Putative Biosynthesis of Talarodioxadione & Talarooxime from Talaromyces stipitatus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fermentation and Extraction
2.2. Isolation of Metabolites
2.3. Structure Elucidation
2.4. Spectroscopic and Spectrometric Data
2.5. Bioinformatics Analysis
3. Results and Discussion
3.1. Structure Elucidation
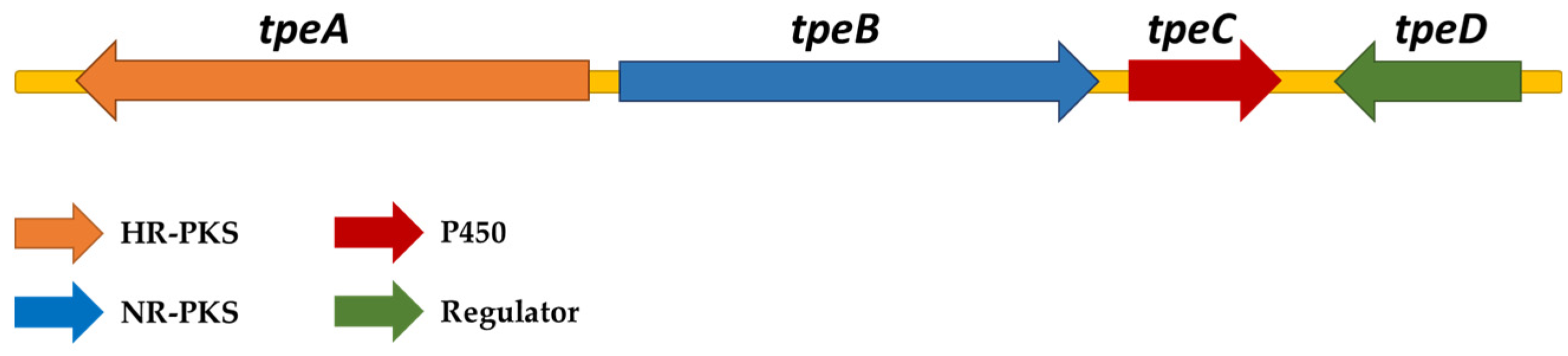
3.2. Identification of Talaromyces Stipitatus Polyesters Biosynthetic Gene Cluster
3.3. The Biosynthesis of Talaromyces Stipitatus Polyesters
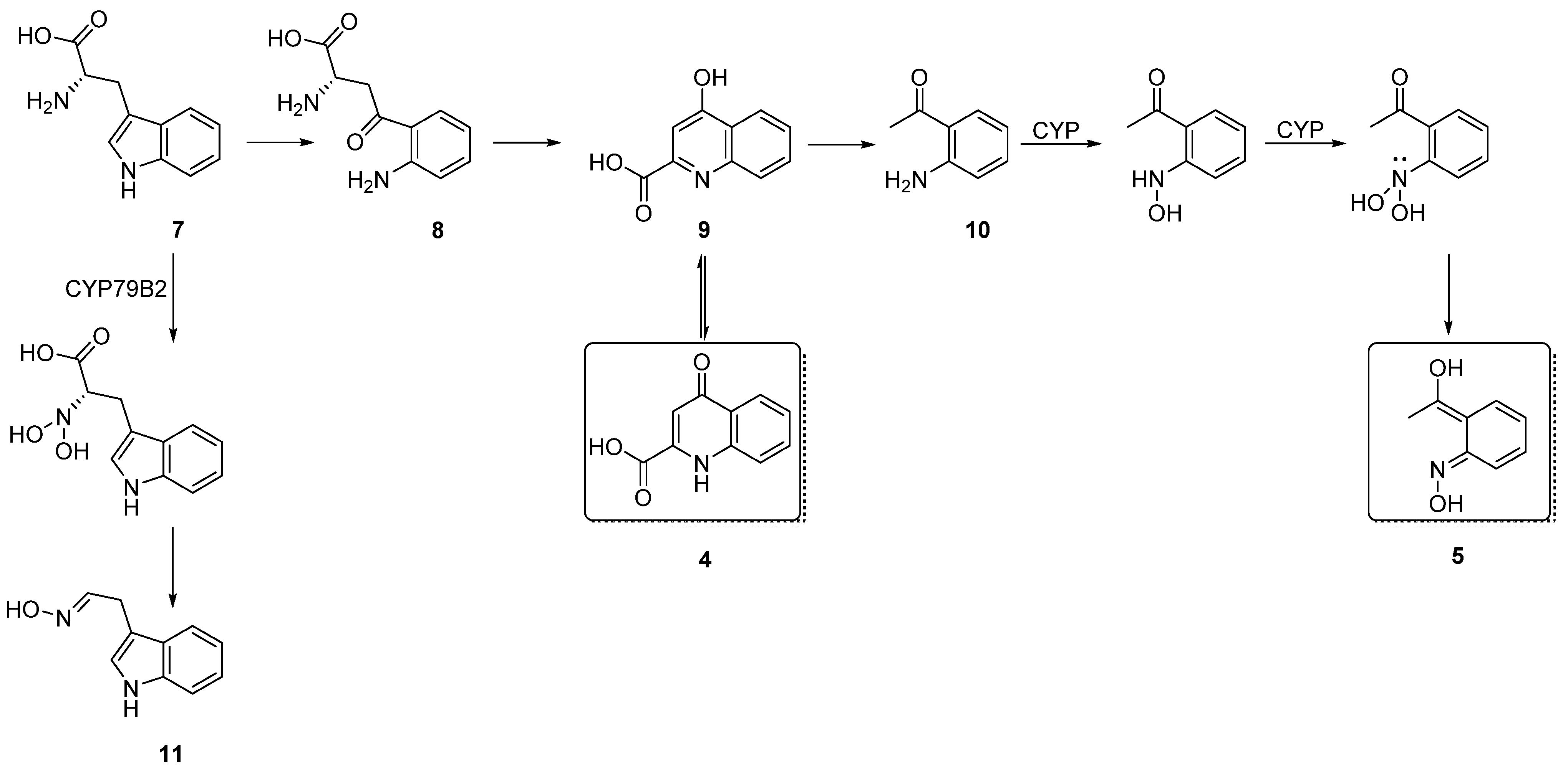
3.4. The Biosynthesis of Talarooxime
4. Conclusions
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Keller, N.P. Fungal secondary metabolism: Regulation, function and drug discovery. Nat. Rev. Microbiol. 2019, 17, 167–180. [Google Scholar] [CrossRef] [PubMed]
- Lan, D.; Wu, B. Chemistry and Bioactivities of Secondary Metabolites from the Genus Talaromyces. Chem. Biodivers. 2020, 17, e2000229. [Google Scholar] [CrossRef] [PubMed]
- Nicoletti, R.; Trincone, A. Bioactive Compounds Produced by Strains of Penicillium and Talaromyces of Marine Origin. Mar. Drugs 2016, 14, 37. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ito, M.; Tsuchida, Y.; Mizoue, K.; Hanada, K. NG-011 and NG-012, novel potentiators of nerve growth factor. II. The structure determination of NG-011 and NG-012. J. Antibiot. 1992, 45, 1566–1572. [Google Scholar] [CrossRef] [Green Version]
- Breinholt, J.; Jensen, G.W.; Nielsen, R.I.; Olsen, C.E.; Frisvad, J.C. Antifungal macrocyclic polylactones from Penicillium verruculosum. J. Antibiot. 1993, 46, 1101–1108. [Google Scholar] [CrossRef] [Green Version]
- Roy, K.; Chatterjee, S.; Deshmukh, S.K.; Vijayakumar, E.K.; Ganguli, B.N.; Fehlhaber, H.W. Orbuticin, a new secondary metabolite from Acremonium butyri. J. Antibiot. 1996, 49, 1186–1187. [Google Scholar] [CrossRef] [Green Version]
