Size-Based Sorting and In Situ Clonal Expansion of Single Cells Using Microfluidics
Abstract
:1. Introduction
2. Materials and Methods
2.1. Device Design and Fabrication
2.2. Numerical Studies of the Flow Field
2.3. Numerical Studies of the Cell Deformation
2.4. Isolation of K562 Cells in Culture Media
2.5. Isolation of k562 Cells in Whole Blood
2.6. In Situ Clonal Expansion of Trapped Single Cells
2.7. Cell Lineage Tracking and Data Analysis
3. Results
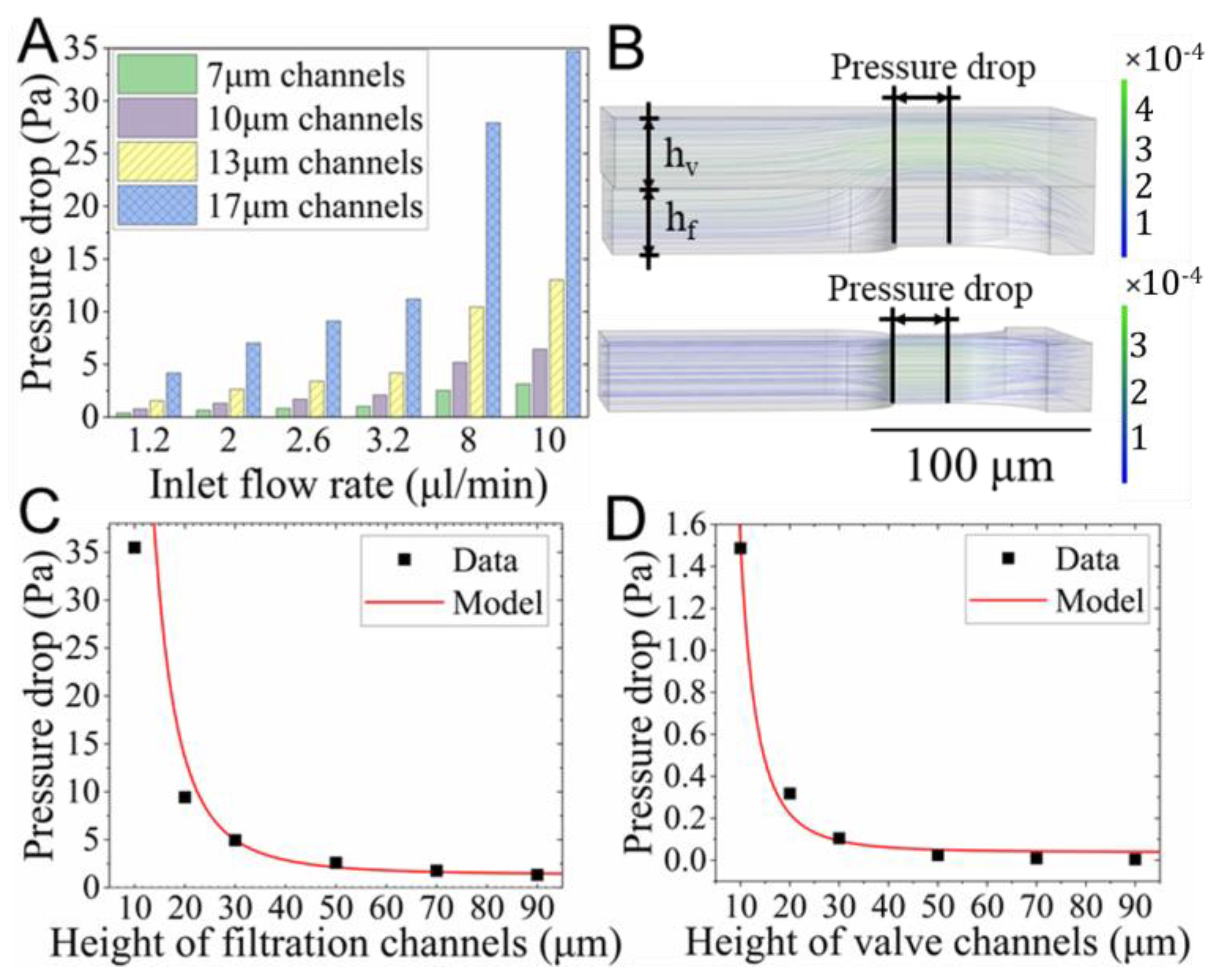
3.1. Numerical Studies of the Flow Field
3.2. Numerical Studies of the Cell Deformation
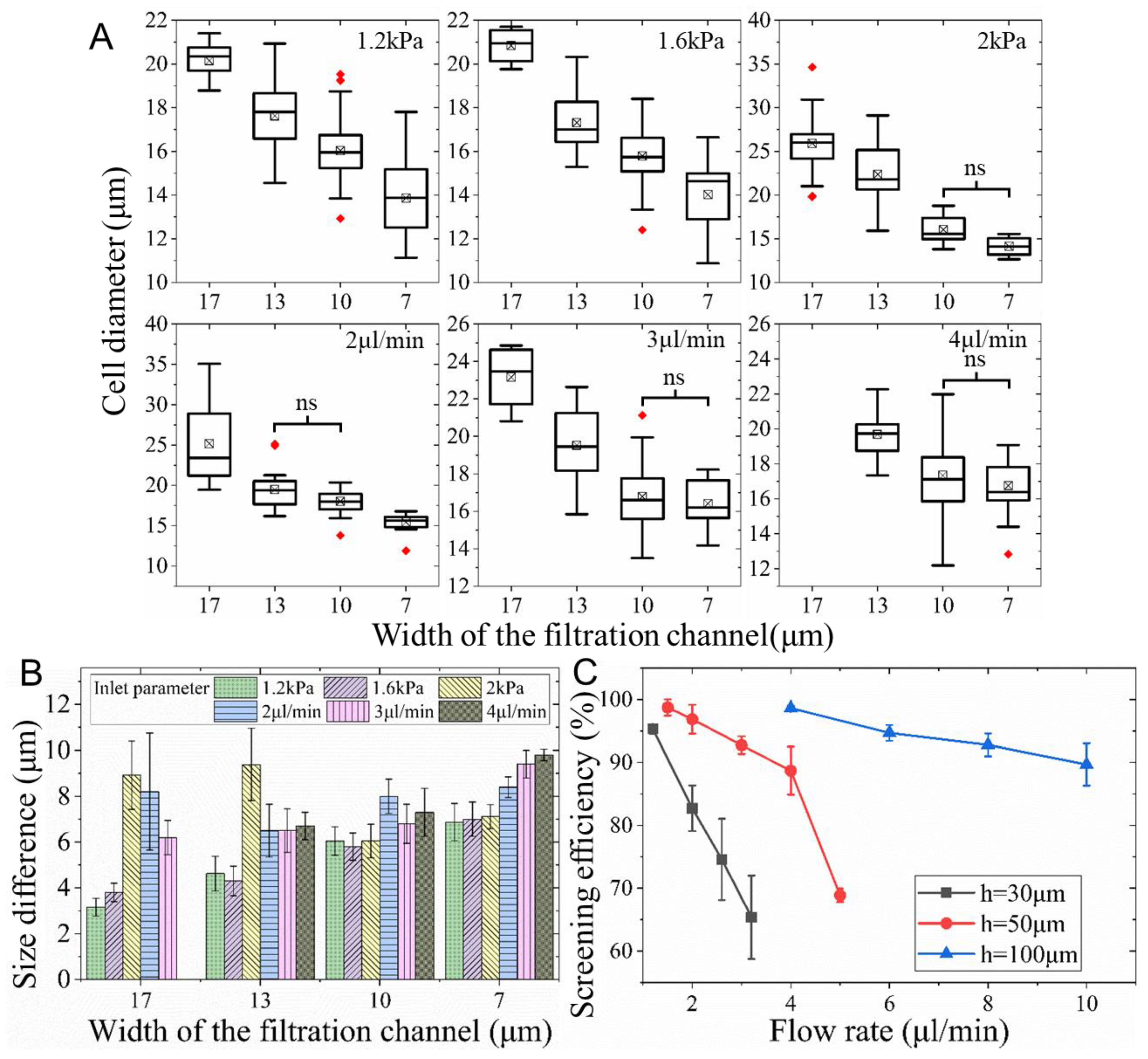
3.3. Isolation of K562 Cells from Culture Media
3.4. Screening of K562 Cells from Whole Blood
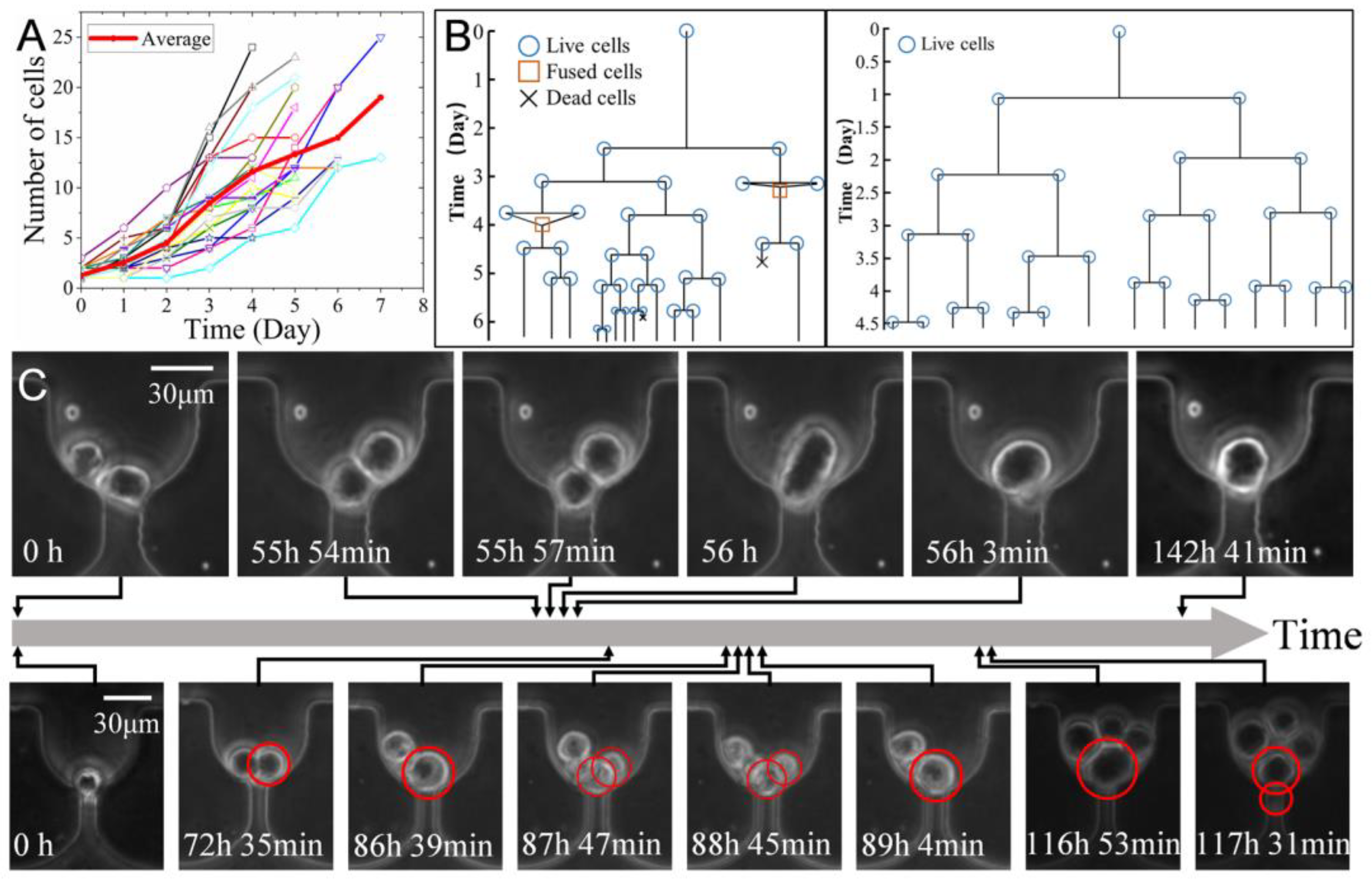
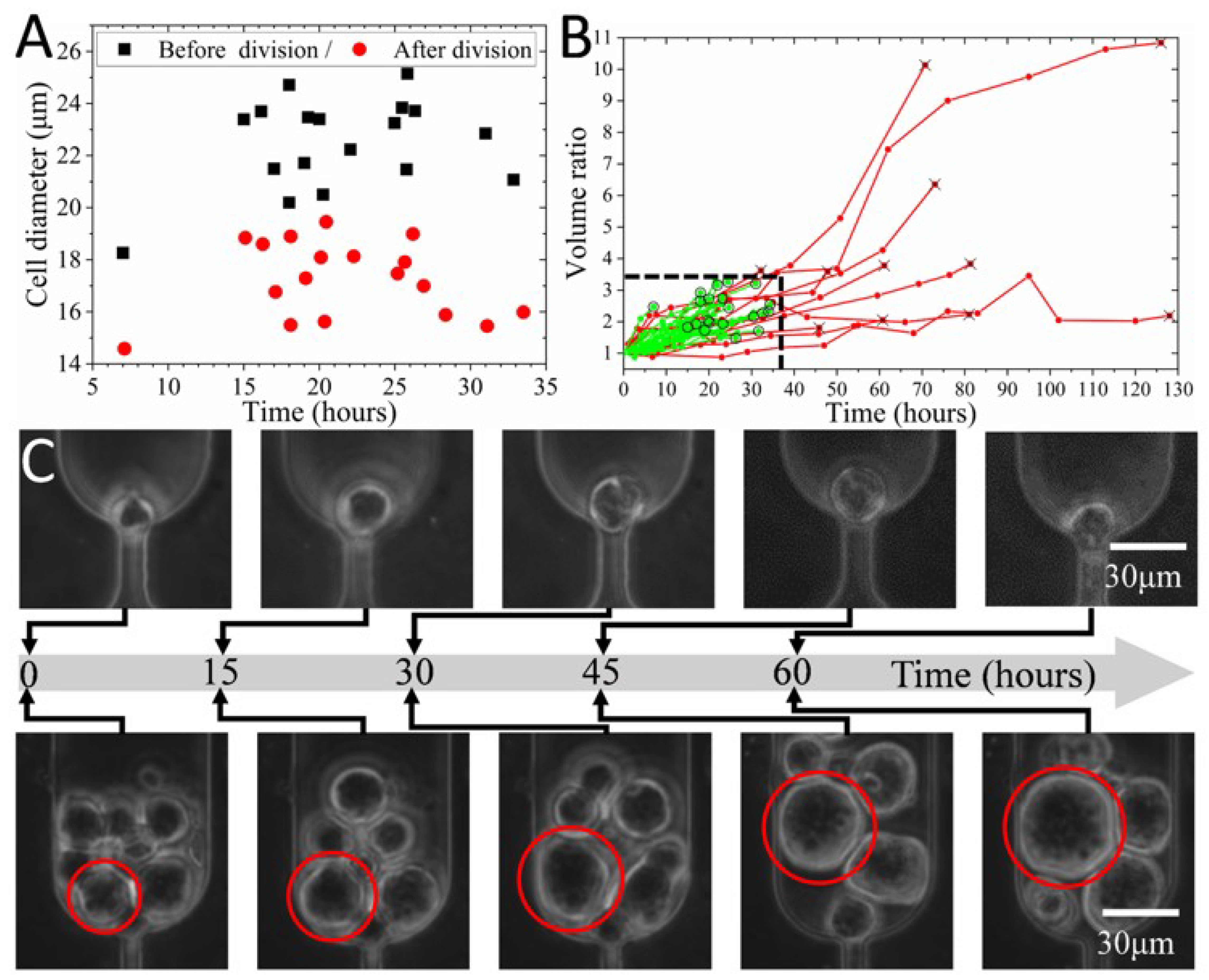
3.5. In Situ Clonal Expansion of Trapped Single Cells
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pang, L.; Ding, J.; Liu, X.-X.; Yuan, H.; Ge, Y.; Fan, J.; Fan, S.-K. Microstructure-based techniques for single-cell manipulation and analysis. TrAC Trends Anal. Chem. 2020, 129, 115940. [Google Scholar] [CrossRef]
- Spiller, D.G.; Wood, C.D.; Rand, D.A.; White, M.R.H. Measurement of single-cell dynamics. Nature 2010, 465, 736–745. [Google Scholar] [CrossRef] [PubMed]
- Titmarsh, D.M.; Chen, H.; Wolvetang, E.J.; Cooper-White, J.J. Arrayed cellular environments for stem cells and regenerative medicine. Biotechnol. J. 2013, 8, 167–179. [Google Scholar] [CrossRef] [PubMed]
- McKenna, A.; Gagnon, J.A. Recording development with single cell dynamic lineage tracing. Development 2019, 146, dev169730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anggraini, D.; Ota, N.; Shen, Y.; Tang, T.; Tanaka, Y.; Hosokawa, Y.; Li, M.; Yalikun, Y. Recent advances in microfluidic devices for single-cell cultivation: Methods and applications. Lab Chip 2022, 22, 1438–1468. