Label-Free Proteomics Reveals the Molecular Mechanism of Subculture Induced Strain Degeneration and Discovery of Indicative Index for Degeneration in Pleurotus ostreatus
Abstract
:1. Introduction
2. Results
2.1. High-Throughput Proteomics Identification and Quantitative Analysis of the Hyphae from P1 and P10 in P. ostreatus
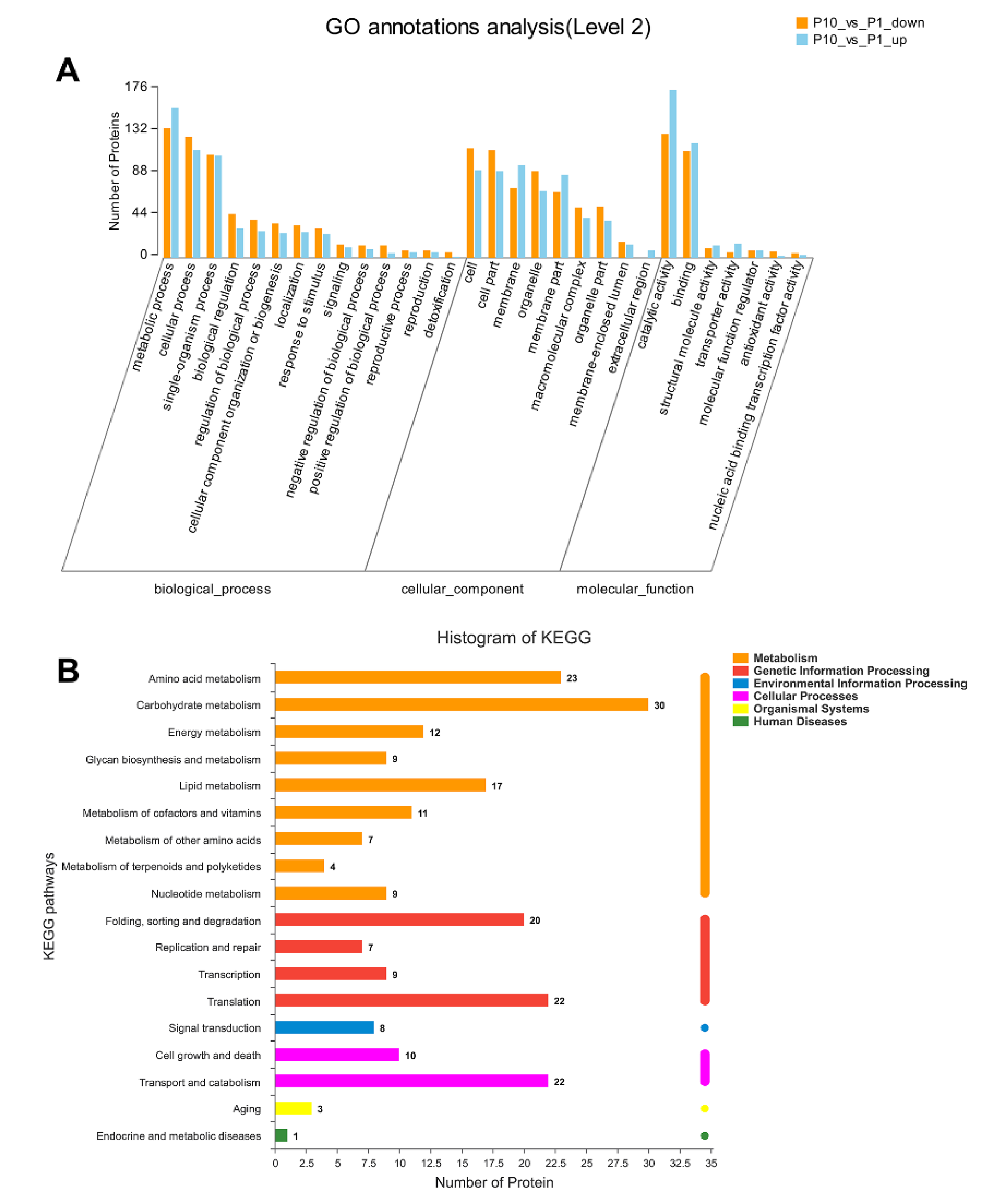
2.2. GO and KEGG Annotation of DEPs in P. ostreatus
2.3. GO Enrichment Analysis Revealed an Asynchronous Repair System in P10 in Response to DNA Damages
2.3.1. Base Excision Repair System Was Activated in the Subcultured Mycelium
2.3.2. Double-Strand Break (DSB) Repair System Impaired in the Subcultured Mycelium
2.4. KEGG Enrichment Analysis Revealed the Cell Cycle Was Stalled in P10
2.5. Enzymatic Activity Assays Showed Consistent Results with the Quantitative Expression of the DEPs
2.6. TUNEL Assay Further Confirmed the Accumulation of DNA Damages from the 1st Generation to the 15th Generation
2.7. Application of the Enzymatic Activity Assay for Strain Degenerative Determination in the Process of P. ostreatus Production
