Journal Description
Journal of Developmental Biology
Journal of Developmental Biology
is an international, peer-reviewed, open access journal on the development of multicellular organisms at the molecule, cell, tissue, organ and whole organism levels published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 27.1 days after submission; acceptance to publication is undertaken in 5.6 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about Journal of Developmental Biology.
Impact Factor:
2.5 (2024);
5-Year Impact Factor:
2.8 (2024)
Latest Articles
Fetal–Fetal and Fetal–Maternal Microchimerism: Insights from Mammalian Placental Biology
J. Dev. Biol. 2026, 14(2), 19; https://doi.org/10.3390/jdb14020019 - 28 Apr 2026
Abstract
►
Show Figures
Feto-maternal microchimerism (Mc) refers to the exchange of cells between the fetus and mother, and fetal–fetal Mc to the exchange between fetuses during pregnancy. This phenomenon occurs across mammalian species, including humans, mice, and cattle. Key data on Mc cells and theoretical considerations
[...] Read more.
Feto-maternal microchimerism (Mc) refers to the exchange of cells between the fetus and mother, and fetal–fetal Mc to the exchange between fetuses during pregnancy. This phenomenon occurs across mammalian species, including humans, mice, and cattle. Key data on Mc cells and theoretical considerations regarding the presence of fetal-derived material, such as trophoblast cells, cell-free fetal DNA (cffDNA), and exosomes in maternal blood are summarized. This review aims to first, synthesize current knowledge on feto-maternal and fetal–fetal Mc across mammals, second, address three core questions: how and where Mc has been demonstrated in animals, what techniques have been used over time to detect fetal-derived material and Mc, and how placental structures influence the frequency of Mc. Finally, it aims to identify gaps in the literature for species such as horses, goats, and pigs. This article concludes that Mc is a widespread phenomenon among mammals, but detection methods and reported frequencies vary significantly by species and placental type. A biological model is presented in this article in which multinucleated trophoblast cells undergo apoptosis, releasing cffDNA that enters the maternal blood circulation after multinucleated trophoblast invasion. Advances in molecular biology technology have improved the ability to detect fetal-derived material, cells, DNA, and exosomes in maternal blood. However, notable research gaps remain for Mc in horses, goats, and pigs, highlighting the need for targeted studies to better understand species-specific patterns or a general biological model.
Full article
Open AccessArticle
Melatonin Receptor 1 and Melatonin Receptor 2 Expression During Human Kidney Development and Their Association with CAKUT
by
Ann-Kathrin Schmitt, Victoria Tjora, Nela Kelam, Marija Jurić Gunjača, Petar Todorović, Clelia Picard, Manel Loche-Dalmon, Katarina Vukojević and Anita Racetin
J. Dev. Biol. 2026, 14(2), 18; https://doi.org/10.3390/jdb14020018 - 15 Apr 2026
Abstract
Background/Objectives: Growing evidence indicates that melatonin contributes to kidney development and function, while disruptions of fetal circadian signaling have been linked to congenital anomalies of the kidney and urinary tract (CAKUT). This study aimed to characterize the developmental and spatial expression patterns of
[...] Read more.
Background/Objectives: Growing evidence indicates that melatonin contributes to kidney development and function, while disruptions of fetal circadian signaling have been linked to congenital anomalies of the kidney and urinary tract (CAKUT). This study aimed to characterize the developmental and spatial expression patterns of melatonin receptors MTNR1A and MTNR1B in normal human fetal kidneys and in CAKUT phenotypes. Methods: This study analyzed 40 human fetal kidney specimens, including healthy controls and CAKUT cases (horseshoe kidneys, duplex kidneys, and dysplastic kidneys), obtained from spontaneous abortions and pregnancy terminations. Samples were classified into developmental phases Ph2–Ph4 according to established morphological criteria. Immunofluorescence staining was used to visualize MTNR1A and MTNR1B expression. Quantitative analysis was performed using ImageJ, measuring the fluorescence area percentage. Statistical comparisons were conducted using a two-way ANOVA. Results: In control kidneys, MTNR1A expression was predominantly observed in glomeruli and interstitial cells and showed a descending trend across developmental stages, whereas MTNR1B was localized to glomeruli and strongly to the apical membranes of tubules, particularly distal tubules, without substantial developmental variation. CAKUT phenotypes exhibited higher expression of both receptors compared to controls. Significant phase-dependent differences in MTNR1A expression were observed in horseshoe, duplex, and dysplastic kidneys. MTNR1B expression decreased across developmental stages in dysplastic kidneys and differed significantly between Ph3 and Ph4 in duplex kidneys. At Ph3, duplex kidneys showed the highest MTNR1B expression. Conclusions: Altered developmental expression patterns of MTNR1A and MTNR1B in CAKUT suggest an association between melatonin signaling and abnormal human kidney development.
Full article
(This article belongs to the Special Issue Developmental Biology of the Kidney: From Molecular Mechanisms to Congenital Disorders)
►▼
Show Figures

