Finding the First Potential Inhibitors of Shikimate Kinase from Methicillin Resistant Staphylococcus aureus through Computer-Assisted Drug Design
Abstract
:1. Introduction
2. Results and Discussion
2.1. Compounds Filtering
2.2. Virtual Screening
2.3. Molecular Dynamics Studies
2.4. Linear Interaction Energy
2.5. ADME-Tox Evaluation
3. Materials and Methods
3.1. Small Molecules Chemical Library
3.2. Docking Studies
3.3. Molecular Dynamics Studies
3.4. Linear Interaction Energy Calculations
3.5. ADME Properties Prediction
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aslam, B.; Wang, W.; Arshad, M.I.; Khurshid, M.; Muzammil, S.; Rasool, M.H.; Nisar, N.A.; Alvi, R.F.; Aslam, M.A.; Qamar, M.U.; et al. Antibiotic resistance: A rundown of a global crisis. Infect. Drug Resist. 2018, 11, 1645–1658. [Google Scholar] [CrossRef] [Green Version]
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Pulcini, C.; Kahlmeter, G.; Kluytmans, J.; Carmeli, Y.; et al. WHO Pathogens Priority List Working Group. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327. [Google Scholar] [CrossRef]
- World Health Organization. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. Available online: http://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf (accessed on 15 August 2019).
- Tong, S.Y.; Davis, J.S.; Eichenberger, E.; Holland, T.L.; Fowler, V.G., Jr. Staphylococcus aureus infections: Epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 2015, 28, 603–661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kourtis, A.P.; Hatfield, K.; Baggs, J.; Mu, Y.; See, I.; Epson, E.; Nadle, J.; Kainer, M.A.; Dumyati, G.; Petit, S.; et al. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus aureus Bloodstream Infections—United States. MMWR Morb. Mortal Wkly. Rep. 2019, 68, 214–219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drew, R.H. Emerging options for treatment of invasive, multidrug-resistant Staphylococcus aureus infections. Pharmacotherapy 2007, 27, 227–249. [Google Scholar] [CrossRef]
- McCallum, N.; Berger-Bächi, B.; Senn, M.M. Regulation of antibiotic resistance in Staphylococcus aureus. Int. J. Med. Microbiol. 2007, 300, 118–129. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M. Multidrug-Resistant Staphylococcus aureus, India, 2013–2015. Emerg. Infect. Dis. 2016, 22, 1666–1667. [Google Scholar] [CrossRef] [Green Version]
- McGuinness, W.A.; Malachowa, N.; DeLeo, F.R. Vancomycin Resistance in Staphylococcus aureus. Yale J. Biol. Med. 2017, 90, 269–281. [Google Scholar]
- Duval, R.E.; Grare, M.; Demoré, B. Fight Against Antimicrobial Resistance: We Always Need New Antibacterials but for Right Bacteria. Molecules 2019, 24, 3152. [Google Scholar] [CrossRef] [Green Version]
- Gajdács, M. The Concept of an Ideal Antibiotic: Implications for Drug Design. Molecules 2019, 24, 892. [Google Scholar] [CrossRef] [Green Version]
- Bentley, R. The shikimate pathway—A metabolic tree with many branches. Crit. Rev. Biochem. Mol. Biol. 1990, 25, 307–384. [Google Scholar] [CrossRef] [PubMed]
- Haslam, E.H. (Ed.) 1—The Shikimate Pathway: Biosynthesis of the Aromatic Amino Acids. In Biosynthesis of Natural Products Series, The Shikimate Pathway; Butterworth-Heinemann: Oxford, UK, 1974; pp. 3–48. ISBN 9780408705691. [Google Scholar]
- Coggins, J.R.; Abell, C.; Evans, L.B.; Frederickson, M.; Robinson, D.A.; Roszak, A.W.; Lapthorn, A.P. Experiences with the shikimate-pathway enzymes as targets for rational drug design. Biochem. Soc. Trans. 2003, 31 Pt 3, 548–552. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.; Simithy, J.; Goodwin, D.C.; Calderón, A.I. Selective Mycobacterium tuberculosis Shikimate Kinase Inhibitors as Potential Antibacterials. Perspect. Med. Chem. 2015, 7, 9–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parish, T.; Stoker, N.G. The common aromatic amino acid biosynthesis pathway is essential in Mycobacterium tuberculosis. Microbiology 2002, 148 Pt 10, 3069–3077. [Google Scholar] [CrossRef] [Green Version]