3. Discussion
3.1. Where Are the DNA Damages in the Cells of Hyphae Coming From?
3.2. Multiple Level and Coordinated Regulation of the Cellular Processes in Response to DNA Damage through Cell Cycle Regulation, MAPK Signaling, Chromatin Remodeling, and Epigenetic Modification
3.3. Effective Ways of Preventing the Strain Degeneration During Mushroom Production
4. Materials and Methods
4.1. Mycelium Subculture and Fruiting Body Culture Conditions
4.2. Protein Extraction, Digestion, and Quantification
4.3. LC-MS/MS Analysis
4.4. Data Processing and Analysis
4.5. Bioinformatic Analysis
4.6. Enzymatic Activity Assay
4.7. TUNEL Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sánchez, C. Cultivation of Pleurotus ostreatus and other edible mushrooms. Appl. Microbiol. Biotechnol. 2010, 85, 1321–1337. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Hu, J.; Li, Y.; Yang, B.; Guan, Y.; Xu, C.; Chen, F.; Chi, J.; Bao, Y. Comparative Proteomic Analysis of Pleurotus ostreatus Reveals Great Metabolic Differences in the Cap and Stipe Development and the Potential Role of Ca(2+) in the Primordium Differentiation. Int. J. Mol. Sci. 2019, 20, 6317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, S.K.; Upadhyay, R.C.; Kamal, S.; Tiwari, M. Mushroom cryopreservation and its effect on survival, yield and genetic stability. Cryo Lett. 2004, 25, 23–32. [Google Scholar]
- Liu, Q.; Wang, F.; Liu, K.; Dong, C. Influence of Strain Preservation Methods on Fruiting Body Growth and Metabolite Production by the Medicinal Mushroom Cordyceps militaris (Ascomycetes). Int. J. Med. Mushrooms 2018, 20, 1003–1011. [Google Scholar] [CrossRef] [PubMed]
- Lou, H.; Lin, J.; Guo, L.; Wang, X.; Tian, S.; Liu, C.; Zhao, Y.; Zhao, R. Advances in research on Cordyceps militaris degeneration. Appl. Microbiol. Biotechnol. 2019, 103, 7835–7841. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.J.; Deng, C.H.; Zhang, L.Y.; Hu, K.H. Molecular analysis and biochemical characteristics of degenerated strains of Cordyceps militaris. Arch. Microbiol. 2017, 199, 939–944. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, Z.; Liu, X.; Cui, B.; Miao, W.; Cheng, W.; Zhao, F. Characteristics Analysis Reveals the Progress of Volvariella volvacea Mycelium Subculture Degeneration. Front. Microbiol. 2019, 10, 2045. [Google Scholar] [CrossRef] [PubMed]
- Magae, Y.; Akahane, K.; Nakamura, K.; Tsunoda, S. Simple colorimetric method for detecting degenerate strains of the cultivated basidiomycete Flammulina velutipes (Enokitake). Appl. Environ. Microbiol. 2005, 71, 6388–6389. [Google Scholar] [CrossRef] [Green Version]
- Leonard, T.J. An inherited “neoplasm” in fungus. Proc. Natl. Acad. Sci. USA 1975, 72, 4626–4630. [Google Scholar] [CrossRef] [Green Version]
- Magae, Y.; Hayashi, N. Double-stranded RNA and virus-like particles in the edible basidiomycete Flammulina velutipes (Enokitake). FEMS Microbiol. Lett. 1999, 180, 331–335. [Google Scholar] [CrossRef]
- Qiu, L.; Li, Y.; Liu, Y.; Gao, Y.; Qi, Y.; Shen, J. Particle and naked RNA mycoviruses in industrially cultivated mushroom Pleurotus ostreatus in China. Fungal Biol. 2010, 114, 507–513. [Google Scholar] [CrossRef] [PubMed]
