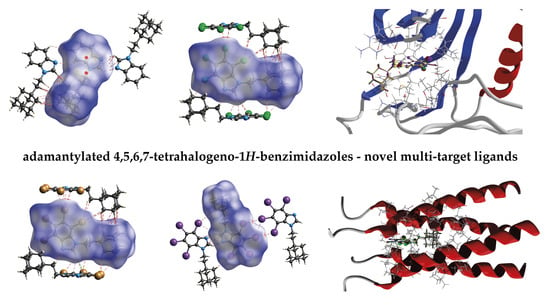
Synthesis and Crystal Structure of Adamantylated 4,5,6,7-Tetrahalogeno-1H-benzimidazoles Novel Multi-Target Ligands (Potential CK2, M2 and SARS-CoV-2 Inhibitors); X-ray/DFT/QTAIM/Hirshfeld Surfaces/Molecular Docking Study
Abstract
:1. Introduction
2. Results and Discussion
2.1. Crystal Packing Analysis
and 3.451 Å (Cl; tClAB) < 3.534 Å (Br; tBrAB)
2.2. Interactions Pattern in the Solid State
2.2.1. Distribution and Types of Interactions—3D HS/2D FP
2.2.2. Electrostatic Nature of Interactions—ESP
2.2.3. Characterization of the Strength of the Interactions—QTAIM
2.3. Chemical Reactivity
2.4. Molecular Docking
2.4.1. Human Casein Kinase 2 (CK2)
2.4.2. Membrane Matrix 2 Protein (M2)
tClAB > tBrAB > tIAB > AB (docking with M2 from 2MUV)
tClAB > tIAB > tBrAB > AB (docking with M2 from 2MUW)
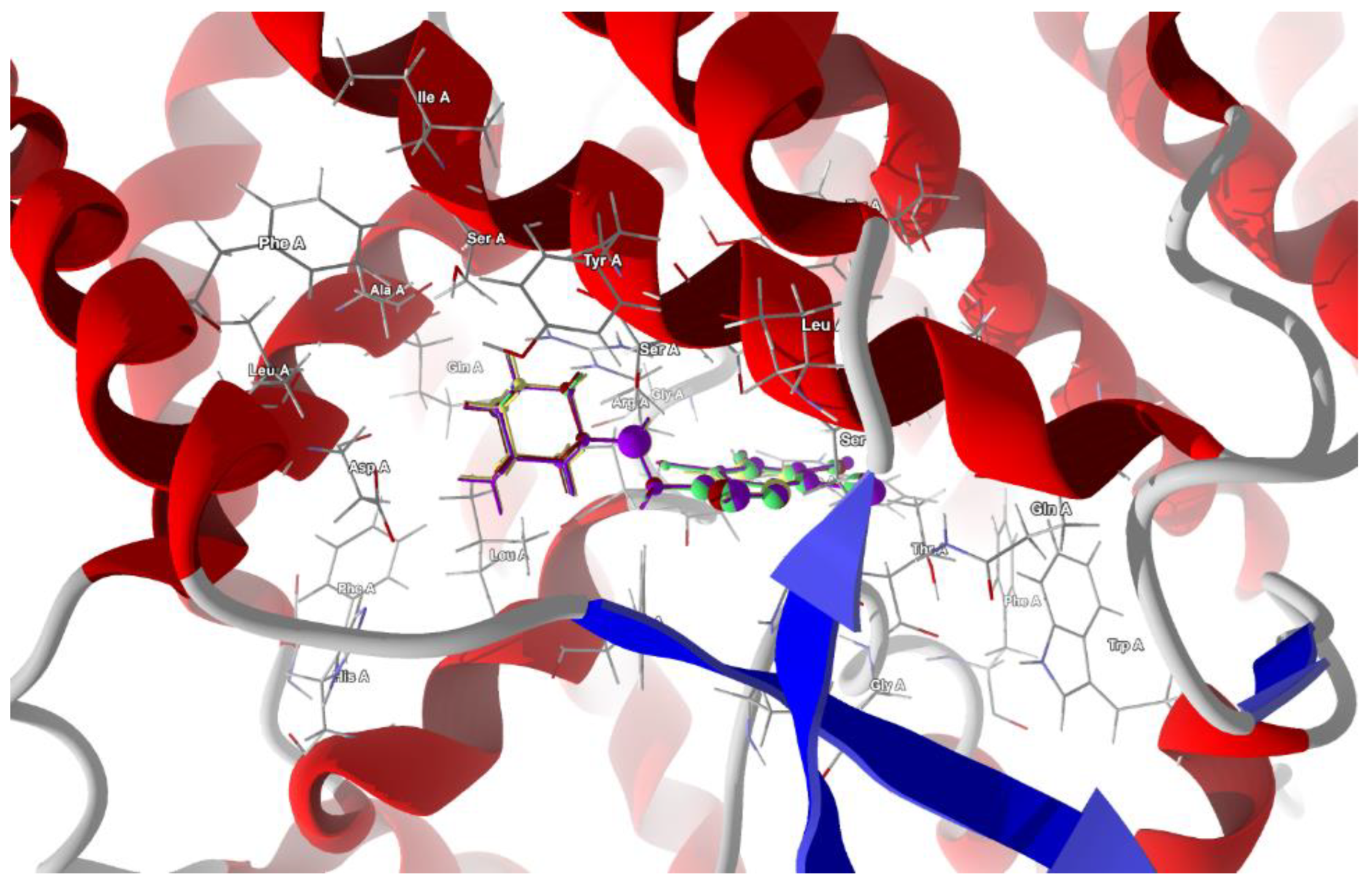
2.4.3. SARS-CoV-2 Spike Glycoprotein (S Protein)
3. Experimental Section
3.1. General Procedure for the Synthesis and Single Crystal Growth
3.2. X-ray Single Crystal Studies and Structure Solution
3.3. CLP/PIXEL
3.4. DFT/QTAIM/RDS/Hirshfeld Surfaces
3.5. Molecular Docking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Zimmermann, G.R.; Lehar, J.; Keith, C.T. Multi-target therapeutics: When the whole is greater than the sum of the parts. Drug Discov. Today 2007, 12, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Gianoncelli, A.; Cozza, G.; Orzeszko, A.; Meggio, F.; Kazimierczuk, Z.; Pinna, L.A. Tetraiodobenzimidazoles are potent inhibitors of protein kinase CK2. Bioorg. Med. Chem. 2009, 17, 7281–7289. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramezani, H.; Mansoori, G.A. Diamondoids as molecular building blocks for nanotechnology. In Molecular Building Blocks for Nanotechnology; Mansoori, G.A., George, T., Assoufid, L., Zhang, G., Eds.; Springer: New York, NY, USA, 2007; Volume 109, pp. 44–71. [Google Scholar] [CrossRef]
- Kryczka, T.; Kazimierczuk, Z.; Kozlowska, M.; Chrapusta, S.J.; Vilpo, L.; Vilpo, J.; Stachnik, K.; Janisz, M.; Grieb, P. Two novel nucleoside ester derivatives of chlorambucil as potential antileukemic prodrugs: A preliminary study. Anti-Cancer Drugs 2007, 18, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.A.; Andrzejewska, M.; Ruzzene, M.; Sarno, S.; Cesaro, L.; Bain, J.; Elliott, M.; Meggio, F.; Kazimierczuk, Z.; Pinna, L.A. Optimization of protein kinase CK2 inhibitors derived from 4,5,6,7-tetrabromobenzimidazole. J. Med. Chem. 2004, 47, 6239–6247. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.A.; Bain, J.; Kazimierczuk, Z.; Sarno, S.; Ruzzene, M.; Di Maira, G.; Elliott, M.; Orzeszko, A.; Cozza, G.; Meggio, F.; et al. The selectivity of inhibitors of protein kinase CK2: An update. Biochem. J. 2008, 415, 353–365. [Google Scholar] [CrossRef] [Green Version]
