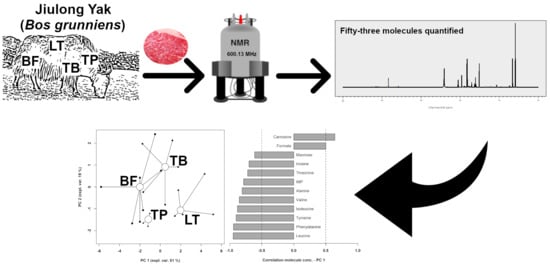
An Untargeted Metabolomics Investigation of Jiulong Yak (Bos grunniens) Meat by 1H-NMR
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Metabolome Analysis
2.3. Statistical Analysis
3. Results
3.1. 1H-NMR Spectra of Yak Raw Meat
3.2. Comparison among TB, BF and LT Muscles
3.3. Comparison between LT and TP Muscles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zi, X.D.; Zhong, G.H.; Wen, Y.L.; Zhong, J.C.; Liu, C.L.; Ni, Y.A.; Yezi, Y.H.; Ashi, M.G. Growth performance, carcass composition and meat quality of Jiulong-yak (Bos grunniens). Asian-Australas. J. Anim. Sci. 2004, 17, 410–414. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, J.; Zhang, D.; Hasegawa, M.; Liu, W.; Zhang, J.; Ma, T.; Cao, C.; Lenstra, J.A.; Zang, X.; et al. The yak genome and adaptation to life at high altitude. Nat. Genet. 2012, 44, 946–949. [Google Scholar]
- Zhang, L.; Wu, J.; Wang, X.; Liu, B.; Ma, B. Isolation of metallothionein genes and in silico structural characterization of their proteins Using molecular modeling from Yak (Bos grunniens). Biochem. Genet. 2012, 50, 585–599. [Google Scholar] [CrossRef]
- Shang, Z.; Liang, J.B.; Long, R.; Wang, H.; Ding, L.; Guo, X. Comparison of Nitrogen Metabolism in Yak (Bos grunniens) and Indigenous Cattle (Bos taurus) on the Qinghai-Tibetan Plateau. Asian-Australas. J. Anim. Sci. 2011, 24, 766–773. [Google Scholar]
- Zhang, Q.; Gong, J.; Wang, X.; Wu, X.; Li, Y.; Ma, Y.; Zhang, Y.; Zhao, X. Molecular cloning, bioinformatics analysis and expression of insulin-like growth factor 2 from tianzhu white yak, Bos grunniens. Int. J. Mol. Sci. 2014, 15, 504–524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, Y.Q.; Wang, G.S.; Feng, J.; Huang, J.Q.; Xu, Y.O.; Jin, S.Y.; Li, Y.P.; Jiang, Z.R.; Zheng, Y.C. Comparison of enzyme activities and gene expression profiling between yak and bovine skeletal muscles. Livest. Sci. 2011, 135, 93–97. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, B.; Yu, Q.; Ji, Q.; Xie, P.; Li, H.; Wang, L.; Zhou, Y.; Li, Y.; Huang, C.; et al. The breed and sex effect on the carcass size performance and meat quality of yak in different muscles. Korean J. Food Sci. Anim. Resour. 2016, 36, 223–229. [Google Scholar] [CrossRef] [Green Version]
- Shi, X.X.; Yu, Q.L.; Han, L.; Tian, J.C.H.; Zhang, L.; Zhang, Y.B.; Guo, Z.H.B. Changes in meat quality characteristics and calpains activities in Gannan Yak (Bos grunniens) meat during post mortem ageing. J. Anim. Vet. Adv. 2013, 12, 363–368. [Google Scholar]
- Rajagopal, K.; Oommen, G.T. Myofibril Fragmentation Index as an Immediate Postmortem Predictor of Buffalo Meat Tenderness. J. Food Process. Preserv. 2015, 39, 1166–1171. [Google Scholar] [CrossRef]
- Jia, J.; Zhang, Q.; Chen, Q.; Zhang, H.; Lin, F.; Zhao, J. Differential expression of proteins in Datong Yak and Chaidamu yellow cattle longissimus lumborum muscles and relation to meat water holding capacity. Kafkas Univ. Vet. Fak. Derg. 2018, 24, 691–700. [Google Scholar]
- Picard, B.; Gagaoua, M. Meta-proteomics for the discovery of protein biomarkers of beef tenderness: An overview of integrated studies. Food Res. Int. 2020, 127, 108739. [Google Scholar] [CrossRef] [PubMed]
- Laghi, L.; Picone, G.; Capozzi, F. Nuclear magnetic resonance for foodomics beyond food analysis. TrAC Trends Anal. Chem. 2014, 59, 93–102. [Google Scholar] [CrossRef]