- Xin, X.; Yin, J.; Zhang, B.; Li, Z.; Zhao, S.; Gui, Z. Genome-wide analysis of DNA methylation in subcultured Cordyceps militaris. Arch. Microbiol. 2019, 201, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.C. Cell degeneration and gill remodelling during basidiocarp development in the fungus Coprinus cinereus. Can. J. Bot. 1991, 69, 1161–1169. [Google Scholar] [CrossRef]
- White, S.; McIntyre, M.; Berry, D.R.; McNeil, B. The autolysis of industrial filamentous fungi. Crit. Rev. Biotechnol. 2002, 22, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Venkatachalam, G.; Surana, U.; Clément, M.V. Replication stress-induced endogenous DNA damage drives cellular senescence induced by a sub-lethal oxidative stress. Nucleic Acids Res. 2017, 45, 10564–10582. [Google Scholar] [CrossRef] [PubMed]
- Tubbs, A.; Nussenzweig, A. Endogenous DNA Damage as a Source of Genomic Instability in Cancer. Cell 2017, 168, 644–656. [Google Scholar] [CrossRef]
- Friedberg, E.C. A brief history of the DNA repair field. Cell Res. 2008, 18, 3–7. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Selby, C.P.; Adar, S.; Adebali, O.; Sancar, A. Molecular mechanisms and genomic maps of DNA excision repair in Escherichia coli and humans. J. Biol. Chem. 2017, 292, 15588–15597. [Google Scholar] [CrossRef] [Green Version]
- Sancar, A. DNA excision repair. Annu. Rev. Biochem. 1996, 65, 43–81. [Google Scholar] [CrossRef]
- Friedberg, E.C. A history of the DNA repair and mutagenesis field: The discovery of base excision repair. DNA Repair 2016, 37, A35–A39. [Google Scholar] [CrossRef]
- DeMott, M.S.; Zigman, S.; Bambara, R.A. Replication protein A stimulates long patch DNA base excision repair. J. Biol. Chem. 1998, 273, 27492–27498. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rahmanian, S.; Taleei, R.; Nikjoo, H. Radiation induced base excision repair (BER): A mechanistic mathematical approach. DNA Repair 2014, 22, 89–103. [Google Scholar] [CrossRef]
- Pearl, L.H. Structure and function in the uracil-DNA glycosylase superfamily. Mutat. Res. 2000, 460, 165–181. [Google Scholar] [CrossRef]
- Hölz, K.; Pavlic, A.; Lietard, J.; Somoza, M.M. Specificity and Efficiency of the Uracil DNA Glycosylase-Mediated Strand Cleavage Surveyed on Large Sequence Libraries. Sci. Rep. 2019, 9, 17822. [Google Scholar] [CrossRef] [PubMed]
- Whitaker, A.M.; Freudenthal, B.D. APE1: A skilled nucleic acid surgeon. DNA Repair 2018, 71, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Pekhale, K.; Haval, G.; Perween, N.; Antoniali, G.; Tell, G.; Ghaskadbi, S.; Ghaskadbi, S. DNA repair enzyme APE1 from evolutionarily ancient Hydra reveals redox activity exclusively found in mammalian APE1. DNA Repair 2017, 59, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Raj, J.; Li, J.; Ha, A.; Hossain, M.A.; Richardson, C.; Mukherjee, P.; Yan, S. APE1 senses DNA single-strand breaks for repair and signaling. Nucleic Acids Res. 2020, 48, 1925–1940. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fortini, P.; Pascucci, B.; Parlanti, E.; Sobol, R.W.; Wilson, S.H.; Dogliotti, E. Different DNA polymerases are involved in the short- and long-patch base excision repair in mammalian cells. Biochemistry 1998, 37, 3575–3580. [Google Scholar] [CrossRef]
- Stucki, M.; Pascucci, B.; Parlanti, E.; Fortini, P.; Wilson, S.H.; Hübscher, U.; Dogliotti, E. Mammalian base excision repair by DNA polymerases delta and epsilon. Oncogene 1998, 17, 835–843. [Google Scholar] [CrossRef] [Green Version]
