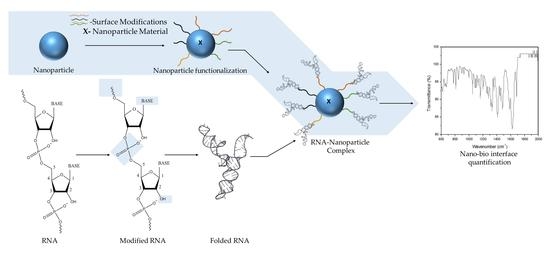
Engineering the RNA-Nanobio Interface
Abstract
:1. Introduction
2. Engineering the RNA-Nanobio Interface
3. Characterizing the RNA-Nanobio Interface
4. RNA-NPC and the Biological Milieu
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Newman, A. Molecular biology: RNA enzymes for RNA splicing. Nature 2001, 413, 695–696. [Google Scholar] [CrossRef] [PubMed]
- Hiller, D.A.; Strobel, S.A. The chemical versatility of RNA. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2011, 366, 2929–2935. [Google Scholar] [CrossRef] [PubMed]
- Serganov, A.; Patel, D.J. Ribozymes, riboswitches and beyond: Regulation of gene expression without proteins. Nat. Rev. Genet. 2007, 8, 776–790. [Google Scholar] [CrossRef] [PubMed]
- Yingling, Y.G.; Shapiro, B.A. Computational Design of an RNA Hexagonal Nanoring and an RNA Nanotube. Nano Lett. 2007, 7, 2328–2334. [Google Scholar] [CrossRef] [PubMed]
- Afonin, K.A.; Bindewald, E.; Yaghoubian, A.J.; Voss, N.; Jacovetty, E.; Shapiro, B.A.; Jaeger, L. In vitro assembly of cubic RNA-based scaffolds designed in silico. Nat. Nanotechnol. 2010, 5, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, L.; Westhof, E.; Leontis, N.B. TectoRNA: Modular assembly units for the construction of RNA nano-objects. Nucleic Acids Res. 2001, 29, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Khaled, A.; Guo, S.; Li, F.; Guo, P. Controllable self-assembly of nanoparticles for specific delivery of multiple therapeutic molecules to cancer cells using RNA nanotechnology. Nano Lett. 2005, 5, 1797–1808. [Google Scholar] [CrossRef] [PubMed]
- Afonin, K.A.; Grabow, W.W.; Walker, F.M.; Bindewald, E.; Dobrovolskaia, M.A.; Shapiro, B.A.; Jaeger, L. Design and self-assembly of siRNA-functionalized RNA nanoparticles for use in automated nanomedicine. Nat. Protoc. 2011, 6, 2022–2034. [Google Scholar] [CrossRef] [PubMed]
- Shu, Y.; Haque, F.; Shu, D.; Li, W.; Zhu, Z.; Kotb, M.; Lyubchenko, Y.; Guo, P. Fabrication of 14 different RNA nanoparticles for specific tumor targeting without accumulation in normal organs. RNA 2013, 19, 767–777. [Google Scholar] [CrossRef] [PubMed]
- Bindewald, E.; Hayes, R.; Yingling, Y.G.; Kasprzak, W.; Shapiro, B.A. RNAJunction: A database of RNA junctions and kissing loops for three-dimensional structural analysis and nanodesign. Nucleic Acids Res. 2008, 36, D392–D397. [Google Scholar] [CrossRef] [PubMed]
- Guo, P.; Zhang, C.; Chen, C.; Garver, K.; Trottier, M. Inter-RNA interaction of phage phi29 pRNA to form a hexameric complex for viral DNA transportation. Mol. Cell 1998, 2, 149–155. [Google Scholar] [CrossRef]
- Guo, S.; Tschammer, N.; Mohammed, S.; Guo, P. Specific delivery of therapeutic RNAs to cancer cells via the dimerization mechanism of phi29 motor pRNA. Hum. Gene Ther. 2005, 16, 1097–1109. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Huang, F.; Guo, P. Construction of folate-conjugated pRNA of bacteriophage phi29 DNA packaging motor for delivery of chimeric siRNA to nasopharyngeal carcinoma cells. Gene Ther. 2006, 13, 814–820. [Google Scholar] [CrossRef] [PubMed]
