.; Croset, M.; Sornay-Rendu, E.; Borel, O.; Chapurlat, R. Association of circulating microRNAs with prevalent and incident knee osteoarthritis in women: The OFELY study. Arthritis Res. Ther. 2020, 22, 2. [Google Scholar] [CrossRef] [Green Version]
- Lao, T.; Le, T. Data Integration Reveals the Potential Biomarkers of Circulating MicroRNAs in Osteoarthritis. Diagnostics 2021, 11, 412. [Google Scholar] [CrossRef]
- Tavallaee, G.; Rockel, J.S.; Lively, S.; Kapoor, M. MicroRNAs in Synovial Pathology Associated with Osteoarthritis. Front. Med. 2020, 7, 376. [Google Scholar] [CrossRef]
- Yu, X.-M.; Meng, H.-Y.; Yuan, X.-L.; Wang, Y.; Guo, Q.-Y.; Peng, J.; Wang, A.-Y.; Lu, S.-B. MicroRNAs’ Involvement in Osteoarthritis and the Prospects for Treatments. Evid.-Based Complement. Altern. Med. 2015, 2015, 236179. [Google Scholar] [CrossRef] [Green Version]
- Xie, Y.; Chen, W.; Zhao, M.; Xu, Y.; Yu, H.; Qin, J.; Li, H. Exploration of Exosomal miRNAs from Serum and Synovial Fluid in Arthritis Patients. Diagnostics 2022, 12, 239. [Google Scholar] [CrossRef]
- Kmiołek, T.; Paradowska-Gorycka, A. miRNAs as Biomarkers and Possible Therapeutic Strategies in Rheumatoid Arthritis. Cells 2022, 11, 452. [Google Scholar] [CrossRef]
- Zhang, M.; Lygrissea, K.; Wanga, J. Role of MicroRNA in Osteoarthritis. J. Arthritis 2017, 6, 239. [Google Scholar] [CrossRef]
- Mihanfar, A.; Shakouri, S.K.; Khadem-Ansari, M.H.; Fattahi, A.; Latifi, Z.; Nejabati, H.R.; Nouri, M. Exosomal miRNAs in osteoarthritis. Mol. Biol. Rep. 2020, 47, 4737–4748. [Google Scholar] [CrossRef]
- Pekáčová, A.; Baloun, J.; Švec, X.; Šenolt, L. Non-coding RNAs in diseases with a focus on osteoarthritis. Wiley Interdiscip. Rev. RNA 2022, e1756. [Google Scholar] [CrossRef]
- Wang, R.; Shiu, H.T.; Lee, W.Y.W. Emerging role of lncRNAs in osteoarthritis: An updated review. Front. Immunol. 2022, 13, 982773. [Google Scholar] [CrossRef]
- Kim, J. Dysregulated circular RNAs and their pathological implications in knee osteoarthritis: Potential novel therapeutic targets and diagnostic biomarkers. All Life 2022, 15, 23–49. [Google Scholar] [CrossRef]

| Upregulated miRs | Downregulated miRs |
|---|---|
| miR-195-5p | miR-335-5p |
| miR-9 | miR-146 |
| miR-25 | miR-149 |
| miR-98 | miR-30 |
| miR-27a | miR-337-3p |
| miR-34b | miR-140 |