- Kodani, Y.; Miyakawa, T.; Komatsu, T.; Tanokura, M. NMR-based metabolomics for simultaneously evaluating multiple determinants of primary beef quality in Japanese Black cattle. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ercolini, D.; Ferrocino, I.; Nasi, A.; Ndagijimana, M.; Vernocchi, P.; La Storia, A.; Laghi, L.; Mauriello, G.; Guerzoni, M.E.; Villani, F. Monitoring of microbial metabolites and bacterial diversity in beef stored under different packaging londitions. Appl. Environ. Microbiol. 2011, 77, 7372–7381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castejón, D.; García-Segura, J.M.; Escudero, R.; Herrera, A.; Cambero, M.I. Metabolomics of meat exudate: Its potential to evaluate beef meat conservation and aging. Anal. Chim. Acta 2015, 901, 1–11. [Google Scholar] [CrossRef]
- Graham, S.F.; Kennedy, T.; Chevallier, O.; Gordon, A.; Farmer, L.; Elliott, C.; Moss, B. The application of NMR to study changes in polar metabolite concentrations in beef longissimus dorsi stored for different periods post mortem. Metabolomics 2010, 6, 395–404. [Google Scholar] [CrossRef]
- Ritota, M.; Casciani, L.; Failla, S.; Valentini, M. HRMAS-NMR spectroscopy and multivariate analysis meat characterisation. Meat Sci. 2012, 92, 754–761. [Google Scholar] [CrossRef]
- Jung, Y.; Lee, J.; Kwon, J.; Lee, K.S.; Ryu, D.H.; Hwang, G.S. Discrimination of the geographical origin of beef by 1H NMR-based metabolomics. J. Agric. Food Chem. 2010, 58, 10458–10466. [Google Scholar] [CrossRef]
- Shintu, L.; Caldarelli, S.; Franke, B.M. Pre-selection of potential molecular markers for the geographic origin of dried beef by HR-MAS NMR spectroscopy. Meat Sci. 2007, 76, 700–707. [Google Scholar] [CrossRef]
- Zanardi, E.; Caligiani, A.; Palla, L.; Mariani, M.; Ghidini, S.; Di Ciccio, P.A.; Palla, G.; Ianieri, A. Metabolic profiling by 1H NMR of ground beef irradiated at different irradiation doses. Meat Sci. 2015, 103, 83–89. [Google Scholar] [CrossRef]
- Luo, X.L.; Tong, Z.B.; Wei, Y.P.; Zhao, X.Q. Meat characteristics of Qinghai yak and semi-wild yak. Anim. Sci. J. 2006, 77, 230–234. [Google Scholar] [CrossRef]
- Wiener, G.; Han, J.; Long, R. The Yak, 2nd ed.; FAO Regional Office for Asia and the Pacific, Ed.; FAO Regional Office for Asia and the Pacific: Bangkok, Thailand, 2003; ISBN 92-5-104965-3. [Google Scholar]
- Zhu, C.; Faillace, V.; Laus, F.; Bazzano, M.; Laghi, L. Characterization of trotter horses urine metabolome by means of proton nuclear magnetic resonance spectroscopy. Metabolomics 2018, 14, 106. [Google Scholar] [CrossRef] [PubMed]
- Kneen, M.A.; Annegarn, H.J. Algorithm for fitting XRF, SEM and PIXE X-ray spectra backgrounds. Nucl. Instrum. Methods Phys. Res. Sect. B Beam Interact. Mater. At. 1996, 109–110, 209–213. [Google Scholar] [CrossRef]
- Liland, K.H.; Almøy, T.; Mevik, B.H. Optimal choice of baseline correction for multivariate calibration of spectra. Appl. Spectrosc. 2010, 64, 1007–1016. [Google Scholar] [CrossRef]
- Dieterle, F.; Ross, A.; Schlotterbeck, G.; Senn, H. Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in1H NMR metabonomics. Anal. Chem. 2006, 78, 4281–4290. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing. Available online: https://www.gbif.org/tool/81287/r-a-language-and-environment-for-statistical-computing (accessed on 15 March 2020).