- Hoeprich, S.; Zhou, Q.; Guo, S.; Shu, D.; Qi, G.; Wang, Y.; Guo, P. Bacterial virus phi29 pRNA as a hammerhead ribozyme escort to destroy hepatitis B virus. Gene Ther. 2003, 10, 1258–1267. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Cui, Y.; Levenson, R.M.; Chung, L.W.K.; Nie, S. In vivo cancer targeting and imaging with semiconductor quantum dots. Nat. Biotechnol. 2004, 22, 969–976. [Google Scholar] [CrossRef] [PubMed]
- Wagner, M.K.; Li, F.; Li, J.; Li, X.-F.; Le, X.C. Use of quantum dots in the development of assays for cancer biomarkers. Anal. Bioanal. Chem. 2010, 397, 3213–3224. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.-Y.; Yeh, H.-C.; Kuroki, M.T.; Wang, T.-H. Single-quantum-dot-based DNA nanosensor. Nat. Mater. 2005, 4, 826–831. [Google Scholar] [CrossRef] [PubMed]
- Bindhu, M.R.; Umadevi, M.; Micheal, M.K.; Arasu, M.V.; Al-Dhabi, N.A. Structural, morphological and optical properties of MgO nanoparticles for antibacterial applications. Mater. Lett. 2016, 166, 19–22. [Google Scholar] [CrossRef]
- Azmath, P.; Baker, S.; Rakshith, D.; Satish, S. Mycosynthesis of silver nanoparticles bearing antibacterial activity. Saudi Pharm. J. 2016, 24, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Alshaer, W.; Hillaireau, H.; Vergnaud, J.; Ismail, S.; Fattal, E. Functionalizing liposomes with anti-CD44 aptamer for selective targeting of cancer cells. Bioconjug. Chem. 2015, 26, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.; Philippi, C.; Loretz, B.; Nafee, N.; Schaefer, U.F.; Friedel, G.; Ammon-Treiber, S.; Griese, E.U.; Lehr, C.M.; Klotz, U.; et al. Tissue slice model of human lung cancer to investigate telomerase inhibition by nanoparticle delivery of antisense 2′-O-methyl-RNA. Int. J. Pharm. 2011, 419, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Xu, S.; Quan, J.; Yung, B.C.; Pang, J.; Zhou, C.; Cho, Y.A.; Zhang, M.; Liu, S.; Muthusamy, N.; et al. CD33-Targeted Lipid Nanoparticles (aCD33LNs) for Therapeutic Delivery of GTI-2040 to Acute Myelogenous Leukemia. Mol. Pharm. 2015, 12, 2010–2018. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Devi, G.; Qu, Q.; Toh, D.-F.K.; Chen, G.; Zhao, Y. Intracellular Delivery of Antisense Peptide Nucleic Acid by Fluorescent Mesoporous Silica Nanoparticles. Bioconjug. Chem. 2014, 25, 1412–1420. [Google Scholar] [CrossRef] [PubMed]
- Farokhzad, O.C.; Jon, S.; Khademhosseini, A.; Tran, T.-N.T.; Lavan, D.A.; Langer, R. Nanoparticle-aptamer bioconjugates: A new approach for targeting prostate cancer cells. Cancer Res. 2004, 64, 7668–7672. [Google Scholar] [CrossRef] [PubMed]
- Javier, D.J.; Nitin, N.; Levy, M.; Ellington, A.; Richards-Kortum, R. Aptamer-Targeted Gold Nanoparticles as Molecular-Specific Contrast Agents for Reflectance Imaging. Bioconjug. Chem. 2008, 19, 1309–1312. [Google Scholar] [CrossRef] [PubMed]