Figure 1
Open AccessReview
Novel Functions and Potential of Ribosomes: From Cellular Transdifferentiation to Applications in Cell-Cultured Foods
by
Shota Inoue, Hiroaki Hatano, Ikko Kawashima and Kunimasa Ohta
J. Dev. Biol. 2026, 14(2), 17; https://doi.org/10.3390/jdb14020017 - 9 Apr 2026
Abstract
►▼
Show Figures
Ribosomes are widely recognized as large intracellular macromolecular complexes responsible for protein synthesis. However, in recent years, numerous studies have revealed that ribosomal proteins possess non-canonical functions beyond translation, including roles in cell fate regulation, development, and disease. This review outlines emerging concepts
[...] Read more.
Ribosomes are widely recognized as large intracellular macromolecular complexes responsible for protein synthesis. However, in recent years, numerous studies have revealed that ribosomal proteins possess non-canonical functions beyond translation, including roles in cell fate regulation, development, and disease. This review outlines emerging concepts surrounding the extracellular functions of ribosomes, with a particular focus on ribosome-induced cellular plasticity and transdifferentiation. Our studies have demonstrated that the incorporation of exogenous ribosomes reprograms somatic cells into a multipotent state and promotes differentiation into multiple lineages. These findings represent an alternative perspective to the conventional view of ribosomes as merely translational components. Furthermore, we discuss the biological significance of factors secreted by ribosome-incorporated cells by integrating the paracrine hypothesis with ribosome-mediated cell fate conversion. Finally, we explore the potential applications of ribosomes in regenerative medicine and cell-cultured food production. By redefining ribosomes as active regulators of cellular identity, this review provides a conceptual framework for understanding ribosome-driven cell fate regulation and its potential applications in sustainable biotechnology.
Full article

Figure 1
Open AccessArticle
Conserved Metanephric Kidney Development and Genome Methylation in Red-Eared Slider Turtle (Trachemys scripta elegans)
by
Bing Jia, Mohamed Milad, Hannah C. Boehler, Adam Guerra, Joshua Mowry, Jessica Hiley, James Kasen Lisonbee, Michael Hafen and Troy Camarata
J. Dev. Biol. 2026, 14(2), 16; https://doi.org/10.3390/jdb14020016 - 7 Apr 2026
Abstract
►▼
Show Figures
Mammals and reptiles possess a metanephric kidney as the terminal renal organ for homeostasis of solutes and waste products. The development of the metanephric kidney has primarily been studied in mammalian model systems. Little is known about the conservation of metanephric kidney formation
[...] Read more.
Mammals and reptiles possess a metanephric kidney as the terminal renal organ for homeostasis of solutes and waste products. The development of the metanephric kidney has primarily been studied in mammalian model systems. Little is known about the conservation of metanephric kidney formation in non-mammalian species such as reptiles. Uniquely, reptiles maintain kidney progenitor cell populations throughout life and continually develop new nephrons, the functional unit of the kidney. The red-eared slider turtle, Trachemys scripta elegans, was utilized to investigate the conservation of reptilian metanephric kidney development. The nephron progenitor cell (NPC) marker, Six2, was detected in whole-mount turtle kidneys in a similar pattern to mammals. However, there were differences in progenitor cell niche morphology where turtle NPC populations formed distinct elongated rows instead of the rosette-like morphology found in the mouse. The pattern of NPC populations in the embryonic turtle kidney was maintained in the adult turtle. Whole-genome bisulfite sequencing was performed on cortical tissue containing the NPC populations from adult turtle kidneys and compared to those of adult mice. Significant conservation of gene methylation was detected in adult cortical tissue between the two species, although unique signatures were detected in turtle samples related to DNA repair and β-catenin signaling. This suggests a high level of conservation of metanephric kidney development at the genetic level.
Full article

Figure 1
Open AccessArticle
Compensatory Serotonin Synthesis and Histone H3 Serotonylation in Preimplantation Embryos Exposed to Maternal Fluoxetine or Monoamine Oxidase Blockade
by
Veronika S. Frolova and Denis A. Nikishin
J. Dev. Biol. 2026, 14(2), 15; https://doi.org/10.3390/jdb14020015 - 3 Apr 2026
Abstract
Serotonin is a critical morphogen in early development, yet the mechanisms regulating its homeostasis in the preimplantation embryo remain unclear, particularly under conditions of maternal antidepressant exposure. Here, we investigated embryonic serotonergic autonomy using mouse models of pharmacological transport blockade (maternal fluoxetine treatment)
[...] Read more.
Serotonin is a critical morphogen in early development, yet the mechanisms regulating its homeostasis in the preimplantation embryo remain unclear, particularly under conditions of maternal antidepressant exposure. Here, we investigated embryonic serotonergic autonomy using mouse models of pharmacological transport blockade (maternal fluoxetine treatment) and in vitro treatment with the monoamine oxidase inhibitor pargyline. We employed immunofluorescence, RT-qPCR, and live-cell imaging to assess metabolic flux, gene expression, and physiological health. We demonstrate that monoamine oxidase functions as a metabolic firewall, progressively maturing from zygote to blastocyst to degrade excess amines. Paradoxically, maternal serotonin transporter blockade triggered significant intracellular serotonin hyper-accumulation in blastocysts, associated with a trend toward a compensatory upregulation of the biosynthetic gene Ddc. While this serotonin overload did not compromise morphology, mitochondrial function, or pluripotency marker expression, it induced a robust epigenetic response. Excess serotonin promoted elevated H3Q5ser immunoreactivity in both nuclear and cytoplasmic compartments via a transglutaminase-dependent mechanism. These findings reveal that the preimplantation embryo possesses a resilient, autonomous serotonergic system capable of compensatory synthesis. However, environmental fluctuations are chemically recorded via transglutaminase-mediated serotonylation, representing an epigenetic mark that warrants further long-term study within the Developmental Origins of Health and Disease (DOHaD) framework.
Full article
(This article belongs to the Special Issue Genetic and Epigenetic Mechanisms in Gametogenesis and Early Development: Insights from Models, Stem Cells, and Human Disorders)
►▼
Show Figures