- Gan, J.; Gu, Y.; Li, Y.; Yan, H.; Ji, X. Crystal structure of Mycobacterium tuberculosis shikimate kinase in complex with shikimic acid and an ATP analogue. Biochemistry 2006, 45, 8539–8545. [Google Scholar] [CrossRef]
- Favela-Candia, A.; Téllez-Valencia, A.; Campos-Almazán, M.; Sierra-Campos, E.; Valdez-Solana, M.; Oria-Hernández, J.; Castillo-Villanueva, A.; Nájera, H.; Avitia-Domínguez, C. Biochemical, Kinetic, and Computational Structural Characterization of Shikimate Kinase from Methicillin-Resistant Staphylococcus aureus. Mol. Biotechnol. 2019, 61, 274–285. [Google Scholar] [CrossRef]
- Hughes, J.P.; Rees, S.; Kalindjian, S.B.; Philpott, K.L. Principles of early drug discovery. Br. J. Pharmacol. 2011, 162, 1239–1249. [Google Scholar] [CrossRef] [Green Version]
- Gimeno, A.; Ojeda-Montes, M.J.; Tomás-Hernández, S.; Cereto-Massagué, A.; Beltrán-Debón, R.; Mulero, M.; Pujadas, G.; Garcia-Vallvé, S. The Light and Dark Sides of Virtual Screening: What Is There to Know? Int. J. Mol. Sci. 2019, 20, 1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartmann, M.D.; Bourenkov, G.P.; Oberschall, A.; Strizhov, N.; Bartunik, H.D. Mechanism of phosphoryl transfer catalyzed by shikimate kinase from Mycobacterium tuberculosis. J. Mol. Biol. 2006, 364, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Blanco, B.; Prado, V.; Lence, E.; Otero, J.M.; Garcia-Doval, C.; van Raaij, M.J.; Llamas-Saiz, A.L.; Lamb, H.; Hawkins, A.R.; González-Bello, C. Mycobacterium tuberculosis Shikimate Kinase Inhibitors: Design and Simulation Studies of the Catalytic Turnover. J. Am. Chem. Soc. 2013, 135, 12366–12376. [Google Scholar] [CrossRef] [PubMed]
- Pereira, J.H.; de Oliveira, J.S.; Canduri, F.; Dias, M.V.; Palma, M.S.; Basso, L.A.; Santos, D.S.; de Azevedo, W.F., Jr. Structure of shikimate kinase from Mycobacterium tuberculosis reveals the binding of shikimic acid. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, D60, 2310–2319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, C.; Zhang, J.; Chen, L.; Chen, K.; Shen, X.; Jiang, H. Discovery of Helicobacter pylori shikimate kinase inhibitors: Bioassay and molecular modeling. Bioorganic Med. Chem. 2007, 15, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Mehra, R.; Rajput, V.S.; Gupta, M.; Chib, R.; Kumar, A.; Wazir, P.; Khan, I.A.; Nargotra, A. Benzothiazole Derivative as a Novel Mycobacterium tuberculosis Shikimate Kinase Inhibitor: Identification and Elucidation of Its Allosteric Mode of Inhibition. J. Chem. Inf. Model. 2016, 56, 930–940. [Google Scholar] [CrossRef] [PubMed]
- Rajput, V.S.; Mehra, R.; Kumar, S.; Nargotra, A.; Singh, P.P.; Khan, I.A. Screening of antitubercular compound library identifies novel shikimate kinase inhibitors of Mycobacterium tuberculosis. Appl. Microbiol. Biotechnol. 2016, 100, 5415–5426. [Google Scholar] [CrossRef] [PubMed]
- Simithy, J.; Fuanta, N.R.; Alturki, M.; Hobrath, J.V.; Wahba, A.E.; Pina, I.; Rath, J.; Hamann, M.T.; DeRuiter, J.; Goodwin, D.C.; et al. Slow-Binding Inhibition of Mycobacterium tuberculosis Shikimate Kinase by Manzamine Alkaloids. Biochemistry 2018, 57, 4923–4933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, W.C.; Chang, Y.N.; Wang, W.C. Structural basis for shikimate-binding specificity of Helicobacter pylori shikimate kinase. J. Bacteriol. 2005, 187, 8156–8163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, Y.; Reshetnikova, L.; Li, Y.; Wu, Y.; Yan, H.; Singh, S.; Ji, X. Crystal structure of shikimate kinase from Mycobacterium tuberculosis reveals the dynamic role of the LID domain in catalysis. J. Mol. Biol. 2002, 319, 779–789. [Google Scholar] [CrossRef]
- Åqvist, J.; Medina, C.; Samuelsson, J.-E. A new method for predicting binding affinity in computer-aided drug design. Protein Eng. Des. Sel. 1994, 7, 385–391. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [Green Version]
- Seul, South Corea: Bioinformatics and Molecular Design Research Center; PreADMET Program. 2004. Available online: http://preadmet.bmdrc.org (accessed on 23 August 2021).