- Rouge, J.L.; Sita, T.L.; Hao, L.; Kouri, F.M.; Briley, W.E.; Stegh, A.H.; Mirkin, C.A. Ribozyme–Spherical Nucleic Acids. J. Am. Chem. Soc. 2015, 137, 10528–10531. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.H.; Cho, Y.W.; Jung, B.; Park, K.; Kim, J.-D. Self-Assembled Nanoparticles of Ribozymes with Poly(ethylene glycol)-b-Poly(L-lysine) Block Copolymers. Jpn. J. Appl. Phys. 2006, 45, 591. [Google Scholar] [CrossRef]
- Kong, L.; Wu, Y.; Alves, C.S.; Shi, X. Efficient delivery of therapeutic siRNA into glioblastoma cells using multifunctional dendrimer-entrapped gold nanoparticles. Nanomedicine 2016, 11, 3103–3115. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.-C.; Zhang, L.; Feng, S.S.; Hong, L.; Zheng, H.L.; Chen, L.L.; Zheng, X.L.; Ye, Y.Q.; Zhao, M.D.; Wang, W.X.; et al. Efficient delivery of Notch1 siRNA to SKOV3 cells by cationic cholesterol derivative-based liposome. Int. J. Nanomed. 2016, 11, 5485–5496. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.; Yu, B.; Wang, X.; Lu, Y.; Schmidt, C.R.; Lee, R.J.; Lee, L.J.; Jacob, S.T.; Ghoshal, K. Cationic lipid nanoparticles for therapeutic delivery of siRNA and miRNA to murine liver tumor. Nanomed. Nanotechnol. Biol. Med. 2013, 9, 1169–1180. [Google Scholar] [CrossRef] [PubMed]
- Arami, S.; Mahdavi, M.; Rashidi, M.R.; Fathi, M.; Hejazi, M.; Samadi, N. Novel polyacrylate-based cationic nanoparticles for survivin siRNA delivery combined with mitoxantrone for treatment of breast cancer. Biologicals 2016, 44, 487–496. [Google Scholar] [CrossRef] [PubMed]
- Fattal, E.; Vauthier, C.; Aynie, I.; Nakada, Y.; Lambert, G.; Malvy, C.; Couvreur, P. Biodegradable polyalkylcyanoacrylate nanoparticles for the delivery of oligonucleotides. J. Control. Release 1998, 53, 137–143. [Google Scholar] [CrossRef]
- Guo, P.; Haque, F. Challenges, Solution, and Perspectives in RNA Nanotechnology. In RNA Nanotechnology and Therapeutics; CRC Press: Boca Raton, FL, USA, 2014; pp. 39–45. [Google Scholar]
- DeLong, R.K.; Reynolds, C.M.; Malcolm, Y.; Schaeffer, A.; Severs, T.; Wanekaya, A. Functionalized gold nanoparticles for the binding, stabilization, and delivery of therapeutic DNA, RNA, and other biological macromolecules. Nanotechnol. Sci. Appl. 2010, 3, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Shaw, B.R.; Sergueev, D.; He, K.; Porter, K.; Summers, J.; Sergueeva, Z.; Rait, V. Boranophosphate backbone: A mimic of phosphodiesters, phosphorothioates, and methyl phosphonates. Methods Enzymol. 2000, 313, 226–257. [Google Scholar] [PubMed]
- Webb, M.S.; Tortora, N.; Cremese, M.; Kozlowska, H.; Blaquiere, M.; Devine, D.V.; Kornbrust, D.J. Toxicity and toxicokinetics of a phosphorothioate oligonucleotide against the c-myc oncogene in cynomolgus monkeys. Antisense Nucleic Acid Drug Dev. 2001, 11, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Guo, S.; Cinier, M.; Shlyakhtenko, L.S.; Shu, Y.; Chen, C.; Shen, G.; Guo, P. Fabrication of Stable and RNase-Resistant RNA Nanoparticles Active in Gearing the Nanomotors for Viral DNA-Packaging. ACS Nano 2011, 5, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.S.; Reed, S.M. Direct visualization of electrophoretic mobility shift assays using nanoparticle-aptamer conjugates. Electrophoresis 2012, 33, 348–351. [Google Scholar] [CrossRef] [PubMed]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA Mimics of Green Fluorescent Protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef] [PubMed]