Figure 1
Open AccessReview
Transformation of the Biological Paradigm in Bone Regeneration: An Integrative Review
by
Diyana Vladova
J. Dev. Biol. 2026, 14(1), 14; https://doi.org/10.3390/jdb14010014 - 11 Mar 2026
Abstract
►▼
Show Figures
Bone tissue is among the most commonly transplanted tissues worldwide. The treatment of critical-sized bone defects remains a significant challenge, as there is currently no universally accepted experimental model or therapeutic standard. Recent advances in fundamental cell biology are driving a paradigm shift
[...] Read more.
Bone tissue is among the most commonly transplanted tissues worldwide. The treatment of critical-sized bone defects remains a significant challenge, as there is currently no universally accepted experimental model or therapeutic standard. Recent advances in fundamental cell biology are driving a paradigm shift in approaches to bone regeneration, highlighting the transformative potential of biofabrication technologies that integrate tissue engineering with personalized regenerative strategies. Three-dimensional (3D) bioprinting technology enables precise control over the architecture and spatial distribution of cellular and biologically active components, facilitating the creation of complex, personalized bone constructs. Central to this process are bioinks and biomaterials that mimic the extracellular matrix (ECM) and provide an optimal microenvironment for cellular function. Despite the substantial body of accumulated data, a comprehensive theoretical framework for functional bone biofabrication has not yet been fully established, emphasizing both the challenges and the innovative potential of the field. This integrative review synthesizes current knowledge on bone biology—from embryogenesis and cell–matrix interactions to molecular and neural regulation—and links it to the opportunities offered by biofabrication. Particular attention is given to bioinks as mediators between cell biology and engineering sciences, as well as to strategies for creating biomimetic ECM, optimizing scaffold design, and guiding future research toward clinically translatable bone regeneration.
Full article

Figure 1
Open AccessReview
SIX3 as a Regulator of Development and Disease
by
Ana Beatriz Matos, Laura Jesus Castro and Torcato Martins
J. Dev. Biol. 2026, 14(1), 13; https://doi.org/10.3390/jdb14010013 - 6 Mar 2026
Abstract
►▼
Show Figures
Transcriptional regulation is pivotal for developmental processes and cell fate specification in homeostasis. One particularly relevant group of transcription factors is the sine oculis homeobox (SIX) family, which is involved in a wide range of molecular processes from development to tissue maintenance. Within
[...] Read more.
Transcriptional regulation is pivotal for developmental processes and cell fate specification in homeostasis. One particularly relevant group of transcription factors is the sine oculis homeobox (SIX) family, which is involved in a wide range of molecular processes from development to tissue maintenance. Within this family, distinct subfamilies exhibit specific DNA-binding preferences and can function as transcriptional activators or repressors. In this review, we focus on the Optix/SIX3–SIX6 subfamily and discuss their roles as transcriptional regulators, as well as the consequences of their deregulation for neuronal and ocular development and for the maintenance of tissue homeostasis. We further examine how SIX3 can act either as a tumour suppressor or as a marker of poor prognosis in different cancer types. Moreover, we summarize recent findings on the role of SIX3 in pancreatic β cells and highlight emerging evidence that SIX2 also contributes to β-cell identity and regulatory stability. Downregulation of SIX2 and SIX3 alters gene regulatory programs associated with β-cell homeostasis and contributes to type 2 diabetes. As accumulating evidence links members of the SIX family to cancer and metabolic disease, it is crucial to characterize how these transcription factors regulate cell identity, with important implications for disease mechanisms and therapeutic strategies.
Full article

Figure 1
Open AccessReview
Genomic Impacts of Biological Exposures
by
Amalia S. Parra and Christopher A. Johnston
J. Dev. Biol. 2026, 14(1), 12; https://doi.org/10.3390/jdb14010012 - 5 Mar 2026
Abstract
►▼
Show Figures
Development and maintenance of complex tissues depends on a number of coordinated steps from early development through adulthood. These processes are fundamentally controlled by highly regulated gene expression patterns. Although critical contributors during development, intrinsic changes in gene expression alone cannot fully explain
[...] Read more.
Development and maintenance of complex tissues depends on a number of coordinated steps from early development through adulthood. These processes are fundamentally controlled by highly regulated gene expression patterns. Although critical contributors during development, intrinsic changes in gene expression alone cannot fully explain the complicated pathways that control tissue homeostasis. Rather, tissues are continuously exposed to extrinsic factors that also influence essential cellular processes. These external environmental factors are collectively known as the exposome. Notably, how different exposures impact gene expression and protein function, as well as how certain exposures lead to disease states, is not well understood. To understand how internal and external factors influence organismal development and homeostasis, it is necessary to consider how genetic and nongenetic components interact to direct critical biochemical pathways. Doing so presents new avenues for precision medicine, understanding disease progression, identifying biological threats, and improving biological security concerns. In this review, we present recent advances in exposure biology, focusing on how these innovations can help identify novel biomarkers to better understand changing exposome components. We also discuss the need to integrate technologies and exposure research to better identify and predict threats.
Full article

