1. Introduction
There are different formulations of the maximum entropy production principle (MEP) [
1,
2,
3,
4,
5,
6,
7,
8]. Applications to biochemical systems leading to some optimal parameters are still rare [
9,
10,
11,
12,
13]. The apparent contradiction between minimum entropy production theorem (MinEP) [
14] and MEP has been examined on a number of occasions [
15,
16]. MinEP is a very special case of the static head steady state near thermodynamic equilibrium when the linear force–flux relationship is a good approximation, net output current vanishes, and the efficiency of free-energy transduction also vanishes [
17]. Free energy is stored but it cannot be used to fuel the synthesis of biological macromolecules, work of biological molecular motors, movement, and growth. The MEP principle has much more general validity both near equilibrium and far from equilibrium. The relationships among proposed MEP principles and their connections to other variational principles from irreversible thermodynamics has been frequently discussed during the last 60 years [
18,
19,
20,
21].
The application of entropy production calculations to bioenergetics and to enzyme kinetics has a special attraction for researchers interested in the question of how physics can provide an insight into life’s ability to maintain its far-from equilibrium structure through interactions with the rest of the universe. Attempts to use MinEP or to question the second law of thermodynamics in the case of photosynthesis resulted in additional controversy without contributing any additional insight to answer this question [
22,
23]. On the other hand, living systems are notoriously complex, and at the same time fragile and robust with the paramount importance of their biological function and evolutionary changes, which subordinates structure to function. It is probably an understatement if we say that this feature of life uniqueness is underexplored by physicists. At the same time, the application of MEP in biology opens many pertinent questions. For instance, Tyler Volk and Olivier Pauluis asked if MEP can predict dissipation details within the system and which fluxes of the system are those that are most likely to be maximized [
24].
In this review, we analyzed the distribution of entropy production contributions among all catalytic steps of enzyme catalytic cycles for very different enzymes, from the triosephosphate isomerase (TIM enzyme), β-lactamases, ATP synthase, to the photochemical cycle of bacteriorhodopsin. We used our extension of Terrel Hill formalism [
25,
26] to the light-absorbing systems [
9] for the calculations, in the case of bacteriorhodopsin. Catalytic cycles of all enzymes were analyzed with and without the application of the maximum trasitional entropy production (MTEP) theorem, which is our second extension of Hill’s formalism [
13]. The MTEP theorem does not apply to the total entropy production but gives the optimal value of rate constants for the corresponding maximal value of entropy production in considered transition. The steady-state is assumed in all cases and appropriately modified Kirchhoff’s laws for biochemical circuits simplified calculations.
We asked the question of how MTEP theorem can be applied to very different examples of enzyme kinetics. Firstly, through examples of cytoplasmic enzymes, we shall illustrate how thermodynamic and kinetic criteria can be unified to obtain a good agreement between theoretical predictions and experimental measurements of kinetic rate constants. We shall also show how MTEP requirements for the last step in the catalytic cycle (the recovery step) can be used to accelerate this rate-limiting step with the concomitant increases in catalytic efficiency, free-energy transduction efficiency, output flux, and overall entropy production. Accordingly, for far-from-equilibrium systems, the relationship between entropy production and efficiency can be such that the increase of former drives the increase of the latter quantity as well. Although the kinetic and thermodynamic relationships we use are quite general, the dissection of all contributions to steady-state entropy production does point toward certain directed nano-currents as being the most important for each particular example of enzymatic free-energy conversion. The optimization of those rate-limiting steps through the MTEP requirement leads to an increase of catalytic efficiency for several enzymes that biochemists described as “perfect enzymes”. When the evolutionary distance is calculated from a putative common ancestor, more evolved enzymes show higher catalytic constants, higher catalytic efficiency, and higher overall entropy production. Bioinformatic analysis is essential for finding this connection between biological and thermodynamic evolution, which holds independently of MTEP optimization or any other entropy production principle or theorem. Secondly, for bacteriorhodopsin embedded in the topologically closed membrane, we examined how the input free energy of absorbed photons can be converted with an optimal efficiency into the electrochemical proton gradient. Power transfer efficiency can reach 90% for large enough secondary force when observed or inferred rate constants are used as input parameters. The MTEP application to the recovery step leads to an additional increase in the output flux and overall entropy production, but such an optimization procedure works better when chosen steady state is compatible with natural bacterial membrane. Similarly, the recovery step of the ATPase catalytic cycle, when ATP has been released from that membrane-bound rotary nanomotor, is also responsible for the largest contribution to total entropy production. All relevant parameters (output ATP flux, the efficiency of free-energy transduction, entropy production due to ATP synthesis) decrease for less than the optimal angular position of the transition state catalytic dwell, while an optimal angular position, as calculated from the MTEP requirement, agrees with the estimate from experiments. Taken together, these in silico results belong to quantitative conformations for the evolution-coupling hypothesis, which postulates that thermodynamic evolution accelerates biological evolution and vice versa [
27].
