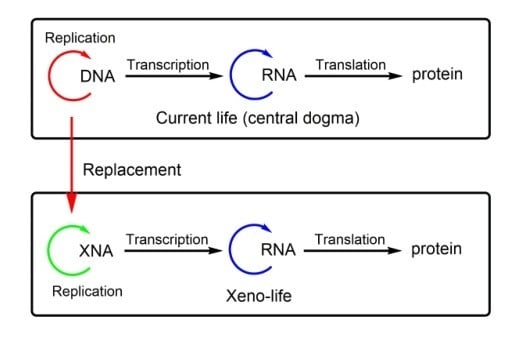
Synthetic Life with Alternative Nucleic Acids as Genetic Materials
Abstract
:1. Introduction
2. XNAs in Synthetic Biology
2.1. Sugar-Modified XNAs
2.2. Phosphate-Modified XNAs
2.3. Sugar- and Phosphate-Modified XNAs
2.4. Base-Modified XNAs
2.5. Unnatural Base Pairs (UBPs)
3. Conclusions and Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Watson, J.D.; Crick, F.H. Genetical implications of the structure of deoxyribonucleic acid. Nature 1953, 171, 964–967. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, M.; Gama-Sosa, M.A.; Carreira, L.H.; Ljungdahl, L.G.; Kuo, K.C.; Gehrke, C.W. DNA methylation in thermophilic bacteria: N4-methylcytosine, 5-methylcytosine, and N6-methyladenine. Nucleic Acids Res. 1985, 13, 1399–1412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Portela, A.; Esteller, M. Epigenetic modifications and human disease. Nat. Biotechnol. 2010, 28, 1057. [Google Scholar] [CrossRef] [PubMed]
- Carell, T.; Kurz, M.Q.; Müller, M.; Rossa, M.; Spada, F. Non-canonical bases in the genome: The regulatory information layer in DNA. Angew. Chem. Int. Ed. 2018, 57, 4296–4312. [Google Scholar] [CrossRef]
- Bilyard, M.K.; Becker, S.; Balasubramanian, S. Natural, modified DNA bases. Curr. Opin. Chem. Biol. 2020, 57, 1–7. [Google Scholar] [CrossRef]
- Weigele, P.; Raleigh, E.A. Biosynthesis and function of modified bases in bacteria and their viruses. Chem. Rev. 2016, 116, 12655–12687. [Google Scholar] [CrossRef]
- Jin, S.-G.; Wu, X.; Li, A.X.; Pfeifer, G.P. Genomic mapping of 5-hydroxymethylcytosine in the human brain. Nucleic Acids Res. 2011, 39, 5015–5024. [Google Scholar] [CrossRef]
- Sánchez-Romero, M.A.; Cota, I.; Casadesús, J. DNA methylation in bacteria: From the methyl group to the methylome. Curr. Opin. Microbiol. 2015, 25, 9–16. [Google Scholar] [CrossRef]
- Warren, R. Modified bases in bacteriophage DNAs. Annu. Rev. Microbiol. 1980, 34, 137–158. [Google Scholar] [CrossRef]
- Stewart, C.R.; Casjens, S.R.; Cresawn, S.G.; Houtz, J.M.; Smith, A.L.; Ford, M.E.; Peebles, C.L.; Hatfull, G.F.; Hendrix, R.W.; Huang, W.M. The genome of Bacillus subtilis bacteriophage SPO1. J. Mol. Biol. 2009, 388, 48–70. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, I.; Marmur, J. Replacement of thymidylic acid by deoxyuridylic acid in the deoxyribonucleic acid of a transducing phage for Bacillus subtilis. Nature 1963, 197, 794–795. [Google Scholar] [CrossRef] [PubMed]
- Price, A.R. Bacteriophage PBS2-induced deoxycytidine triphosphate deaminase in Bacillus subtilis. J. Virol. 1974, 14, 1314–1317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehta, A.P.; Li, H.; Reed, S.A.; Supekova, L.; Javahishvili, T.; Schultz, P.G. Replacement of Thymidine by a Modified Base in the Escherichia coli Genome. J. Am. Chem. Soc. 2016, 138, 7272–7275. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehta, A.P.; Li, H.; Reed, S.A.; Supekova, L.; Javahishvili, T.; Schultz, P.G. Replacement of 2′-Deoxycytidine by 2′-Deoxycytidine Analogues in the E. coli Genome. J. Am. Chem. Soc. 2016, 138, 14230–14233. [Google Scholar] [CrossRef] [Green Version]
- Ghadessy, F.J.; Ramsay, N.; Boudsocq, F.; Loakes, D.; Brown, A.; Iwai, S.; Vaisman, A.; Woodgate, R.; Holliger, P. Generic expansion of the substrate spectrum of a DNA polymerase by directed evolution. Nat. Biotechnol. 2004, 22, 755–759. [Google Scholar] [CrossRef] [PubMed]