- Sterling, T.; Irwin, J.J. ZINC 15—Ligand Discovery for Everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Limban, C.; Nuţă, D.C.; Chiriţă, C.; Negreș, S.; Arsene, A.L.; Goumenou, M.; Karakitsios, S.P.; Tsatsakis, A.M.; Sarigiannis, D.A. The use of structural alerts to avoid the toxicity of pharmaceuticals. Toxicol. Rep. 2018, 5, 943–953. [Google Scholar] [CrossRef] [PubMed]
- Sander, T.; Freyss, J.; von Korff, M.; Rufener, C. DataWarrior: An open-source program for chemistry aware data visualization and analysis. J. Chem. Inf. Model. 2015, 55, 460–473. [Google Scholar] [CrossRef]
- Schrödinger Suite Release 2019-4; Schrödinger, LLC: New York, NY, USA, 2019.
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Murphy, R.B.; Repasky, M.P.; Frye, L.L.; Greenwood, J.R.; Halgren, T.A.; Sanschagrin, P.C.; Mainz, D.T. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Pall, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High Performance Molecular Simulations through Multi-Level Parallelism from Laptops to Supercomputers. SoftwareX 2015, 1, 19–25. [Google Scholar] [CrossRef] [Green Version]
- Huang, J.; MacKerell, A.D. CHARMM36 All-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 2013, 34, 2135–2145. [Google Scholar] [CrossRef] [Green Version]
- Zoete, V.; Cuendet, M.A.; Grosdidier, A.; Michielin, O. SwissParam: A fast force field generation tool for small organic molecules. J. Comput. Chem. 2011, 32, 2359–2368. [Google Scholar] [CrossRef]
- Boonstra, S.; Onck, P.R.; van der Giessen, E. CHARMM TIP3P Water Model Suppresses Peptide Folding by Solvating the Unfolded State. J. Phys. Chem. B 2016, 120, 3692–3698. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef] [Green Version]
- Parrinello, M.; Rahman, A. Polymorphic Transitions in Single Crystals: A New Molecular Dynamics Method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Punkvang, A.; Saparpakorn, P.; Hannongbua, S.; Wolschann, P.; Beyer, A.; Pungpo, P. Investigating the structural basis of arylamides to improve potency against M. tuberculosis strain through molecular dynamics simulations. Eur. J. Med. Chem. 2010, 45, 5585–5593. [Google Scholar] [CrossRef] [PubMed]
- Aguayo-Ortiz, R.; Méndez-Lucio, O.; Medina-Franco, J.L.; Castillo, R.; Yépez-Mulia, L.; Hernández-Luis, F.; Hernández-Campos, A. Towards the identification of the binding site of benzimidazoles to β-tubulin of Trichinella spiralis: Insights from computational and experimental data. J. Mol. Graph. Model. 2013, 41, 12–19. [Google Scholar] [CrossRef] [PubMed]






| Compound | Zinc ID | Structure | Docking Score (kcal/mol) | Interacting Residues (Distance Cut-Off 4.0Å) |