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Sun, J.; Wolvetang, E.; Cooper-White, J. High-throughput, deterministic single cell trapping and long-term clonal cell culture in microfluidic devices. Lab Chip 2015, 15, 1072–1083. [Google Scholar] [CrossRef]
- Schreier, S.; Triampo, W. The Blood Circulating Rare Cell Population. What Is It and What Is It Good for? Cells 2020, 9, 790. [Google Scholar] [CrossRef] [Green Version]
- Franken, B.; de Groot, M.R.; Mastboom, W.J.B.; Vermes, I.; van der Palen, J.; Tibbe, A.G.J.; Terstappen, L.W.M.M. Circulating tumor cells, disease recurrence and survival in newly diagnosed breast cancer. Breast Cancer Res. 2012, 14, R133. [Google Scholar] [CrossRef] [Green Version]
- Glia, A.; Deliorman, M.; Sukumar, P.; Janahi, F.K.; Samara, B.; Brimmo, A.T.; Qasaimeh, M.A. Herringbone Microfluidic Probe for Multiplexed Affinity-Capture of Prostate Circulating Tumor Cells. Adv. Mater. Technol. 2021, 6, 2100053. [Google Scholar] [CrossRef]
- Shi, J.; Tong, L.; Tong, W.; Chen, H.; Lan, M.; Sun, X.; Zhu, Y. Current progress in long-term and continuous cell metabolite detection using microfluidics. TrAC Trends Anal. Chem. 2019, 117, 263–279. [Google Scholar] [CrossRef]
- Khoo, B.L.; Grenci, G.; Lim, Y.B.; Lee, S.C.; Han, J.; Lim, C.T. Expansion of patient-derived circulating tumor cells from liquid biopsies using a CTC microfluidic culture device. Nat. Protoc. 2018, 13, 34–58. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Shiratsuchi, H.; Lin, J.; Chen, G.; Reddy, R.M.; Azizi, E.; Fouladdel, S.; Chang, A.C.; Lin, L.; Jiang, H.; et al. Expansion of CTCs from early stage lung cancer patients using a microfluidic co-culture model. Oncotarget 2014, 5, 12383–12397. [Google Scholar] [CrossRef] [Green Version]