We have listed below in
Table 1 examples of MTEP optimizations considered in this study. In all of the presented examples, we took into account that enzymes cannot break time-reversal symmetry. For each transition between enzyme functional states in the forward direction, we assumed that the enzyme is capable to catalyze the back reaction as well. This dynamic equilibrium does not imply the absence of dissipation for the considered transition.
2. MTEP Theorem for Transitions between Enzyme Functional States
In this section, we shall repeat the derivation of the MTEP theorem for transitions between enzyme functional states as obtained by ŽBL and DJ [
28]. We consider the enzyme in a steady state that can exist in any finite number of discrete functional states and can cycle among these states by going through several transitions. Generally, the affinity and the flux for a given transition can be expressed in terms of known or assumed forward and backward rate constants
and
as
where
is the Boltzmann constant, T is temperature, n is number of moles, and
is affinity felt between the species
i and
j of a single macromolecule, while
is the net flux for the
i-
j transition. The number of moles n will be taken to be one in the following text. Stationary probabilities
and
of the
i-th and
j-th state, respectively, are the fraction of macromolecular units in states
i and
j. Within the Hill’s diagram method [
25,
26] we associate each transition between two states with entropy production, which can be calculated starting from the flux and the corresponding affinity as [
14]:
Expression (3) allows either a linear or non-linear relation between affinity and flux [
21,
25]. The linear relation between affinity and flux is valid for systems very close to equilibrium. However, for far from equilibrium biochemical processes, the non-linear relations between the affinities and fluxes have to be used.
The stationary probability
of the
i-th state that can be expressed through the directional diagrams
, including the rate constant
, and diagrams
, excluding the rate constant
. Accordingly, one may write
and obtain the entropy production in a given transition in terms of the forward kinetic constant in that transition as
Here,
is the sum of the directional diagrams of all states where
and
are the sums of all state directional diagrams with or without line describing the
ij transition, respectively. Note that each entropy production density
for considered
i-
j transition (transitional entropy production) is a function of all the rate constants in an arbitrary kinetic process. In entropy production (6) and sum
, we expressed the backward constant as
/
, where
Kij is the equilibrium constant for the
ij transition. For a given cycle between functional states, the product of all equilibrium constants defines the overall equilibrium constant
K, and thus the overall thermodynamic force (or affinity)
. Examples of diagrams are shown in
Figure 1, which will be discussed in the next section.
Results from previous research [
28] show that there is a unique maximum for the entropy production of any given transition with respect to variation in the corresponding forward rate constant when we restrict our modeling to fixed values of equilibrium constants for all transitions and assume known values of kinetic constants in all other transitions. This is because the associated transition flux and affinity are, respectively, monotonically increasing and decreasing functions of the forward rate constant
. In other words, there is a simple trade-off between thermodynamic flux and force. Since the equilibrium constant for the
ij transition is constrained to the fixed value, the backward rate constant is calculated from the forward rate constant through expression
/
after the optimization procedure. With constant
Kij, only coordinated change is possible in corresponding forward and backward kinetic constants.