- Kunkel, T.A.; Bebenek, K. DNA replication fidelity. Annu. Rev. Biochem. 2000, 69, 497–529. [Google Scholar] [CrossRef] [Green Version]
- Chen, T.; Romesberg, F.E. Directed polymerase evolution. FEBS Lett. 2014, 588, 219–229. [Google Scholar] [CrossRef] [Green Version]
- Houlihan, G.; Arangundy-Franklin, S.; Holliger, P. Exploring the chemistry of genetic information storage and propagation through polymerase engineering. Acc. Chem. Res. 2017, 50, 1079–1087. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Holliger, P. The XNA world: Progress towards replication and evolution of synthetic genetic polymers. Curr. Opin. Chem. Biol. 2012, 16, 245–252. [Google Scholar] [CrossRef]
- Eremeeva, E.; Herdewijn, P. Reprint of: Non Canonical Genetic Material. Curr. Opin. Biotechnol. 2019. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Taylor, A.I.; Cozens, C.; Abramov, M.; Renders, M.; Zhang, S.; Chaput, J.C.; Wengel, J.; Peak-Chew, S.-Y.; McLaughlin, S.H.; et al. Synthetic genetic polymers capable of heredity and evolution. Science 2012, 336, 341–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tarköy, M.; Leumann, C. Synthesis and Pairing Properties of Decanucleotides from (3′ S, 5′ R)-2′-Deoxy-3′, 5′-ethanoβ-D-ribofuranosyladenine and-thymine. Angew. Chem. Int. Ed. 1993, 32, 1432–1434. [Google Scholar] [CrossRef]
- Koshkin, A.A.; Nielsen, P.; Meldgaard, M.; Rajwanshi, V.K.; Singh, S.K.; Wengel, J. LNA (locked nucleic acid): An RNA mimic forming exceedingly stable LNA: LNA duplexes. J. Am. Chem. Soc. 1998, 120, 13252–13253. [Google Scholar] [CrossRef]
- Obika, S.; Nanbu, D.; Hari, Y.; Andoh, J.; Morio, K.; Doi, T.; Imanishi, T. Stability and structural features of the duplexes containing nucleoside analogues with a fixed N-type conformation, 2′-O, 4′-C-methyleneribonucleosides. Tetrahedron Lett. 1998, 39, 5401–5404. [Google Scholar] [CrossRef]
- Augustyns, K.; Van Aerschot, A.; Urbanke, C.; Herdewijn, P. Influence of the Incorporation of 1-(2, 3-Dideoxy-β-D-Erythro-Hexopyranosyl)-Thymine on the Enzymatic Stability and Base-Pairing Properties of Oligodeoxynucleotides. Bull. Soc. Chim. Belg. 1992, 101, 119–130. [Google Scholar] [CrossRef]
- Eschenmoser, A. Etiology of potentially primordial biomolecular structures: From vitamin B12 to the nucleic acids and an inquiry into the chemistry of life’s origin: A retrospective. Angew. Chem. Int. Ed. 2011, 50, 12412–12472. [Google Scholar] [CrossRef]
- Eschenmoser, A.; Dobler, M. Warum Pentose-und nicht Hexose-Nucleinsäuren?? Teil I. Einleitung und Problemstellung, Konformationsanalyse für Oligonucleotid-Ketten aus 2′, 3′-Dideoxyglucopyranosyl-Bausteinen (‘Homo-DNS’) sowie Betrachtungen zur Konformation von A-und B-DNS. Helv. Chim. Acta 1992, 75, 218–259. [Google Scholar] [CrossRef]
- Hendrix, C.; Rosemeyer, H.; Verheggen, I.; Van Aerschot, A.; Seela, F.; Herdewijn, P. 1′, 5′-Anhydrohexitol oligonucleotides: Synthesis, base pairing and recognition by regular oligodeoxyribonucleotides and oligoribonucleotides. Chem. Eur. J. 1997, 3, 110–120. [Google Scholar] [CrossRef]
- Schöning, K.-U.; Scholz, P.; Guntha, S.; Wu, X.; Krishnamurthy, R.; Eschenmoser, A. Chemical etiology of nucleic acid structure: The α-threofuranosyl-(3′→ 2′) oligonucleotide system. Science 2000, 290, 1347–1351. [Google Scholar] [CrossRef]