- Arai, M.; Tomoda, H.; Okuda, T.; Wang, H.; Tabata, N.; Masuma, R.; Yamaguchi, Y.; Omura, S. Funicone-related compounds, potentiators of antifungal miconazole activity, produced by Talaromycesflavus FKI-0076. J. Antibiot. 2002, 55, 172–180. [Google Scholar] [CrossRef] [Green Version]
- He, J.-W.; Mu, Z.-Q.; Gao, H.; Chen, G.-D.; Zhao, Q.; Hu, D.; Sun, J.-Z.; Li, X.-X.; Li, Y.; Liu, X.-Z. New polyesters from Talaromyces flavus. Tetrahedron 2014, 70, 4425–4430. [Google Scholar] [CrossRef]
- Ruanglek, V.; Chokpaiboon, S.; Rattanaphan, N.; Madla, S.; Auncharoen, P.; Bunyapaiboonsri, T.; Isaka, M. Menisporopsin B, a new polyester from the seed fungus Menisporopsis theobromae BCC 4162. J. Antibiot. 2007, 60, 748–751. [Google Scholar] [CrossRef]
- Zhang, J.-L.; Wang, W.-J.; Xu, X.-M.; Li, D.-Y.; Hua, H.-M.; Ma, E.-L.; Li, Z.-L. From macrocyclic to linear and further: Naturally degradable polyesters from the fungus Ascotricha sp. ZJ-M-5. Tetrahedron 2016, 72, 4895–4901. [Google Scholar] [CrossRef]
- Wu, Z.; Liu, D.; Huang, J.; Proksch, P.; Zhu, K.; Lin, W. Hansforesters A–M, polyesters from the sponge-associated fungus Hansfordia sinuosae with antibacterial activities. RSC Adv. 2018, 8, 39756–39768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sultan, S. Bioactive Polyesters in Marine and Plant Endophytic Fungi. In Polyester; IntechOpen: London, UK, 2012. [Google Scholar]
- Zhai, M.M.; Li, J.; Jiang, C.X.; Shi, Y.P.; Di, D.L.; Crews, P.; Wu, Q.X. The Bioactive Secondary Metabolites from Talaromyces species. Nat. Prod. Bioprospecting 2016, 6, 1–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heinis, C. Drug discovery: Tools and rules for macrocycles. Nat. Chem. Biol. 2014, 10, 696–698. [Google Scholar] [CrossRef] [PubMed]
- Driggers, E.M.; Hale, S.P.; Lee, J.; Terrett, N.K. The exploration of macrocycles for drug discovery--an underexploited structural class. Nat. Rev. Drug Discov. 2008, 7, 608–624. [Google Scholar] [CrossRef] [PubMed]