The condition for the extreme value of entropy production (6) in considered
ij transition
gives the equation for the optimal value of the forward kinetic constant
The extreme value of the entropy production in the
ij transition for the optimal value of the forward rate constant denoted
, is then
This is the maximal value of the entropy production (6) as its second derivative is negative for the optimal value of the forward kinetic constant. The above result for entropy production maximum in a given transition is valid for kinetic schemes containing multiple cycle diagrams also when we impose the same condition of fixed equilibrium constants Kij and known values of kinetic constants in all other transitions.
Entropy production for a chosen transition is just the component of total entropy production, which is associated with a conformational transition between two functionally important enzyme states. In addition to the directional movement of enzyme atoms and amino acid residues, a conformational transition can include substrates, products, and other small molecules, atoms, or elementary particles entering or exiting the reaction pathway. In our formalism, this is taken into account in an implicit manner through second-order rate constants. We must note that maximal transitional entropy production σij for the transition between enzyme functional states i and j does not ensure that maximal total entropy production has been reached for a chosen steady-state system at a constant temperature.
4. MTEP Theorem Optimization of Transition State Parameters for ATPase
The integral membrane protein F
0F
1-ATP synthase (ATPase), found universally in chloroplasts, bacteria, and mitochondria, couples transmembrane proton translocation to ATP synthesis/hydrolysis. ATPase is composed of two rotary motors: electrical F
0 and chemical F
1 [
41]. They are coupled by elastic power transmission resulting in high efficiency of ATP synthesis. The kinetic reaction scheme for the F
0 motor takes into account the torsional angle and elastic energy [
42]. This kinetic model of ATPase [
10], shown in
Figure 5, consists of five functional states: empty (O:) or binding either ATP (O:ATP), ADP (O:ADP), P
i (O:P), or P
i and ADP together (O:P.ADP). The net flux
(given by expression (2)) of the transition from O:P.ADP to O:ATP equals the net rate of ATP synthesis (number of ATP molecules produced per enzyme per second), with associated forward and backward kinetic constants k
syn and k
hyd, respectively. It is the M transition from
Figure 5 for which the application of the MTEP theorem is particularly interesting, since it leads to free energy conversion from proton gradient into ATP synthesis. The stationary probability of the
i-th state) (i = O:, O:ATP, O:ADP, O:P, O:P.ADP),
can be obtained using Hill’s diagram method [
25]. From MTEP theorem application to transition M, obtained optimal
values for all transitions are listed in
Table 2 together with values of entropy production and flux in each transition. States O:ATP and O:ADP have the largest stationary probabilities. As is seen from
Table 2, the greatest contribution to the total entropy production comes from the recovery step, the T transition (from O:ATP to O:), when ATP is released. This transition, together with the M transition, has the highest flux equal to the net ATP synthesis flux in a steady state. We note that the optimal kinetic constant k
syn found from the MTEP application coincides with the corresponding one found by information entropy maximization. Furthermore, MTEP and maximum entropy (MaxEnt) modeling predict high optimal efficiency for the percentage of free-energy storage: E
out/E
in = 69%. MaxEnt and MTEP optimizations for transition state parameters are in agreement with an empirical estimate about optimal angular position of about 72
o for the ATP-binding transition [
10,
42]. Therefore, joint MaxEnt and MTEP optimization results are consistent with the observed design of spinach chloroplast ATP-ase and predict an optimal working regime for this nanomotor near the inflection point of a force–flux relationship when current J is maximally sensitive to changes in the protonmotive force Δ
µH+.
