Journal Description
Pathogens
Pathogens
is an international, peer-reviewed, open access journal on pathogens and pathogen-host interactions published monthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, CaPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 14.1 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Pathogens include: Parasitologia and Bacteria.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
3.3 (2024);
5-Year Impact Factor:
3.6 (2024)
Latest Articles
Rainfall-Driven Mobilisation of Clinically Relevant Burkholderia pseudomallei in a Groundwater-Connected Urban Creek, Northern Australia
Pathogens 2026, 15(3), 276; https://doi.org/10.3390/pathogens15030276 (registering DOI) - 3 Mar 2026
Abstract
Burkholderia pseudomallei is a saprophytic environmental bacterium and the causative agent of melioidosis, a serious opportunistic infection in tropical regions, including northern Australia. Infection occurs following environmental exposure via percutaneous inoculation, ingestion, or inhalation; however, the environmental reservoirs and transmission pathways responsible for
[...] Read more.
Burkholderia pseudomallei is a saprophytic environmental bacterium and the causative agent of melioidosis, a serious opportunistic infection in tropical regions, including northern Australia. Infection occurs following environmental exposure via percutaneous inoculation, ingestion, or inhalation; however, the environmental reservoirs and transmission pathways responsible for human disease remain poorly defined. Groundwater has been implicated as a potential source of infection, but the factors influencing the persistence and mobility of B. pseudomallei in surface waters in North Queensland are not well understood. Water samples were collected from a groundwater-connected seasonal creek in Townsville, North Queensland, over a 12-month period encompassing wet and dry seasons. Samples were cultured on Ashdown agar and confirmed as B. pseudomallei by qPCR. Multi-locus sequence typing (MLST) was performed using targeted allele sequencing on the Oxford Nanopore MinION platform. Eighteen of 59 water samples were culture-positive for B. pseudomallei. Detection occurred exclusively in turbid, flowing water following ≥30 mm of rainfall and was observed in both wet and dry seasons. MLST of 48 isolates identified 18 sequence types, including 12 novel types. Six sequence types matched previously reported Townsville clinical isolates. These findings indicate that groundwater from a connected urban creek may function as a mobile reservoir for clinically relevant B. pseudomallei strains under specific hydrological and climatic conditions, highlighting rainfall-driven processes as key drivers of environmental exposure risk.
Full article
(This article belongs to the Section Bacterial Pathogens)
►
Show Figures
Open AccessArticle
Characterization of the Gut Microbiome of Patients with Clostridioides difficile Infection and Healthy Individuals in Greece
by
Dimitra Mougiou, Georgia Gioula, Lemonia Skoura, Fani Minti, Theodoros Karampatakis, Dimitrios Malandris, Konstantinos Pelekoudas and Melania Kachrimanidou
Pathogens 2026, 15(3), 275; https://doi.org/10.3390/pathogens15030275 (registering DOI) - 3 Mar 2026
Abstract
►▼
Show Figures
Background/Objectives: The gut microbiome plays an important role in the colonization of an individual by Clostridioides difficile and in the development of Clostridioides difficile infection (CDI). The main purpose of this study was to compare the gut microbiomes of patients with CDI and
[...] Read more.
Background/Objectives: The gut microbiome plays an important role in the colonization of an individual by Clostridioides difficile and in the development of Clostridioides difficile infection (CDI). The main purpose of this study was to compare the gut microbiomes of patients with CDI and healthy individuals. Methods: We prospectively included 48 individuals: 32 patients with CDI and 16 healthy individuals. Microbiomes were analyzed by sequencing the hypervariable regions of the 16S rRNA gene using an Ion GeneStudio™ S5 System. Further statistical analysis of microbiome data was performed with the open-source programming language R version 3.5.2. Results: Among the CDI patients, Firmicutes and Proteobacteria were the most abundant phyla, while Enterobacteriaceae and Enterococcaceae were the most abundant families. Genus-level analysis showed that Enterococcus was the dominant genus in CDI patients; in contrast, in healthy individuals, Faecalibacterium was the most abundant. The MaAsLin2 tool revealed that members of the family Enterococcaceae and the genus Enterococcus were more abundant in patients with CDI than in healthy individuals. Alpha and beta diversity did not reveal differences between the two study groups. Conclusions: We observed differences in microbiome patterns between healthy individuals and CDI patients that were consistent with the literature. Further studies are needed.
Full article
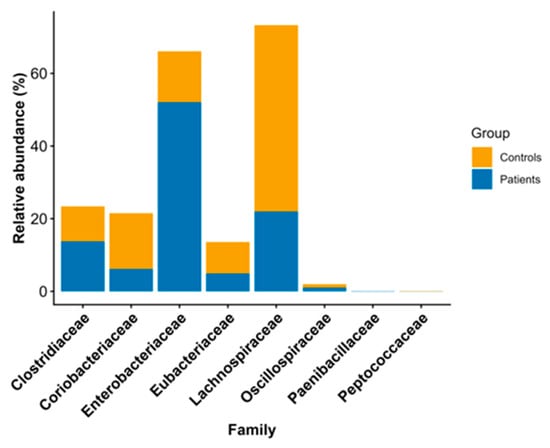
Figure 1
Open AccessArticle
Prevalence, Virulence Determinants, and Antimicrobial Resistance of Staphylococcus aureus and Escherichia coli Isolated from Computer Devices Used by Staff and Students at a Northern Thailand University
by
Sorawit Upakut, Achiraya Siriphap, Ornampai Japa, Pathumwan Watsing, Peerapat Bunpak, Aacharaporn Ta-In, Cholthicha Inmanee, Chutamas Thepmalee, Nittiya Suwannasom and Krissana Khoothiam
Pathogens 2026, 15(3), 274; https://doi.org/10.3390/pathogens15030274 - 3 Mar 2026
Abstract
Computer devices in university settings are frequently shared and repeatedly handled, making them potential reservoirs for pathogenic bacteria. This study aimed to investigate the prevalence, virulence determinants, and antimicrobial resistance profiles of Staphylococcus aureus and Escherichia coli isolated from computer devices used by
[...] Read more.