- Lin, D.; Chen, X.; Liu, Y.; Lin, Z.; Luo, Y.; Fu, M.; Yang, N.; Liu, D.; Cao, J. Microgel Single-Cell Culture Arrays on a Microfluidic Chip for Selective Expansion and Recovery of Colorectal Cancer Stem Cells. Anal. Chem. 2021, 93, 12628–12638. [Google Scholar] [CrossRef] [PubMed]
- García Alonso, D.; Yu, M.; Qu, H.; Ma, L.; Shen, F. Advances in Microfluidics-Based Technologies for Single Cell Culture. Adv. Biosyst. 2019, 3, 1900003. [Google Scholar] [CrossRef] [PubMed]
- Yin, D.; Shi, A.; Zhou, B.; Wang, M.; Xu, G.; Shen, M.; Zhu, X.; Shi, X. Efficient Capture and Separation of Cancer Cells Using Hyaluronic Acid-Modified Magnetic Beads in a Microfluidic Chip. Langmuir 2022, 38, 11080–11086. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wu, L.; Nie, W.; Huang, L.; Zhang, J.; Li, F.; Xie, H.-Y. Biomimetic Microfluidic System for Fast and Specific Detection of Circulating Tumor Cells. Anal. Chem. 2019, 91, 15726–15731. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.-T.; Hadlock, T.; Lo, T.-W.; Purcell, E.; Mutukuri, A.; Fouladdel, S.; Raguera, M.D.S.; Fairbairn, H.; Murlidhar, V.; Durham, A.; et al. Dual-Isolation and Profiling of Circulating Tumor Cells and Cancer Exosomes from Blood Samples with Melanoma Using Immunoaffinity-Based Microfluidic Interfaces. Adv. Sci. 2020, 7, 2001581. [Google Scholar] [CrossRef]
- He, G.; Yang, C.; Feng, J.; Wu, J.; Zhou, L.; Wen, R.; Huang, S.; Wu, Q.; Liu, F.; Chen, H.-J.; et al. Hierarchical Spiky Microstraws-Integrated Microfluidic Device for Efficient Capture and In Situ Manipulation of Cancer Cells. Adv. Funct. Mater. 2019, 29, 1806484. [Google Scholar] [CrossRef]
- Chen, H.; Rosengarten, G.; Li, M.; Nordon, R.E. Design of microdevices for long-term live cell imaging. J. Micromech. Microeng. 2012, 22, 065033. [Google Scholar] [CrossRef]