- Wang, J.; Verbeure, B.; Luyten, I.; Lescrinier, E.; Froeyen, M.; Hendrix, C.; Rosemeyer, H.; Seela, F.; Van Aerschot, A.; Herdewijn, P. Cyclohexene nucleic acids (CeNA): Serum stable oligonucleotides that activate RNase H and increase duplex stability with complementary RNA. J. Am. Chem. Soc. 2000, 122, 8595–8602. [Google Scholar] [CrossRef]
- Noronha, A.M.; Wilds, C.J.; Lok, C.-N.; Viazovkina, K.; Arion, D.; Parniak, M.A.; Damha, M.J. Synthesis and Biophysical Properties of Arabinonucleic Acids (ANA): Circular Dichroic Spectra, Melting Temperatures, and Ribonuclease H Susceptibility of ANA.RNA Hybrid Duplexes. Biochemistry 2000, 39, 7050–7062. [Google Scholar] [CrossRef] [PubMed]
- Wilds, C.J.; Damha, M.J. 2′-Deoxy-2′-fluoro-β-D-arabinonucleosides and oligonucleotides (2′ F-ANA): Synthesis and physicochemical studies. Nucleic Acids Res. 2000, 28, 3625–3635. [Google Scholar] [CrossRef]
- Zhang, L.; Peritz, A.; Meggers, E. A simple glycol nucleic acid. J. Am. Chem. Soc. 2005, 127, 4174–4175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, S.K.; Koshkin, A.A.; Wengel, J.; Nielsen, P. LNA (locked nucleic acids): Synthesis and high-affinity nucleic acid recognition. Chem. Commun. 1998, 455–456. [Google Scholar] [CrossRef]
- Joyce, G.; Inoue, T.; Orgel, L. Non-enzymatic template-directed synthesis on RNA random copolymers: Poly (C, U) templates. J. Mol. Biol. 1984, 176, 279–306. [Google Scholar] [CrossRef]
- Adamala, K.; Szostak, J.W. Nonenzymatic template-directed RNA synthesis inside model protocells. Science 2013, 342, 1098–1100. [Google Scholar] [CrossRef] [Green Version]
- Rose, K.M.; Ferreira-Bravo, I.A.; Li, M.; Craigie, R.; Ditzler, M.A.; Holliger, P.; DeStefano, J.J. Selection of 2′-Deoxy-2′-Fluoroarabino Nucleic Acid (FANA) Aptamers That Bind HIV-1 Integrase with Picomolar Affinity. ACS Chem. Biol. 2019, 14, 2166–2175. [Google Scholar] [CrossRef]
- Ferreira-Bravo, I.A.; Cozens, C.; Holliger, P.; DeStefano, J.J. Selection of 2′-deoxy-2′-fluoroarabinonucleotide (FANA) aptamers that bind HIV-1 reverse transcriptase with picomolar affinity. Nucleic Acids Res. 2015, 43, 9587–9599. [Google Scholar]
- Larsen, A.C.; Dunn, M.R.; Hatch, A.; Sau, S.P.; Youngbull, C.; Chaput, J.C. A general strategy for expanding polymerase function by droplet microfluidics. Nat. Commun. 2016, 7, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Dunn, M.R.; Otto, C.; Fenton, K.E.; Chaput, J.C. Improving polymerase activity with unnatural substrates by sampling mutations in homologous protein architectures. ACS Chem. Biol. 2016, 11, 1210–1219. [Google Scholar] [CrossRef]
- Mei, H.; Liao, J.-Y.; Jimenez, R.M.; Wang, Y.; Bala, S.; McCloskey, C.; Switzer, C.; Chaput, J.C. Synthesis and evolution of a threose nucleic acid aptamer bearing 7-deaza-7-substituted guanosine residues. J. Am. Chem. Soc. 2018, 140, 5706–5713. [Google Scholar] [CrossRef] [PubMed]