Computer devices in university settings are frequently shared and repeatedly handled, making them potential reservoirs for pathogenic bacteria. This study aimed to investigate the prevalence, virulence determinants, and antimicrobial resistance profiles of Staphylococcus aureus and Escherichia coli isolated from computer devices used by staff and students at a university in Northern Thailand. A total of 400 computer devices were sampled, with each device defined as a single sampling unit comprising both the keyboard and computer mouse. Bacterial identification was performed using PCR, while staphylococcal enterotoxin (se) genes and diarrheagenic E. coli (DEC)-associated virulence genes were detected by PCR. Antimicrobial susceptibility was assessed using the disk diffusion method. Overall, 74 (18.5%) S. aureus isolates and 6 (1.5%) E. coli isolates were recovered. The highest prevalence of S. aureus was observed among personal-use student computer devices (29%; p < 0.001), whereas E. coli was most frequently detected on public-use staff computer devices (4%). Among S. aureus isolates, 24.3% (18/74) carried at least one se gene, with sec being the most prevalent (13.5%). Half of the E. coli isolates harbored the astA gene. Low resistance rates (<10%) were observed among S. aureus; however, four isolates (5.4%) were classified as MRSA, three of which exhibited multidrug resistance. All E. coli isolates were resistant to ampicillin, and 50% displayed multidrug-resistant phenotypes. These findings suggest that computer devices can act as occasional reservoirs of potentially pathogenic and antimicrobial-resistant bacteria in university environments.
Full article
(This article belongs to the Special Issue Advanced Antimicrobial Agents: Combatting Multi-Drug Resistant Bacterial Infections)
►▼
Show Figures

Figure 1
Open AccessReview
Trypanosoma vivax in Water Buffaloes (Bubalus bubalis): A Host-Centered Synthesis of Pathogenesis, Epidemiology, Diagnosis, and Integrated Control with Implications for Tropical Production Systems
by
André de Medeiros Costa Lins, Dryelle Vieira de Oliveira Brandão, Fernanda Monik Silva Martins, Aline Maia Silva, Henrique dos Anjos Bonjardim and Felipe Masiero Salvarani
Pathogens 2026, 15(3), 273; https://doi.org/10.3390/pathogens15030273 - 3 Mar 2026
Abstract
Trypanosoma vivax is a hemoprotozoan parasite of major veterinary importance affecting domestic ungulates in Africa and the Americas. While traditionally addressed within cattle-centered paradigms, accumulating evidence indicates that water buffaloes (Bubalus bubalis) are both clinically susceptible and epidemiologically significant hosts. This
[...] Read more.
Trypanosoma vivax is a hemoprotozoan parasite of major veterinary importance affecting domestic ungulates in Africa and the Americas. While traditionally addressed within cattle-centered paradigms, accumulating evidence indicates that water buffaloes (Bubalus bubalis) are both clinically susceptible and epidemiologically significant hosts. This structured narrative review provides a host-centered synthesis of global evidence on T. vivax infection in buffaloes, integrating pathogenesis, transmission biology, epidemiology, diagnostics, chemotherapy, and integrated control. The analysis encompasses literature from 2000 to 2025 and incorporates seminal experimental studies published prior to 2000 that established buffalo susceptibility and reservoir competence. Evidence from cyclical (tsetse-mediated) and mechanical transmission systems is comparatively interpreted to clarify host–parasite dynamics. The Amazon biome is discussed as a model system for high-density buffalo production under mechanical vector pressure, offering case-based contextualization without geographic restriction. Particular attention is given to immunopathological mechanisms, chronic low-parasitemia carriage, diagnostic sensitivity in subclinical infections, emerging trypanocide resistance, and ecological constraints on vector control. Controversies and buffalo-specific knowledge gaps are highlighted throughout. By adopting a buffalo-centered analytical framework, this review supports translational diagnostics, targeted surveillance, and sustainable control strategies for trypanosomiasis in tropical livestock systems.
Full article
(This article belongs to the Topic Advances in Infectious and Parasitic Diseases of Animals)
Open AccessArticle
Beyond the Mutation Abyss: Revisiting SARS-CoV-2 Receptor-Binding Domain Evolution from ACE2 Binding Optimization to Immune Epitope Remodeling
by
Omar A. Soliman, Yasmine Shahine, Daniel Baecker and Ahmed Noby Amer
Pathogens 2026, 15(3), 272; https://doi.org/10.3390/pathogens15030272 - 3 Mar 2026
Abstract
The SARS-CoV-2 Omicron variant and its descendants accumulated unprecedented numbers of spike substitutions yet remained transmissible, implying compensatory mechanisms that preserve entry while eroding humoral immunity. We analyzed 32 variants for sequence-level mutation, physicochemical profiling, and epitope disruption; 25 had growth-advantage estimates, and
[...] Read more.
The SARS-CoV-2 Omicron variant and its descendants accumulated unprecedented numbers of spike substitutions yet remained transmissible, implying compensatory mechanisms that preserve entry while eroding humoral immunity. We analyzed 32 variants for sequence-level mutation, physicochemical profiling, and epitope disruption; 25 had growth-advantage estimates, and 18 underwent molecular dynamics/MM-PBSA simulations. We applied a systems-virology framework to the SARS-CoV-2 receptor-binding domain (RBD), integrating immunodominance-weighted epitope conservation (567 B-cell and 97 T-cell epitopes) across variants (Wuhan-Hu-1 to KP.3) with molecular dynamics, molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) binding energetics, and deep mutational scanning (DMS) benchmarking. B-cell epitope conservation declined from a median of 72.7% in pre-Omicron variants to 28.8% in BA.1 and 10.6% in KP.3, and was strongly inversely associated with a breakthrough-infection proxy (Spearman ρ = −0.8246, p < 0.001), whereas RBD T-cell epitopes remained comparatively conserved (91.5% to 87.2%). Despite the loss of the ancestral K417–ACE2 D30 salt bridge, Omicron reconfigured the interface via alternative electrostatic contacts (Q493R–E35 and Q498R–D38), producing compensatory interactions captured by MM-PBSA, but with only modest agreement with DMS affinity changes (r = 0.682, p = 0.007), consistent with enthalpy–entropy compensation. Finally, mutation tolerance shifted toward stronger epistatic buffering in Omicron (two-fold higher epistasis than pre-Omicron; p = 0.0093), enabling extensive antigenic change without structural collapse. Together, these results support a multi-objective evolutionary strategy—epitope erosion, interface rewiring, and epistatic compensation—that can be operationalized to prioritize emerging lineages for surveillance and to inform vaccine designs that emphasize conserved T-cell targets.