- Swennenhuis, J.F.; Tibbe, A.G.J.; Stevens, M.; Katika, M.R.; van Dalum, J.; Duy Tong, H.; van Rijn, C.J.M.; Terstappen, L.W.M.M. Self-seeding microwell chip for the isolation and characterization of single cells. Lab Chip 2015, 15, 3039–3046. [Google Scholar] [CrossRef]
- Nguyen, A.; Khoo, W.H.; Moran, I.; Croucher, P.I.; Phan, T.G. Single Cell RNA Sequencing of Rare Immune Cell Populations. Front. Immunol. 2018, 9, 1553. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Jin, F.; Zhou, M.; Jiang, Y. Recent advances in single cell manipulation and biochemical analysis on microfluidics. Analyst 2019, 144, 766–781. [Google Scholar] [CrossRef] [PubMed]
- Lagus, T.P.; Edd, J.F. A review of the theory, methods and recent applications of high-throughput single-cell droplet microfluidics. J. Phys. D Appl. Phys. 2013, 46, 114005. [Google Scholar] [CrossRef]
- Zhu, Z.; Yang, C.J. Hydrogel Droplet Microfluidics for High-Throughput Single Molecule/Cell Analysis. Acc. Chem. Res. 2017, 50, 22–31. [Google Scholar] [CrossRef]
- Tan, W.-H.; Takeuchi, S. A trap-and-release integrated microfluidic system for dynamic microarray applications. Proc. Natl. Acad. Sci. USA 2007, 104, 1146–1151. [Google Scholar] [CrossRef] [Green Version]
- Kobel, S.; Valero, A.; Latt, J.; Renaud, P.; Lutolf, M. Optimization of microfluidic single cell trapping for long-term on-chip culture. Lab Chip 2010, 10, 857–863. [Google Scholar] [CrossRef] [Green Version]
- Hong, S.; Pan, Q.; Lee, L.P. Single-cell level co-culture platform for intercellular communication. Integr. Biol. 2012, 4, 374–380. [Google Scholar] [CrossRef]
- Jin, S.H.; Jang, S.-C.; Lee, B.; Jeong, H.-H.; Jeong, S.-G.; Lee, S.S.; Kim, K.P.; Lee, C.-S. Monitoring of chromosome dynamics of single yeast cells in a microfluidic platform with aperture cell traps. Lab Chip 2016, 16, 1358–1365. [Google Scholar] [CrossRef]