- Taylor, A.I.; Pinheiro, V.B.; Smola, M.J.; Morgunov, A.S.; Peak-Chew, S.; Cozens, C.; Weeks, K.M.; Herdewijn, P.; Holliger, P. Catalysts from synthetic genetic polymers. Nature 2015, 518, 427–430. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Ngor, A.K.; Nikoomanzar, A.; Chaput, J.C. Evolution of a general RNA-cleaving FANA enzyme. Nat. Commun. 2018, 9, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, H.; Zhang, S.; Chaput, J.C. Darwinian evolution of an alternative genetic system provides support for TNA as an RNA progenitor. Nat. Chem. 2012, 4, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Verheggen, I.; Van Aerschot, A.; Van Meervelt, L.; Rozenski, J.; Wiebe, L.; Snoeck, R.; Andrei, G.; Balzarini, J.; Claes, P. Synthesis, biological evaluation, and structure analysis of a series of new 1, 5-anhydrohexitol nucleosides. J. Med. Chem. 1995, 38, 826–835. [Google Scholar] [CrossRef]
- Pezo, V.; Liu, F.W.; Abramov, M.; Froeyen, M.; Herdewijn, P.; Marlière, P. Binary Genetic Cassettes for Selecting XNA-Templated DNA Synthesis In Vivo. Angew. Chem. Int. Ed. 2013, 52, 8139–8143. [Google Scholar] [CrossRef] [Green Version]
- Inoue, N.; Minakawa, N.; Matsuda, A. Synthesis and properties of 4′-ThioDNA: Unexpected RNA-like behavior of 4′-ThioDNA. Nucleic Acids Res. 2006, 34, 3476–3483. [Google Scholar] [CrossRef]
- Inoue, N.; Shionoya, A.; Minakawa, N.; Kawakami, A.; Ogawa, N.; Matsuda, A. Amplification of 4′-ThioDNA in the presence of 4′-Thio-dTTP and 4′-Thio-dCTP, and 4′-ThioDNA-directed transcription in vitro and in mammalian cells. J. Am. Chem. Soc. 2007, 129, 15424–15425. [Google Scholar] [CrossRef]
- Maruyama, H.; Furukawa, K.; Kamiya, H.; Minakawa, N.; Matsuda, A. Transcription of 4′-thioDNA templates to natural RNA in vitro and in mammalian cells. Chem. Commun. 2015, 51, 7887–7890. [Google Scholar] [CrossRef] [Green Version]
- Huang, Z.; Schneider, K.C.; Benner, S.A. Building blocks for oligonucleotide analogs with dimethylene sulfide, sulfoxide, and sulfone groups replacing phosphodiester linkages. J. Org. Chem. 1991, 56, 3869–3882. [Google Scholar] [CrossRef]
- Lackey, D.B.; Patel, J. Biochemical synthesis of chirally pure Rp oligonucleotide phosphorothioates. Biotechnol. Lett. 1997, 19, 475–478. [Google Scholar] [CrossRef]
- Lelyveld, V.S.; Zhang, W.; Szostak, J.W. Synthesis of phosphoramidate-linked DNA by a modified DNA polymerase. Proc. Natl. Acad. Sci. USA 2020, 117, 7276–7283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gryaznov, S.; Chen, J.-K. Oligodeoxyribonucleotide N3′. fwdarw. P5′ Phosphoramidates: Synthesis and Hybridization Properties. J. Am. Chem. Soc. 1994, 16, 3143–3144. [Google Scholar] [CrossRef]
- El-Sagheer, A.H.; Sanzone, A.P.; Gao, R.; Tavassoli, A.; Brown, T. Biocompatible artificial DNA linker that is read through by DNA polymerases and is functional in Escherichia coli. Proc. Natl. Acad. Sci. USA 2011, 108, 11338–11343. [Google Scholar] [CrossRef] [Green Version]
- Rejman, D.; Snášel, J.; Liboska, R.; Točík, Z.; Pačes, O.; Králíková, Š.; Rinnová, M.; Koiš, P.; Rosenberg, I. Oligonucleotides with isopolar phosphonate internucleotide linkage: A new perspective for antisense compounds? Nucleosides Nucleotides Nucleic Acids 2001, 20, 819–823. [Google Scholar] [CrossRef]
- Westheimer, F.H. Why nature chose phosphates. Science 1987, 235, 1173–1178. [Google Scholar] [CrossRef]
- Benner, S.A.; Hutter, D. Phosphates, DNA, and the search for nonterrean life: A second generation model for genetic molecules. Bioorg. Chem. 2002, 30, 62–80. [Google Scholar] [CrossRef] [Green Version]
- Roughton, A.L.; Portmann, S.; Benner, S.A.; Egli, M. Crystal structure of a dimethylene sulfone-linked ribodinucleotide analog. J. Am. Chem. Soc. 1995, 117, 7249–7250. [Google Scholar] [CrossRef]