5. Light-Activated Creation of the Protonmotive Force, Dissipation, and Free Energy Transduction Efficiency. The Example of Bacteriorhodopsin
For us, absorption of a photon can trigger an eye–brain communication we call vision. For certain bacteria, photon triggers a protein quake, charge separation, electric field builds up, and photosynthesis. Although separated by a billion years of evolutionary gap, the same protein type and the same chromophore are responsible for both outcomes. It is an integral membrane protein with seven membrane-spanning helices, one of them covalently connected to the retinal chromophore. When acting as a photon detector in human rod cells, the protein is named rhodopsin. When performing the first step of photosynthesis for
Halobacterium salinarium, the protein is named bacteriorhodopsin. Bacteriorhodopsin (bR) is the simplest solution nature found for a light-activated proton pump, which can easily perform proton active transport against the electrochemical proton gradient [
43]. In its natural membrane environment, bacteriorhodopsin dissipates more than 85% of photon free energy. One can ask, why such a low efficiency of light power conversion to proton-motive power? We have seen that converting proton-motive power into ATP synthesis by an ATP-synthase is a more efficient process. Higher efficiency can be easily achieved if the stronger electric field is created, that is, greater than a minimal field of about 130 mV needed to put into rotation the ATP-synthase rotary motor for producing ATP. Photon free energy for photons with a wavelength of about 570 nm, which bacteriorhodopsin prefers to absorb, is quite high and more than enough to create a much stronger electric field. The problem with too strong an electric field is that it will cause a dielectric breakdown of a plasma membrane and cellular death.
H. salinarium can develop a maximal electric field of about 280 mV [
44]. Still, assuming that bacteriorhodopsins can be incorporated in much more robust artificial membranes, we can examine in simulations the cases when weak, strong, and super-strong secondary force is developed corresponding to the membrane potential of −195, −278, and −1185 mV, respectively. The first value of −195 mV is quite common for the membrane potential of bacteria, archaea, and mitochondria, and it is identical to one we used in earlier simulations [
9]. The second value of −278 mV was also used by us earlier [
45] as similar to measured maximal value for membrane potential established by
H. salinarium, while the third and highest value of membrane potential, equal to −1.185 V, corresponds to maximal efficiency of free energy conversion, which is slightly higher than 70%. Corresponding
Table 3 values are expressed in kJ/mol as respectively −18.84, −26.86, and −123 kJ/mol. We used estimated kinetic and thermodynamic parameters data for bR [
46] and performed the simulations designed to answer several questions:
Which transition step, out of seven Ti steps (
Figure 6a), is associated with the greatest entropy production?
What is the rate limiting step among all Ti transitions involved in a complex interplay of retinal, protein atoms, and water molecule movements, resulting in a proton pumping and charge separation [
47]?
When MTEP theorem is used to optimize each transition, is there a single catalytic step for which photochemical quantum yield, the efficiency of free energy conversion, and total entropy production all exhibit increased optimal values with respect to values obtained without optimization?
In references, [
46] and [
48] experiments and modeling were used to construct the kinetic model illustrated in
Figure 6. In order to apply irreversible thermodynamics to the initial photon absorption step, we introduced the excited state bR
*, light activated transition L from ground to excited state, and non-radiative transition D back to ground state (
Figure 6b). For details, see our earlier papers [
9,
45] about this extension of Hill’s formalism [
25] to the light-absorbing systems. Proton transfer and charge separation take place in the productive T1 to T7 pathway. In a reversible model of van Stokkum and Lozier [
46], all thermodynamic and kinetic parameters (equilibrium constant, forward, and reverse kinetic constants) have been estimated for T2 to T6 transitions. They estimated the forward constant k
7 in the last recovery transition (T7) as k
7 = 700 s
−1. To get
Table 3 values, we used that and other estimated parameters by van Stokkum and Lozier for pH = 7, while τ = 4 ps time-constant estimate by Nango et al. [
48] was used to calculate the forward constant k
1 as k
1 = 2.5 × 10
11 s
−1. With a choice of K
7 = 2 × 10
7 for recovery transition equilibrium constant, the remaining constants for the T1 transition could be easily calculated from the requirement that the product of all equilibrium constants in the charge separation cycle must be equal to exp(X
sec/k
BT) [
25]. We have chosen the kinetic constant k
d for non-radiative D transition as k
d = 10
8 s
−1 and the light-absorption rate α
01 = 100 s
−1 by following our 2003 choice [
9] for modeling bacteriorhodopsin photocycle with system being at room temperature T = 298.16 K. Equilibrium constants in the light cycle L–D are found as K
L = exp(hv/k
BT
R) and K
D = exp(hν/k
BT), where T
R is an effective temperature, which is higher than T and increases with increased light absorption rate α
01 and increased light intensity I = J
L [
9].