Figure 1
Open AccessReview
Origins of Avian Hyperactive Mitochondria, Genome Compaction, and Air-Sac Physiology in Early Theropods During the Carnian Pluvial Episode
by
Takumi Satoh
J. Dev. Biol. 2026, 14(1), 11; https://doi.org/10.3390/jdb14010011 - 2 Mar 2026
Abstract
►▼
Show Figures
Extant birds and the earliest dinosaurs may share fundamental metabolic features essential for aerobic exercise, suggesting that the extraordinary physical performance typical of avian species originated when dinosaurs first appeared during the Carnian Pluvial Episode (CPE). This physiological adaptation is complemented by hyperactive
[...] Read more.
Extant birds and the earliest dinosaurs may share fundamental metabolic features essential for aerobic exercise, suggesting that the extraordinary physical performance typical of avian species originated when dinosaurs first appeared during the Carnian Pluvial Episode (CPE). This physiological adaptation is complemented by hyperactive mitochondria that exhibit high oxygen consumption and low reactive oxygen species production. Molecular genomics of fossils, the so-called “Jurassic Genome,” indicates that these early dinosaurs possessed compact genomes, 50–60% the size of the human genome, and small cells, implying a highly stringent metabolic regime. We suggest that hyperactive mitochondria, closely associated with compact genomes and small cells, drive theropod adaptation to the hot, dry, and hypoxic environments of the Late Triassic period, ultimately enabling their ecological dominance. Early dinosaurs such as Herrerasaurus are hypothesized to have possessed advanced physiological traits shared with modern birds, including hyperactive mitochondria, compact genomes, small cells, and a developing air-sac system. Collectively, these features most likely may have contributed to exceptional metabolic capacity, locomotor performance, and adaptation to the harsh environment of the CPE.
Full article

Graphical abstract
Open AccessSystematic Review
Evolutionary Restructuring and Systematic Review of the NBPF Gene Family: Comparative Genomics, Functional Divergence, and Disease-Linked Pathways
by
Manuel Escalona and Rosa Roy
J. Dev. Biol. 2026, 14(1), 10; https://doi.org/10.3390/jdb14010010 - 24 Feb 2026
Abstract
►▼
Show Figures
The Neuroblastoma Breakpoint Family (NBPF) consists of 23 genes, 9 of which are pseudogenes, and is characterized by extensive duplication events and species-specific diversification in Homo sapiens, as well as by the presence of a unique protein domain known as Olduvai (also
[...] Read more.
The Neuroblastoma Breakpoint Family (NBPF) consists of 23 genes, 9 of which are pseudogenes, and is characterized by extensive duplication events and species-specific diversification in Homo sapiens, as well as by the presence of a unique protein domain known as Olduvai (also referred to as DUF1220 or the NBPF domain). Previous studies have attempted to define subfamilies based on the presence of HLS triplet domains; however, this classification has become increasingly unclear with the identification of additional NBPF members. The family remains poorly understood, and the functions of many genes are still unknown, although several have been hypothesized to play key roles in cell proliferation and developmental processes, particularly in neural and skeletal tissues. In this study, we systematically analyzed all available data on the NBPF gene family using the PRISMA-S methodology to infer the biological functions in which these genes may be involved. We also generated multiple phylogenetic trees to support the creation of coherent subfamilies and to correlate the origin of each subfamily with homologous genes in our last common ancestor with the Pan genus, providing what we believe to be one of the most comprehensive phylogenetic reconstructions including all currently annotated NBPF members. Through comparative genomic and phylogenetic analyses, we propose that the NBPF may have originated from a duplication of the PDE4DIP gene, with NBPF26 representing the ancestral member from which the remaining NBPF genes diverged via lineage-specific segmental duplications. In this systematic review and comparative genomic study, we present the first integrative synthesis of our knowledge of the NBPF, encompassing its evolutionary origins, structural dynamics, expression across tissues, and clinical associations.
Full article