Full article
(This article belongs to the Special Issue Antimicrobial Resistance in the Post-COVID Era: A Silent Pandemic)
►▼
Show Figures

Figure 1
Open AccessArticle
Effect of trans-Cinnamaldehyde on Adhesion and Other Virulence Factors of Methicillin-Resistant Staphylococcus aureus
by
Barbara Kot, Kamila Wierzchowska, Agata Grużewska, Elżbieta Anna Trafny, Małgorzata Stępińska and Małgorzata Witeska
Pathogens 2026, 15(3), 271; https://doi.org/10.3390/pathogens15030271 - 3 Mar 2026
Abstract
Methicillin-resistant Staphylococcus aureus (MRSA) produces virulence factors and causes hard-to-treat infections. This study aimed to evaluate the effect of trans-cinnamaldehyde (TC) on the selected virulence factors of MRSA: adhesion to host plasma and extracellular matrix proteins, protease, DNase and esterase production, and
[...] Read more.
Methicillin-resistant Staphylococcus aureus (MRSA) produces virulence factors and causes hard-to-treat infections. This study aimed to evaluate the effect of trans-cinnamaldehyde (TC) on the selected virulence factors of MRSA: adhesion to host plasma and extracellular matrix proteins, protease, DNase and esterase production, and hemolytic activity. Our results showed that TC at ½ MBIC (Minimum Biofilm Inhibition Concentration) of 240 µg/mL or 60 µg/mL, depending on the isolate, significantly reduced MRSA adhesion. Inhibition varied between isolates, ranging from 26.1% to 41.3% (fibrinogen), 18.2% to 34.9% (elastin), 26.5% to 32.4% (laminin), and 17.1% to 30.5% (collagen). TC at ½ MIC (Minimum Inhibitory Concentration) of 30 µg/mL also significantly inhibited MRSA enzyme production, and reduced hemolytic activity (by 80.0–83.1%, depending on the isolate). TC may be an alternative to antibiotics for combating infections caused by S. aureus, as it not only reduces bacterial survival in the host but also reduces S. aureus virulence at subinhibitory concentrations. TC at higher concentrations exhibits cytotoxicity in human fibroblasts, limiting its topical use. Therefore, to exploit TC’s antibacterial potential, it is necessary to identify substances that act synergistically with it, enabling reduced effective doses.
Full article
(This article belongs to the Special Issue Challenges of Multidrug-Resistant Bacteria and Drug Development of Antibacterials)
►▼
Show Figures
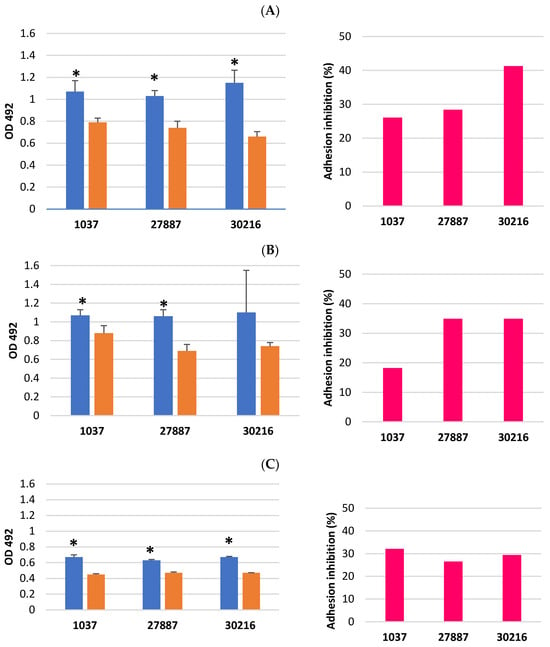
Figure 1
Open AccessReview
Plastic-Derived Pollutants as Emerging Modifiers of Viral Diseases
by
Fatima Hisam, Ramina Kordbacheh, Ebenezer Senu, Spandan Mukherjee, Jon Sin and Erica L. Sanchez
Pathogens 2026, 15(3), 270; https://doi.org/10.3390/pathogens15030270 (registering DOI) - 3 Mar 2026
Abstract
Plastic pollutants, including phthalates, bisphenol A (BPA), per- and polyfluoroalkyl substances (PFAS), and microplastics (MPs), are increasingly recognized as emerging environmental cofactors that intersect with infectious disease dynamics. These compounds, once considered inert, can alter immune function, reshape host–pathogen interactions, and directly influence
[...] Read more.
Plastic pollutants, including phthalates, bisphenol A (BPA), per- and polyfluoroalkyl substances (PFAS), and microplastics (MPs), are increasingly recognized as emerging environmental cofactors that intersect with infectious disease dynamics. These compounds, once considered inert, can alter immune function, reshape host–pathogen interactions, and directly influence viral survival and transmission. In this review, we compile current evidence on the chemistry, environmental occurrence, and biological activity of major plastic-associated pollutants with emphasis on their role in viral infections. Phthalates such as di(2-ethylhexyl) phthalate (DEHP) and its metabolite MEHP modulate innate immune signaling and have been shown to exacerbate infections, including Dengue and Coxsackievirus B3. Other DEHP-like phthalates, such as dibutyl phthalate (DBP), exhibit consistent infection-enhancing effects, while high molecular weight or cyclical phthalates such as polyvinyl acetate phthalate (PVAP) display conflicting results in their modulation of viral infections. BPA, widely detected in human tissues, acts through endocrine and immune disruption, worsening viral myocarditis, and altering influenza outcomes. PFAS, persistent “forever chemicals,” reshape adaptive immune responses and are associated with increased susceptibility, viral persistence, or severity of infection of herpesvirus (HCMV, EBV, HSV-1), hepatitis virus, and influenza infection. Microplastics represent a distinct risk by acting as physical carriers for viruses and bacteria, stabilizing viral RNA, enhancing host cell uptake, and skewing immune responses. Together, these pollutants extend beyond toxicology into virology, providing novel insights into how environmental exposures converge with viral pathogenesis. We highlight mechanistic advances and critical knowledge gaps and propose future directions for integrating environmental health and infectious disease research.
Full article
(This article belongs to the Section Viral Pathogens)
►▼
Show Figures

Figure 1
Open AccessArticle
Pro-Inflammatory Response of Bovine Lung Explant Induced by Mycoplasma mycoides subsp. mycoides
by
Leruo Keokilwe, Giovanni Di Teodoro, Marta Di Federico, Massimo Ancora, Ivanka Krasteva, Gianluca Orsini, Cesare Camma, Fabrizia Perletta, Chiara Di Pancrazio, Mirella Luciani, Chandapiwa Marobela-Raborokgwe, Massimo Scacchia and Flavio Sacchini
Pathogens 2026, 15(3), 269; https://doi.org/10.3390/pathogens15030269 - 3 Mar 2026
Abstract
Contagious bovine pleuropneumonia (CBPP) is a significant respiratory disease in cattle caused by Mycoplasma mycoides subsp. mycoides (Mmm). A better understanding of the pathogenesis of CBPP and the immune response of the host to infection will assist in the development of
[...] Read more.