|---|---|---|---|---|
| C1 | 000737165696 |  | −3.905 | Arg61 2, Gly82 1, Arg120 1, 2, Arg138 1, Met14 4, Asp37 4, Glu41 4, Thr47 4, Ile48 4, Phe49 4, Phe52 4, Gly81 4, Gly83 4, Asp116 4, His119 4, Pro121 4, Asn122 4 |
| C2 | 000019366016 |  | −3.846 | Lys18 2, Arg61 2, Arg120 2, Arg138 2, Met14 4, Gly15 4, Lys18 4, Asp37 4, Ile48 4, Phe52 4, Phe60 4, Glu64 4, Thr80 4, Gly81 4, Gly82 4, Gly83 4, Pro121 4, Asn122 4 |
| C3 | 000653035164 S enantiomer |  | −3.451 | Asp35 1, Asp37 1, Arg120 1, Arg138 1, Phe13 4, Met14 4, Lys18 4, Ser19 4, Ile36 4, Glu41 4, Ile48 4, Phe52 4, Arg61 4, Ala79 4, Thr80 4, Gly81 4, Gly82 4, Gly83 4, Asp116 4, His119 4, Pro121 4, Asn122 4 |
| C4 | 000000197090 |  | −3.406 | Glu41 1,2, Gly82 1, Asp116 2, Arg120 1, Met14 4, Gly15 4, Thr16 4, Gly17 4, Lys18 4, Ser19 4, Asp35 4, Asp37 4, Ser38 4, Ile48 4, Pro49 4, Phe52 4, Phe60 4, Arg61 4, Ala79 4, Thr80 4, Gly81 4, Gly83 4, His119 4, Pro121 4, Asn122 4, Arg138 4 |
| C5 | 001153862505 R enantiomer | 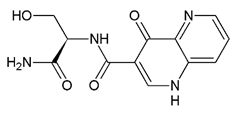 | −3.286 | Lys18 2, Arg61 1, Glu64 1, Gly82 1, Arg120 3, Met14 4, Ser19 4, Asp35 4, Asp37 4, Ile48 4, Phe52 4, Phe60 4, Thr80 4, Gly81 4, Gly83 4, Ile84 4, Pro121 4, Asn122 4, Arg138 4 |
| 0 ns | 20 ns | 40 ns | 60 ns | 80 ns | 100 ns | |
|---|---|---|---|---|---|---|
| C1 | * Arg61, Gly82, Arg138 | * Gly15, Lys18 | * Gly15, Lys18, Arg61 | * Gly15, Lys18, Arg138 | * Gly15, Lys18, Arg138 | * Gly15, Lys18, Arg138 |
| C2 | * Lys18, Arg61, Gly82, Arg120 Asn122, Arg138 | * Lys18, Arg61, Gly82 | * Lys18, Arg61, Gly82, Asn115 | * Lys18, Gly82, Lys126 | * Asn122, Lys 126, Arg138 | * Thr127 |
| C3 | * Lys18, Asp37, Arg120, Arg138 | * Ser19, Asp35, Gly82, Arg120, Asn124 | * Ser19, Asp35, Arg120, Asn124 | * Asp35, Arg120, Ala123 | * Ser19, Asp35, Arg61 | * Ser19, Asp35, Arg120, Asn122 |
| C4 | *Ser19, Glu41, Arg120 | *Glu41, Gly82 | *Glu41, Arg120 | *Asp37, Glu41 | *Glu41, Gly82 | *Ser19, Glu41 , |
| Energy (Kcal/mol) | |||||
|---|---|---|---|---|---|
| Complex | (VLJ)Bound | (VLJ)Free | (VCL)Bound | (VCL)Free | ΔGbind kcal/mol |
| SaSK-C1 | −21.39 | −5.46 | −107.92 | −70.41 | −21.62 |
| SaSK-C2 | −14.45 | 1.96 | −65.15 | −160.90 | 48.41 |
| SaSK-C3 | −13.93 | −4.69 | −6.74 | −8.58 | 0.8 |
| SaSK-C4 | −23.25 | −14.70 | −28.31 | −43.56 | −37.47 |
| C1 | C2 | C3 | C4 | |
|---|---|---|---|---|
| MW (g/mol) | 266.24 | 272.30 | 328.34 | 335.4 |
| RB | 4 | 4 | 5 | 3 |
| HBA | 9 | 8 | 7 | 6 |
| HBD | 0 | 0 | 4 | 1 |
| MR | 58.9 | 59.83 | 81.04 | 102.64 |
| TPSA (Å2) | 124.04 | 137.64 | 134.01 | 78.88 |
| cLogP | 0.47 | −2.60 | 0.68 | 2.96 |