- Dolgosheina, E.V.; Jeng, S.C.Y.; Panchapakesan, S.S.S.; Cojocaru, R.; Chen, P.S.K.; Wilson, P.D.; Hawkins, N.; Wiggins, P.A.; Unrau, P.J. RNA Mango Aptamer-Fluorophore: A Bright, High-Affinity Complex for RNA Labeling and Tracking. ACS Chem. Biol. 2014, 9, 2412–2420. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.; Bhattarai, N.; Sun, C.; Zhang, M. Functionalized Nanoparticles with Long-Term Stability in Biological Media. Small Weinh. Bergstr. Ger. 2009, 5, 1637–1641. [Google Scholar] [CrossRef] [PubMed]
- Song, W.-J.; Du, J.-Z.; Sun, T.-M.; Zhang, P.-Z.; Wang, J. Gold Nanoparticles Capped with Polyethyleneimine for Enhanced siRNA Delivery. Small 2010, 6, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Nafee, N.; Taetz, S.; Schneider, M.; Schaefer, U.F.; Lehr, C.-M. Chitosan-coated PLGA nanoparticles for DNA/RNA delivery: Effect of the formulation parameters on complexation and transfection of antisense oligonucleotides. Nanomed. Nanotechnol. Biol. Med. 2007, 3, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Famulok, M.; Szostak, J.W. Stereospecific recognition of tryptophan agarose by in vitro selected RNA. J. Am. Chem. Soc. 1992, 114, 3990–3991. [Google Scholar] [CrossRef]
- Connell, G.J.; Illangesekare, M.; Yarus, M. Three small ribooligonucleotides with specific arginine sites. Biochemistry (Mosc.) 1993, 32, 5497–5502. [Google Scholar] [CrossRef]
- Famulok, M. Molecular Recognition of Amino Acids by RNA-Aptamers: An L-Citrulline Binding RNA Motif and Its Evolution into an L-Arginine Binder. J. Am. Chem. Soc. 1994, 116, 1698–1706. [Google Scholar] [CrossRef]
- Geiger, A.; Burgstaller, P.; von der Eltz, H.; Roeder, A.; Famulok, M. RNA Aptamers That Bind l-Arginine with Sub-Micromolar Dissociation Constants and High Enantioselectivity. Nucleic Acids Res. 1996, 24, 1029–1036. [Google Scholar] [CrossRef] [PubMed]
- Tseng, Y.-L.; Liu, J.-J.; Hong, R.-L. Translocation of Liposomes into Cancer Cells by Cell-Penetrating Peptides Penetratin and Tat: A Kinetic and Efficacy Study. Mol. Pharmacol. 2002, 62, 864–872. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.J.; Saltzman, W.M. Enhanced siRNA delivery into cells by exploiting the synergy between targeting ligands and cell-penetrating peptides. Biomaterials 2011, 32, 6194–6203. [Google Scholar] [CrossRef] [PubMed]
- Niu, Z.; Li, Y. Removal and Utilization of Capping Agents in Nanocatalysis. Chem. Mater. 2014, 26, 72–83. [Google Scholar] [CrossRef]