- Matsukura, M.; Shinozuka, K.; Zon, G.; Mitsuya, H.; Reitz, M.; Cohen, J.S.; Broder, S. Phosphorothioate analogs of oligodeoxynucleotides: Inhibitors of replication and cytopathic effects of human immunodeficiency virus. Proc. Natl. Acad. Sci. USA 1987, 84, 7706–7710. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Jiang, S.; Deng, Z.; Dedon, P.C.; Chen, S. DNA phosphorothioate modification—A new multi-functional epigenetic system in bacteria. Fems Microbiol. Rev. 2019, 43, 109–122. [Google Scholar] [CrossRef] [Green Version]
- Zhou, X.; He, X.; Liang, J.; Li, A.; Xu, T.; Kieser, T.; Helmann, J.D.; Deng, Z. A novel DNA modification by sulphur. Mol. Microbiol. 2005, 57, 1428–1438. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Yao, F.; Zhou, X.; Deng, Z.; You, D. A novel host-specific restriction system associated with DNA backbone S-modification in Salmonella. Nucleic Acids Res. 2010, 38, 7133–7141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gryaznov, S.M.; Lloyd, D.H.; Chen, J.-K.; Schultz, R.G.; DeDionisio, L.A.; Ratmeyer, L.; Wilson, W.D. Oligonucleotide N3′--> P5′ phosphoramidates. Proc. Natl. Acad. Sci. USA 1995, 92, 5798–5802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, S.; Blain, J.C.; Zielinska, D.; Gryaznov, S.M.; Szostak, J.W. Fast and accurate nonenzymatic copying of an RNA-like synthetic genetic polymer. Proc. Natl. Acad. Sci. USA 2013, 110, 17732–17737. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Flaherty, D.K.; Zhou, L.; Szostak, J.W. Nonenzymatic template-directed synthesis of mixed-sequence 3′-NP-DNA up to 25 nucleotides long inside model protocells. J. Am. Chem. Soc. 2019, 141, 10481–10488. [Google Scholar] [CrossRef] [Green Version]
- Lelyveld, V.S.; O’Flaherty, D.K.; Zhou, L.; Izgu, E.C.; Szostak, J.W. DNA polymerase activity on synthetic N3′→ P5′ phosphoramidate DNA templates. Nucleic Acids Res. 2019, 47, 8941–8949. [Google Scholar] [CrossRef]
- Nguyen, H.; Abramov, M.; Eremeeva, E.; Herdewijn, P. In Vivo Expression of Genetic Information from Phosphoramidate-DNA. ChemBioChem 2020, 21, 272–278. [Google Scholar] [CrossRef]
- Wolfe, J.L.; Kawate, T.; Belenky, A.; Stanton Jr, V. Synthesis and polymerase incorporation of 5′-amino-2′, 5′-dideoxy-5′-N-triphosphate nucleotides. Nucleic Acids Res. 2002, 30, 3739–3747. [Google Scholar] [CrossRef] [Green Version]
- Renders, M.; Emmerechts, G.; Rozenski, J.; Krecmerová, M.; Holý, A.; Herdewijn, P. Enzymatic synthesis of phosphonomethyl oligonucleotides by therminator polymerase. Angew. Chem. 2007, 119, 2553–2556. [Google Scholar] [CrossRef]
- Renders, M.; Lievrouw, R.; Krecmerová, M.; Holý, A.; Herdewijn, P. Enzymatic polymerization of phosphonate nucleosides. ChemBioChem 2008, 9, 2883–2888. [Google Scholar] [CrossRef] [Green Version]
- Kukwikila, M.; Gale, N.; El-Sagheer, A.H.; Brown, T.; Tavassoli, A. Assembly of a biocompatible triazole-linked gene by one-pot click-DNA ligation. Nat. Chem. 2017, 9, 1089. [Google Scholar] [CrossRef] [PubMed]
- Birts, C.N.; Sanzone, A.P.; El-Sagheer, A.H.; Blaydes, J.P.; Brown, T.; Tavassoli, A. Transcription of click-linked DNA in human cells. Angew. Chem. Int. Ed. 2014, 53, 2362–2365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, C.; Cozens, C.; Jaziri, F.; Rozenski, J.; Marechal, A.; Dumbre, S.; Pezo, V.; Marlière, P.; Pinheiro, V.B.; Groaz, E.; et al. Phosphonomethyl oligonucleotides as backbone-modified artificial genetic polymers. J. Am. Chem. Soc. 2018, 140, 6690–6699. [Google Scholar] [CrossRef] [Green Version]