According to van Stokkum and Lozier [
46], when T1 transition is not considered, most free energy is dissipated in the recovery step. This is confirmed by our calculations of entropy production for each step in the charge separation pathway when picoseconds relaxation from excited state bR
* to K
590 spectroscopic state and associated entropy production σ
1 is not considered.
After examining all results, we concluded that only the MTEP application to recovery step (transition T7) for the kinetic model of the bacteriorhodopsin photocycle leads to an additional increase in the output flux and overall entropy production, with respect to values obtained without optimization (
Table 3). We note that state probability p
2 of the excited state bR
* is very small because the transition from state 2 to state 3 is very fast with large forward kinetic constant k
1 and correspondingly large equilibrium constant, while thermal relaxation to the ground state bR is also fast for the D transition. In order to choose representative secondary forces for the construction of
Table 3 and
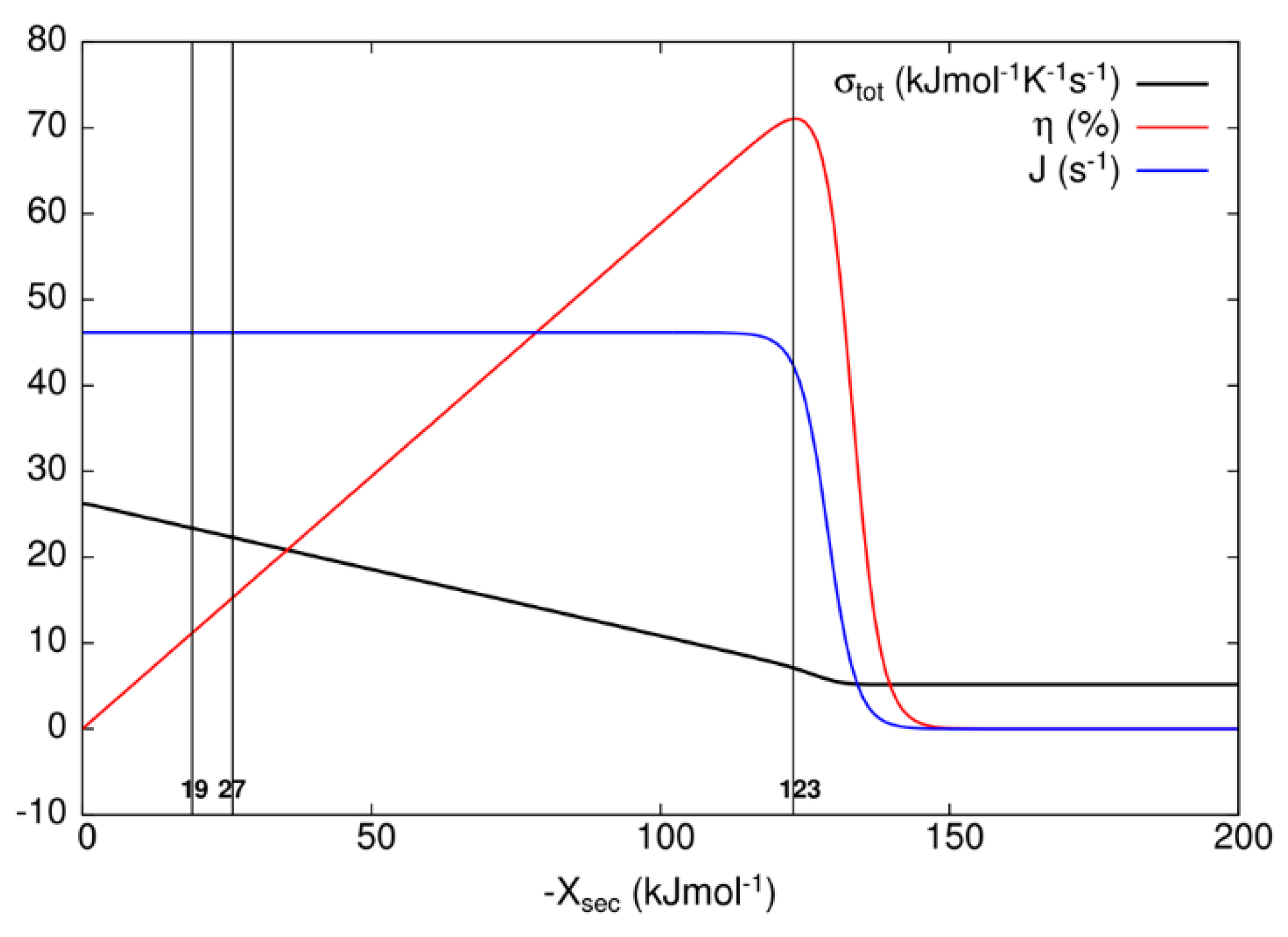
Figure 7 for the kinetic model of the bacteriorhodopsin photocycle, we have investigated how flux J, entropy production σ
tot, and efficiency η vary as the secondary force X
sec varies. As it is seen from
Figure 7, flux stays almost constant and entropy production decreases as the efficiency increases from zero to its maximal value when one varies secondary force from zero to −123 kJ/mol. As one further varies secondary force, flux and efficiency fall to zero and entropy production falls to a finite, almost constant, low value. When oriented bacteriorhodopsins are incorporated in robust bioelectronic devices, a high efficiency can be reached [
49], with an additional bonus of smaller entropy production, that is, considerably less heating with a very small decrease in produced proton current. Another bR-based bioelectronic application is in the field of volumetric optical memory [
50]. It is using different means to channel the same recovery transition T7, which we found to be critical for the optimization, into a branched pathway enabling writing, reading, and erasing information. From our analysis, it follows that the MTEP optimization for the T1 transition can increase the O state occupancy and efficiency of a branched recovery pathway (not shown).