- Woehrle, G.H.; Brown, L.O.; Hutchison, J.E. Thiol-Functionalized, 1.5-nm Gold Nanoparticles through Ligand Exchange Reactions: Scope and Mechanism of Ligand Exchange. J. Am. Chem. Soc. 2005, 127, 2172–2183. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Xin, H.; Ren, Q.; Gu, J.; Zhu, L.; Du, F.; Feng, C.; Xie, Y.; Sha, X.; Fang, X. Nanoparticles of 2-deoxy-D-glucose functionalized poly(ethylene glycol)-co-poly(trimethylene carbonate) for dual-targeted drug delivery in glioma treatment. Biomaterials 2014, 35, 518–529. [Google Scholar] [CrossRef] [PubMed]
- El-Brolossy, T.A.; Abdallah, T.; Mohamed, M.B.; Abdallah, S.; Easawi, K.; Negm, S.; Talaat, H. Shape and size dependence of the surface plasmon resonance of gold nanoparticles studied by Photoacoustic technique. Eur. Phys. J. Spec. Top. 2008, 153, 361–364. [Google Scholar] [CrossRef]
- Buchman, Y.K.; Lellouche, E.; Zigdon, S.; Bechor, M.; Michaeli, S.; Lellouche, J.-P. Silica nanoparticles and polyethyleneimine (PEI)-mediated functionalization: A new method of PEI covalent attachment for siRNA delivery applications. Bioconjug. Chem. 2013, 24, 2076–2087. [Google Scholar] [CrossRef] [PubMed]
- Speranza, G.; Abdallah, T.; Mohamed, M.B.; Abdallah, S.; Easawi, K.; Negm, S.; Talaat, H. XPS Characterization of Iron Oxide and Gold Nanoparticles for Tumor Care. Adv. Sci. Technol. 2010, 76, 165–170. [Google Scholar] [CrossRef]
- Hurst, M.N.; DeLong, R.K. Two-Dimensional Fluorescence Difference Spectroscopy to Characterize Nanoparticles and their Interactions. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Bujold, K.E.; Hsu, J.C.C.; Sleiman, H.F. Optimized DNA “Nanosuitcases” for Encapsulation and Conditional Release of siRNA. J. Am. Chem. Soc. 2016, 138, 14030–14038. [Google Scholar] [CrossRef] [PubMed]
- Ezzat, K.; Aoki, Y.; Koo, T.; McClorey, G.; Benner, L.; Coenen-Stass, A.; O’Donovan, L.; Lehto, T.; Garcia-Guerra, A.; Nordin, J.; et al. Self-Assembly into Nanoparticles Is Essential for Receptor Mediated Uptake of Therapeutic Antisense Oligonucleotides. Nano Lett. 2015, 15, 4364–4373. [Google Scholar] [CrossRef] [PubMed]
- Pi, F.; Zhang, H.; Li, H.; Thiviyanathan, V.; Gorenstein, D.G.; Sood, A.K.; Guo, P. RNA nanoparticles harboring annexin A2 aptamer can target ovarian cancer for tumor-specific doxorubicin delivery. Nanomed. Nanotechnol. Biol. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.J.; Saltzman, W.M. Polymer nanoparticle-mediated delivery of microRNA inhibition and alternative splicing. Mol. Pharm. 2012, 9, 1481–1488. [Google Scholar] [CrossRef] [PubMed]
- Ekin, A.; Karatas, O.F.; Culha, M.; Ozen, M. Designing a gold nanoparticle-based nanocarrier for microRNA transfection into the prostate and breast cancer cells. J. Gene Med. 2014, 16, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Binzel, D.W.; Shu, Y.; Li, H.; Sun, M.; Zhang, Q.; Shu, D.; Guo, B.; Guo, P. Specific Delivery of miRNA for High Efficient Inhibition of Prostate Cancer by RNA Nanotechnology. Mol. Ther. J. Am. Soc. Gene Ther. 2016, 24, 1267–1277. [Google Scholar] [CrossRef] [PubMed]