- Nickel, W.; Austermann, S.; Bialek, G.; Grosse, F. Interactions of azidothymidine triphosphate with the cellular DNA polymerases alpha, delta, and epsilon and with DNA primase. J. Biol. Chem. 1992, 267, 848–854. [Google Scholar]
- Shcherbakova, P.V.; Pavlov, Y.I. 3’-->5’ exonucleases of DNA polymerases epsilon and delta correct base analog induced DNA replication errors on opposite DNA strands in Saccharomyces cerevisiae. Genetics 1996, 142, 717–726. [Google Scholar]
- Morrison, A.; Sugino, A. The 3’-->5’ exonucleases of both DNA polymerases delta and epsilon participate in correcting errors of DNA replication in Saccharomyces cerevisiae. Mol. Gen. Genet. 1994, 242, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Raji, H.; Hartsuiker, E. Double-strand break repair and homologous recombination in Schizosaccharomyces pombe. Yeast 2006, 23, 963–976. [Google Scholar] [CrossRef] [PubMed]
- Jeggo, P.A.; Löbrich, M. DNA double-strand breaks: Their cellular and clinical impact? Oncogene 2007, 26, 7717–7719. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, D.O.; Alt, F.W. DNA double strand break repair and chromosomal translocation: Lessons from animal models. Oncogene 2001, 20, 5572–5579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Agarwal, S.; Tafel, A.A.; Kanaar, R. DNA double-strand break repair and chromosome translocations. DNA Repair 2006, 5, 1075–1081. [Google Scholar] [CrossRef]
- Ceccaldi, R.; Rondinelli, B.; D’Andrea, A.D. Repair Pathway Choices and Consequences at the Double-Strand Break. Trends Cell Biol. 2016, 26, 52–64. [Google Scholar] [CrossRef] [Green Version]
- Krenning, L.; van den Berg, J.; Medema, R.H. Life or Death after a Break: What Determines the Choice? Mol. Cell 2019, 76, 346–358. [Google Scholar] [CrossRef]
- Her, J.; Bunting, S.F. How cells ensure correct repair of DNA double-strand breaks. J. Biol. Chem. 2018, 293, 10502–10511. [Google Scholar] [CrossRef] [Green Version]
- Mao, P.; Liu, J.; Zhang, Z.; Zhang, H.; Liu, H.; Gao, S.; Rong, Y.S.; Zhao, Y. Homologous recombination-dependent repair of telomeric DSBs in proliferating human cells. Nat. Commun. 2016, 7, 12154. [Google Scholar] [CrossRef] [Green Version]
- Arnoult, N.; Correia, A.; Ma, J.; Merlo, A.; Garcia-Gomez, S.; Maric, M.; Tognetti, M.; Benner, C.W.; Boulton, S.J.; Saghatelian, A.; et al. Regulation of DNA repair pathway choice in S and G2 phases by the NHEJ inhibitor CYREN. Nature 2017, 549, 548–552. [Google Scholar] [CrossRef] [Green Version]
- Pannunzio, N.R.; Watanabe, G.; Lieber, M.R. Nonhomologous DNA end-joining for repair of DNA double-strand breaks. J. Biol. Chem. 2018, 293, 10512–10523. [Google Scholar] [CrossRef] [Green Version]
- Bouwman, B.A.M.; Crosetto, N. Endogenous DNA Double-Strand Breaks during DNA Transactions: Emerging Insights and Methods for Genome-Wide Profiling. Genes 2018, 9, 632. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lindahl, T.; Barnes, D.E. Repair of endogenous DNA damage. Cold Spring Harb. Symp. Quant. Biol. 2000, 65, 127–133. [Google Scholar] [CrossRef]