Graphical abstract
Open AccessArticle
The Influence of Fluidic Flow Stress on the Development of the Secondary Palate
by
Masayo Nagata, Satoru Hayano, Ziyi Wang, Takahiro Kosami and Hiroshi Kamioka
J. Dev. Biol. 2026, 14(1), 9; https://doi.org/10.3390/jdb14010009 - 12 Feb 2026
Abstract
►▼
Show Figures
Craniofacial development is orchestrated by a finely regulated interplay of numerous genes and signaling pathways. Palatogenesis proceeds through a complex, stepwise process, in which endogenous mechanical stresses within tissues have been implicated. However, the impact of exogenous fluidic flow mechanical stress derived from
[...] Read more.
Craniofacial development is orchestrated by a finely regulated interplay of numerous genes and signaling pathways. Palatogenesis proceeds through a complex, stepwise process, in which endogenous mechanical stresses within tissues have been implicated. However, the impact of exogenous fluidic flow mechanical stress derived from maternal movement on palatal development remains unclear. In this study, we investigated the effect of exogenous fluidic flow mechanical stress on palatal morphogenesis, focusing on the horizontal outgrowth of palatal shelves after elevation. Palatal tissues dissected from mouse embryos were subjected to organ culture with or without mechanical loading (loaded and unloaded groups, respectively). Stress magnitude was quantified by calculating wave energy, and morphometric and molecular analyses were performed. Compared with the unloaded group, palatal shelves in the loaded group showed significant increases in thickness and volume, accompanied by enhanced cell proliferation, nuclear translocation of YAP and β-catenin, and upregulation of the osteogenic markers Osterix and Osteocalcin. No significant difference in apoptosis was observed. These findings indicate that exogenous mechanical stress promotes cell proliferation and osteogenic differentiation through the Hippo and WNT/β-catenin pathways in palate explants. Our results suggest that moderate maternal movement-induced mechanical stress contributes to normal palatogenesis, providing new insights into the mechanisms underlying cleft palate.
Full article

Figure 1
Open AccessArticle
Vestigial-like 4 Regulates Neurogenesis and Neural Crest Formation During Xenopus Development
by
Pierre Thiébaud, Emilie Simon, François Moisan, Sandrine Fedou, Hamid-Reza Rezvani and Nadine Thézé
J. Dev. Biol. 2026, 14(1), 8; https://doi.org/10.3390/jdb14010008 - 11 Feb 2026
Abstract
►▼
Show Figures
VESTIGIAL-LIKE proteins constitute a family of evolutionarily conserved proteins that act as cofactors in regulating gene expression through their binding to TEAD transcription factors. Among the four members of this family in vertebrates, VESTIGIAL-LIKE 4 has emerged as a tumor suppressor that competes
[...] Read more.
VESTIGIAL-LIKE proteins constitute a family of evolutionarily conserved proteins that act as cofactors in regulating gene expression through their binding to TEAD transcription factors. Among the four members of this family in vertebrates, VESTIGIAL-LIKE 4 has emerged as a tumor suppressor that competes with YAP in binding TEADs, thus inhibiting the HIPPO pathway downstream of YAP. Nevertheless, very few studies have addressed its function during early vertebrate development. Here, we used gain- and loss-of-function strategies to investigate the role of vestigial-like 4 during Xenopus laevis development. Our data show that vestigial-like 4 is a key regulator of neurogenesis and neural crest formation. In embryos depleted of vestigial-like 4, neurogenesis is severely impaired, and neither neurog1 nor neurod1 is able to stimulate neurogenesis. Vestigial-like 4 is also required for neural crest formation through pax3 and sox9 regulation, and this property does not necessarily require its interaction with tead. Collectively, our findings demonstrate that vestigial-like 4 is an important regulator of neurogenesis and neural crest formation. Although vestigial-like 4 can bind to tead proteins in the embryo, its function does not depend solely on this interaction, suggesting a complex level of regulation with which vestigial-like 4 regulates early steps in development and differentiation.
Full article

Figure 1
Open AccessArticle
Functional State of Lampbrush Chromosomes in Early Vitellogenic Oocytes of Hibernating Frogs Rana temporaria
by
Nadya V. Ilicheva and Olga I. Podgornaya
J. Dev. Biol. 2026, 14(1), 7; https://doi.org/10.3390/jdb14010007 - 2 Feb 2026
Abstract
►▼
Show Figures
Lampbrush chromosomes (LBCs) are a feature of amphibian oocytes and are typically associated with high levels of transcription during active oocyte growth. However, their state during winter hibernation has not been studied. Here, we investigated LBCs in early vitellogenic oocytes (early stage 4)
[...] Read more.
Lampbrush chromosomes (LBCs) are a feature of amphibian oocytes and are typically associated with high levels of transcription during active oocyte growth. However, their state during winter hibernation has not been studied. Here, we investigated LBCs in early vitellogenic oocytes (early stage 4) of the grass frog Rana temporaria during winter hibernation. We found that the chromosomes retained their lampbrush morphology, and the phosphorylated form of RNA polymerase II resided on the lateral loops. Transcription on the lateral loops was reduced but detectable at cold conditions and significantly increased when the oocytes were transferred at room temperature. Satellite S1a transcripts were detected at the lateral loops of the chromosomes by RNA FISH. The possible significance of maintaining chromosomes in the lampbrush form during hibernation is discussed.
Full article

