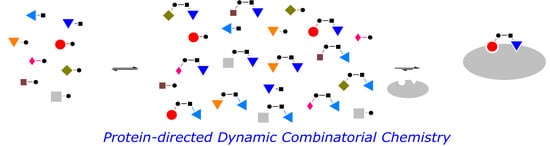
Protein-Directed Dynamic Combinatorial Chemistry: A Guide to Protein Ligand and Inhibitor Discovery
Abstract
:1. Introduction
2. The Protein-Directed DCC Method
2.1. General Concept
2.2. Composition of a Dynamic Combinatorial Library
2.3. Reversible Reactions and Reaction Conditions for Protein-Directed DCC
2.4. Protein Template
2.5. Analytical Techniques
3. Summary and Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| 2OG | 2-Oxoglutarate |
| ATP | Adenosine triphosphate |
| DCC | Dynamic combinatorial chemistry |
| DCL | Dynamic combinatorial library |
| DNA | Deoxyribonucleic acid |
| FITC | Fluorescein isothiocyanate |
| GABA | γ-Animobytyric acid |
| GAT1 | γ-Animobytyric acid transporter 1 |
| HEWL | Hen egg-white lysozyme |
| HIF | Hypoxia inducible factor |
| HPLC | High-performance liquid chromatography |
| HSP | Heat shock protein |
| HTS | High-throughput screening |
| JMJD2 | Jumonji domain 2 |
| LC-MS | Liquid chromatography-mass spectrometry |
| NMR | Nuclear magnetic resonance |
| NOE | Nuclear Overhauser effect |
| PHD2 | Prolyl hydroxylase domain 2 |
| STD | Saturation transfer difference |
| TPR | Tetratricopeptide repeat |
| WaterLOGSY | Water ligand-observed via gradient spectroscopy |
References
- Drews, J. Drug discovery: A historical perspective. Science 2000, 287, 1960–1964. [Google Scholar] [CrossRef] [PubMed]
- Fox, S.; Farr-Jones, S.; Sopchak, L.; Boggs, A.; Wang Nicely, H.; Khoury, R.; Biros, M. High-throughput screening: Update on practices and success. J. Biomol. Screen. 2006, 11, 864–869. [Google Scholar] [CrossRef] [PubMed]
- Dove, A. Screening for content—The evolution of high throughput. Nat. Biotechnol. 2003, 21, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Swinney, D.C.; Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 2011, 10, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Gribbon, P.; Andreas, S. High-throughput drug discovery: What can we expect from HTS? Drug Discov. Today 2005, 10, 17–22. [Google Scholar] [CrossRef]
- Mayr, L.M.; Fuerst, P. The future of high-throughput screening. J. Biomol. Screen. 2008, 13, 443–448. [Google Scholar] [CrossRef] [PubMed]
- Mayr, L.M.; Bojanic, D. Novel trends in high-throughput screening. Curr. Opin. Pharmacol. 2009, 9, 580–588. [Google Scholar] [CrossRef] [PubMed]
- Macarron, R.; Banks, M.N.; Bojanic, D.; Burns, D.J.; Cirovic, D.A.; Garyantes, T.; Green, D.V.S.; Hertzberg, R.P.; Janzen, W.P.; Paslay, J.W.; et al. Impact of high-throughput screening in biomedical research. Nat. Rev. Drug Discov. 2011, 10, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Bleicher, K.H.; Böhm, H.-J.; Müller, K.; Alanine, A.I. A guide to drug discovery: Hit and lead generation: Beyond high-throughput screening. Nat. Rev. Drug Discov. 2003, 2, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Lehn, J.-M. Dynamic combinatorial chemistry and virtual combinatorial libraries. Chem. Eur. J. 1999, 5, 2455–2463. [Google Scholar] [CrossRef]
- Klekota, B.; Miller, B.L. Dynamic diversity and small-molecule evolution: A new paradigm for ligand identification. Trends Biotechnol. 1999, 17, 205–209. [Google Scholar] [CrossRef]