- Pagano, M.A.; Meggio, F.; Ruzzene, M.; Andrzejewska, M.; Kazimierczuk, Z.; Pinna, L.A. 2-Dimethylamino-4,5,6,7-tetrabromo-1H-benzimidazole: A novel powerful and selective inhibitor of protein kinase CK2. Biochem. Biophys. Res. Commun. 2004, 321, 1040–1044. [Google Scholar] [CrossRef]
- Starcevic, K.; Kralj, M.; Ester, K.; Sabol, I.; Grce, M.; Pavelic, K.; Karminski-Zamola, G. Synthesis, antiviral and antitumor activity of 2-substituted-5-amidino-benzimidazoles. Bioorg. Med. Chem. 2007, 15, 4419–4426. [Google Scholar] [CrossRef] [Green Version]
- Andries, K.; Moeremans, M.; Gevers, T.; Willebrords, R.; Sommen, C.; Lacrampe, J.; Janssens, F.; Wyde, P.R. Substituted benzimidazoles with nanomolar activity against respiratory syncytial virus. Antivir. Res. 2003, 60, 209–219. [Google Scholar] [CrossRef]
- Palma, A.M.D.; Heggermont, W.; Leyssen, P.; Pürstinger, G.; Wimmer, E.; Clercq, E.D.; Rao, A.; Monforte, A.-M.; Chimirri, A.; Neyts, J. Anti-enterovirus activity and structure–activity relationship of a series of 2,6-dihalophenyl-substituted 1H,3H-thiazolo [3,4-a]benzimidazoles. Biochem. Biophys. Res. Commun. 2007, 353, 628–632. [Google Scholar] [CrossRef]
- Li, Y.; Kataoka, M.; Tatsuta, M.; Yasoshima, K.; Yura, T.; Urbahns, K.; Kiba, A.; Yamamoto, N.; Gupta, J.B.; Hashimoto, K. Benzimidazole derivatives as novel nonpeptide luteinizing hormone-releasing hormone (LHRH) antagonists. Part 2: Benzimidazole-5-sulfonamides. Bioorg. Med. Chem. Lett. 2005, 15, 805–807. [Google Scholar] [CrossRef]
- Li, G.R.; Liu, J.; Pan, Q.; Song, Z.B.; Luo, F.L.; Wang, S.R.; Zhang, X.L.; Zhou, X. Synthesis and anti-HIV activity of [ddN]-[ddN] dimers and benzimidazole nucleoside dimers. Chem. Biodivers. 2009, 6, 2200–2208. [Google Scholar] [CrossRef] [PubMed]
- Kazimierczuk, Z.; Upcroft, J.A.; Upcroft, P.; Gorska, A.; Starosciak, B.; Laudy, A. Synthesis, antiprotozoal and antibacterial activity of nitro- and halogeno-substituted benzimidazole derivatives. Acta Biochim. Pol. 2002, 49, 185–195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, S.; Kim, J.S.; Sim, S.P.; Liu, A.; Pilch, D.S.; Liu, L.F.; LaVoie, E.J. Heterocyclic bibenzimidazole derivatives as topoisomerase I inhibitors. Bioorg. Med. Chem. Lett. 2000, 10, 719–723. [Google Scholar] [CrossRef]