- Bunnak, W.; Wonnapinij, P.; Sriboonlert, A.; Lazarus, C.M.; Wattana-Amorn, P. Heterologous biosynthesis of a fungal macrocyclic polylactone requires only two iterative polyketide synthases. Org. Biomol. Chem. 2019, 17, 374–379. [Google Scholar] [CrossRef]
- Wattana-amorn, P.; Juthaphan, P.; Sirikamonsil, M.; Sriboonlert, A.; Simpson, T.J.; Kongkathip, N. Biosynthetic origins of menisporopsin A. J. Nat. Prod. 2013, 76, 1235–1237. [Google Scholar] [CrossRef]
- Davison, J.; al Fahad, A.; Cai, M.; Song, Z.; Yehia, S.Y.; Lazarus, C.M.; Bailey, A.M.; Simpson, T.J.; Cox, R.J. Genetic, molecular, and biochemical basis of fungal tropolone biosynthesis. Proc. Natl. Acad. Sci. USA 2012, 109, 7642–7647. [Google Scholar] [CrossRef] [Green Version]
- al Fahad, A.; Abood, A.; Simpson, T.J.; Cox, R.J. The biosynthesis and catabolism of the maleic anhydride moiety of stipitatonic acid. Angew. Chem. Int. Ed. Engl. 2014, 53, 7519–7523. [Google Scholar] [CrossRef]
- Zang, Y.; Genta-Jouve, G.; Sun, T.A.; Li, X.; Didier, B.; Mann, S.; Mouray, E.; Larsen, A.K.; Escargueil, A.E.; Nay, B.; et al. Unexpected talaroenamine derivatives and an undescribed polyester from the fungus Talaromyces stipitatus ATCC10500. Phytochemistry 2015, 119, 70–75. [Google Scholar] [CrossRef]
- Nierman, W.C.; Fedorova-Abrams, N.D.; Andrianopoulos, A. Genome Sequence of the AIDS-Associated Pathogen Penicillium marneffei (ATCC18224) and Its Near Taxonomic Relative Talaromyces stipitatus (ATCC10500). Genome Announc. 2015, 3, e01559-14. [Google Scholar] [CrossRef] [Green Version]
- Krohn, K.; Bahramsari, R.; Flörke, U.; Ludewig, K.; Kliche-Spory, C.; Michel, A.; Aust, H.J.; Draeger, S.; Schulz, B.; Antus, S. Dihydroisocoumarins from fungi: Isolation, structure elucidation, circular dichroism and biological activity. Phytochemistry 1997, 45, 313–320. [Google Scholar] [CrossRef]