- Karan, C.; Miller, B.L. Dynamic diversity in drug discovery: Putting small-molecule evolution to work. Drug Discov. Today 2000, 5, 67–75. [Google Scholar] [CrossRef]
- Lehn, J.-M.; Eliseev, A.V. Dynamic combinatorial chemistry. Science 2001, 291, 2331–2332. [Google Scholar] [CrossRef] [PubMed]
- Ramström, O.; Lehn, J.-M. Drug discovery by dynamic combinatorial libraries. Nat. Rev. Drug Discov. 2002, 1, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Rowan, S.J.; Cantrill, S.J.; Cousins, G.R.L.; Sanders, J.K.M.; Stoddart, J.F. Dynamic covalent chemistry. Angew. Chem. Int. Ed. 2002, 41, 898–952. [Google Scholar] [CrossRef]
- Hochgürtel, M.; Lehn, J.-M. Dynamic combinatorial diversity in drug discovery. In Fragment-Based Approaches in Drug Discovery; Jahnke, W., Erlanson, D.A., Eds.; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, German, 2006; Chapter 16; pp. 341–364. [Google Scholar]
- Greaney, M.F.; Bhat, V.T. Protein-directed dynamic combinatorial chemistry. In Dynamic Combinatorial Chemistry: In Drug Discovery, Bioinorganic Chemistry, and Materials Sciences; Miller, B.L., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010; Chapter 2; pp. 43–82. [Google Scholar]
- Mondal, M.; Hirsch, A.K.H. Dynamic combinatorial chemistry: A tool to facilitate the identification of inhibitors for protein targets. Chem. Soc. Rev. 2015, 44, 2455–2488. [Google Scholar] [CrossRef] [PubMed]
- Rees, D.C.; Congreve, M.; Murray, C.W.; Carr, R. Fragment-based lead discovery. Nat. Rev. Drug Discov. 2004, 3, 660–672. [Google Scholar] [CrossRef] [PubMed]
- Hajduk, P.J.; Greer, J. A decade of fragment-based drug design: Strategic advances and lessons learned. Nat. Rev. Drug Discov. 2007, 6, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Congreve, M.; Chessari, G.; Tisi, D.; Woodhead, A.J. Recent developments in fragment-based drug discovery. J. Med. Chem. 2008, 51, 3661–3680. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.W.; Rees, D.C. The rise of fragment-based drug discovery. Nat. Chem. 2009, 1, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Scott, D.E.; Coyne, A.G.; Hudson, S.A.; Abell, C. Fragment-based approaches in drug discovery and chemical biology. Biochemistry 2012, 51, 4990–5003. [Google Scholar] [CrossRef] [PubMed]
- Huc, I.; Lehn, J.-M. Virtual combinatorial libraries: Dynamic generation of molecular and supramolecular diversity by self-assembly. Proc. Natl. Acad. Sci. USA 1997, 94, 2106–2110. [Google Scholar] [CrossRef] [PubMed]
- Demetriades, M.; Leung, I.K.H.; Chowdhury, R.; Chan, M.C.; McDonough, M.A.; Yeoh, K.K.; Tian, Y.-M.; Claridge, T.D.W.; Ratcliffe, P.J.; Woon, E.C.Y.; et al. Dynamic combinatorial chemistry employing boronic acids/boronate esters leads to potent oxygenase inhibitors. Angew. Chem. Int. Ed. 2012, 51, 6672–6675. [Google Scholar] [CrossRef] [PubMed]
- Rose, N.R.; Woon, E.C.Y.; Kingham, G.L.; King, O.N.F.; Mecinović, J.; Clifton, I.J.; Ng, S.S.; Talib-Hardy, J.; Oppermann, U.; McDonough, M.A.; et al. Selective inhibitors of the JMJD2 histone demethylases: Combined nondenaturing mass spectrometric screening and crystallographic approaches. J. Med. Chem. 2010, 53, 1810–1818. [Google Scholar] [CrossRef] [PubMed]