- Luo, M.; Groaz, E.; Froeyen, M.; Pezo, V.; Jaziri, F.; Leonczak, P.; Schepers, G.; Rozenski, J.; Marlière, P.; Herdewijn, P. Invading Escherichia coli Genetics with a Xenobiotic Nucleic Acid Carrying an Acyclic Phosphonate Backbone (ZNA). J. Am. Chem. Soc. 2019, 141, 10844–10851. [Google Scholar] [CrossRef] [PubMed]
- Holý, A. Antiviral acyclic nucleoside phosphonates structure activity studies. Antivir. Res. 2006, 71, 248–253. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Froeyen, M.; Kempeneers, V.; Pannecouque, C.; Wang, J.; Busson, R.; De Clercq, E.; Herdewijn, P. Deoxythreosyl phosphonate nucleosides as selective anti-HIV agents. J. Am. Chem. Soc. 2005, 127, 5056–5065. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, P.E.; Egholm, M.; Berg, R.H.; Buchardt, O. Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 1991, 254, 1497–1500. [Google Scholar] [CrossRef]
- Sharma, C.; Awasthi, S.K. Versatility of peptide nucleic acids (PNA s): Role in chemical biology, drug discovery, and origins of life. Chem. Biol. Drug Des. 2017, 89, 16–37. [Google Scholar] [CrossRef]
- Nelson, K.E.; Levy, M.; Miller, S.L. Peptide nucleic acids rather than RNA may have been the first genetic molecule. Proc. Natl. Acad. Sci. USA 2000, 97, 3868–3871. [Google Scholar] [CrossRef] [Green Version]
- Böhler, C.; Nielsen, P.E.; Orgel, L.E. Template switching between PNA and RNA oligonucleotides. Nature 1995, 376, 578–581. [Google Scholar] [CrossRef]
- Singhal, A.; Nielsen, P.E. Cross-catalytic peptide nucleic acid (PNA) replication based on templated ligation. Org. Biomol. Chem. 2014, 12, 6901–6907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rosenbaum, D.M.; Liu, D.R. Efficient and sequence-specific DNA-templated polymerization of peptide nucleic acid aldehydes. J. Am. Chem. Soc. 2003, 125, 13924–13925. [Google Scholar] [CrossRef] [PubMed]
- Benner, S.A. Understanding nucleic acids using synthetic chemistry. Acc. Chem. Res. 2004, 37, 784–797. [Google Scholar] [CrossRef] [PubMed]
- Cadet, J.; Wagner, J.R. DNA base damage by reactive oxygen species, oxidizing agents, and UV radiation. Cold Spring Harb. Perspect. Biol. 2013, 5, a012559. [Google Scholar] [CrossRef]
- Kunkel, T.A. DNA replication fidelity. J. Biol. Chem. 2004, 279, 16895–16898. [Google Scholar] [CrossRef] [Green Version]
- Meek, K.N.; Rangel, A.E.; Heemstra, J.M. Enhancing aptamer function and stability via in vitro selection using modified nucleic acids. Methods 2016, 106, 29–36. [Google Scholar] [CrossRef] [Green Version]
- Rohloff, J.C.; Gelinas, A.D.; Jarvis, T.C.; Ochsner, U.A.; Schneider, D.J.; Gold, L.; Janjic, N. Nucleic acid ligands with protein-like side chains: Modified aptamers and their use as diagnostic and therapeutic agents. Mol. Ther. Nucleic Acids 2014, 3, e201. [Google Scholar] [CrossRef]
- Hollenstein, M. Expanding the catalytic repertoire of DNAzymes by modified nucleosides. J. Chim. 2011, 65, 770–775. [Google Scholar] [CrossRef]
- Sidorov, A.V.; Grasby, J.A.; Williams, D.M. Sequence-specific cleavage of RNA in the absence of divalent metal ions by a DNAzyme incorporating imidazolyl and amino functionalities. Nucleic Acids Res. 2004, 32, 1591–1601. [Google Scholar] [CrossRef] [Green Version]