- Krizsan, K.; Almasi, E.; Merenyi, Z.; Sahu, N.; Viragh, M.; Koszo, T.; Mondo, S.; Kiss, B.; Balint, B.; Kues, U.; et al. Transcriptomic atlas of mushroom development reveals conserved genes behind complex multicellularity in fungi. Proc. Natl. Acad. Sci. USA 2019, 116, 7409–7418. [Google Scholar] [CrossRef] [Green Version]
- Kiss, E.; Hegedus, B.; Viragh, M.; Varga, T.; Merenyi, Z.; Koszo, T.; Balint, B.; Prasanna, A.N.; Krizsan, K.; Kocsube, S.; et al. Comparative genomics reveals the origin of fungal hyphae and multicellularity. Nat. Commun. 2019, 10, 4080. [Google Scholar] [CrossRef] [Green Version]
- Varga, T.; Krizsan, K.; Foldi, C.; Dima, B.; Sanchez-Garcia, M.; Sanchez-Ramirez, S.; Szollosi, G.J.; Szarkandi, J.G.; Papp, V.; Albert, L.; et al. Megaphylogeny resolves global patterns of mushroom evolution. Nat. Ecol. Evol. 2019, 3, 668–678. [Google Scholar] [CrossRef] [Green Version]
- Kues, U. Life history and developmental processes in the basidiomycete Coprinus cinereus. Microbiol. Mol. Biol. Rev. 2000, 64, 316–353. [Google Scholar] [CrossRef] [Green Version]
- Chen, A.; Wang, Y.; Shao, Y.; Huang, B. A Novel Technique for Rejuvenation of Degenerated Caterpillar Medicinal Mushroom, Cordyceps militaris (Ascomycetes), a Valued Traditional Chinese Medicine. Int. J. Med. Mushrooms 2017, 19, 87–91. [Google Scholar] [CrossRef]
- Longhese, M.P.; Foiani, M.; Muzi-Falconi, M.; Lucchini, G.; Plevani, P. DNA damage checkpoint in budding yeast. EMBO J. 1998, 17, 5525–5528. [Google Scholar] [CrossRef] [Green Version]
- Foiani, M.; Pellicioli, A.; Lopes, M.; Lucca, C.; Ferrari, M.; Liberi, G.; Muzi Falconi, M.; Plevani, P. DNA damage checkpoints and DNA replication controls in Saccharomyces cerevisiae. Mutat. Res. 2000, 451, 187–196. [Google Scholar] [CrossRef]
- Borisova, M.E.; Wagner, S.A.; Beli, P. Mass Spectrometry-Based Proteomics for Quantifying DNA Damage-Induced Phosphorylation. Methods Mol. Biol. 2017, 1599, 215–227. [Google Scholar] [PubMed]
- Zannini, L.; Delia, D.; Buscemi, G. CHK2 kinase in the DNA damage response and beyond. J. Mol. Cell Biol. 2014, 6, 442–457. [Google Scholar] [CrossRef] [Green Version]
- Min, S.; Choi, Y.W.; Yun, H.; Jo, S.; Ji, J.H.; Cho, H. Post-Translational Regulation of the RSF1 Chromatin Remodeler under DNA Damage. Mol. Cells 2018, 41, 127–133. [Google Scholar]
- Dutto, I.; Scalera, C.; Prosperi, E. CREBBP and p300 lysine acetyl transferases in the DNA damage response. Cell Mol. Life Sci. 2018, 75, 1325–1338. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhu, W.G.; Xu, X. Ubiquitin-like modifications in the DNA damage response. Mutat. Res. 2017, 803–805, 56–75. [Google Scholar] [CrossRef]
- Lin, D.; Luo, Y.; Song, Y. The post-transcriptional regulation of the DNA damage response. Yi Chuan Hered. 2014, 36, 309–315. [Google Scholar]
- Carter, R.J.; Parsons, J.L. Base Excision Repair, a Pathway Regulated by Posttranslational Modifications. Mol. Cell Biol. 2016, 36, 1426–1437. [Google Scholar] [CrossRef] [Green Version]
- Haedens, V.; Malagnac, F.; Silar, P. Genetic control of an epigenetic cell degeneration syndrome in Podospora anserina. Fungal Genet. Biol. 2005, 42, 564–577. [Google Scholar] [CrossRef]
- Zhao, X.; Wei, C.; Li, J.; Xing, P.; Li, J.; Zheng, S.; Chen, X. Cell cycle-dependent control of homologous recombination. Acta Biochim. Biophys. Sin. 2017, 49, 655–668. [Google Scholar] [CrossRef] [Green Version]