- Abbanat, D.; Leighton, M.; Maiese, W.; Jones, E.B.; Pearce, C.; Greenstein, M. Cell wall active antifungal compounds produced by the marine fungus hypoxylon oceanicum LL-15G256. I. Taxonomy and fermentation. J. Antibiot. 1998, 51, 296–302. [Google Scholar] [CrossRef] [PubMed]
- al-Khalil, S.; Alkofahi, A.; el-Eisawi, D.; al-Shibib, A. Transtorine, a new quinoline alkaloid from Ephedra transitoria. J. Nat. Prod. 1998, 61, 262–263. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Letunic, I.; Khedkar, S.; Bork, P. SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res. 2021, 49, D458–D460. [Google Scholar] [CrossRef]
- Smith, S.; Tsai, S.C. The type I fatty acid and polyketide synthases: A tale of two megasynthases. Nat. Prod. Rep. 2007, 24, 1041–1072. [Google Scholar] [CrossRef] [Green Version]
- Khosla, C.; Tang, Y.; Chen, A.Y.; Schnarr, N.A.; Cane, D.E. Structure and mechanism of the 6-deoxyerythronolide B synthase. Annu. Rev. Biochem. 2007, 76, 195–221. [Google Scholar] [CrossRef]
- Kimber, M.S.; Martin, F.; Lu, Y.; Houston, S.; Vedadi, M.; Dharamsi, A.; Fiebig, K.M.; Schmid, M.; Rock, C.O. The structure of (3R)-hydroxyacyl-acyl carrier protein dehydratase (FabZ) from Pseudomonas aeruginosa. J. Biol. Chem. 2004, 279, 52593–52602. [Google Scholar] [CrossRef] [Green Version]
- Keatinge-Clay, A. Crystal structure of the erythromycin polyketide synthase dehydratase. J. Mol. Biol. 2008, 384, 941–953. [Google Scholar] [CrossRef] [Green Version]
- Fisch, K.M. Biosynthesis of natural products by microbial iterative hybrid PKS–NRPS. RSC Adv. 2013, 3, 18228–18247. [Google Scholar] [CrossRef] [Green Version]
- Mayorga, M.E.; Timberlake, W.E. The developmentally regulated Aspergillus nidulans wA gene encodes a polypeptide homologous to polyketide and fatty acid synthases. Mol. Gen. Genet. MGG 1992, 235, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Takano, Y.; Kubo, Y.; Shimizu, K.; Mise, K.; Okuno, T.; Furusawa, I. Structural analysis of PKS1, a polyketide synthase gene involved in melanin biosynthesis in Colletotrichum lagenarium. Mol. Gen. Genet. MGG 1995, 249, 162–167. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Leonard, T.J. Sterigmatocystin biosynthesis in Aspergillus nidulans requires a novel type I polyketide synthase. J. Bacteriol. 1995, 177, 4792–4800. [Google Scholar] [CrossRef] [Green Version]
- Horsman, M.E.; Hari, T.P.; Boddy, C.N. Polyketide synthase and non-ribosomal peptide synthetase thioesterase selectivity: Logic gate or a victim of fate? Nat. Prod. Rep. 2016, 33, 183–202. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhou, T.; Zhang, S.; Xuan, L.J.; Zhan, J.; Molnár, I. Thioesterase domains of fungal nonreducing polyketide synthases act as decision gates during combinatorial biosynthesis. J. Am. Chem. Soc. 2013, 135, 10783–10791. [Google Scholar] [CrossRef] [Green Version]
- Bunnak, W.; Winter, A.J.; Lazarus, C.M.; Crump, M.P.; Race, P.R.; Wattana-Amorn, P. SAXS reveals highly flexible interdomain linkers of tandem acyl carrier protein-thioesterase domains from a fungal nonreducing polyketide synthase. FEBS Lett. 2021, 595, 133–144. [Google Scholar] [CrossRef]
- Thongkawphueak, T.; Winter, A.J.; Williams, C.; Maple, H.J.; Soontaranon, S.; Kaewhan, C.; Campopiano, D.J.; Crump, M.P.; Wattana-Amorn, P. Solution Structure and Conformational Dynamics of a Doublet Acyl Carrier Protein from Prodigiosin Biosynthesis. Biochemistry 2021, 60, 219–230. [Google Scholar] [CrossRef]
- Cox, R.J.; Skellam, E.J. Fungal Non-Reducing Polyketide Synthases; Elsevier: Oxford, UK, 2020. [Google Scholar]
- He, J.W.; Gao, H.; Liu, X.Z.; Yao, X.S. Two new polyesters from wetland soil-derived fungus Talaromyces flavus. China J. Chin. Mater. Med. 2015, 40, 3347–3351. [Google Scholar]