- Andrzejewska, M.; Yepez-Mulia, L.; Tapia, A.; Cedillo-Rivera, R.; Laudy, A.E.; Starosciak, B.J.; Kazimierczuk, Z. Synthesis, and antiprotozoal and antibacterial activities of S-substituted 4,6-dibromo- and 4,6-dichloro-2-mercaptobenzimidazoles. Eur. J. Pharm. Sci. 2004, 21, 323–329. [Google Scholar] [CrossRef]
- Klimesova, V.; Koci, J.; Waisser, K.; Kaustova, J. New benzimidazole derivatives as antimycobacterial agents. Farmaco 2002, 57, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Serafin, B.; Borkowska, G.; Glowczyk, J.; Kowalska, I.; Rump, S. Potential antihypertensive benzimidazole derivatives. Pol. J. Prarmacol. Pharm. 1989, 41, 89–96. [Google Scholar]
- Tsukamoto, G.; Yoshino, K.; Kohno, T.; Ohtaka, H.; Kagaya, H.; Ito, K. Synthesis and antiinflammatory activity of some 2-(substituted-pyridinyl)benzimidazoles. J. Med. Chem. 1980, 23, 734–738. [Google Scholar] [CrossRef]
- Sondhi, S.M.; Rajvanshi, S.; Johar, M.; Bharti, N.; Azam, A.; Singh, A.K. Anti-inflammatory, analgesic and antiamoebic activity evaluation of pyrimido[1,6-a]benzimidazole derivatives synthesized by the reaction of ketoisothiocyanates with mono and diamines. Eur. J. Med. Chem. 2002, 37, 835–843. [Google Scholar] [CrossRef]
- Maryanoff, B.E.; Ho, W.; McComsey, D.F.; Reitz, A.B.; Grous, P.P.; Nortey, S.O.; Shank, R.P.; Dubinsky, B.; Taylor, R.J., Jr.; Gardocki, J.F. Potential Anxiolytic Agents. Pyrido[1,2-a]benzimidazoles: A New Structural Class of Ligands for the Benzodiazepine Binding Site on GABA-A Receptors. J. Med. Chem. 1995, 38, 16–20. [Google Scholar] [CrossRef]
- Mederski, W.W.K.R.; Dorsch, D.; Anzali, S.; Gleitz, J.; Cezanne, B.; Tsaklakidis, C. Halothiophene benzimidazoles as P1 surrogates of inhibitors of blood coagulation factor Xa. Bioorg. Med. Chem. Lett. 2004, 14, 3763–3769. [Google Scholar] [CrossRef]
- Beaulieu, C.; Wang, Z.; Denis, D.; Greig, G.; Lamontagne, S.; O’Neill, G.; Slipetz, D.; Wang, J. Benzimidazoles as new potent and selective DP antagonists for the treatment of allergic rhinitis. Bioorg. Med. Chem. Lett. 2004, 14, 3195–3199. [Google Scholar] [CrossRef]
- Li, Y.F.; Wang, G.F.; Luo, Y.; Huang, W.G.; Tang, W.; Feng, C.L.; Shi, L.P.; Ren, Y.D.; Zuo, J.P.; Lu, W. Identification of 1-isopropylsulfonyl-2-amine benzimidazoles as a new class of inhibitors of hepatitis B virus. Eur. J. Med. Chem. 2007, 42, 1358–1364. [Google Scholar] [CrossRef] [PubMed]
- Ayhan-Kilcigil, G.; Kus, C.; Coban, T.; Can-Eke, B.; Iscan, M. Synthesis and antioxidant properties of novel benzimidazole derivatives. J. Enzyme Inhib. Med. Chem. 2004, 19, 129–135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bathini, P.; Kameshwari, L.; Vijaya, N. Antidiabetic effect of 2 nitro-benzimidazole in alloxan induced diabetic rats. Int. J. Basic Clin. Pharmacol. 2013, 2, 814–818. [Google Scholar] [CrossRef]