- Di Carlo, D.; Aghdam, N.; Lee, L.P. Single-Cell Enzyme Concentrations, Kinetics, and Inhibition Analysis Using High-Density Hydrodynamic Cell Isolation Arrays. Anal. Chem. 2006, 78, 4925–4930. [Google Scholar] [CrossRef]
- Dura, B.; Dougan, S.K.; Barisa, M.; Hoehl, M.M.; Lo, C.T.; Ploegh, H.L.; Voldman, J. Profiling lymphocyte interactions at the single-cell level by microfluidic cell pairing. Nat. Commun. 2015, 6, 5940. [Google Scholar] [CrossRef]
- Liu, P.Y.; Chin, L.K.; Ser, W.; Ayi, T.C.; Yap, P.H.; Bourouina, T.; Leprince-Wang, Y. An optofluidic imaging system to measure the biophysical signature of single waterborne bacteria. Lab Chip 2014, 14, 4237–4243. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.J.; Yobas, L.; Lee, G.Y.H.; Ong, C.N.; Lim, C.T. Microdevice for the isolation and enumeration of cancer cells from blood. Biomed. Microdevices 2009, 11, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Chou, C.-K.; Xia, X.; Hung, M.-C.; Qin, L. Block-Cell-Printing for live single-cell printing. Proc. Natl. Acad. Sci. USA 2014, 111, 2948–2953. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mu, H.-Y.; Ou, Y.-C.; Chuang, H.-N.; Lu, T.-J.; Jhan, P.-P.; Hsiao, T.-H.; Huang, J.-H. Triple Selection Strategy for In Situ Labeling of Circulating Tumor Cells with High Purity and Viability toward Preclinical Personalized Drug Sensitivity Analysis. Adv. Biosyst. 2020, 4, 2000013. [Google Scholar] [CrossRef]
- Rosenberg, R.; Gertler, R.; Friederichs, J.; Fuehrer, K.; Dahm, M.; Phelps, R.; Thorban, S.; Nekarda, H.; Siewert, J.R. Comparison of two density gradient centrifugation systems for the enrichment of disseminated tumor cells in blood. Cytometry 2002, 49, 150–158. [Google Scholar] [CrossRef]