- Dakwar, G.R.; Braeckmans, K.; Demeester, J.; Ceelen, W.; Smedt, S.C.D.; Remaut, K. Disregarded Effect of Biological Fluids in siRNA Delivery: Human Ascites Fluid Severely Restricts Cellular Uptake of Nanoparticles. ACS Appl. Mater. Interfaces 2015, 7, 24322–24329. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, C.C.; Payne, C.K. Nanoparticle–Cell Interactions: Molecular Structure of the Protein Corona and Cellular Outcomes. Acc. Chem. Res. 2014, 47, 2651–2659. [Google Scholar] [CrossRef] [PubMed]
- Gerasimova, Y.V.; Erchenko, I.A.; Shakirov, M.M.; Godovikova, T.S. Interaction of human serum albumin and its clinically relevant modification with oligoribonucleotides. Bioorg. Med. Chem. Lett. 2008, 18, 4511–4514. [Google Scholar] [CrossRef] [PubMed]
- Yogasundaram, H.; Bahniuk, M.S.; Singh, H.; Aliabadi, H.M.; Uludaǧ, H.; Unsworth, L.D. BSA Nanoparticles for siRNA Delivery: Coating Effects on Nanoparticle Properties, Plasma Protein Adsorption, and In Vitro siRNA Delivery. Int. J. Biomater. 2012. [Google Scholar] [CrossRef] [PubMed]
| RNA+ Modification | NP+ Modification | RNA-Nanobio Characterization | Biological Significance | Ref. | |
|---|---|---|---|---|---|
| Anti-Sense Oligonucleotides (ASO) | - | Liposome DOTAP:DOPE-DOC-PEI 1 | 1H NMR, FT-IR, Z-potential (ζ) | Selectively target CD33 positive Kasumi-1 cells Greater transfection efficiency Highest downregulation of R2 Treated mice-increased tumor inhibition | [22] |
| 2′-O-Met, FAM | Polymer PLGA + Chitosan | AFM (Atomic Force Microscopy) | Increased cellular uptake by primary lung cancer cells and fibroblasts. Inhibition of telomerase activity Decreased cytotoxicity | [21] | |
| PNA | Inorganic Mesoporous Silica (MSNP) + FITC + SS | SEM, TEM, FT-IR, Z-potential, MALDI-TOF MS 2, UV Abs. | Higher cellular uptake Directed delivery into cytoplasm of HeLa cells Silencing of Bcl-2 protein expression Low toxicity | [23] | |
| TcDNA + PS And B-PPMO | Nucleic Acid Self-Forming micelle nanoparticle | TEM, DLS (Dynamic Light Scatter), Critical Micelle Concentration (CMC), Z-potential | ASOs had increased inclination to self-assemble into nanoparticles Fucoidin Sulfate and Dextran sulfate competitively inhibited B-PPMO uptake by Class A scavenger receptor subtypes (SCARAs) | [58] | |
| Aptamer | Apt1 (anti-CD44)+ 2′-F | Liposome DPPC:Cholesterol:DSPE+ Maleimide + PEG | Z-potential, Gel electrophoresis | Enhanced CD44 binding Increased cellular uptake by Lung (A549) and Breast Cancer (MDA-MB-231) Cell lines 3. Constant IL-6, IL-1β, IL-8—does not induce inflammatory response | [20] |
| anti-PSMA 4 | Polymer PLA-PEG-COOH | Z-potential, SEM | PMSA specific Increase in efficiency and selectivity | [24] | |
| anti-PSMA A9 | Inorganic Gold+ Thiol+ hexa(ethylene glycol) | DLS, Fluorescence | Selectively labels PSMA positive cells (LNCaP) Stability preserved | [25] | |
| Endo28 5 Aptamer | Nucleic Acid pRNA-3WJ | Serum Stability with polyacrylamide gel, Z-potential, DLS | RNA-NP showed increased binding to AnnexinA2 expressing cancer cells-IGROV-1 RNA-NP w/dox specifically delivers to AnnexinA2positive cell lines. Enhanced AnnexinA2 tumor selective binding | [59] | |