| miR-30b | miR-146a |
| miR-140 | miR-27a-5p |
| miR-23a- 3p | miR-329 |
| miR-24-3p | miR-655 |
| miR-27a-3p | miR-708-3p |
| miR-27b-3p | miR-934 |
| miR-29c-3p | miR-101-5p |
| miR-34a-5p | miR-20 |
| miR-186-5p | miR-27b-3p |
| miR-483a-5p | miR-26a-5p |
| miR-103 | miR-27a |
| miR-146a | miR-107 |
| miR-218-5p | miR-222 |
| miR-449a | miR-320c |
| miR-324-5p | miR-26a |
| miR-1236 | miR-26b |
| miR-16-5p | miR-93-5p |
| miR-29b | miR-377-3p |
| miR-455-3p | miR-675-3p |
| miR-16 | |
| miR-132 | |
| miR-33b-3p | |
| miR-140-3p | |
| miR-671-3p | |
| miR-140-5p | |
| miR-142-5p | |
| miR-197 |
| Apoptosis | Induced Effect | Signaling Pathway | Reference No. |
|---|---|---|---|
| miR-195-5p | mRNA and protein levels of IL-1β, IL-6 and TNF-α were significantly enhanced | mir-195-5p activated the lPS-induced repression of the Wnt/β-catenin signaling pathway and activation of the nuclear factor (NF)-κB signaling pathway in ATDC5 cells | [5] |
| miR-335-5p | significantly reduces the expression of inflammatory factors (IL-1β, IL-6, and TNF-α) | significantly enhanced expression levels of the autophagy-related genes encoding the autophagy-related proteins Beclin-1, ATG5, and ATG7 | [6] |
| miR-9 | reduces the IL-1β mediated production of TNF-α and reduces basal and IL-1β-induced MMP13 protein release | PXR/RXR activation, G-protein coupled receptor (GPCR) signaling and Wnt/b-catenin signaling | [7] |
| miR-145 | miR-145 up-regulation decreases LPS-induced inflammatory injury in ATDC5 cells through SAL | functioned in LPS-induced injury by blocking NF-κB and p38MAPK signaling pathways | [9] |
| miR-337-3p | - | miR-337-3p in OA increased PTEN expression and thus PI3K/AKT signaling pathway was blocked by PTEN | [13] |
| miR-27b-3p | TNF-αand IL-6 levels repressed by the silencing of PVT1 were significantly enhanced by knockdown of miR-27b-3p in C28/I2 cells treated by IL-1β | miR-27b-3p decreased cell viability that was promoted by the silencing of PVT1 in IL-1β-treated C28/I2 cells, miR-27b-3p abrogated the promoting effect of PVT1 knockdown on autophagy in C28/I2 cells challenged by IL-1β | [20] |
| miR-125b | - | miR-125b noticeably alleviated the LPS+pc-THRIL-induced JAK1/STAT3 and NF-κB pathways activation | [23] |
| miR-26a-5p | - | - | [24] |