Figure 1
Open AccessArticle
Female Aging Affects Coilin Pattern in Mouse Cumulus Cells
by
Alexey S. Anisimov, Dmitry S. Bogolyubov and Irina O. Bogolyubova
J. Dev. Biol. 2026, 14(1), 6; https://doi.org/10.3390/jdb14010006 - 15 Jan 2026
Abstract
►▼
Show Figures
Cumulus cells (CCs) are a distinct population of granulosa cells (GCs) that surround the developing and ovulated mammalian oocyte. The features of their structural organization and the expression pattern of key genes significantly affect oocyte viability. Changes in the functional activity of the
[...] Read more.
Cumulus cells (CCs) are a distinct population of granulosa cells (GCs) that surround the developing and ovulated mammalian oocyte. The features of their structural organization and the expression pattern of key genes significantly affect oocyte viability. Changes in the functional activity of the nucleus are often expressed in changes in the structure of nuclear bodies (NBs), including Cajal bodies (CBs). The diagnostic protein of CBs is coilin, which maintains their structural integrity. Using fluorescent and electron microscopy, we examined maternal aging-associated changes in coilin pattern in mouse CCs. We found that older mice had a decrease in the number of coilin-positive bodies, while external transcriptome data analysis revealed no significant changes in Coil and Smn1 gene expression. We hypothesized that the age-related dynamics of coilin-containing bodies are determined not by changes in the expression level of key components of these bodies, but by age-related changes in CC metabolism. Considering that CCs are a by-product of IVF protocols, making them available for analysis in sufficient quantities, age-related changes in the number and size of coilin-positive NBs in CCs may serve as a promising biomarker for assessing ovarian functional aging.
Full article

Graphical abstract
Open AccessArticle
Influence of Obstructive Uropathy on Cyst Formation and Nephrogenesis: Insights from a Fetal Lamb Model
by
Kohei Kawaguchi, Takuya Kawaguchi, Juma Obayashi, Yasuji Seki, Kunihide Tanaka, Kei Ohyama, Junki Koike, Shigeyuki Furuta, Kevin C. Pringle and Hiroaki Kitagawa
J. Dev. Biol. 2026, 14(1), 5; https://doi.org/10.3390/jdb14010005 - 9 Jan 2026
Abstract
Obstructive uropathy (OU) during fetal development induces a fetal cystic dysplastic kidney. The mechanisms of cyst formation and the onset of renal dysfunction remain unclear. Determining whether nephrogenic potential persists during fetal life may suggest whether early intervention could preserve renal development. We
[...] Read more.
Obstructive uropathy (OU) during fetal development induces a fetal cystic dysplastic kidney. The mechanisms of cyst formation and the onset of renal dysfunction remain unclear. Determining whether nephrogenic potential persists during fetal life may suggest whether early intervention could preserve renal development. We aimed to evaluate residual nephrogenic activity in fetal cystic dysplastic kidneys using β-catenin and CD10 immunostaining, and to assess whether the site of obstruction influences cystogenesis. After appropriate approval, 20 timed-gestation fetal lambs had OU created at 60 days. Males underwent urethral and urachal ligation (n = 8, 3 lost), and females underwent unilateral ureteric ligation (n = 8, 1 lost). Fetuses were sacrificed at 80 days (n = 6) and 140 days (term, n = 10), comparing kidneys with normal controls of the same gestational age using immunohistochemical staining for β-catenin and CD10. Developing fetal cystic dysplastic kidneys were identified at 80 days. β-catenin staining showed the absence of granular cytoplasmic expression in cystic regions, indicating arrested nephrogenesis. In male models, cysts originated exclusively from proximal tubules. Female models exhibited mixed proximal and distal tubular involvement. CD10 staining confirmed the loss of proximal tubular markers. Renal development remained arrested at term. Cyst formation disrupts renal development early in gestation, which persists until term. Differences in cystogenesis between the models suggest that the site of obstruction influences pathogenic mechanisms.
Full article
(This article belongs to the Special Issue Developmental Biology of the Kidney: From Molecular Mechanisms to Congenital Disorders)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Discovery of New Markers for Haemogenic Endothelium and Haematopoietic Progenitors in the Mouse Yolk Sac
by
Guillermo Diez-Pinel, Alessandro Muratore, Christiana Ruhrberg and Giovanni Canu
J. Dev. Biol. 2026, 14(1), 4; https://doi.org/10.3390/jdb14010004 - 6 Jan 2026
Abstract
►▼
Show Figures
Erythro-myeloid progenitors (EMPs) originate from the haemogenic endothelium in the yolk sac via an endothelial-to-haematopoietic transition (EHT) to generate blood and immune cells that support embryo development. Yet, the transitory nature of EHT and the limited availability of molecular markers have constrained our
[...] Read more.
Erythro-myeloid progenitors (EMPs) originate from the haemogenic endothelium in the yolk sac via an endothelial-to-haematopoietic transition (EHT) to generate blood and immune cells that support embryo development. Yet, the transitory nature of EHT and the limited availability of molecular markers have constrained our understanding of the origin, identity, and differentiation dynamics of EMPs. Here, we have refined the annotation of yolk sac haemato-vascular populations in publicly available single-cell RNA sequencing (scRNAseq) datasets from mouse embryos to identify novel molecular markers of haemogenic endothelium and EMPs. By sub-clustering key cell populations followed by pseudotime analysis, we refined cluster annotations and then reconstructed differentiation trajectories. Subsequent differential gene expression analysis between clusters identified novel cell surface markers for haemogenic endothelial cells (Fxyd5 and Scarf1) and EMPs (Fcer1g, Tyrobp, and Mctp1). Further, we have identified candidate signalling and metabolic pathways that may regulate yolk sac haematopoietic emergence and differentiation. The specificity of FXYD5, SCARF1, and FCER1G for haemogenic endothelium and EMPs was validated by immunostaining of the mouse yolk sac. These insights into the transcriptional dynamics in the yolk sac should support future investigation of EHT and haematopoietic differentiation during early mammalian development.
Full article