- Schlingmann, G.; Milne, L.; Carter, G. Isolation and identification of antifungal polyesters from the marine fungus Hypoxylon oceanicum LL-15G256. Tetrahedron 2002, 58, 6825–6835. [Google Scholar] [CrossRef]
- Cervenka, I.; Agudelo, L.Z.; Ruas, J.L. Kynurenines: Tryptophan’s metabolites in exercise, inflammation, and mental health. Science 2017, 357, eaaf9794. [Google Scholar] [CrossRef] [Green Version]
- Badawy, A.A. Kynurenine Pathway of Tryptophan Metabolism: Regulatory and Functional Aspects. Int. J. Tryptophan Res. 2017, 10, 1–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaseda, H.; Noguchi, T.; Kido, R. Biosynthetic routes to 2-aminoacetophenone and 2-amino-3-hydroxyacetophenone. J. Biochem. 1973, 74, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Celenza, J.L. Metabolism of tyrosine and tryptophan-new genes for old pathways. Curr. Opin. Plant Biol. 2001, 4, 234–240. [Google Scholar] [CrossRef]
- Mikkelsen, M.D.; Hansen, C.H.; Wittstock, U.; Halkier, B.A. Cytochrome P450 CYP79B2 from Arabidopsis catalyzes the conversion of tryptophan to indole-3-acetaldoxime, a precursor of indole glucosinolates and indole-3-acetic acid. J. Biol. Chem. 2000, 275, 33712–33717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Platten, M.; Wick, W.; Van den Eynde, B.J. Tryptophan catabolism in cancer: Beyond IDO and tryptophan depletion. Cancer Res. 2012, 72, 5435–5440. [Google Scholar] [CrossRef] [Green Version]
- Vujkovic-Cvijin, I.; Dunham, R.M.; Iwai, S.; Maher, M.C.; Albright, R.G.; Broadhurst, M.J.; Hernandez, R.D.; Lederman, M.M.; Huang, Y.; Somsouk, M.; et al. Dysbiosis of the gut microbiota is associated with HIV disease progression and tryptophan catabolism. Sci. Transl. Med. 2013, 5, 193ra191. [Google Scholar] [CrossRef] [Green Version]
- Thomas, T.; Stefanoni, D.; Reisz, J.A.; Nemkov, T.; Bertolone, L.; Francis, R.O.; Hudson, K.E.; Zimring, J.C.; Hansen, K.C.; Hod, E.A.; et al. COVID-19 infection alters kynurenine and fatty acid metabolism, correlating with IL-6 levels and renal status. JCI Insight 2020, 5, e140327. [Google Scholar] [CrossRef]