- Schnell, J.R.; Chou, J.J. Structure and mechanism of the M2 proton channel of influenza A virus. Nature 2008, 451, 591–595. [Google Scholar] [CrossRef] [Green Version]
- Alves Galvao, M.G.; Rocha Crispino Santos, M.A.; Alves da Cunha, A.J. Amantadine and rimantadine for influenza A in children and the elderly. Cochrane Database Syst. Rev. 2012, 1, Cd002745. [Google Scholar] [CrossRef]
- Stylianakis, I.; Kolocouris, A.; Kolocouris, N.; Fytas, G.; Foscolos, G.B.; Padalko, E.; Neyts, J.; De Clercq, E. Spiro[pyrrolidine-2,2′-adamantanes]: Synthesis, anti-influenza virus activity and conformational properties. Bioorg. Med. Chem. Lett. 2003, 13, 1699–1703. [Google Scholar] [CrossRef]
- Stamatiou, G.; Foscolos, G.B.; Fytas, G.; Kolocouris, A.; Kolocouris, N.; Pannecouque, C.; Witvrouw, M.; Padalko, E.; Neyts, J.; De Clercq, E. Heterocyclic rimantadine analogues with antiviral activity. Bioorg. Med. Chem. 2003, 11, 5485–5492. [Google Scholar] [CrossRef]
- Fytas, C.; Kolocouris, A.; Fytas, G.; Grigoris, Z.; Valmas, C.; Basler, C.F. Influence of an Additional Amino Group on the Potency of Aminoadamantanes Against Influenza Virus A. II—Synthesis of Spiropiperazines and In-Vitro Activity Against Influenza A H3N2 Virus. Bioorg. Chem. 2010, 38, 247–251. [Google Scholar] [CrossRef] [Green Version]
- Sheu, T.G.; Fry, A.M.; Garten, R.J.; Deyde, V.M.; Shwe, T.; Bullion, L.; Peebles, P.J.; Li, Y.; Klimov, A.I.; Gubareva, L.V. Dual resistance to adamantanes and oseltamivir among seasonal influenza A(H1N1) viruses: 2008–2010. J. Infects. Dis. 2011, 203, 13–17. [Google Scholar] [CrossRef]
- Garcia, V.; Aris-Brosou, S. Comparative dynamics and distribution of influenza drug resistance acquisition to protein M2 and neuraminidase inhibitors. Mol. Biol. Evol. 2014, 31, 355–363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lepiller, Q.; Soulier, E.; Li, Q.; Lambotin, M.; Barths, J.; Fuchs, D.; Stoll-Keller, F.; Liang, T.J.; Barth, H. Antiviral and Immunoregulatory Effects of Indoleamine-2,3-Dioxygenase in Hepatitis C Virus Infection. J. Innate Immun. 2015, 7, 530–544. [Google Scholar] [CrossRef] [PubMed]
- Duleu, S.; Mangas, A.; Poulletier de Gannes, F.; Tranchant, M.C.; Geffard, M. Circulating antibodies to conjugated tryptophan derivatives of the IDO pathway in amyotrophic lateral sclerosis, Alzheimer’s, Parkinson’s and multiple sclerosis patients. Immuno-Anal. Biol. Spéc. 2008, 23, 27–34. [Google Scholar] [CrossRef]
- Grelak, R.P.; Clark, R.; Stump, J.M.; Vernier, V.G. Amantadine-dopamine interaction: Possible mode of action in Parkinsonism. Science 1970, 169, 203–204. [Google Scholar] [CrossRef]