Figure 1
Open AccessReview
The Interplay of One-Carbon Metabolism, Mitochondrial Function, and Developmental Programming in Ruminant Livestock
by
Kazi Sarjana Safain, Kendall C. Swanson and Joel S. Caton
J. Dev. Biol. 2026, 14(1), 3; https://doi.org/10.3390/jdb14010003 - 3 Jan 2026
Abstract
Maternal nutrition during gestation profoundly influences fetal growth, organogenesis, and long-term offspring performance through developmental programming. Among the molecular mechanisms responsive to maternal nutrient availability, one-carbon metabolism plays a central role by integrating folate, methionine, choline, and vitamin B12 pathways that regulate
[...] Read more.
Maternal nutrition during gestation profoundly influences fetal growth, organogenesis, and long-term offspring performance through developmental programming. Among the molecular mechanisms responsive to maternal nutrient availability, one-carbon metabolism plays a central role by integrating folate, methionine, choline, and vitamin B12 pathways that regulate methylation, nucleotide synthesis, and antioxidant defense. These processes link maternal nutritional status to epigenetic remodeling, cellular proliferation, and redox balance during fetal development. Mitochondria act as nutrient sensors that translate maternal metabolic cues into bioenergetic and oxidative signals, shaping tissue differentiation and metabolic flexibility. Variations in maternal diet have been associated with shifts in fetal amino acid, lipid, and energy metabolism, suggesting adaptive responses to constrained intrauterine environments. This review focuses on the molecular interplay between one-carbon metabolism, mitochondrial function, and metabolomic adaptation in developmental programming of ruminant livestock. Understanding these mechanisms offers opportunities to design precision nutritional strategies that enhance fetal growth, offspring productivity, and long-term resilience in livestock production systems.
Full article
(This article belongs to the Special Issue Mitochondrial Function and Dysfunction in Developmental Biology Metabolism)
►▼
Show Figures
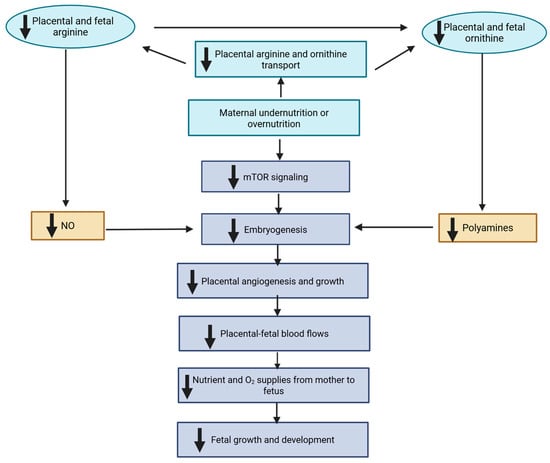
Figure 1
Open AccessArticle
A Zebrafish Seizure Model of cblX Syndrome Reveals a Dose-Dependent Response to mTor Inhibition
by
Claudia B. Gil, David Paz, Briana E. Pinales, Victoria L. Castro, Claire E. Perucho, Annalise Gonzales, Giulio Francia, Sepiso K. Masenga, Antentor Hinton, Jr. and Anita M. Quintana
J. Dev. Biol. 2026, 14(1), 2; https://doi.org/10.3390/jdb14010002 - 25 Dec 2025
Abstract
►▼
Show Figures
Mutations in the transcriptional co-factor HCFC1 cause methylmalonic aciduria and homocystinemia, cblX type (cblX) (MIM#309541), non-syndromic X-linked intellectual disability (XLID), and focal epilepsy. Zebrafish studies have revealed increased activation of the Akt/mTor signaling pathway after mutation of hcfc1a, one ortholog
[...] Read more.
Mutations in the transcriptional co-factor HCFC1 cause methylmalonic aciduria and homocystinemia, cblX type (cblX) (MIM#309541), non-syndromic X-linked intellectual disability (XLID), and focal epilepsy. Zebrafish studies have revealed increased activation of the Akt/mTor signaling pathway after mutation of hcfc1a, one ortholog of HCFC1. mTOR hyperactivation is linked to seizures, and its inhibition alleviates epilepsy in other preclinical models. We hypothesized that mTor overactivity in hcfc1a mutant zebrafish increases seizure susceptibility and/or severity. We employed a two-concentration model of the seizure-inducing agent, pentylenetetrazol (PTZ), with or without pretreatment of the mTor inhibitor, torin1. Mutation of hcfc1a did not alter the response to PTZ at sub-optimal concentrations, and the pharmaceutical inhibition of mTor using the compound Torin1 reduced response to 1 µM PTZ, but only in a dose-dependent manner. Higher doses of mTor inhibition did not reduce the seizure response in mutant larvae but were effective in wildtype siblings. These data suggest that inhibition of mTor in an hcfc1a-deficient background leads to a reaction that differs from the traditional response observed in wildtype siblings. Collectively, we present a model that can be used to test dose–response and the development of combinatorial treatment approaches in a high-throughput manner.
Full article