- Murray, J.M.; Carr, A.M. Integrating DNA damage repair with the cell cycle. Curr. Opin. Cell Biol. 2018, 52, 120–125. [Google Scholar] [CrossRef] [Green Version]
- Hu, Z.; Cools, T.; De Veylder, L. Mechanisms Used by Plants to Cope with DNA Damage. Annu. Rev. Plant Biol. 2016, 67, 439–462. [Google Scholar] [CrossRef] [PubMed]
- Van den Berg, J.; Manjón, A.G.; Kielbassa, K.; Feringa, F.M.; Freire, R.; Medema, R.H. A limited number of double-strand DNA breaks is sufficient to delay cell cycle progression. Nucleic Acids Res. 2018, 46, 10132–10144. [Google Scholar] [CrossRef]
- Kastan, M.B.; Bartek, J. Cell-cycle checkpoints and cancer. Nature 2004, 432, 316–323. [Google Scholar] [CrossRef]
- Seeber, A.; Dion, V.; Gasser, S.M. Checkpoint kinases and the INO80 nucleosome remodeling complex enhance global chromatin mobility in response to DNA damage. Genes Dev. 2013, 27, 1999–2008. [Google Scholar] [CrossRef] [Green Version]
- Adam, S.; Polo, S.E. Chromatin Dynamics during Nucleotide Excision Repair: Histones on the Move. Int. J. Mol. Sci. 2012, 13, 11895–11911. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.-H. Chromatin Remodeling and Epigenetic Regulation in Plant DNA Damage Repair. Int. J. Mol. Sci. 2019, 20, 4093. [Google Scholar] [CrossRef] [Green Version]
- Goellner, E.M. Chromatin remodeling and mismatch repair: Access and excision. DNA Repair 2020, 85, 102733. [Google Scholar] [CrossRef]
- Pandita, T.K.; Christine, R. Chromatin remodeling finds its place in the DNA double-strand break response. Nucleic Acids Res. 2009, 37, 1363–1377. [Google Scholar] [CrossRef]
- Bao, Y.; Shen, X. Chromatin remodeling in DNA double-strand break repair. Curr. Opin. Genet. Dev. 2007, 17, 126–131. [Google Scholar] [CrossRef]
- Hauer, M.H.; Seeber, A.; Singh, V.; Thierry, R.; Sack, R.; Amitai, A.; Kryzhanovska, M.; Eglinger, J.; Holcman, D.; Owen-Hughes, T.; et al. Histone degradation in response to DNA damage enhances chromatin dynamics and recombination rates. Nat. Struct. Mol. Biol. 2017, 24, 99–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Copenhaver, G.P.; Yu, S.; Teng, Y.; Waters, R.; Reed, S.H. How Chromatin Is Remodelled during DNA Repair of UV-Induced DNA Damage in Saccharomyces cerevisiae. PLoS Genet. 2011, 7, e1002124. [Google Scholar]
- Nair, N.; Shoaib, M.; Sørensen, C.S. Chromatin Dynamics in Genome Stability: Roles in Suppressing Endogenous DNA Damage and Facilitating DNA Repair. Int. J. Mol. Sci. 2017, 18, 1486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meas, R.; Wyrick, J.J.; Smerdon, M.J. Nucleosomes Regulate Base Excision Repair in Chromatin. Mutat. Res. 2019, 780, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Hinz, J.M.; Czaja, W. Facilitation of base excision repair by chromatin remodeling. DNA Repair 2015, 36, 91–97. [Google Scholar] [CrossRef] [Green Version]
- Ling, E.; Yin, J.; Xin, X.; Weng, Y.; Gui, Z. Transcriptome-wide analysis reveals the progress of Cordyceps militaris subculture degeneration. PLoS ONE 2017, 12, e0186279. [Google Scholar]
- Wiśniewski, J.R. Quantitative Evaluation of Filter Aided Sample Preparation (FASP) and Multienzyme Digestion FASP Protocols. Anal. Chem. 2016, 88, 5438–5443. [Google Scholar] [CrossRef] [Green Version]








| Accession | Description | P10/P1 | p Value |