- Woon, E.C.Y.; Demetriades, M.; Bagg, E.A.L.; Aik, W.; Krylova, S.M.; Ma, J.H.Y.; Chan, M.; Walport, L.J.; Wegman, D.W.; Dack, K.N.; et al. Dynamic combinatorial mass spectrometry leads to inhibitors of a 2-oxoglutarate-dependent nucleic acid demethylase. J. Med. Chem. 2012, 55, 2173–2184. [Google Scholar] [CrossRef] [PubMed]
- Mondal, M.; Radeva, N.; Köster, H.; Park, A.; Potamitis, C.; Zervou, M.; Klebe, G.; Hirsch, A.K.H. Structure-based design of inhibitors of the aspartic protease endothiapepsin by exploiting dynamic combinatorial chemistry. Angew. Chem. Int. Ed. 2014, 53, 3259–3263. [Google Scholar] [CrossRef] [PubMed]
- Lehn, J.-M.; Ramström, O. Generation and Screening of a Dynamic Combinatorial Library. WO 20,010,164,605, 7 September 2001. [Google Scholar]
- Steeneck, C.; Eliseev, A.V.; Hochguertel, M.; Kroth, H. A Method of Forming a Dynamic Combinatorial Library Using a Scaffold. WO 2,003,034,071, 24 April 2003. [Google Scholar]
- Cancilla, M.T.; He, M.M.; Viswanathan, N.; Simmons, R.L.; Taylor, M.; Fung, A.D.; Cao, K.; Erlanson, D.A. Discovery of an Aurora kinase inhibitor through site-specific dynamic combinatorial chemistry. Bioorg. Med. Chem. Lett. 2008, 18, 3978–3981. [Google Scholar] [CrossRef] [PubMed]
- Erlanson, D.A.; Wells, J.A.; Braisted, A.C. Tethering: Fragment-based drug discovery. Annu. Rev. Biophys. Biomol. Struct. 2004, 33, 199–223. [Google Scholar] [CrossRef] [PubMed]
- Ingerman, L.A.; Cuellar, M.E.; Waters, M.L. A small molecule receptor that selectively recognizes trimethyl lysine in a histone peptide with native protein-like affinity. Chem. Commun. 2010, 46, 1839–1841. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, M.; Shumacher, I.; Cohen-Luria, R.; Ashkenasy, G. Dynamic combinatorial libraries of artificial repeat proteins. Bioorg. Med. Chem. 2013, 21, 3450–3457. [Google Scholar] [CrossRef] [PubMed]
- Bunyapaiboonsri, T.; Ramström, O.; Lohmann, S.; Lehn, J.-M.; Peng, L.; Goeldner, M. Dynamic deconvolution of a pre-equilibrated dynamic combinatorial library of acetylcholinesterase inhibitors. ChemBioChem 2001, 2, 438–444. [Google Scholar] [CrossRef]
- Hochgürtel, M.; Kroth, H.; Piecha, D.; Hofmann, M.W.; Nicolau, C.; Krause, S.; Schaaf, O.; Sonnenmoser, G.; Eliseev, A.V. Target-induced formation of neuraminidase inhibitors from in vitro virtual combinatorial libraries. Proc. Natl. Acad. Sci. USA 2002, 99, 3382–3387. [Google Scholar] [CrossRef] [PubMed]
- Hochgürtel, M.; Niesinger, R.; Kroth, H.; Piecha, D.; Hofmann, M.W.; Krause, S.; Schaaf, O.; Nicolau, C.; Eliseev, A.V. Ketones as building blocks for dynamic combinatorial libraries: Highly active neuraminidase inhibitors generated via selective pressure of the biological target. J. Med. Chem. 2003, 46, 356–358. [Google Scholar] [CrossRef] [PubMed]
- Zameo, S.; Vauzeilles, B.; Beau, J.-M. Dynamic combinatorial chemistry: Lysozyme selects an aromatic motif that mimics a carbohydrate residue. Angew. Chem. Int. Ed. 2005, 44, 965–969. [Google Scholar] [CrossRef] [PubMed]