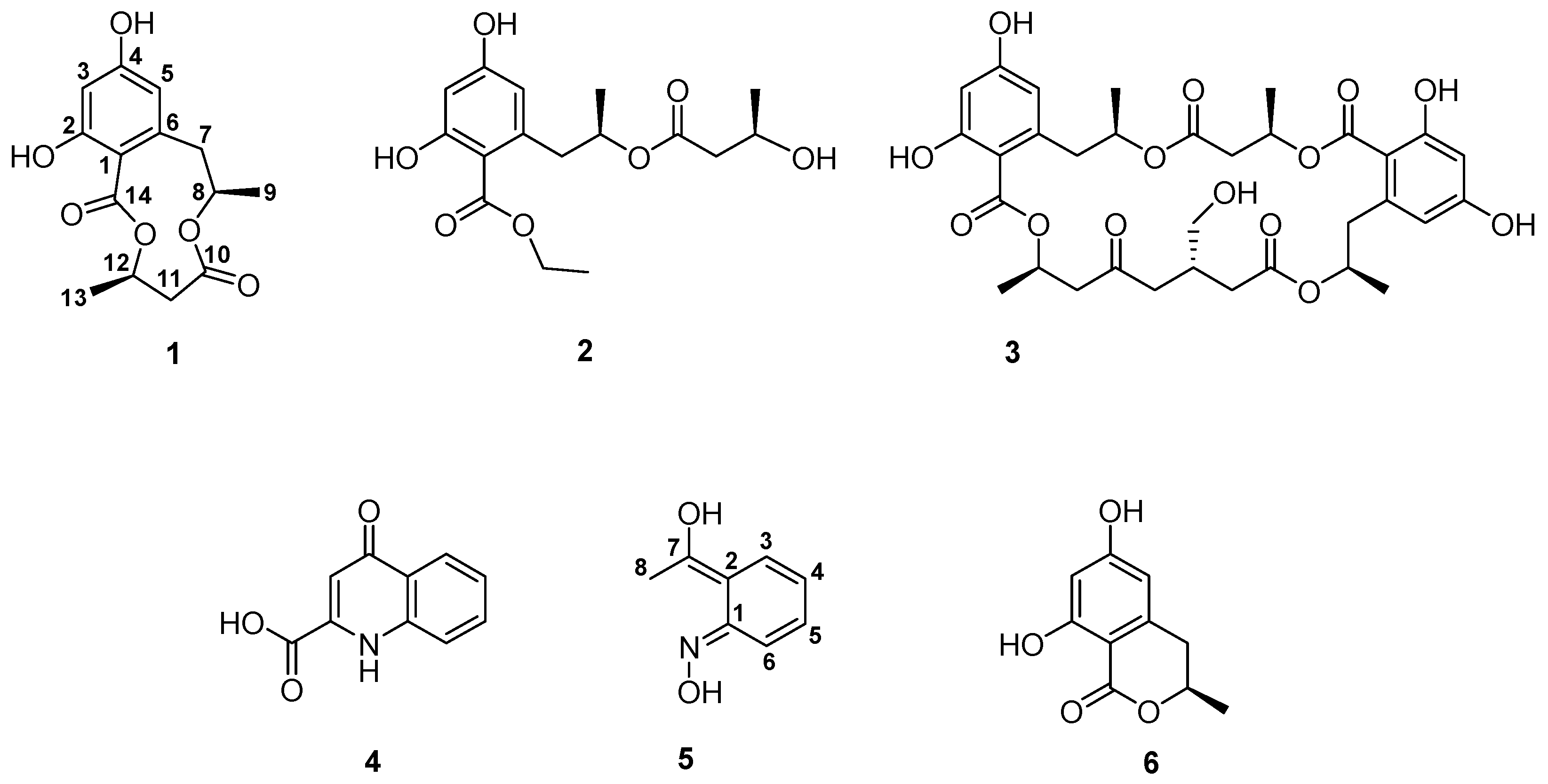
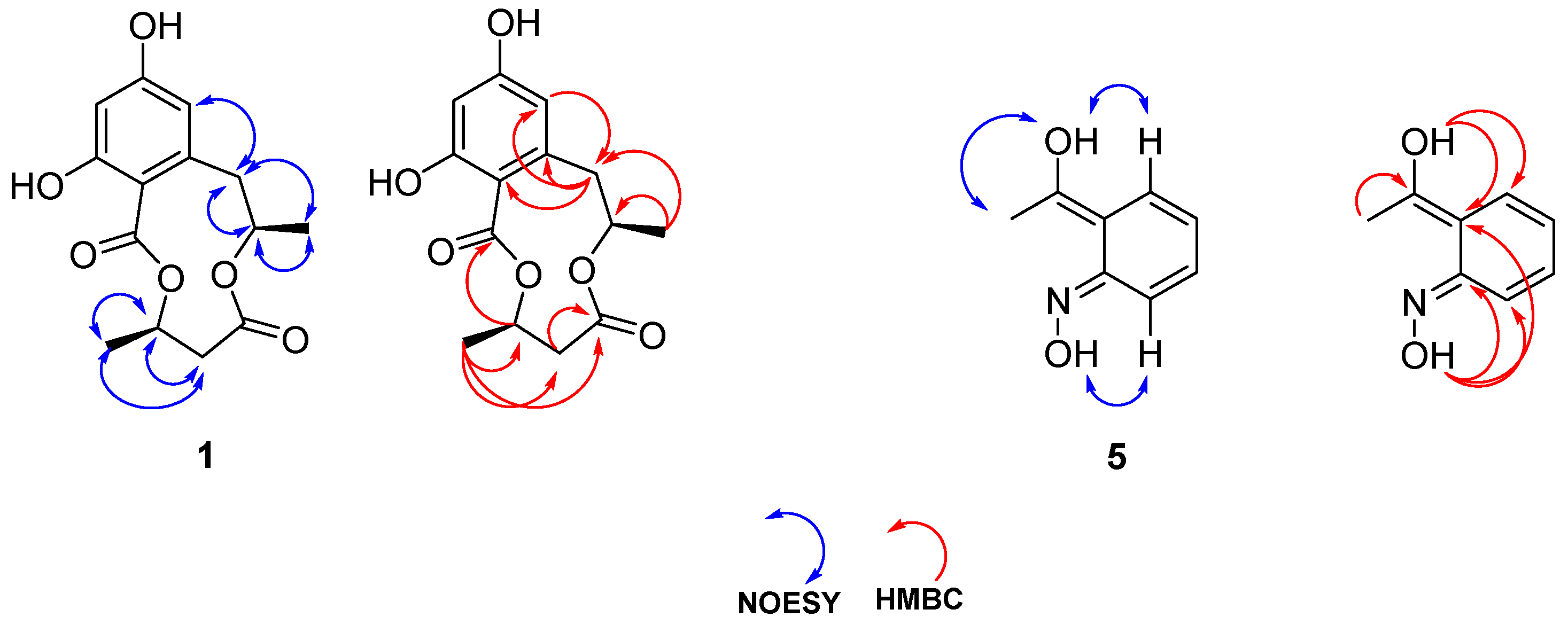

| Position | δc (ppm) | δH (ppm), (m, J Values in Hz, nH) |
|---|---|---|
| 1 | 110.6 | --- |
| 2 | 160.2 | |
| 3 | 102.5 | 6.37 (s, 1H) |
| 4 | 159.6 | --- |
| 5 | 113.0 | 6.19 (d, 2.2, 1H) |
| 6 | 139.3 | --- |
| 7 | 39.3 | 3.66 a (dd, 14.3, 3.8, 1H), 2.51–2.47 b (m, 2H) |
| 8 | 70.9 | 5.30 (dt, 6.9, 3.5, 1H) |
| 9 | 18.2 | 1.03 (d, 6.7, 3H) |
| 10 | 171.2 | --- |
| 11 | 41.3 | 2.86 a (t, 11.3, 1H), 2.51–2.47 b (m, 2H) |
| 12 | 74.5 | 5.20 (ddd, 10.8, 6.8, 3.8 Hz, 1H) |
| 13 | 21.0 | 1.68 (d, 6.6, 3H) |
| 14 | 170.7 | --- |
| Gene | Gene Location | Protein Accession No. | Size | Function | Conserved Domain |
|---|---|---|---|---|---|
| tpeA | TSTA_008130 | XP_002488696 | 2521 aa | HR-PKS | KS-MAT-DH-ER-KR-ACP |
| tpeB | TSTA_008140 | XP_002488697 | 2200 aa | NR-PKS | SAT-KS-MAT-PT-ACP1-ACP2-TE |
| tpeC | TSTA_008150 | XP_002488698 | 507 aa | Oxygenase | Cytochrome P450 (CYP) |
| tpeD | TSTA_008170 | XP_002488702 | 846 aa | Regulator | BED zinc finger (ZnF_BED) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
al Fahad, A.J. Putative Biosynthesis of Talarodioxadione & Talarooxime from Talaromyces stipitatus. Molecules 2022, 27, 4473. https://doi.org/10.3390/molecules27144473
al Fahad AJ. Putative Biosynthesis of Talarodioxadione & Talarooxime from Talaromyces stipitatus. Molecules. 2022; 27(14):4473. https://doi.org/10.3390/molecules27144473
Chicago/Turabian Styleal Fahad, Ahmed J. 2022. "Putative Biosynthesis of Talarodioxadione & Talarooxime from Talaromyces stipitatus" Molecules 27, no. 14: 4473. https://doi.org/10.3390/molecules27144473
APA Styleal Fahad, A. J. (2022). Putative Biosynthesis of Talarodioxadione & Talarooxime from Talaromyces stipitatus. Molecules, 27(14), 4473. https://doi.org/10.3390/molecules27144473