6. Discussion
By using a classic combination of irreversible thermodynamics and enzyme kinetics [
25,
26], we examined the question of how thermodynamic evolution is connected to biological evolution. With our extensions of Hill’s formalism (MTEP theorem, light-absorbing systems), it is possible to identify those rate-limiting transitions that are leading to an increase in total entropy production after MTEP optimization, to check if optimized rate constants are comparable to measured rate constants, and also to see if enzyme catalytic efficiency can be increased. It is also possible to examine the structure-dynamic and structure-function connection from a fresh outlook. All examined examples have in common that enzyme structure enables the transfer of protons among several critical amino acid residues and water molecules, resulting in a directed nano-current of protons. Proton flux would not exist without imposed external force that is keeping the whole system safely away from thermodynamic equilibrium. For simpler cytoplasmic enzymes, a substrate–product pair is maintained in a homeostatic disequilibrium. Nonequilibrium substrate and product concentrations are responsible for chemical affinity as a single driving force producing the flux of product molecules. This is the case with TIM enzyme and β-lactamases. More complex enzymes are usually membrane-embedded proteins capable of converting the primary force–flux couple into a secondary force–flux couple. This is the case for ATP-ase and bacteriorhodopsin. Entropy production calculations can already lead to the identification of rate-limiting steps among all transitions without performing any optimization if a complete set of rate constants is known, but MTEP theorem helps for frequent cases when some rate constant is not known. The last catalytic step, that is, the recovery of the enzyme to its original ground state, is often at the same time the rate-limiting step and the cause for the highest dissipation among all other transitions.