- Zameo, S.; Vauzeilles, B.; Beau, J.-M. Direct composition analysis of a dynamic library of imines in an aqueous medium. Eur. J. Org. Chem. 2006, 2006, 5441–5444. [Google Scholar] [CrossRef]
- Valade, A.; Urban, D.; Beau, J.-M. Target-assisted selection of galatosyltransferase binders from dynamic combinatorial libraries. An unexpected solution with restricted amounts of the enzyme. ChemBioChem 2006, 7, 1023–1027. [Google Scholar] [CrossRef] [PubMed]
- Bugaut, A.; Toulmé, J.-J.; Rayner, B. SELEX and dynamic combinatorial chemistry interplay for the selection of conjugated RNA aptamers. Org. Biomol. Chem. 2006, 4, 4082–4088. [Google Scholar] [CrossRef] [PubMed]
- Bunyapaiboonsri, T.; Ramström, H.; Ramström, O.; Haiech, J.; Lehn, J.-M. Generation of bis-cationic heterocyclic inhibitors of Bacillus subtilis HPr kinase/phosphatase from a ditopic dynamic combinatorial library. J. Med. Chem. 2003, 46, 5803–5811. [Google Scholar] [CrossRef] [PubMed]
- Valade, A.; Urban, D.; Beau, J.-M. Two galatosyltransferases’ selection of different binders from the same uridine-based dynamic combinatorial library. J. Comb. Chem. 2007, 9, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Congreve, M.S.; Davis, D.J.; Devine, L.; Granata, C.; O’Reilly, M.; Wyatt, P.G.; Jhoti, H. Detection of ligands from a dynamic combinatorial library by X-ray crystallography. Angew. Chem. Int. Ed. 2003, 42, 4479–4482. [Google Scholar] [CrossRef] [PubMed]
- Ramström, O.; Lohmann, S.; Bunyapaiboonsri, T.; Lehn, J.-M. Dynamic combinatorial carbohydrate libraries: Probing the binding site of the concanavalin A lectin. Chem. Eur. J. 2004, 10, 1711–1715. [Google Scholar] [CrossRef] [PubMed]
- Sindelar, M.; Wanner, K.T. Library screening by means of mass spectrometry (MS) binding assays-exemplarily demonstrated for a pseudostatic library addressing γ-aminobutyric acid (GABA) transporter 1 (GAT1). ChemMedChem 2012, 7, 1678–1690. [Google Scholar] [CrossRef] [PubMed]
- Sindelar, M.; Lutz, T.A.; Petrera, M.; Wanner, K.T. Focused pseudostatic hydrazone libraries screened by mass spectrometry binding assay: Optimizing affinities toward γ-aminobutyric acid transporter 1. J. Med. Chem. 2013, 56, 1323–1340. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, S.-A. Direct screening of a dynamic combinatorial library using mass spectrometry. J. Am. Soc. Mass Spectrom. 2006, 17, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, A. Dynamic mixtures and combinatorial libraries: Imines as probes for molecular evolution at the interface between chemistry and biology. Org. Biomol. Chem. 2009, 7, 3195–3204. [Google Scholar] [CrossRef] [PubMed]
- Ramström, O.; Lehn, J.-M. In situ generation and screening of a dynamic combinatorial carbohydrate library against concanavalin A. ChemBioChem 2000, 1, 41–48. [Google Scholar] [CrossRef]
- Milanesi, L.; Hunter, C.A.; Sedelnikova, S.E.; Waltho, J.P. Amplification of bifunctional ligands for calmodulin from a dynamic combinatorial library. Chem. Eur. J. 2006, 12, 1081–1087. [Google Scholar] [CrossRef] [PubMed]