Contagious bovine pleuropneumonia (CBPP) is a significant respiratory disease in cattle caused by Mycoplasma mycoides subsp. mycoides (Mmm). A better understanding of the pathogenesis of CBPP and the immune response of the host to infection will assist in the development of novel interventions to prevent disease progression. In this study, bovine lung explants (BLEs) were exposed to Mmm to investigate the upregulation and release of early inflammatory cytokines, mediators and receptors following tissue infection. Immunomodulatory molecules indicative of cell activation were investigated by immunoblotting on the BLEs and the tissue culture supernatants, and quantitative real-time PCR (RTq-PCR) was performed on the BLEs to determine the fold change in the expression of the respective mRNA. Immunoblotting indicated the production of inflammatory cytokines, mediators and receptors in Mmm-infected BLEs; however, this contrasted strongly with the mRNA expression profile, which did not show any significant fold increase. Infection of the BLEs with Mmm stimulated the production of some pro-inflammatory cytokines and mediators, including IL-1β, COX-2, 5-LOX and iNOS. Toll-like receptor proteins TLR2 and TLR4 were detected solely in the tissue culture supernatant of Mmm-infected BLEs. These receptors are considered to be involved in the recognition of Mmm by BLE tissue cells, thus triggering intracellular pathways that produce specific inflammatory cytokines and mediators, initiating the inflammatory response.
Full article
(This article belongs to the Special Issue Mycoplasmas in Respiratory Tract Infections of Cattle: 2nd Edition)
►▼
Show Figures
Figure 1
Open AccessArticle
Acomys cahirinus seurati as a Potential Reservoir Host of Leishmania major
by
Sergei Karlin, Barbora Bečvářová, Kamal Eddine Benallal, Tomáš Bečvář, Ghania Mezai, Mounir Zaabar, Mohamed Mefissel, Petr Volf and Jovana Sádlová
Pathogens 2026, 15(3), 268; https://doi.org/10.3390/pathogens15030268 - 3 Mar 2026
Abstract
►▼
Show Figures
Cutaneous leishmaniasis (CL) caused by Leishmania major is a zoonotic disease transmitted by phlebotomine sand flies. Identification of reservoir hosts is critical for understanding transmission and guiding control. While Psammomys obesus and Meriones shawi are recognized reservoirs in North Africa, the role of
[...] Read more.
Cutaneous leishmaniasis (CL) caused by Leishmania major is a zoonotic disease transmitted by phlebotomine sand flies. Identification of reservoir hosts is critical for understanding transmission and guiding control. While Psammomys obesus and Meriones shawi are recognized reservoirs in North Africa, the role of other rodents remains unclear. We experimentally assessed the reservoir potential of Acomys cahirinus seurati from southeastern Algeria. Animals were intradermally infected, and clinical signs and infectiousness to sand flies were monitored. Parasite persistence in skin and visceral tissues was measured by quantitative PCR. All animals developed localized, self-limiting lesions. Seven of sixteen individuals transmitted parasites to Phlebotomus papatasi between 10 and 25 weeks post-infection. Parasites were mostly restricted to the inoculation site, with limited dissemination to contralateral pinnae and hindpaws, and rare presence in spleen or liver. Notably, some animals were infectious without visible lesions, indicating that ulceration is not required for transmission. These findings show that A. cahirinus seurati can maintain L. major infection for up to 6 months and transmit parasites to sand flies, fulfilling key criteria of a reservoir host. Thus, this species may contribute to CL transmission in endemic foci lacking classical reservoirs, emphasizing the need to consider alternative rodent hosts in surveillance and control programs in North Africa.
Full article
Graphical abstract
Open AccessCommunication
Characterization of Salmonella spp. Based on CRISPR PCR and FT-IR Approaches: A Pilot Study in Northern Italy
by
Clara Tramuta, Irene Floris, Monica Pitti, Giulia Federica Cazzaniga, Miriam Cordovana and Daniela Manila Bianchi
Pathogens 2026, 15(3), 267; https://doi.org/10.3390/pathogens15030267 - 2 Mar 2026
Abstract
The aim of the present study was to employ a genotypic method (CRISPR PCR) and a phenotypic method (FT-IR combined with Linear Discriminant Analysis, LDA) for the rapid identification and discrimination of major virulent Salmonella serovars. Specifically, a total of 24 Salmonella Enteritidis,
[...] Read more.
The aim of the present study was to employ a genotypic method (CRISPR PCR) and a phenotypic method (FT-IR combined with Linear Discriminant Analysis, LDA) for the rapid identification and discrimination of major virulent Salmonella serovars. Specifically, a total of 24 Salmonella Enteritidis, 24 Salmonella Typhimurium, 24 S. Typhimurium 4,5,12:i:-, and 14 Salmonella Infantis strains, previously serotyped according to the Kaufmann–White scheme, were analyzed. CRISPR PCR generated serotype-specific amplicons: 850 bp for S. Enteritidis, 700 bp and 2000 bp for S. Typhimurium, 1300 bp and 1500 bp for S. Typhimurium 4,5,12:i:-, and 1000 bp and 1900 bp for S. Infantis. FT-IR spectra of all serotypes were analyzed by LDA, which revealed a clear separation of three serotype-defined groups: S. Enteritidis, S. Infantis, and a group comprising S. Typhimurium and its monophasic variant 4,5,12:i:-. Further discrimination based on CRISPR-defined classes was observed within each serovar: two classes for S. Typhimurium, two for S. Typhimurium 4,5,12:i:-, and three for S. Infantis. Overall, the combined CRISPR PCR and FT-IR-LDA approach provides preliminary evidence supporting the potential application of a combined approach for the epidemiological surveillance of Salmonella.
Full article
(This article belongs to the Special Issue Advances in Salmonella Epidemiology and Pathogenesis)
►▼
Show Figures

Figure 1
Open AccessArticle
Identification of TgENT1 as the TgUUT1 Uracil/Uridine Transporter of Toxoplasma gondii
by
Hamza A. A. Elati, Mariana Ferriera Silva, Lilach Sheiner and Harry P. de Koning
Pathogens 2026, 15(3), 266; https://doi.org/10.3390/pathogens15030266 - 2 Mar 2026
Abstract
The protozoan pathogen Toxoplasma gondii is responsible for toxoplasmosis, a disease that can be deadly in immunocompromised patients and the developing fetus during pregnancy. Current treatments are widely considered to be suboptimal. We have recently reported that 5-fluoropyrimidines have highly promising anti-toxoplasmosis effects
[...] Read more.