Graphical abstract
Open AccessArticle
The Epithelial Egg Tooth of the Chicken Shares Protein Markers with the Embryonic Subperiderm and Feathers
by
Attila Placido Sachslehner, Julia Steinbinder, Claudia Hess, Veronika Mlitz and Leopold Eckhart
J. Dev. Biol. 2026, 14(1), 1; https://doi.org/10.3390/jdb14010001 - 22 Dec 2025
Abstract
The epithelial egg tooth is used by birds to open the eggshell for hatching. This ectodermal structure consists of a multilayered periderm and a hard cornified portion, the caruncle or actual egg tooth. Here, we determined the protein composition of the egg tooth
[...] Read more.
The epithelial egg tooth is used by birds to open the eggshell for hatching. This ectodermal structure consists of a multilayered periderm and a hard cornified portion, the caruncle or actual egg tooth. Here, we determined the protein composition of the egg tooth of the chicken and compared the proteins to markers of other epithelia identified in previous studies. The egg tooth and the upper beak of chicken embryos of Hamburger and Hamilton (HH) stage 44 were subjected to mass spectrometry-based proteomics. We found that scaffoldin, a marker of the embryonic periderm and the feather sheath, was enriched in the egg tooth relative to the beak. Likewise, Epidermal Differentiation protein containing DPCC Motifs (EDDM) and Epidermal Differentiation protein starting with a MTF motif and rich in Histidine (EDMTFH), which had previously been characterized as markers of the subperiderm on embryonic scutate scales and the barbs of feathers, were also enriched in the egg tooth. The expression of EDDM and EDMTFH was confirmed RT-PCR analysis. Our data suggest that the epithelial egg tooth is related to the subperiderm and feathers, a hypothesis with potentially important implications for the evolution of the avian integument.
Full article
(This article belongs to the Special Issue Feature Papers from Journal of Developmental Biology Reviewers, 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessReview
Pathophysiology and Management of Placenta Accreta Spectrum
by
Lana Shteynman, Genevieve Monanian, Gilberto Torres, Giancarlo Sabetta, Deborah M. Li, Zhaosheng Jin, Tiffany Angelo, Bahaa E. Daoud and Morgane Factor
J. Dev. Biol. 2025, 13(4), 45; https://doi.org/10.3390/jdb13040045 - 10 Dec 2025
Cited by 1
Abstract
►▼
Show Figures
Placenta Accreta Spectrum (PAS) disorders, including placenta accreta, increta, and percreta, are serious obstetric conditions characterized by abnormal placental adherence to the uterine wall. With increasing incidence, PAS poses significant risks, primarily through massive hemorrhage during or after delivery, often necessitating hysterectomy. Key
[...] Read more.
Placenta Accreta Spectrum (PAS) disorders, including placenta accreta, increta, and percreta, are serious obstetric conditions characterized by abnormal placental adherence to the uterine wall. With increasing incidence, PAS poses significant risks, primarily through massive hemorrhage during or after delivery, often necessitating hysterectomy. Key risk factors include prior cesarean sections, uterine surgery, and placenta previa diagnosis. In this review, we will examine the pathophysiology of PAS, with a focus on the mechanisms underlying abnormal trophoblast invasion and defective decidualization. We will highlight the role of uterine scarring, extracellular matrix remodeling, dysregulated signaling pathways, and immune and vascular alterations in disrupting the maternal-fetal interface, ultimately predisposing to morbid placentation and delivery complications. We will also discuss the life-threatening complications of PAS, such as shock and multi-organ failure, which require urgent multidisciplinary intensive care, as well as the optimization of management through preoperative planning and intraoperative blood loss control to reduce maternal morbidity and mortality.
Full article

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics

Special Issues
Special Issue in
JDB
Genetic and Epigenetic Mechanisms in Gametogenesis and Early Development: Insights from Models, Stem Cells, and Human Disorders
Guest Editors: Lon J. van Winkle, Philip Iannaccone, Rebecca Jean RyznarDeadline: 15 August 2026
Special Issue in
JDB
Mechanisms of Morphogenesis, Degeneration, and Regeneration
Guest Editors: Tsutomu Nohno, Hideyo Ohuchi, Naoyuki WadaDeadline: 16 September 2026
Special Issue in
JDB
Mitochondrial Function and Dysfunction in Developmental Biology Metabolism
Guest Editor: Simon J. ConwayDeadline: 15 October 2026
Special Issue in
JDB
Feature Papers in Journal of Developmental Biology 2026
Guest Editor: Simon J. ConwayDeadline: 31 December 2026
Topical Collections
Topical Collection in
JDB
Hedgehog Signaling in Embryogenesis
Collection Editors: Henk Roelink, Kay Grobe