- Danieli, B.; Giardini, A.; Lesma, G.; Passarella, D.; Peretto, B.; Sacchetti, A.; Silvani, A.; Pratesi, G.; Zunino, F. Thiocolchicine-podophyllotoxin conjugates: Dynamic libraries based on disulfide exchange reaction. J. Org. Chem. 2006, 71, 2848–2853. [Google Scholar] [CrossRef] [PubMed]
- Scott, D.E.; Dawes, G.J.; Ando, M.; Abell, C.; Ciulli, A. A fragment-based approach to probing adenosine recognition sites by using dynamic combinatorial chemistry. ChemBioChem 2009, 10, 2772–2779. [Google Scholar] [CrossRef] [PubMed]
- Pei, Z.; Larsson, R.; Aastrup, T.; Anderson, H.; Lehn, J.-M.; Ramström, O. Quartz crystal microbalance bioaffinity sensor for rapid identification of glycosyldisulfide lectin inhibitors from a dynamic combinatorial library. Biosens. Bioelectron. 2006, 22, 42–48. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Pei, Z.; Siebert, H.-C.; Ramström, O.; Gabius, H.-J. Glycosyldisulfides from dynamic combinatorial libraries as O-glycoside mimetics for plant and endogenous lectins: Their reactivities in solid-phase and cell assays and conformational analysis by molecular dynamics simulations. Bioorg. Med. Chem. 2006, 14, 6314–6326. [Google Scholar] [CrossRef] [PubMed]
- Liénard, B.M.R.; Selevsek, N.; Oldham, N.J.; Schofield, C.J. Combined mass spectrometry and dynamic chemistry approach to identify metalloenzyme inhibitors. ChemMedChem 2007, 2, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Liénard, B.M.R.; Hüting, R.; Lassaux, P.; Galleni, M.; Frére, J.-M.; Schofield, C.J. Dynamic combinatorial mass spectrometry leads to metallo-β-lactamase inhibitors. J. Med. Chem. 2008, 51, 684–688. [Google Scholar] [CrossRef] [PubMed]
- Shi, B.; Stevenson, R.; Campopiano, D.J.; Greaney, M.F. Discovery of glutathione S-transferase inhibitors using dynamic combinatorial chemistry. J. Am. Chem. Soc. 2006, 128, 8459–8467. [Google Scholar] [CrossRef] [PubMed]
- Caraballo, R.; Dong, H.; Ribeiro, J.P.; Jiménez-Barbero, J.; Ramström, O. Direct STD NMR identification of β-galactosidase inhibitors from a virtual dynamic hemithioacetal system. Angew. Chem. Int. Ed. 2010, 49, 589–539. [Google Scholar] [CrossRef] [PubMed]
- Caraballo, R.; Sakulsombat, M.; Ramström, O. Towards dynamic drug design: Identification and optimisation of β-galactosidase inhibitors from a dynamic hemithioacetal system. ChemBioChem 2010, 11, 1600–1606. [Google Scholar] [CrossRef] [PubMed]
- Leung, I.K.H.; Demetriades, M.; Hardy, A.P.; Lejeune, C.; Smart, T.J.; Szöllössi, A.; Kawamura, A.; Schofield, C.J.; Claridge, T.D.W. Reporter ligand NMR screening method for 2-oxoglutarate oxygenase inhibitors. J. Med. Chem. 2013, 56, 547–555. [Google Scholar] [CrossRef] [PubMed]
- Leung, I.K.H.; Brown, T., Jr.; Schofield, C.J.; Claridge, T.D.W. An approach to enzyme inhibition employing reversible boronate ester formation. Med. Chem. Commun. 2011, 2, 390–395. [Google Scholar] [CrossRef]