- Eremeeva, E.; Abramov, M.; Margamuljana, L.; Rozenski, J.; Pezo, V.; Marlière, P.; Herdewijn, P. Chemical morphing of DNA containing four noncanonical bases. Angew. Chem. Int. Ed. 2016, 55, 7515–7519. [Google Scholar] [CrossRef] [PubMed]
- Eremeeva, E.; Abramov, M.; Margamuljana, L.; Herdewijn, P. Base-modified nucleic acids as a powerful tool for synthetic biology and biotechnology. Chem. Eur. J. 2017, 23, 9560–9576. [Google Scholar] [CrossRef] [PubMed]
- Krueger, A.T.; Lu, H.; Lee, A.H.; Kool, E.T. Synthesis and properties of size-expanded DNAs: Toward designed, functional genetic systems. Acc. Chem. Res. 2007, 40, 141–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krueger, A.T.; Peterson, L.W.; Chelliserry, J.; Kleinbaum, D.J.; Kool, E.T. Encoding phenotype in bacteria with an alternative genetic set. J. Am. Chem. Soc. 2011, 133, 18447–18451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chelliserrykattil, J.; Lu, H.; Lee, A.H.; Kool, E.T. Polymerase Amplification, Cloning, and Gene Expression of Benzo-Homologous “yDNA” Base Pairs. ChemBioChem 2008, 9, 2976–2980. [Google Scholar] [CrossRef]
- Mehta, A.P.; Wang, Y.; Reed, S.A.; Supekova, L.; Javahishvili, T.; Chaput, J.C.; Schultz, P.G. Bacterial genome containing chimeric DNA–RNA sequences. J. Am. Chem. Soc. 2018, 140, 11464–11473. [Google Scholar] [CrossRef]
- Gavette, J.V.; Stoop, M.; Hud, N.V.; Krishnamurthy, R. RNA–DNA chimeras in the context of an RNA world transition to an RNA/DNA world. Angew. Chem. Int. Ed. 2016, 55, 13204–13209. [Google Scholar] [CrossRef]
- Hanawalt, P. [95] Preparation of 5-bromouracil-labeled DNA. Methods Enzym. 1967, 12, 702–708. [Google Scholar]
- Bick, M.D.; Davidson, R.L. Total substitution of bromodeoxyuridine for thymidine in the DNA of a bromodeoxyuridine-dependent cell line. Proc. Natl. Acad. Sci. USA 1974, 71, 2082–2086. [Google Scholar] [CrossRef] [Green Version]
- Marlière, P.; Patrouix, J.; Döring, V.; Herdewijn, P.; Tricot, S.; Cruveiller, S.; Bouzon, M.; Mutzel, R. Chemical evolution of a bacterium’s genome. Angew. Chem. Int. Ed. 2011, 50, 7109–7114. [Google Scholar] [CrossRef] [Green Version]
- Hamashima, K.; Kimoto, M.; Hirao, I. Creation of unnatural base pairs for genetic alphabet expansion toward synthetic xenobiology. Curr. Opin. Chem. Biol. 2018, 46, 108–114. [Google Scholar] [CrossRef]
- Dien, V.T.; Morris, S.E.; Karadeema, R.J.; Romesberg, F.E. Expansion of the genetic code via expansion of the genetic alphabet. Curr. Opin. Chem. Biol. 2018, 46, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Benner, S.A.; Karalkar, N.B.; Hoshika, S.; Laos, R.; Shaw, R.W.; Matsuura, M.; Fajardo, D.; Moussatche, P. Alternative Watson–Crick synthetic genetic systems. Cold Spring Harb. Perspect. Biol. 2016, 8, a023770. [Google Scholar] [CrossRef]
- Switzer, C.; Moroney, S.E.; Benner, S.A. Enzymatic incorporation of a new base pair into DNA and RNA. J. Am. Chem. Soc. 1989, 111, 8322–8323. [Google Scholar] [CrossRef]