Repeated calculations for evolutionary distances of three lactamases have shown the robustness of previously found proportionality among distances from a common ancestor, catalytic constant, catalytic efficiency, and total entropy production [
13]. This result follows from a straightforward application of Hill’s formalism for entropy production calculations [
25] in combination with a bioinformatic analysis without recourse to any optimization technique. The acylation and deacylation steps with concomitant proton shuttles are the rate limiting for β-lactamases PC1, RTEM, and Lac1 and the most important contributors to overall entropy production. When these two steps are optimized by using the MTEP theorem, the catalytic activity (the turnover number) can be increased from one to two orders of magnitude (
Figure 4). In practice, this can be achieved by finding mutations which can lower the activation energies for proton shuttles, that is, by finding the specific transition state mutants for accelerating the acylation and deacylation catalytic steps. It is not something we want to do of course, because the evolvability of β-lactamases has already led to the worldwide spread of multidrug-resistant bacterial pathogens. This happened despite common opinion among biochemists that wild-type lactamases are nearly perfect enzymes. However, “super-lactamases” predicted by the application of MTEP theorem are uncomfortably close to or even inside the forbidden diffusion-limit region. Still, this simple application of irreversible thermodynamics to enzyme kinetics led us directly to the main driving engine for enzyme evolution: the transition-state mutations for rate-limiting steps. In the examples of β-lactamases and TIM enzyme entropy production, calculations can identify those crucial catalytic steps, which are at the same time the most efficient in performing catalysis and in dissipating free-energy gradients.
For ATP-ase kinetic scheme (
Figure 5), we have achieved the best agreement between MTEP and MaxENT predictions and experimental findings with the caveat (DJ personal communication with Oliver Pänke and Bernd Rumberg) that empirical estimate for the relative angular position of the catalytic dwell κ (corresponding to angular position 72
o) was not very accurate. Also, in the ATP-ase kinetic scheme, the recovery step when ATP molecule is released from the F
1 rotor is accompanied with the highest contribution to overall entropy production (
Table 2). ATP-as nanomotors are biologically very old inventions. Both respiration and initial photosynthetic steps converge toward creating protonmotive force, which can be used by ATP-ase to convert spontaneous inward-directed proton nano currents first into rotation of their stator subunits, then into elastic energy and finally into the pushing together of ADP and inorganic phosphate to create ATP molecule without the hindrance of water molecules. This principle is observed by all ATP-ase molecular motors including the ATP-ase from spinach chloroplasts studied by us [
10] and ATP-ase from
Halobacterium salinarium. The efficiency of converting protonmotive force free energy into free energy of far-from-equilibrium ADP–ATP concentrations is truly amazing. It was estimated to be close to 70% and calculated by us as the optimal value of 69% as the result of MTEP and MaxENT maximization.
When known and estimated values for kinetic constants are used to calculate the contribution to entropy production of all transitions in the 8-state bacteriorhodopsin photocycle (
Table 3), the last recovery step is responsible for the highest contribution, but only when developed electrochemical proton gradient becomes too high for the photosynthetic cell to maintain the integrity of cytoplasmic membrane. It will surely experience a dielectric breakdown at a considerably lower value of the transmembrane electric field. For more realistic values of membrane potential, the contribution of the bR* to K transition T1 (
Figure 6) is more important for total entropy production. In
Halobacterium salinarium, the membrane-embedded proton pumps of bacteriorhodopsin and ATP-ase are coupled together through proton flux in the simplest photosynthetic circuit created by natural evolution. Coming back to the efficiency of power transduction by bacteriorhodopsin, we can see that in the hypothetical case of engineered membrane capable of withholding the membrane potential of −1.185 volts (corresponding to −123 kJ/mol secondary force for the maximal efficiency of 71%, see
Table 3), an overall photosynthetic efficiency would be close to 50%. Naturally evolved photosynthetic organelles, cells, and organisms do not need such a high efficiency for producing ATP molecules, but our civilization has an urgent need to use renewable free energy coming from our sun in the most efficient way possible. If these proton pumps can provide an inspiration for how to achieve such a goal, the bonus will be a decreased level of entropy production, because with the higher secondary force, we are actually approaching something similar to the static head state for nonlinear force–flux relationships, as can be seen from
Figure 7 and earlier calculations [
15].