| miR-107 | - | miR-107 inhibited the activation of AKT/mTOR and NF-κB pathway | [25] |
| miR-218-5p | - | The expression of PIK3C2A, Akt, mTOR and S6 was downregulated, while 24 4EBP1 was upregulated | [26] |
| miR-222 | - | - | [27] |
| miR-140-5p | miR-140-5p reduced the expression of HMGB1 protein, p-AKT (Ser473) and p-PI3K in IL-1β-induced chondrocytes | inhibited the PI3K/ AKT signaling pathway and suppressed the progression of OA through targeting HMGB1 | [28] |
| miR-1236 | - | miR-1236 overexpression promotes OA by inhibiting proliferation and induces apoptosis of chondrocytes, partially through PIK3R3 repression. | [29] |
| miR-26b | - | - | [30] |
| miR-93-5p | Overexpression of miR-93-5p was found to significantly inhibit IL-1β–induced chondrocyte apoptosis. In general, IL-1β enhances the expression of matrix-degrading enzymes (MMP3 and MMP13), which in turn degrade the ECM | miR-93-5p exerted its actions in chondrocytes partially through suppression of TCF4 expression | [31] |
| miR-675-3p | miR-675-3p overexpression inhibited the IL-6 and IL-8 expression that was caused by IL-1b stimulation | miR-675-3p mimic markedly attenuated the increase in the GNG5 expression levels | [32] |
| miR-29b | - | - | [33] |
| miR-455-3p | miR-455-3p significantly inhibited the viability of CHON-001 cells IL-1β-induced apoptosis of CHON-001 was significantly increased by up-regulation of miR-455-3p | miR-455-3p directly regulated COL2A1 expression through binding its 3′UTR sequence | [34] |
| miR-142-5p | miR-142-5p overexpression resulted in significantly decreased levels of IL-1β and TNF-α and increased levels of IL-10 | miR-142-5p overexpression downregulated CXCR4 expression | [35] |
| Inflammation | Proliferation |
|---|---|
| miR-195-5p * | miR-195-5p * |
| miR-335-5p * | miR-335-5p * |
| miR-9 * | miR-140-5p * |
| miR-98 | miR-337-3p * |
| mir-146 | miR-26a-5p * |
| miR-145 * | miR-103 |
| miR-140-5p | miR-20 |
| miR-140-3p | miR-218-5p * |
| miR-146a | miR-320c |
| miR-155 | miR-1236 * |
| miR-27b-3p * | miR-26a |
| miR-125b * | miR-26b * |
| miR-27a | miR-377-3p |
| miR-140-5p * | miR-675-3p * |
| miR-20 | miR-29b * |
| miR-377-3p | miR-197 |
| miR-455-3p * | |
| miR-142-5p * | |
| miR-197 | |
| miR-23a-3p | |
| miR-24-3p | |
| miR-29c-3p | |
| miR-186-5p |