- Huber, T.J.; Dietrich, D.E.; Emrich, H.M. Possible use of amantadine in depression. Pharmacopsychiatry 1999, 32, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Takikawa, O.; Truscott, R.J.; Fukao, M.; Miwa, S. Age-related nuclear cataract and indoleamine 2,3-dioxygenase-initiated tryptophan metabolism in the human lens. Adv. Exp. Med. Biol. 2003, 527, 277–285. [Google Scholar] [CrossRef]
- Uyttenhove, C.; Pilotte, L.; Theate, I.; Stroobant, V.; Colau, D.; Parmentier, N.; Boon, T.; Van den Eynde, B.J. Evidence for a tumoral immune resistance mechanism based on tryptophan degradation by indoleamine 2,3-dioxygenase. Nat. Med. 2003, 9, 1269–1274. [Google Scholar] [CrossRef]
- Inaba, T.; Ino, K.; Kajiyama, H.; Yamamoto, E.; Shibata, K.; Nawa, A.; Nagasaka, T.; Akimoto, H.; Takikawa, O.; Kikkawa, F. Role of the immunosuppressive enzyme indoleamine 2,3-dioxygenase in the progression of ovarian carcinoma. Gynecol. Oncol. 2009, 115, 185–192. [Google Scholar] [CrossRef]
- Dawson, M.I.; Fontana, J.A. The peptidomimetic, 1-adamantyl-substituted, and flex-het classes of retinoid-derived molecules: Structure-activity relationships and retinoid receptor-independent anticancer activities. Mini-Rev. Med. Chem. 2010, 10, 455–491. [Google Scholar] [CrossRef]
- Maurin, J.K.; Lasek, W.; Górska, A.; Świtaj, T.; Jakubowska, A.B.; Kazimierczuk, Z. Adamantylsulfanyl- and N-Adamantylcarboxamido-Derivatives of Heterocycles and Phenoles: Synthesis, Crystal Structure, Tumor Necrosis Factor-α Production-Enhancing Properties, and Theoretical Considerations. Chem. Biodivers. 2004, 1, 1498–1512. [Google Scholar] [CrossRef]
- Schmidt, S.V.; Schultze, J.L. New Insights into IDO Biology in Bacterial and Viral Infections. Front. Immunol. 2014, 5, 384. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, J.; Obando, D.; Liao, V.; Lifa, T.; Codd, R. The many faces of the adamantyl group in drug design. Eur. J. Med. Chem. 2011, 46, 1949–1963. [Google Scholar] [CrossRef] [PubMed]
- Kreutzberger, A.; Schröders, H.-H. ur synthese und antiviruswirkung von adamantyl-(1)- thioharnstoffen. Tetrahedron Lett. 1969, 10, 5101–5104. [Google Scholar] [CrossRef]
- Fort, R.C.; Schleyer, P.v.R. Adamantane: Consequences of the Diamondoid Structure. Chem. Rev. 1964, 64, 277–300. [Google Scholar] [CrossRef]