The protozoan pathogen Toxoplasma gondii is responsible for toxoplasmosis, a disease that can be deadly in immunocompromised patients and the developing fetus during pregnancy. Current treatments are widely considered to be suboptimal. We have recently reported that 5-fluoropyrimidines have highly promising anti-toxoplasmosis effects and are internalized by the parasite by a high-affinity uracil/uridine transporter, TgUUT1. Here, we attempt to identify the gene encoding this transport protein. The only nucleoside or nucleobase family identified in the T. gondii genome was the Equilibrative Nucleoside Transporter (ENT) family, with four members. Of these, TgAT1 is known to be purine-specific, and deletion of the TgENT2 and TgENT3 genes, either separately or jointly, did not affect uridine transport or sensitivity to 5-fluoropyrimidines. In contrast, depletion of TgENT1, an essential gene, resulted in a significant reduction in the uptake of both uracil and uridine but not of the amino acid tryptophan. Moreover, expression of TgENT1 in a Leishmania mexicana cell line with low endogenous uracil uptake rates significantly increased uracil uptake for these cells. We conclude that it is highly probable that TgENT1 encodes the T. gondii uracil/uridine transporter. On the basis of our previous results, we infer that TgENT1 likely also mediates the uptake of 5-fluoropyrimidines.
Full article
(This article belongs to the Special Issue Toxoplasmosis in Humans and Animals)
►▼
Show Figures
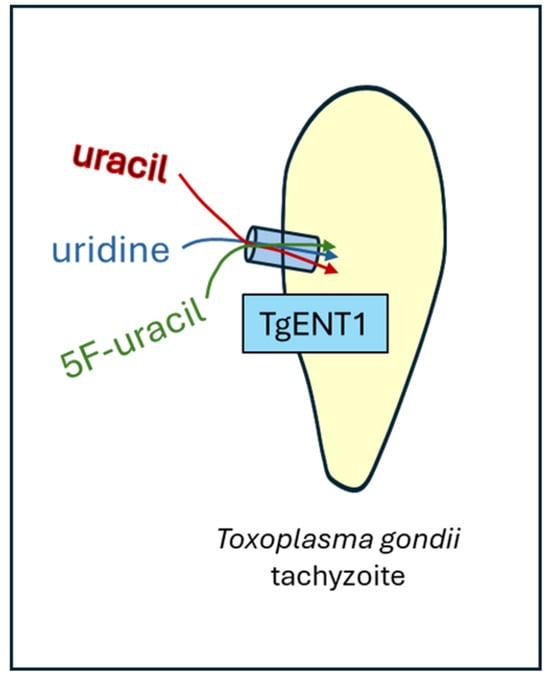
Graphical abstract
Open AccessArticle
The Exploration of Therapeutic Antivirals for Human Papillomavirus in the Last 40 Years: Bibliometric Research
by
Zixiao Jiang, Liangrui Jin, Chengjun Wu, Zhenqing Li, Zhangrong Lou and Peng Qu
Pathogens 2026, 15(3), 265; https://doi.org/10.3390/pathogens15030265 - 2 Mar 2026
Abstract
Human papillomavirus (HPV) is a well-known carcinogenic DNA virus, responsible for about 4% of all cancer cases globally. Effective antiviral treatments for those who are already infected with HPV are still in their early stages, despite the fact that prophylactic vaccinations have shown
[...] Read more.
Human papillomavirus (HPV) is a well-known carcinogenic DNA virus, responsible for about 4% of all cancer cases globally. Effective antiviral treatments for those who are already infected with HPV are still in their early stages, despite the fact that prophylactic vaccinations have shown impressive success in preventing new infections. Effective treatments for HPV-related malignancies are also hampered by the fact that current articles address a wide spectrum of pathways but lack thorough systematic studies. In this work, we use bibliometric techniques to examine research trends and innovative approaches in the development of HPV antivirals over the last 40 years. Our results are intended to offer insightful information and direct future research into effective antiviral treatments for HPV-induced cancers.
Full article
(This article belongs to the Special Issue Epidemiology, Diagnostics, and Therapeutics of High-Risk Human Papillomavirus (HPV) Infections)
►▼
Show Figures

Figure 1
Open AccessArticle
Integrated Laboratory Evaluation of Rift Valley Fever Virus Antibodies Using the Competitive ELISA and Virus Neutralization Test
by
Ommer M. Dafalla, Abdullah A. Alashor, Mohammed O. Hussien, Elsiddig M. Noureldin, Tellal B. Ageep, Mohammed A. Najmi, Mohamed S. Mohamed, Ali A. Hakami, Saleh A. Alrashedi, Fisal A. Bushlaibi and Fahad N. Abukhalil
Pathogens 2026, 15(3), 264; https://doi.org/10.3390/pathogens15030264 - 2 Mar 2026
Abstract
Background: Rift Valley fever virus (RVFV) is a significant mosquito-borne zoonotic virus with high public health and veterinary importance in Africa and the Middle East. Reliable diagnostic assays for detecting antibodies and assessing their functional neutralizing capacity are essential for surveillance programs, vaccine
[...] Read more.