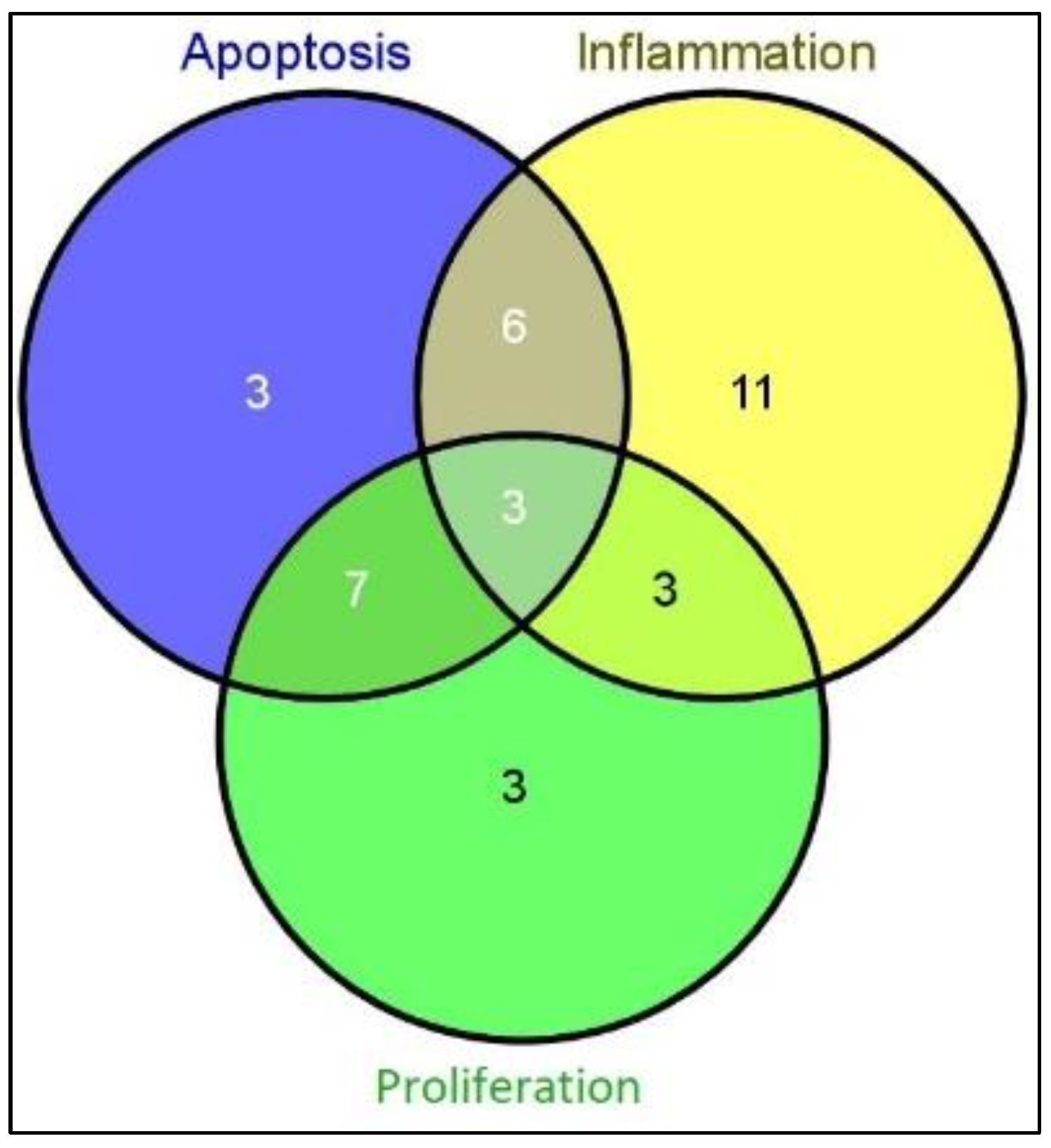
| Apoptosis (Total) | Inflammation (Total) | Proliferation (Total) | Total miRs in All Processes |
| 19 | 23 | 16 | 58 |
| Only apoptosis | Only inflammation | Only proliferation | Total miRs |
| 3 | 11 | 3 | 17 |
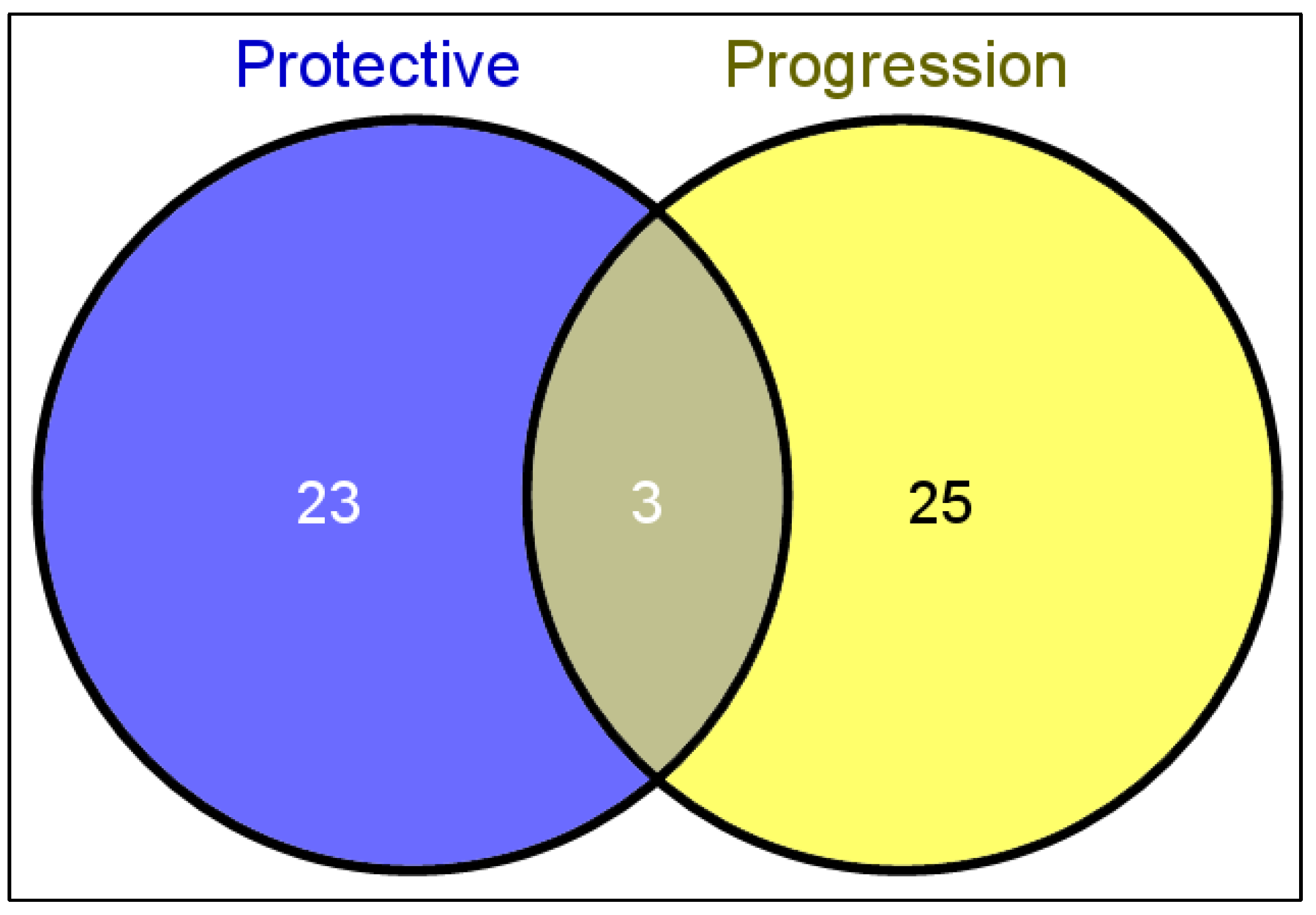
| miRs with Protective Roles in OA | Effective Role | miRs with Destructive Roles in OA | Effective Role |
|---|---|---|---|
| miR-195-5p | inhibits chondrocyte apoptosis and the inflammatory response | miR-9 | - |
| miR-335-5p | significantly increases cell viability and autophagy-related factors, and reduces inflammatory mediators | miR-145 | miR-145 down-regulation increases LPS-induced inflammatory injury |
| miR-337-3p | inhibits the apoptosis of OA chondrocytes | miR-140 | - |
| miR-140-5p | inhibits inflammation mediators and cartilage degradation and upregulates chondrogenic proteins | miR-30b | promotes cartilage degradation |
| miR-146a | reduces the expression levels of inflammatory genes and catabolic proteases in nucleus pulposus | miR-23a-3p | inflammatory stimulus |
| miR-27a-5p | - | miR-483-5p | severe cartilage damage, cartilage matrix degeneration, and increased chondrocyte hypertrophy |
| miR-20 | inhibits chondrocyte proliferation and autophagy | miR-125b | promotes inflammatory injury |
| miR-103 | - | miR-26a-5p | - |
| miR-222 | induces osteoblast proliferation and attenuates cardiomyocyte hypertrophy | miR-27a | increases the proliferation, apoptosis and inflammation of chondrocytes |
| miR-26a | regulates chondrocytes apoptosis | miR-107 | inhibits cell apoptosis and promotes autophagy in OA chondrocytes |
| miR-26b | regulates chondrocytes apoptosis | miR-140-5p | increases cartilage degradation |
| miR-93-5p | attenuates chondrocyte apoptosis and cartilage degradation | miR-146a | increases and promotes inflammation |
| miR-377-3p | alleviates chondrocyte apoptosis and cartilage degradation | miR-218-5p | promotes cartilage destruction |
| miR-16 | - | miR-320c | promotes cartilage destruction |
| miR-132 | - | miR-449a | promotes chondrocytes degradation |
| miR-33b-3p | - | miR-324-5p | - |
| miR-140-3p | protective against OA development via inhibition of inflammation and oxidative stress, enhancement of autophagy | miR-1236 | induces cell apoptosis in chondrocytes, suppresses cell proliferation. |