| microRNA | miR-122 mimic | Liposome DODMA-eggPC-chol-PEG | Z-potential | Preferentially taken up by tumor cell lines Down-regulation miR-122 genes in tumor tissue. Z-potential in different pH-colloidal stability | [30] |
| Anti-miR-155 PMO and PNA | Polymer PLGA+ argCPP | BCA Assay 6, Flow Cytometry, TEM | Inhibition of miRNA Altered splicing to produce Mcl-1S isoform opposed to Mcl-1L isoform | [60] | |
| miR-145 | Inorganic Gold+ thiol oligo | Z-potential, UV spectrophotometry | Overexpression of ectopic miR-145 in PC3 and MCF7 cell types. Efficient delivery system | [61] | |
| Anti-miR-21 | Nucleic Acid pRNA-3WJ with PMSA aptamer+ Cy5/Alexa647 | Z-potential, hydrodynamic diameter, Temperature Gradient Gel Electrophoresis (TGGE) | Specific delivery of anti-miR-21 to LNCaP-FGC (PMSA+) cells Delivery of anti-miR-21 is achieved through PMSA aptamer binding Increase in Caspase III indicating cell death Specific targeting and accumulation of RNA-NP to xenograft tumor Low toxicity profiles in kidney and liver Increase in PTEN and PDCD4 tumor suppressor genes | [62] | |
| Ribozymes | Rzs | Polymer PEG-b-PLL | DLS, TEM, EtBr Displacement Assay 5, 1H NMR, Electrophoretic Mobility Shift | Stable complexes formed Stable in RNAse-rich environment | [27] |
| MGMT 7+ Fluorescein+ 5′Cy5 | Inorganic Gold SNA | Gel elec, RT-PCR, DLS, Z-potential | Cleaved MGMT substrate Sustained stability in harsh enzymatic environment Knocked down MGMT in T98G glioma cells Increase in Caspase-3/7 activity | [26] | |
| siRNA | Notch1-homo-siRNA-FAM | Liposome DMAPA | Z-potential, TEM, Gel electrophoresis | Protect against RNase A in serum No SKOV3 cell cytotoxicity Increased cellular uptake Increased percentage of apoptotic cells | [29] |
| anti-survivin siRNA | Polymer PA-PEI + Fe3O4 Magnetic NP | FT-IR, Z-potential, Gel retardation assay | Increased cell uptake Increased knockdown of survivin gene Increased apoptosis: 3-fold | [31] | |
| VEGF siRNA/ B-cell lymphoma siRNA | Inorganic Gold+ RGD-PEG-COOH Dendrimer | Gel retardation assay, DLS, Z-potential | Delivered specifically to integrin-overexpressing cells Induce specific silencing of genes High transfection efficiency Down-regulation of VEGF and Bcl-2 | [28] | |
| FASE siRNA+ Bcl-xl/2 strand 8+ Cy3 | Nucleic Acid DNA Nanocube-prism+ Cy5 | UV-visible melting, UV-visible spectrophotometry for stability, DLS, | Controlled release of siRNA Enhanced stability in DMEM+ serum Increased half-life of 12 hours | [57] | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Murthy, V.; Delong, R.K. Engineering the RNA-Nanobio Interface. Bioengineering 2017, 4, 13. https://doi.org/10.3390/bioengineering4010013
Murthy V, Delong RK. Engineering the RNA-Nanobio Interface. Bioengineering. 2017; 4(1):13. https://doi.org/10.3390/bioengineering4010013
Chicago/Turabian StyleMurthy, Vaibhav, and Robert K. Delong. 2017. "Engineering the RNA-Nanobio Interface" Bioengineering 4, no. 1: 13. https://doi.org/10.3390/bioengineering4010013