- Ji, S.; Cao, W.; Yu, Y.; Xu, H. Dynamic diselenide bonds: Exchange reaction induced by visible light without catalysis. Angew. Chem. Int. Ed. 2014, 53, 6781–6785. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, B.; Sørensen, A.; Gotfredsen, H.; Pittelkow, M. Dynamic combinatorial chemistry with diselenides and disulfides in water. Chem. Commun. 2014, 50, 3716–3718. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, S.-A.; Bornaghi, L.F. Fragment-based drug discovery of carbonic anhydrase II inhibitors by dynamic combinatorial chemistry utilizing alkene cross metathesis. Bioorg. Med. Chem. 2006, 14, 3275–3284. [Google Scholar] [CrossRef] [PubMed]
- Kern, F.T.; Wanner, K.T. Generation and screening of oxime libraries addressing the neuronal GABA transporter GAT1. ChemMedChem 2015, 10, 396–410. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Whitesides, G.M. Selenols catalyze the interchange reactions of dithiols and disulfides in water. J. Org. Chem. 1991, 56, 6931–6933. [Google Scholar] [CrossRef]
- Caldwell, K.A.; Tappel, A.L. Acceleration of sulfhydryl oxidations by selenocystine. Arch. Biochem. Biophys. 1965, 112, 196–200. [Google Scholar] [CrossRef]
- Reddavide, F.V.; Lin, W.; Lehnert, S.; Zhang, Y. DNA-encoded dynamic combinatorial chemical libraries. Angew. Chem. Int. Ed. 2015, 54, 7924–7928. [Google Scholar] [CrossRef] [PubMed]
- Melkko, S.; Scheuermann, J.; Dumelin, C.E.; Neri, D. Encoded self-assembling chemical libraries. Nat. Biotechnol. 2004, 22, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Clark, M.A.; Acharya, R.A.; Arico-Muendel, C.C.; Belyanskaya, S.L.; Benjamin, D.R.; Carlson, N.R.; Centrella, P.A.; Chiu, C.H.; Creaser, S.P.; Cuozzo, J.W.; et al. Design, synthesis and selection of DNA-encoded small-molecule libraries. Nat. Chem. Biol. 2009, 5, 647–654. [Google Scholar] [CrossRef] [PubMed]








| Compound(s) | KD/μM | IC50/μM |
|---|---|---|
 Boronic acid ‘scaffold ligand’ | 24.8 | 126 |
 Boronate ester conjugate | 18.2 * | N/A |
 Boronate ester conjugate | 3.0 * | N/A |
 Boronate ester conjugate | 0.8 * | N/A |
 First generation stable analogue | 7.0 | >500 |
 First generation stable analogue | 1.6 | 107 |
 Second generation stable analogue | 0.5 | 0.017 |
 Second generation stable analogue | 0.8 | 0.013 |
| Peptide | rac-A2B (KD/μM) | meso-A2B (KD/μM) |
|---|---|---|
| H3 K9Me3 | 25 ± 3 | 28 ± 4 |
| H3 K9Me2 | 58 ± 10 | 73 ± 9 |
| H3 K9 Me | 166 ± 50 | not determined |
| H3 K9 | >1200 | not determined |
| Reaction | Scheme | Notes |
|---|---|---|
| Imine formation |  |
|
| Hydrazone formation |  |
|
| Thiol-disulphide exchange |  |
|
| Thiol-enone exchange |  |
|
| Hemithioacetal reaction |  |
|
| Boronate ester formation |  |
|
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, R.; Leung, I.K.H. Protein-Directed Dynamic Combinatorial Chemistry: A Guide to Protein Ligand and Inhibitor Discovery. Molecules 2016, 21, 910. https://doi.org/10.3390/molecules21070910
Huang R, Leung IKH. Protein-Directed Dynamic Combinatorial Chemistry: A Guide to Protein Ligand and Inhibitor Discovery. Molecules. 2016; 21(7):910. https://doi.org/10.3390/molecules21070910
Chicago/Turabian StyleHuang, Renjie, and Ivanhoe K. H. Leung. 2016. "Protein-Directed Dynamic Combinatorial Chemistry: A Guide to Protein Ligand and Inhibitor Discovery" Molecules 21, no. 7: 910. https://doi.org/10.3390/molecules21070910