| miR-671-3p | - | miR-16-5p | - |
| miR-675-3p | inhibits apoptosis, matrix degradation and inflammation of chondrocytes. | miR-29b | facilitates chondrocyte apoptosis |
| miR-142-5p | attenuates chondrocyte apoptosis and cartilage degradation | miR-34a-5p | promotes cartilage damage |
| miR-197 | promotes chondrocyte proliferation, increases migration, and inhibits inflammation | miR-27b-3p | promotes cartilage damage |
| miR-329 | - | miR-455-3p | enhances chondrocytes apoptosis and inflammation |
| miR-655 | - | miR-98 | promotes inflammation |
| miR-708-3p | - | miR-140-3p | promotes inflammation |
| miR-934 | - | miR-24-3p | promotes inflammation |
| miR-146 | negative regulator of immune responses | miR-29c-3p | promotes inflammation |
| miR-186-5p | promotes inflammation | ||
| miR-155 | - |
| MiRs | Regulation (Across Other Studies) | Regulation (Found in Our Review) | Reference |
|---|---|---|---|
| miR-140 | ↑ | ↑↓ | [66] |
| miR-146a | ↑ | ↑↓ | [66] |
| miR-26a * | ↓ | ↓ | [66] |
| miR-26b * | ↓ | ↓ | [66] |
| miR-27a | ↓ | ↑↓ | [66] |
| miR-27b | ↓ | ↑↓ | [66] |
| miR-9 | ↓ | ↑ | [66] |
| mir-140 | ↑ | ↑↓ | [67] |
| miR-9 * | ↑ | ↑ | [67] |
| miR-26a | ↑ | ↓ | [67] |
| miR-146a | ↑ | ↑↓ | [67] |
| miR-27a | ↓ | ↑↓ | [67] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stanciugelu, S.I.; Homorogan, C.; Selaru, C.; Patrascu, J.M.; Patrascu, J.M.; Stoica, R.; Nitusca, D.; Marian, C. Osteoarthritis and microRNAs: Do They Provide Novel Insights into the Pathophysiology of This Degenerative Disorder? Life 2022, 12, 1914. https://doi.org/10.3390/life12111914
Stanciugelu SI, Homorogan C, Selaru C, Patrascu JM, Patrascu JM, Stoica R, Nitusca D, Marian C. Osteoarthritis and microRNAs: Do They Provide Novel Insights into the Pathophysiology of This Degenerative Disorder? Life. 2022; 12(11):1914. https://doi.org/10.3390/life12111914
Chicago/Turabian StyleStanciugelu, Stefan Iulian, Claudia Homorogan, Cosmin Selaru, Jenel Marian Patrascu, Jenel Marian Patrascu, Raymond Stoica, Diana Nitusca, and Catalin Marian. 2022. "Osteoarthritis and microRNAs: Do They Provide Novel Insights into the Pathophysiology of This Degenerative Disorder?" Life 12, no. 11: 1914. https://doi.org/10.3390/life12111914