- Box, G.E.P.; Cox, D.R. An Analysis of Transformations. J. R. Stat. Soc. Ser. B 2018, 26, 211–243. [Google Scholar] [CrossRef]
- Abadie, V.; Kim, S.M.; Lejeune, T.; Palanski, B.A.; Ernest, J.D.; Tastet, O.; Voisine, J.; Discepolo, V.; Marietta, E.V.; Hawash, M.B.F.; et al. IL-15, gluten and HLA-DQ8 drive tissue destruction in coeliac disease. Nature 2020, 578, 600–604. [Google Scholar] [CrossRef]
- Wang, H.; Zhong, H.; Hou, R.; Ayala, J.; Liu, G.; Yuan, S.; Yan, Z.; Zhang, W.; Liu, Y.; Cai, K.; et al. A Diet Diverse in Bamboo Parts is Important for Giant Panda (Ailuropoda melanoleuca) Metabolism and Health. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Hubert, M.; Rousseeuw, P.J.; Vanden Branden, K. ROBPCA: A new approach to robust principal component analysis. Technometrics 2005, 47, 64–79. [Google Scholar] [CrossRef]
- Li, C.; Soufan, O.; Chong, J.; Xia, J.; Bourque, G.; Li, S.; Caraus, I.; Wishart, D.S. MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018, 46, W486–W494. [Google Scholar]
- Marcolini, E.; Babini, E.; Bordoni, A.; Di Nunzio, M.; Laghi, L.; Maczó, A.; Picone, G.; Szerdahelyi, E.; Valli, V.; Capozzi, F. Bioaccessibility of the Bioactive Peptide Carnosine during in Vitro Digestion of Cured Beef Meat. J. Agric. Food Chem. 2015, 63, 4973–4978. [Google Scholar] [CrossRef] [PubMed]
- Bordoni, A.; Picone, G.; Babini, E.; Vignali, M.; Danesi, F.; Valli, V.; Di Nunzio, M.; Laghi, L.; Capozzi, F. NMR comparison of in vitro digestion of Parmigiano Reggiano cheese aged 15 and 30 months. Magn. Reson. Chem. 2011, 49, S61–S70. [Google Scholar] [CrossRef] [PubMed]
- Salek, R.M.; Neumann, S.; Schober, D.; Hummel, J.; Billiau, K.; Kopka, J.; Correa, E.; Reijmers, T.; Rosato, A.; Tenori, L.; et al. COordination of Standards in MetabOlomicS (COSMOS): Facilitating integrated metabolomics data access. Metabolomics 2015, 11, 1587–1597. [Google Scholar] [CrossRef] [PubMed]
- Lawrie, R.A.; Pomeroy, R.W.; Williams, D.R. Studies in the muscles of meat animals. IV. Comparative composition of muscles from ‘doppelender’ and normal sibling heifers. J. Agric. Sci. 1964, 62, 89–92. [Google Scholar] [CrossRef]
- Rao, M.V.; Gault, N.F.S. The influence of fibre-type composition and associated biochemical characteristics on the acid buffering capacities of several beef muscles. Meat Sci. 1989, 26, 5–18. [Google Scholar] [CrossRef]