| Lipinski rules violations | 0 | 0 | 0 | 0 |
| Water Solubility | ||||
| LogS | −1.06 | 2.84 | −1.0 | −1.87 |
| Class | Very soluble | Highly soluble | Very soluble | Very soluble |
| Druglikeness | ||||
| Ghose | Yes | 1 violation: WLOGP < −0.4 | 1 violation: WLOGP<-0.4 | No; 1 violation: WLOGP < −0.4 |
| Veber | Yes | Yes | Yes | Yes |
| Egan | Yes | 1 violation: TPSA > 131.6 | 1 violation: TPSA > 131.6 | Yes |
| Muegge | Yes | 1 violation: XLOGP3 < −2 | Yes | Yes |
| Bioavailability Score | 0.56 | 0.55 | 0.55 | 0.55 |
| Medicinal Chemistry | ||||
| PAINS | No alerts | No alerts | No alerts | No alerts |
| Brenk | No alerts | 1 alert: sulfonic_acid_2 | No alerts | No alerts |
| Leadlikeness | Yes | Yes | No | Yes |
| Synthetic accessibility | 2.21 | 3.12 | 3.19 | 3.16 |
| C1 | C2 | C3 | C4 | |
|---|---|---|---|---|
| BBBP | 0.0488584 | 0.0466103 | 0.037606 | 0.0466103 |
| CaCo-2 (nm/s) | 4.65788 | 2.24237 | 0.373322 | 2.24237 |
| HIA (%) | 64.622234 | 58.373794 | 69.411618 | 58.373794 |
| MDCK (nm/s) | 1.05816 | 354.049 | 0.591682 | 354.049 |
| In vitro P-glycoprotein inhibition | Non | Non | Non | Non |
| PPB (%) | 65.392204 | 45.848496 | 39.567509 | 45.848496 |
| Water solubility in pure water (mg/L) | 9254.72 | 2.96157e + 006 | 3141.21 | 2.96157e + 006 |
| In vitro skin permeability (logKp, cm/h) | −2.71782 | −2.46455 | −4.61446 | −2.46455 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rios-Soto, L.; Téllez-Valencia, A.; Sierra-Campos, E.; Valdez-Solana, M.; Cisneros-Martínez, J.; Gómez Palacio-Gastélum, M.; Castillo-Villanueva, A.; Avitia-Domínguez, C. Finding the First Potential Inhibitors of Shikimate Kinase from Methicillin Resistant Staphylococcus aureus through Computer-Assisted Drug Design. Molecules 2021, 26, 6736. https://doi.org/10.3390/molecules26216736
Rios-Soto L, Téllez-Valencia A, Sierra-Campos E, Valdez-Solana M, Cisneros-Martínez J, Gómez Palacio-Gastélum M, Castillo-Villanueva A, Avitia-Domínguez C. Finding the First Potential Inhibitors of Shikimate Kinase from Methicillin Resistant Staphylococcus aureus through Computer-Assisted Drug Design. Molecules. 2021; 26(21):6736. https://doi.org/10.3390/molecules26216736
Chicago/Turabian StyleRios-Soto, Lluvia, Alfredo Téllez-Valencia, Erick Sierra-Campos, Mónica Valdez-Solana, Jorge Cisneros-Martínez, Marcelo Gómez Palacio-Gastélum, Adriana Castillo-Villanueva, and Claudia Avitia-Domínguez. 2021. "Finding the First Potential Inhibitors of Shikimate Kinase from Methicillin Resistant Staphylococcus aureus through Computer-Assisted Drug Design" Molecules 26, no. 21: 6736. https://doi.org/10.3390/molecules26216736