- Gascoyne, P.; Shim, S. Isolation of Circulating Tumor Cells by Dielectrophoresis. Cancers 2014, 6, 545–579. [Google Scholar] [CrossRef] [Green Version]
- Pødenphant, M.; Ashley, N.; Koprowska, K.; Mir, K.U.; Zalkovskij, M.; Bilenberg, B.; Bodmer, W.; Kristensen, A.; Marie, R. Separation of cancer cells from white blood cells by pinched flow fractionation. Lab Chip 2015, 15, 4598–4606. [Google Scholar] [CrossRef] [Green Version]
- Ribeiro-Samy, S.; Oliveira, M.I.; Pereira-Veiga, T.; Muinelo-Romay, L.; Carvalho, S.; Gaspar, J.; Freitas, P.P.; López-López, R.; Costa, C.; Diéguez, L. Fast and efficient microfluidic cell filter for isolation of circulating tumor cells from unprocessed whole blood of colorectal cancer patients. Sci. Rep. 2019, 9, 8032. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Chen, C.; Bai, S.; Gao, Y.; Metcalfe, G.; Cheng, W.; Zhu, Y. Multiplexed detection of cancer biomarkers using a microfluidic platform integrating single bead trapping and acoustic mixing techniques. Nanoscale 2018, 10, 20196–20206. [Google Scholar] [CrossRef]
- Luo, Z.Y.; Xu, F.; Lu, T.J.; Bai, B.F. Direct Numerical Simulation of Single Leukocyte Deformation in Microchannel Flow for Disease Diagnosis. J. Med. Syst. 2011, 35, 869–876. [Google Scholar] [CrossRef]
- Zhang, X.; Chen, X.; Tan, H. On the thin-film-dominated passing pressure of cancer cell squeezing through a microfluidic CTC chip. Microfluid. Nanofluid. 2017, 21, 146. [Google Scholar] [CrossRef]
- Jin, Q.; Verdier, C.; Singh, P.; Aubry, N.; Chotard-Ghodsnia, R.; Duperray, A. Migration and deformation of leukocytes in pressure driven flows. Mech. Res. Commun. 2007, 34, 411–422. [Google Scholar] [CrossRef] [Green Version]