It is argued in the paper by Jennings et al. [
51] that plant photosystem I, which performs primary charge separation in around 40 ps, does so with minimal entropy production. While the transfer of photon free energy into bond twisting (in the case of bacteriorhodopsin) or electron–hole excitation (for plant photosynthesis) is the obligatory first step (in femtoseconds) of quantum nature, which is associated with very low dissipation, the biologically most relevant step is the light-activated proton transport. Proton pumping creates the protonmotive force. The ratio of output protonmotive power to input power provided by photons is biologically much more relevant quantity than high quantum efficiency, promoted by Jennings et al. [
51] as the evidence for minimal or even negative entropy production [
23]. Indeed, within restrictions of our model for bacteriorhodopsin photocycle, the efficiency of free energy transduction in the biological range of membrane potentials does not exceed 16%, while the photochemical yield or quantum efficiency does not decrease below 99%. Lower than 16% efficiency is easy to realize with an increased dissipation rate (
Figure 7). Similar behavior is observed by Baiesi and Maes [
52], that is, efficiency decreases when external constraints are changed to move the system away from some optimal (low) dissipation level.
In bioenergetics, it is generally accepted that living cells are superior entropy producers with respect to an equivalent volume of some average star, like our sun. The dissipation is tightly coupled to accurate signaling [
53], sensory adaptation [
54], many regulatory feedback cycles in biochemistry, and kinetic proofreading [
55,
56]. One can ask the question: is life a constant struggle against the tendency to produce entropy or not? [
57]. The connection between evolution and increased entropy production is not restricted to the living world. More complex structures emerge with a greater distance from equilibrium for many different open systems. Evolution of galaxies, stars, planets, life, society, and machines is connected to the slow or the fast increase in energy rate density with time [
58]. It is well known that black holes are central players in the evolution of galaxies, and also the most important contributors to entropy increase in the universe [
59]. Dynamic description of complex biochemical and physical systems, including oxidative phosphorylation [
60], metabolic networks [
61], and the earth’s global climate [
62] is in accordance with maximization of entropy production.
The MTEP requirement may be useful as a simple optimization method founded in physics to study the evolutionary optimization of enzymatic reactions. We can assume that the ATP-synthase evolved in accordance with the MTEP theorem and with the statistical principle of maximum Shannon’s entropy [
10]. Shannon’s information entropy of discrete enzyme states increased in accordance with increased evolutionary distance for β-lactamases. However, unlike the maximal entropy production for transitions between functional states, the maximal Shannon’s information entropy cannot always be found if we do not apply additional restrictions. As an example, MTEP optimization in the recovery T7 transition for the kinetic model of the bacteriorhodopsin photocycle did not result in increased Shannon’s information entropy. In
Table 4 we have compared the calculations of Shannon’s entropy and overall entropy production for β-lactamases, TIM enzyme, ATP-ase, and bR, in the case of input parameters (kinetic constants), were estimated from experiments, and when their optimal values were obtained after MTEP optimization for the rate-limiting catalytic transition. The TIM enzyme obviously works not so far from equilibrium, but its Shannon entropy is far from the theoretical maximum for the 4-state kinetic model. The ATP-ase and bacteriorhodopsin are less amenable than β-lactamases to MTEP optimization capable of increasing kinetic constants. This may be due to billions of years which evolution has had to perfect these proton pumps.
Several conclusions follow from found connections between entropy production and enzyme performance parameters honed by the forces of natural selection. Firstly, the dissection of entropy production contributions for evolutionary related enzymes cycling among functional states is not a direct application of the MEP principle. The MEP principle applicability is restricted to small-time intervals and volume elements [
21], while gains in catalytic activity, biological complexity, and corresponding increases in entropy production density over eons [
13,
63] are not restrained within these bounds. The MTEP theorem application can still be helpful for considerations of how natural or human design can increase overall entropy production and at the same time, improve the steady-state operation of bionanomachines. Secondly, we can conclude that the application of the MTEP theorem in enzyme kinetics and bioenergetics produced three main insights: (a) an intimate connection exists among far-from-equilibrium-situation, nonlinear force–flux relationships, concomitant increases in entropy production, and efficiency increase of free-energy transduction, (b) entropy production calculations for each transition between functional states helps to identify the rate-limiting steps among all enzymatic transitions leading to product formation, and (c) evolutionary distance calculations support the evolution-coupling hypothesis [
27] when correlated to concomitant catalytic efficiency and entropy production increase.