- Boasso, A.; Shearer, G.M. How does indoleamine 2,3-dioxygenase contribute to HIV-mediated immune dysregulation. Curr. Drug Metab. 2007, 8, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Orzeszko, B.; Świtaj, T.; Jakubowska-Mućka, A.B.; Lasek, W.; Orzeszko, A.; Kazimierczuk, Z. Tumor necrosis factor-α production-enhancing properties of novel adamantylalkylthio derivatives of some heterocyclic compounds. Z. Naturforsch. B 2005, 60, 471–475. [Google Scholar] [CrossRef]
- Van Derpoorten, K.; Balzarini, J.; De Clercq, E.; Poupaert, J.H. Anti-HIV activity of N-1-adamantyl-4-aminophthalimide. Biomed. Pharmacother. 1997, 51, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Kasloff, S.B.; Pizzuto, M.S.; Silic-Benussi, M.; Pavone, S.; Ciminale, V.; Capua, I. Oncolytic activity of avian influenza virus in human pancreatic ductal adenocarcinoma cell lines. J. Virol. 2014, 88, 9321–9334. [Google Scholar] [CrossRef] [Green Version]
- Fink, K.; Nitsche, A.; Neumann, M.; Grossegesse, M.; Eisele, K.H.; Danysz, W. Amantadine Inhibits SARS-CoV-2 In Vitro. Viruses 2021, 13, 539. [Google Scholar] [CrossRef]
- Oh, C.K.; Nakamura, T.; Beutler, N.; Zhang, X.; Piña-Crespo, J.; Talantova, M.; Ghatak, S.; Trudler, D.; Carnevale, L.N.; McKercher, S.R.; et al. Targeted protein S-nitrosylation of ACE2 inhibits SARS-CoV-2 infection. Nat. Chem. Biol. 2022. [Google Scholar] [CrossRef]
- Van Rikxoort, M.; Michaelis, M.; Wolschek, M.; Muster, T.; Egorov, A.; Seipelt, J.; Doerr, H.W.; Cinatl, J., Jr. Oncolytic effects of a novel influenza A virus expressing interleukin-15 from the NS reading frame. PLoS ONE 2012, 7, e36506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bergmann, M.; Romirer, I.; Sachet, M.; Fleischhacker, R.; Garcia-Sastre, A.; Palese, P.; Wolff, K.; Pehamberger, H.; Jakesz, R.; Muster, T. A genetically engineered influenza A virus with ras-dependent oncolytic properties. Cancer Res. 2001, 61, 8188–8193. [Google Scholar] [PubMed]
- Balzarini, J.; Orzeszko, B.; Maurin, J.K.; Orzeszko, A. Synthesis and anti-HIV studies of 2-adamantyl-substituted thiazolidin-4-ones. Eur. J. Med. Chem. 2007, 42, 993–1003. [Google Scholar] [CrossRef] [PubMed]
- Bader, R. Atoms in Molecules: A Quantum Theory; International Series of Monographs on Chemistry; Oxford University Press: New York, NY, USA, 1994. [Google Scholar]
- Johnson, E.R.; Keinan, S.; Mori-Sánchez, P.; Contreras-García, J.; Cohen, A.J.; Yang, W. Revealing Noncovalent Interactions. J. Am. Chem. Soc. 2010, 132, 6498–6506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Spackman, M.A.; Jayatilaka, D. Hirshfeld surface analysis. CrystEngComm 2009, 11, 19. [Google Scholar] [CrossRef]
- Spackman, M.A.; McKinnon, J.J. Fingerprinting intermolecular interactions in molecular crystals. CrystEngComm 2002, 4, 378. [Google Scholar] [CrossRef]