|---|---|---|---|
| A0A067NEP6 | Laccase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = LACC10 PE = 3 SV = 1 | 40.59462 | 1.83 × 10−6 |
| A0A067NLM3 | Laccase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = LACC2 PE = 3 SV = 1 | 19.24952 | 1.69 × 10−6 |
| A0A067NMI4 | Small subunit of laccase POXA3a OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1067572 PE = 4 SV = 1 | 13.86408 | 0.001334 |
| A0A067NQH1 | Laccase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = LACC6 PE = 3 SV = 1 | 2.9906 | 2.97 × 10−5 |
| A0A067NW26 | Alpha-galactosidase (Fragment) OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1035175 PE = 3 SV = 1 | 6.665047 | 0.000484 |
| A0A067NTX6 | Alpha-galactosidase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_51341 PE = 3 SV = 1 | 5.675188 | 0.000174 |
| A0A067NVN8 | Alpha-galactosidase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1111105 PE = 3 SV = 1 | 1.706567 | 0.025545 |
| A0A067NN26 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_170071 PE = 3 SV = 1 | 13.35758 | 6.89 × 10−5 |
| A0A067NDF5 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1047336 PE = 3 SV = 1 | 1.457164 | 0.032554 |
| A0A067NDV5 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_46151 PE = 3 SV = 1 | 1.775427 | 0.014483 |
| A0A067NFA3 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1051283 PE = 3 SV = 1 | 1.893255 | 0.039742 |
| A0A067NL60 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1078816 PE = 3 SV = 1 | 3.875206 | 0.002788 |
| A0A067NLN4 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1047807 PE = 3 SV = 1 | 1.67987 | 0.000325 |
| A0A067NQW6 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_160636 PE = 3 SV = 1 | 2.516542 | 0.005202 |
| A0A067NZ51 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1040351 PE = 3 SV = 1 | 2.404623 | 5.04 × 10−7 |
| A0A067P113 | Carboxylic ester hydrolase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1091241 PE = 3 SV = 1 | 2.192162 | 0.000768 |
| A0A067NHY5 | Catalase OS = Pleurotus ostreatus PC15 OX = 1137138 GN = PLEOSDRAFT_1090819 PE = 3 SV = 1 | 0.337985 | 1.12 × 10−5 |
Sample Availability: Samples of the P1 and P10 mycelium are available from the authors. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, W.; Hu, J.; Chi, J.; Li, Y.; Yang, B.; Hu, W.; Chen, F.; Xu, C.; Chai, L.; Bao, Y. Label-Free Proteomics Reveals the Molecular Mechanism of Subculture Induced Strain Degeneration and Discovery of Indicative Index for Degeneration in Pleurotus ostreatus. Molecules 2020, 25, 4920. https://doi.org/10.3390/molecules25214920
Zhu W, Hu J, Chi J, Li Y, Yang B, Hu W, Chen F, Xu C, Chai L, Bao Y. Label-Free Proteomics Reveals the Molecular Mechanism of Subculture Induced Strain Degeneration and Discovery of Indicative Index for Degeneration in Pleurotus ostreatus. Molecules. 2020; 25(21):4920. https://doi.org/10.3390/molecules25214920
Chicago/Turabian StyleZhu, Weiwei, Jinbo Hu, Jingliang Chi, Yang Li, Bing Yang, Wenli Hu, Fei Chen, Chong Xu, Linshan Chai, and Yongming Bao. 2020. "Label-Free Proteomics Reveals the Molecular Mechanism of Subculture Induced Strain Degeneration and Discovery of Indicative Index for Degeneration in Pleurotus ostreatus" Molecules 25, no. 21: 4920. https://doi.org/10.3390/molecules25214920