Background: Rift Valley fever virus (RVFV) is a significant mosquito-borne zoonotic virus with high public health and veterinary importance in Africa and the Middle East. Reliable diagnostic assays for detecting antibodies and assessing their functional neutralizing capacity are essential for surveillance programs, vaccine monitoring, and outbreak preparedness. Objective: This study evaluates and compares the analytical performance of a competitive enzyme-linked immunosorbent assay (cELISA) and a virus neutralization test (VNT) for detecting RVFV antibodies in vaccinated sheep sera, establishing an integrated laboratory workflow for virus titration, serological detection, and functional neutralization. Methods: Twenty serum samples were collected from sheep pre-vaccination and one month post-vaccination with Smithburn live attenuated RVFV vaccine. Sera were tested using a commercial multispecies RVFV competitive ELISA to detect antibodies specific to the viral nucleocapsid protein. Viral titration was conducted in Vero cells, and 50% tissue culture infective dose (TCID50/0.1 mL) was calculated using the Reed and Muench method. VNT was performed at 24, 48, 72, and 96 h after infection with different viral doses (102 to 105 TCID50/0.1 mL), and the neutralizing ability of serial serum dilutions (1:2 to 1:1024) was tested. Compared with the control, protection was determined by cytopathic effect (CPE) inhibition. Results: ELISA revealed robust antibody signals up to a 1:32 dilution, with signal-to-noise (S/N) < 40%, whereas for higher dilutions, antibody detection became inconclusive or negative. Virus titration was performed to verify a stock concentration of 106.5 TCID50/0.1 mL. The VNT exhibited time- and dose-dependent kinetics; high protection rates (≥97) were observed at 1:2–1:8 dilutions against 102–103 TCID50/0.1 mL challenge doses; however, neutralizing efficacy decreased significantly at higher viral loads and higher serum dilutions. While cELISA and VNT results correlated strongly at low serum dilutions, the cELISA showed decreased sensitivity at dilutions ≥ 1:64, where the VNT remained capable of detecting functional neutralizing activity. Conclusions/Discussion: The results demonstrate that while both assays correlate well at high antibody concentrations, they diverge at lower concentrations. This discrepancy highlights the functional difference between binding antibodies (N-protein) and neutralizing antibodies (Gn/Gc glycoproteins). Consequently, the cELISA is ideal for rapid screening, whereas the VNT is indispensable for confirming functional immunity. Integrating both assays provides a more accurate immunological profile for RVFV surveillance and vaccine evaluation.
Full article
(This article belongs to the Section Viral Pathogens)
►▼
Show Figures

Figure 1
Open AccessArticle
Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate
by
Verónica I. Hernández-Ramírez, Hugo Varela-Rodríguez, Luis Varela-Rodríguez, Francisco Sierra-López, Daniela Eloísa San Juan-Mora, José Daniel Morales-Mora, Daniela Falcón-Navarrete, Carlos Osorio-Trujillo, Jacqueline Ríos-López, Itzel Berenice Rodríguez-Mera, María Maricela Carrasco-Yépez and Patricia Talamás-Rohana
Pathogens 2026, 15(3), 263; https://doi.org/10.3390/pathogens15030263 - 2 Mar 2026
Abstract
Acanthamoeba castellanii, an opportunistic free-living amoeba, causes severe infections including Acanthamoeba keratitis. This exploratory study evaluated whether three non-steroidal anti-inflammatory drugs (NSAIDs)—acetylsalicylic acid, ibuprofen, and diclofenac (100 µM)—modulate pathogenicity-related processes in A. castellanii and explored the involvement of a gp63-like protein during
[...] Read more.
Acanthamoeba castellanii, an opportunistic free-living amoeba, causes severe infections including Acanthamoeba keratitis. This exploratory study evaluated whether three non-steroidal anti-inflammatory drugs (NSAIDs)—acetylsalicylic acid, ibuprofen, and diclofenac (100 µM)—modulate pathogenicity-related processes in A. castellanii and explored the involvement of a gp63-like protein during encystment and adhesion. Trophozoites were continuously exposed to each drug and analyzed for adhesion, migration on host-derived discontinuous brain micropatterns, encystment efficiency, and parasite-induced cytoskeletal remodeling in MDCK epithelial cells. In silico docking was performed to assess potential drug–protein interactions. Drug exposure reduced adhesion with maximal inhibition at 60 min. After 1 h, migration decreased by 49%, 64%, and 38%, and encystment was reduced by 50%, 85%, and up to 90%, respectively, in cultures treated with acetylsalicylic acid, ibuprofen, and diclofenac. Co-incubation with untreated trophozoites lowered actin fluorescence to approximately 50%, whereas drug-treated co-cultures preserved fluorescence near control levels. Colocalization analysis showed increased spatial overlap between gp63-like protein and F-actin in cysts (~40%) and migrating trophozoites (~20%) compared with non-stimulated forms (~3.8%). Collectively, these findings suggest that NSAID-sensitive pathways influence host interaction, migration, and encystment in A. castellanii and allow for the proposal of gp63-like protein as a putative molecular component of the NSAIDs sensitive pathways.
Full article
(This article belongs to the Special Issue XXth International Meeting on the Biology and Pathogenicity of Free-Living Amoebae (FLAM 2025))
►▼
Show Figures

Graphical abstract
Open AccessArticle
In Vivo Confocal Microscopy as a Prognostic Indicator in Acanthamoeba Keratitis: Insights from a Retrospective Study
by
Yiping Han, Yuan Wei, Qiankun Chen, Jinding Pang, Qingquan Shi and Qingfeng Liang
Pathogens 2026, 15(3), 262; https://doi.org/10.3390/pathogens15030262 - 2 Mar 2026
Abstract
Background: To assess longitudinal changes in the in vivo confocal microscopy (IVCM) features during Acanthamoeba keratitis (AK) treatment and develop a prognostic model. Methods: This retrospective study included 59 AK patients who underwent IVCM at baseline and 1 and 3 months. Fourteen morphological
[...] Read more.
Background: To assess longitudinal changes in the in vivo confocal microscopy (IVCM) features during Acanthamoeba keratitis (AK) treatment and develop a prognostic model. Methods: This retrospective study included 59 AK patients who underwent IVCM at baseline and 1 and 3 months. Fourteen morphological features covering pathogen-related characteristics, cyst arrangement patterns, and inflammatory markers were compared between good and poor prognosis groups, which were defined based on clinical outcomes including corneal perforation, the need for therapeutic keratoplasty, or final best-corrected visual acuity (BCVA) ≤ 0.05. Prognostic modeling was performed exclusively using baseline IVCM features and applied univariable and Firth-corrected multivariable logistic regression with collinearity assessment and clinical filtering, followed by 5-fold cross-validation. Results: Among 59 AK patients, 45 (76.3%) had a good prognosis and 14 (23.7%) had a poor prognosis. Poor prognosis eyes showed a higher prevalence of double-walled cysts, trophozoites, and clustered cysts, along with higher cyst density and deeper stromal invasion. In contrast, good-prognosis eyes had more target-like cysts, immature dendritic cells, and mature dendritic cells. Clustered cysts independently predicted poor prognosis (OR = 2.98), whereas target-like cysts (OR = 0.26) and mature dendritic cells (OR = 0.37) were protective (AUC = 0.883; all p < 0.05). Conclusions: IVCM provides a quantitative tool for early outcome prediction and individualized management. Higher cyst burden, clustered cysts, and persistent stromal involvement indicated poorer prognosis, whereas target-like cysts and mature dendritic cells indicated better prognosis.