- Preetha, A.; Huilgol, N.; Banerjee, R. Interfacial properties as biophysical markers of cervical cancer. Biomed Pharm. 2005, 59, 491–497. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Peng, Z.; Lin, S.-C.S.; Geri, M.; Li, S.; Li, P.; Chen, Y.; Dao, M.; Suresh, S.; Huang, T.J. Cell separation using tilted-angle standing surface acoustic waves. Proc. Natl. Acad. Sci. USA 2014, 111, 12992–12997. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altay, R.; Yapici, M.K.; Koşar, A. A Hybrid Spiral Microfluidic Platform Coupled with Surface Acoustic Waves for Circulating Tumor Cell Sorting and Separation: A Numerical Study. Biosensors 2022, 12, 171. [Google Scholar] [CrossRef]
- Zhou, C.; Yue, P.; Feng, J.J. Simulation of Neutrophil Deformation and Transport in Capillaries using Newtonian and Viscoelastic Drop Models. Ann. Biomed. Eng. 2007, 35, 766–780. [Google Scholar] [CrossRef]
- Cornwell, J.A.; Li, J.; Mahadevan, S.; Draper, J.S.; Joun, G.L.; Zoellner, H.; Asli, N.S.; Harvey, R.P.; Nordon, R.E. TrackPad: Software for semi-automated single-cell tracking and lineage annotation. SoftwareX 2020, 11, 100440. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Meth. 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Chen, Z.; Zhu, Y.; Xu, D.; Alam, M.M.; Shui, L.; Chen, H. Cell elasticity measurement using a microfluidic device with real-time pressure feedback. Lab Chip 2020, 20, 2343–2353. [Google Scholar] [CrossRef]
- Chen, H.; Cao, B.; Sun, B.; Cao, Y.; Yang, K.; Lin, Y.-S. Highly-sensitive capture of circulating tumor cells using micro-ellipse filters. Sci. Rep. 2017, 7, 610. [Google Scholar] [CrossRef]
- O’Connell, K.E.; Mikkola, A.M.; Stepanek, A.M.; Vernet, A.; Hall, C.D.; Sun, C.C.; Yildirim, E.; Staropoli, J.F.; Lee, J.T.; Brown, D.E. Practical murine hematopathology: A comparative review and implications for research. Comp. Med. 2015, 65, 96–113. [Google Scholar] [PubMed]