- Flores, M.; Alasnier, C.; Aristoy, M.C.; Navarro, J.L.; Gandemer, G.; Toldrá, F. Activity of aminopeptidase and lipolytic enzymes in five skeletal muscles with various oxidative patterns. J. Sci. Food Agric. 1996, 70, 127–130. [Google Scholar] [CrossRef]
- Suzuki, Y.; Ito, O.; Mukai, N.; Takahashi, H.; Takamatsu, K. High level of skeletal muscle carnosine contributes to the latter half of exercise performance during 30-s maximal cycle ergometer sprinting. Jpn. J. Physiol. 2002, 52, 199–205. [Google Scholar] [CrossRef] [Green Version]
- Aristoy, M.C.; Toldrá, F. Histidine dipeptides HPLC-based test for the detection of mammalian origin proteins in feeds for ruminants. Meat Sci. 2004, 67, 211–217. [Google Scholar] [CrossRef]
- Mora, L.; Sentandreu, M.Á.; Toldrá, F. Contents of creatine, creatinine and carnosine in porcine muscles of different metabolic types. Meat Sci. 2008, 79, 709–715. [Google Scholar] [CrossRef]
- Ma, D.; Kim, Y.H.B.; Cooper, B.; Oh, J.H.; Chun, H.; Choe, J.H.; Schoonmaker, J.P.; Ajuwon, K.; Min, B. Metabolomics Profiling to Determine the Effect of Postmortem Aging on Color and Lipid Oxidative Stabilities of Different Bovine Muscles. J. Agric. Food Chem. 2017, 65, 6708–6716. [Google Scholar] [CrossRef]
- Odessey, R.; Khairallah, E.A.; Goldbert, A.L. Origin and possible significance of alanine production by skeletal muscle. J. Biol. Chem. 1974, 249, 7623–7629. [Google Scholar] [PubMed]
- Goldstein, L.; Newsholme, E.A. The formation of alanine from amino acids in diaphragm muscle of the rat. Biochem. J. 1976, 154, 555–558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boldyrev, A.; Abe, H. Metabolic transformation of neuropeptide carnosine modifies its biological activity. Cell. Mol. Neurobiol. 1999, 19, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, A.; Tsuneishi, E.; Takimoto, Y. Analysis of ATP and Its Breakdown Products in Beef by Reversed-Phase HPLC. J. Food Sci. 1989, 54, 1169–1172. [Google Scholar] [CrossRef]
- Dannert, R.D.; Pearson, A.M. Concentration of Inosine 5′-Monophosphate in Meat. J. Food Sci. 1967, 32, 49–52. [Google Scholar] [CrossRef]
- Koutsidis, G.; Elmore, J.S.; Oruna-Concha, M.J.; Campo, M.M.; Wood, J.D.; Mottram, D.S. Water-soluble precursors of beef flavour. Part II: Effect of post-mortem conditioning. Meat Sci. 2008, 79, 270–277. [Google Scholar] [CrossRef]
- Djenane, D.; Martínez, L.; Sánchez-Escalante, A.; Beltrán, J.A.; Roncalés, P. Antioxidant effect of carnosine and carnitine in fresh beef steaks stored under modified atmosphere. Food Chem. 2004, 85, 453–459. [Google Scholar] [CrossRef]