Full article
(This article belongs to the Section Parasitic Pathogens)
►▼
Show Figures

Figure 1
Open AccessArticle
Antimicrobial Resistance in Acinetobacter baumannii Isolated from Ready-to-Eat Foods in Saudi Arabia
by
Eman Marzouk and Adil Abalkhail
Pathogens 2026, 15(3), 261; https://doi.org/10.3390/pathogens15030261 - 1 Mar 2026
Abstract
Acinetobacter baumannii is widely recognized as a problematic pathogen in healthcare settings due to its ability to acquire resistance to multiple antimicrobial agents. However, less attention has been given to its presence outside hospitals. In this cross-sectional, laboratory-based surveillance study, we investigated the
[...] Read more.
Acinetobacter baumannii is widely recognized as a problematic pathogen in healthcare settings due to its ability to acquire resistance to multiple antimicrobial agents. However, less attention has been given to its presence outside hospitals. In this cross-sectional, laboratory-based surveillance study, we investigated the occurrence of A. baumannii in ready-to-eat (RTE) foods sold at retail outlets in four cities of the Al-Qassim region, Saudi Arabia, during a single season. A total of 240 RTE food samples were analyzed using culture-based and molecular approaches for species confirmation, and antimicrobial susceptibility profiles were determined. A. baumannii was identified in 19 samples (7.9%), spanning several food categories. Most isolates showed resistance to multiple antimicrobial classes, and 16 met the criteria for multidrug resistance (MDR). Among the confirmed isolates, blaOXA-23-like was detected in 16 (84.2%), blaOXA-24/40-like in 2 (10.5%), and blaOXA-58-like in 1 (5.3%). Resistance to fluoroquinolones, tetracyclines, and aminoglycosides was common, and OXA-type carbapenemase genes were detected in 16 isolates. These findings indicate that RTE foods can represent non-clinical environments in which MDR A. baumannii may be detected. Including food sources in antimicrobial resistance surveillance may therefore strengthen our understanding of the ecology of this pathogen within a One Health framework.
Full article
(This article belongs to the Special Issue Acinetobacter baumannii: An Emerging Pathogen)
►▼
Show Figures

Figure 1
Open AccessArticle
New Insights into the Combined Antiviral Effect of Extracts from Nerium oleander and Boswellia sacra Against Respiratory Syncytial Virus: A Preliminary Report
by
Rebecca Piras, Luca Virdis, Valeria Manca, Marta Cogoni, Vanessa Palmas, Matthew G. Donadu, Aldo Manzin, Giuseppina Sanna and Luay Rashan
Pathogens 2026, 15(3), 260; https://doi.org/10.3390/pathogens15030260 - 1 Mar 2026
Abstract
In recent years, the emergence of drug-resistant pathogens and the limitations of current therapies have highlighted the need for innovative strategies to combat emerging viral infections. Natural compounds, derived from plants, are playing an increasingly significant role in the research of novel and
[...] Read more.
In recent years, the emergence of drug-resistant pathogens and the limitations of current therapies have highlighted the need for innovative strategies to combat emerging viral infections. Natural compounds, derived from plants, are playing an increasingly significant role in the research of novel and effective therapies. Boswellia sacra, a frankincense-producing tree widely distributed in Yemen and Oman, and Nerium oleander, a common ornamental and medicinal plant, are examples of plants with well-documented antimicrobial properties. Their extracts have demonstrated good activity against a wide range of infections, which is attributed to the anti-inflammatory and immunomodulatory compounds they contain. Based on these findings, we assessed, in vitro, the broad-spectrum antiviral activity of combined extracts obtained from Boswellia sacra and Nerium oleander. The extract mixture NOBS7(1) was found to be active against the respiratory virus RSV, Herpesvirus simplex type 1, and Coxsackievirus 5. Furthermore, a combination of cell-based assays was performed to provide additional insights into their potential mechanism of action.
Full article
(This article belongs to the Special Issue Antiviral Strategies Against Human Respiratory Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
Assessment of Systemic Inflammation as a Tool for Estimating the Risk of Death by Visceral Leishmaniasis
by
Ingridi de Souza Sene, Vladimir Costa Silva, Débora Cavalcante Brás, Dorcas Lamounier Costa, Gabriel Reis Ferreira and Carlos Henrique Nery Costa
Pathogens 2026, 15(3), 259; https://doi.org/10.3390/pathogens15030259 - 28 Feb 2026
Abstract
Background: Visceral leishmaniasis (VL) is a life-threatening protozoan disease prevalent in tropical and subtropical regions and a frequent coinfection among people living with HIV. Early identification of patients at high risk of death may reduce case-fatality. This study evaluated the post-test prognostic value
[...] Read more.
Background: Visceral leishmaniasis (VL) is a life-threatening protozoan disease prevalent in tropical and subtropical regions and a frequent coinfection among people living with HIV. Early identification of patients at high risk of death may reduce case-fatality. This study evaluated the post-test prognostic value of C-reactive protein (CRP) and interleukin-6 (IL-6) as biomarkers of mortality in VL. Methods: A retrospective hospital-based cohort of 101 VL patients was analyzed. CRP and IL-6 concentrations at admission were correlated with clinical findings, the Kala-Cal® prognostic score, and in-hospital mortality. Results: Eight patients died, most presenting with hemorrhagic manifestations. At admission, 87.1% of patients had both biomarkers above the predefined cut-offs. CRP and IL-6 levels were markedly elevated in patients with hemorrhage or fatal outcomes. The AUC was 0.85 for CRP and 0.87 for IL-6, with no significant difference between markers. Optimal prognostic cut-offs were 150 mg/L for CRP and 90 pg/mL for IL-6. Conclusions: In this sample, CRP and IL-6 showed good prognostic performance in VL. In patients with low initial clinical risk, positive biomarker results substantially increased the probability of death. When combined with Kala-Cal®, these markers may improve risk stratification and guide referral decisions.
Full article
(This article belongs to the Section Parasitic Pathogens)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Rift Valley Fever Virus Exposure in Camels and Horses Across Northern Nigeria Livestock Markets
by
David Odion Ehizibolo, Olumuyiwa Oyekan, Nicodemus Mkpuma, Habibu Haliru, Ibrahim Garba, Isa Zayyad Turaki, Elizabeth Ene Williams, Agom Danmarwa, Abdullahi Mohammed, Musa Abdullahi Muhammad, Mansur Abubakar, Corrie Brown and Bonto Faburay
Pathogens 2026, 15(3), 258; https://doi.org/10.3390/pathogens15030258 - 28 Feb 2026
Abstract
Rift Valley Fever (RVF) is a neglected vector-borne zoonotic disease of significant veterinary and public health concern in Sub-Saharan Africa. This study investigated the seroprevalence of Rift Valley Fever Virus (RVFV) exposure and associated risk factors among camels and horses marketed in northern
[...] Read more.