- Zhang, L.; Yang, Z.; Sefah, K.; Bradley, K.M.; Hoshika, S.; Kim, M.-J.; Kim, H.-J.; Zhu, G.; Jimènez, E.; Cansiz, S.; et al. Evolution of functional six-nucleotide DNA. J. Am. Chem. Soc. 2015, 137, 6734–6737. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoshika, S.; Leal, N.A.; Kim, M.-J.; Kim, M.-S.; Karalkar, N.B.; Kim, H.-J.; Bates, A.M.; Watkins, N.E.; SantaLucia, H.A.; Meyer, A.J.; et al. Hachimoji DNA and RNA: A genetic system with eight building blocks. Science 2019, 363, 884–887. [Google Scholar] [CrossRef] [PubMed]
- Schweitzer, B.A.; Kool, E.T. Hydrophobic, non-hydrogen-bonding bases and base pairs in DNA. J. Am. Chem. Soc. 1995, 117, 1863–1872. [Google Scholar] [CrossRef] [Green Version]
- Okamoto, I.; Miyatake, Y.; Kimoto, M.; Hirao, I. High fidelity, efficiency and functionalization of Ds–Px unnatural base pairs in PCR amplification for a genetic alphabet expansion system. ACS Synth. Bio. 2016, 5, 1220–1230. [Google Scholar] [CrossRef]
- Malyshev, D.A.; Dhami, K.; Lavergne, T.; Chen, T.; Dai, N.; Foster, J.M.; Corrêa, I.R.; Romesberg, F.E. A semi-synthetic organism with an expanded genetic alphabet. Nature 2014, 509, 385–388. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Ptacin, J.L.; Fischer, E.C.; Aerni, H.R.; Caffaro, C.E.; San Jose, K.; Feldman, A.W.; Turner, C.R.; Romesberg, F.E. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 2017, 551, 644–647. [Google Scholar] [CrossRef] [Green Version]
- Hirao, I.; Kimoto, M. Unnatural base pair systems toward the expansion of the genetic alphabet in the central dogma. roc. Jpn. Acad. Ser. B 2012, 88, 345–367. [Google Scholar] [CrossRef] [Green Version]
- Kimoto, M.; Yamashige, R.; Matsunaga, K.-I.; Yokoyama, S.; Hirao, I. Generation of high-affinity DNA aptamers using an expanded genetic alphabet. Nat. Biotechnol. 2013, 31, 453. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lamb, B.M.; Feldman, A.W.; Zhou, A.X.; Lavergne, T.; Li, L.; Romesberg, F.E. A semisynthetic organism engineered for the stable expansion of the genetic alphabet. Proc. Natl. Acad. Sci. USA 2017, 114, 1317–1322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fischer, E.C.; Hashimoto, K.; Zhang, Y.; Feldman, A.W.; Dien, V.T.; Karadeema, R.J.; Adhikary, R.; Ledbetter, M.P.; Krishnamurthy, R.; Romesberg, F.E. New codons for efficient production of unnatural proteins in a semisynthetic organism. Nat. Chem. Biol. 2020, 16, 570–576. [Google Scholar] [CrossRef] [PubMed]
- Budisa, N.; Kubyshkin, V.; Schmidt, M. Xenobiology: A Journey towards Parallel Life Forms. ChemBioChem 2020, 21, 1–5. [Google Scholar]
- Schmidt, M.; Pei, L.; Budisa, N. Xenobiology: State-of-the-Art, Ethics, and Philosophy of New-to-Nature Organisms. In Synthetic Biology–Metabolic Engineering; Springer: Cham, Switzerland, 2017; pp. 301–315. [Google Scholar]
- Schmidt, M. Xenobiology: A new form of life as the ultimate biosafety tool. Bioessays 2010, 32, 322–331. [Google Scholar] [CrossRef] [PubMed] [Green Version]










© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nie, P.; Bai, Y.; Mei, H. Synthetic Life with Alternative Nucleic Acids as Genetic Materials. Molecules 2020, 25, 3483. https://doi.org/10.3390/molecules25153483
Nie P, Bai Y, Mei H. Synthetic Life with Alternative Nucleic Acids as Genetic Materials. Molecules. 2020; 25(15):3483. https://doi.org/10.3390/molecules25153483
Chicago/Turabian StyleNie, Peng, Yanfen Bai, and Hui Mei. 2020. "Synthetic Life with Alternative Nucleic Acids as Genetic Materials" Molecules 25, no. 15: 3483. https://doi.org/10.3390/molecules25153483