- Purchas, R.W.; Rutherfurd, S.M.; Pearce, P.D.; Vather, R.; Wilkinson, B.H.P. Concentrations in beef and lamb of taurine, carnosine, coenzyme Q 10, and creatine. Meat Sci. 2004, 66, 629–637. [Google Scholar] [CrossRef]
- Williams, P. Nutritional composition of red meat. Nutr. Diet. 2007, 64, S113–S119. [Google Scholar] [CrossRef] [Green Version]



| Triceps brachii (TB) | Biceps femoris (BF) | Longissimus thoracis (LT) | p | |
|---|---|---|---|---|
| Alanine | 4.76 × 10−3 (1.33 × 10−3) a 1 | 5.31 × 10−3 (4.77 × 10−4) a | 3.14 × 10−3 (9.50 × 10−4) b | 0.003 |
| Carnosine | 1.23 × 10−2 (1.65 × 10−3) b | 1.60 × 10−2 (8.17 × 10−3) b | 2.09 × 10−2 (1.84 × 10−3) a | 0.017 |
| Isoleucine | 1.81 × 10−4 (3.39 × 10−5) ab | 2.33 × 10−4 (6.07 × 10−5) a | 1.69 × 10−4 (4.61 × 10−5) b | 0.012 |
| Leucine | 3.54 × 10−4 (7.22 × 10−5) ab | 4.70 × 10−4 (9.67 × 10−5) a | 3.06 × 10−4 (9.42 × 10−5) b | 0.007 |
| Phenylalanine | 2.48 × 10−4 (7.34 × 10−5) ab | 3.37 × 10−4 (7.77 × 10−5) a | 2.39 × 10−4 (4.65 × 10−5) b | 0.013 |
| Threonine | 3.25 × 10−4 (1.88 × 10−4) a | 4.27 × 10−4 (1.55 × 10−4) a | 2.49 × 10−4 (4.85 × 10−5) b | 0.008 |
| Tyrosine | 1.86 × 10−4 (2.99 × 10−5) ab | 2.02 × 10−4 (5.03 × 10−5) a | 1.60 × 10−4 (3.46 × 10−5) b | 0.011 |
| Valine | 3.28 × 10−4 (4.58 × 10−5) ab | 3.99 × 10−4 (7.97 × 10−5) a | 3.03 × 10−4 (7.23 × 10−5) b | 0.019 |
| Mannose | 2.73 × 10−4 (4.26 × 10−5) ab | 3.11 × 10−4 (9.82 × 10−5) a | 2.35 × 10−4 (3.08 × 10−5) b | 0.039 |
| Formate | 7.43 × 10−5 (7.88 × 10−6) ab | 6.94 × 10−5 (1.85 × 10−5) b | 8.34 × 10−5 (5.95 × 10−6) a | 0.012 |
| IMP | 1.39 × 10−4 (1.74 × 10−4) ab | 4.30 × 10−4 (2.29 × 10−4) a | 1.20 × 10−4 (3.44 × 10−5) b | 0.031 |
| Inosine | 1.73 × 10−4 (1.35 × 10−4) ab | 3.81 × 10−4 (1.89 × 10−4) a | 1.76 × 10−4 (4.33 × 10−5) b | 0.021 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, C.; Petracci, M.; Li, C.; Fiore, E.; Laghi, L. An Untargeted Metabolomics Investigation of Jiulong Yak (Bos grunniens) Meat by 1H-NMR. Foods 2020, 9, 481. https://doi.org/10.3390/foods9040481
Zhu C, Petracci M, Li C, Fiore E, Laghi L. An Untargeted Metabolomics Investigation of Jiulong Yak (Bos grunniens) Meat by 1H-NMR. Foods. 2020; 9(4):481. https://doi.org/10.3390/foods9040481
Chicago/Turabian StyleZhu, Chenglin, Massimiliano Petracci, Cheng Li, Enrico Fiore, and Luca Laghi. 2020. "An Untargeted Metabolomics Investigation of Jiulong Yak (Bos grunniens) Meat by 1H-NMR" Foods 9, no. 4: 481. https://doi.org/10.3390/foods9040481