Rift Valley Fever (RVF) is a neglected vector-borne zoonotic disease of significant veterinary and public health concern in Sub-Saharan Africa. This study investigated the seroprevalence of Rift Valley Fever Virus (RVFV) exposure and associated risk factors among camels and horses marketed in northern Nigeria. A total of 1117 animals were sampled, comprising camels (812) and horses (305), across three major livestock markets (Maigatari, Maiduguri, and Illela). The overall seroprevalence was 18.8% (95% CI: 16.6–21.2%), with a striking six-fold disparity: camels showed a prevalence of 24.4% (95% CI: 21.6–27.4%), while horses exhibited only 3.9% (95% CI: 2.1–7.0%). Significant geographic clustering was observed, with Illela camels recording the highest prevalence (34.8%) compared to those in Maigatari (20.3%) and Maiduguri (20.2%). There were no significant associations with age or sex among camels. However, in horses, females were significantly more likely to test positive than males (OR = 0.27, 95% CI: 0.07–0.97). These findings demonstrate endemic RVFV circulation in Nigerian livestock, highlighting species- and location-specific differences, and underscore the zoonotic risks within regional and transboundary livestock trade networks.
Full article
(This article belongs to the Section Emerging Pathogens)
Open AccessArticle
Dynamics of Interleukin-9 Producing Lymphocytes in Strongyloides ratti-Infected Mice
by
Wiebke Hartmann, Lennart Heepmann, Lara Linnemann, Paula Licona-Limon, Florent Colomb, Tania Frangova, Henry J. McSorley and Minka Breloer
Pathogens 2026, 15(3), 257; https://doi.org/10.3390/pathogens15030257 - 28 Feb 2026
Abstract
Helminths infect a quarter of the human population and are controlled in the frame of a canonical type-2 immune response. Interleukin-9 is a cytokine with pleiotropic functions during type-2 immunity that can be produced by many different cells. Accumulating evidence suggest that IL-9
[...] Read more.
Helminths infect a quarter of the human population and are controlled in the frame of a canonical type-2 immune response. Interleukin-9 is a cytokine with pleiotropic functions during type-2 immunity that can be produced by many different cells. Accumulating evidence suggest that IL-9 is of particular relevance in controlling intestinal helminth infections. Using mice infected with the parasitic nematode Strongyloides ratti, we showed previously that ejection from the intestine depends on IL-9 and IL-9-mediated activation of mucosal mast cells. Here we use IL-9 reporter mice to identify the relevant cellular sources of IL-9 in vivo. We report that predominantly CD4+ T cells and group 2 innate lymphoid cells (ILC2s) produced IL-9 in S. ratti-infected or IL-33-treated mice. Interestingly, the IL-33-mediated induction of IL-9 and subsequent mast cell degranulation was modulated by concurrent S. ratti infection. While the IL-33-mediated expansion of IL-9-producing ILC2s was supressed by S. ratti infection, IL-9-producing CD4+ T cells were proportionally increased. Finally, we show that S. ratti-derived E/S products interfered with IL-9 production by BM-derived ILC2 in vitro. In conclusion, we have identified that ILC2 and CD4+ T cells produce IL-9 during S. ratti infection, and that ILC2 responses are suppressed by S. ratti products.
Full article
(This article belongs to the Special Issue Immunology Insights: How the Immune System Battles Parasitic Infections)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Pathogens Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Animals, Arthropoda, Insects, Vaccines, Veterinary Sciences, Pathogens
Ticks and Tick-Borne Pathogens: 2nd Edition
Topic Editors: Alina Rodriguez-Mallon, Alejandro Cabezas-CruzDeadline: 31 March 2026
Topic in
Biomolecules, Cells, IJMS, Pathogens, Vaccines, Biologics, Microorganisms, Biomedicines
Advances in Vaccines and Antimicrobial Therapy—2nd Edition
Topic Editors: Raffaele D’Amelio, Roberto PaganelliDeadline: 30 September 2026
Topic in
JoF, Microbiology Research, Microorganisms, Pathogens
Pathophysiology and Clinical Management of Fungal Infections
Topic Editors: Allan J. Guimarães, Marcos de Abreu AlmeidaDeadline: 30 November 2026
Topic in
Infectious Disease Reports, Insects, IJERPH, Pathogens, TropicalMed, Zoonotic Diseases
Vector-Borne Disease Spatial Epidemiology, Disease Ecology, and Zoonoses
Topic Editors: Chad L. Cross, Louisa Alexandra MessengerDeadline: 31 December 2026

Conferences
Special Issues
Special Issue in
Pathogens
The Role of Environment in Vector-Borne Disease Transmission
Guest Editors: Sarah Gunter, Morgan JibowuDeadline: 10 March 2026
Special Issue in
Pathogens
Toxoplasmosis in Immunocompromised and Immunocompetent Host: Recent Updates About the Epidemiology, Diagnosis, Clinical Manifestations and Management
Guest Editor: Despina Contopoulos-IoannidisDeadline: 10 March 2026
Special Issue in
Pathogens
Zoonotic Vector-Borne Infectious Diseases: The One Health Perspective
Guest Editor: Edward B. BreitschwerdtDeadline: 10 March 2026
Special Issue in
Pathogens
Pathogen Dynamics and Airway Microbiome in Pulmonary Diseases: From Conservative Management to Interventional Therapies
Guest Editors: Jing Gao, Chutian ZhangDeadline: 15 March 2026
Topical Collections
Topical Collection in
Pathogens
Advances in Tick Research
Collection Editors: Alejandro Cabezas-Cruz, Ladislav Šimo, James J. Valdés, Dasiel Obregón
Topical Collection in
Pathogens
Bovine Leukemia Virus Infection
Collection Editors: Tasia M. (Taxis) Kendrick, Paul C. Bartlett
Topical Collection in
Pathogens
New Insights into Bacterial Pathogenesis
Collection Editor: Carmelo Biondo
Topical Collection in
Pathogens
Novel Strategies on Antiviral Drug Discovery Against Human Diseases
Collection Editors: Jun Wang, Richard Y. Zhao, Lin Li