- Rutherford, T.; Clegg, J.B.; Higgs, D.R.; Jones, R.W.; Thompson, J.; Weatherall, D.J. Embryonic erythroid differentiation in the human leukemic cell line K562. Proc. Natl. Acad. Sci. USA 1981, 78, 348–352. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dura, B.; Liu, Y.; Voldman, J. Deformability-based microfluidic cell pairing and fusion. Lab Chip 2014, 14, 2783–2790. [Google Scholar] [CrossRef] [PubMed]
- Karatekin, E.; Rothman, J.E. Fusion of single proteoliposomes with planar, cushioned bilayers in microfluidic flow cells. Nat. Protoc. 2012, 7, 903–920. [Google Scholar] [CrossRef]
- Leroy, H.; Han, M.; Woottum, M.; Bracq, L.; Bouchet, J.; Xie, M.; Benichou, S. Virus-Mediated Cell-Cell Fusion. Int. J. Mol. Sci. USA 2020, 21, 9644. [Google Scholar] [CrossRef]
- Rems, L.; Ušaj, M.; Kandušer, M.; Reberšek, M.; Miklavčič, D.; Pucihar, G. Cell electrofusion using nanosecond electric pulses. Sci. Rep. 2013, 3, 3382. [Google Scholar] [CrossRef] [Green Version]
- Migita, S.; Funakoshi, K.; Tsuya, D.; Yamazaki, T.; Taniguchi, A.; Sugimoto, Y.; Hanagata, N.; Ikoma, T. Cell cycle and size sorting of mammalian cells using a microfluidic device. Anal. Methods 2010, 2, 657. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, H.; Meng, H.; Chen, Z.; Wang, T.; Chen, C.; Zhu, Y.; Jin, J. Size-Based Sorting and In Situ Clonal Expansion of Single Cells Using Microfluidics. Biosensors 2022, 12, 1100. https://doi.org/10.3390/bios12121100
Chen H, Meng H, Chen Z, Wang T, Chen C, Zhu Y, Jin J. Size-Based Sorting and In Situ Clonal Expansion of Single Cells Using Microfluidics. Biosensors. 2022; 12(12):1100. https://doi.org/10.3390/bios12121100
Chicago/Turabian StyleChen, Huaying, Haixu Meng, Zhenlin Chen, Tong Wang, Chuanpin Chen, Yonggang Zhu, and Jing Jin. 2022. "Size-Based Sorting and In Situ Clonal Expansion of Single Cells Using Microfluidics" Biosensors 12, no. 12: 1100. https://doi.org/10.3390/bios12121100