- Murray, J.S.; Politzer, P. The electrostatic potential: An overview. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2011, 1, 153–163. [Google Scholar] [CrossRef]
- Gavezzotti, A. Efficient computer modeling of organic materials. The atom-atom, Coulomb-London-Pauli (AA-CLP) model for intermolecular electrostatic-polarization, dispersion and repulsion energies. New J. Chem. 2011, 35, 1360–1368. [Google Scholar] [CrossRef]
- Gavezzotti, A. Non-conventional bonding between organic molecules. The ‘halogen bond’ in crystalline systems. Mol. Phys. 2008, 106, 1473–1485. [Google Scholar] [CrossRef] [Green Version]
- Latosińska, J.N.; Latosińska, M.; Seliger, J.; Žagar, V.; Kazimierczuk, Z. An Insight into Prototropism and Supramolecular Motifs in Solid-State Structure of Allopurinol, Hypoxanthine, Xanthine and Uric Acid. A 1H-14N NQDR Spectroscopy and Hybrid DFT/QTAIM and Hirshfeld Surface-Based Study. J. Phys. Chem. 2014, 118, 10837–10853. [Google Scholar] [CrossRef]
- Latosińska, J.N.; Latosińska, M.; Seliger, J.; Žagar, V.; Maurin, J.K.; Orzeszko, A.; Kazimierczuk, Z. Structural Study of Selected Polyhalogenated Benzimidazoles (Protein Kinase CK2 Inhibitors) by Nuclear Quadrupole Double Resonance, X-ray, and Density Functional Theory. J. Phys. Chem. A 2010, 114, 563–575. [Google Scholar] [CrossRef]
- Latosińska, J.N.; Latosińska, M.; Olejniczak, G.A.; Seliger, J.; Žagar, V. Topology of the Interactions Pattern in Pharmaceutically Relevant Polymorphs of Methylxanthines (Caffeine, Theobromine, and Theophiline): Combined Experimental (1H–14N Nuclear Quadrupole Double Resonance) and Computational (DFT and Hirshfeld-Based) Study. J. Chem. Inf. Model. 2014, 54, 2570–2584. [Google Scholar] [CrossRef]
- Latosińska, J.N.; Latosińska, M.; Szafrański, M.; Seliger, J.; Žagar, V. Polymorphism and Thermal Stability of Natural Active Ingredients. 3,3′-Diindolylmethane (Chemopreventive and Chemotherapeutic) Studied by a Combined X-ray, 1H–14N NMR-NQR, Differential Scanning Calorimetry, and Solid-State DFT/3D HS/QTAIM/RDS Computational Approach. Cryst. Growth Des. 2016, 16, 4336–4348. [Google Scholar] [CrossRef]
- Latosińska, J.N.; Latosińska, M.; Maurin, J.K.; Orzeszko, A.; Kazimierczuk, Z. Quantum-Chemical Insight into Structure–Reactivity Relationship in 4,5,6,7-Tetrahalogeno-1H-benzimidazoles: A Combined X-ray, DSC, DFT/QTAIM, Hirshfeld Surface-Based, and Molecular Docking Approach. J. Phys. Chem. A 2014, 118, 2089–2106. [Google Scholar] [CrossRef]
- Sarno, S.; Papinutto, E.; Franchin, C.; Bain, J.; Elliott, M.; Meggio, F.; Kazimierczuk, Z.; Orzeszko, A.; Zanotti, G.; Battistutta, R.; et al. ATP site-directed inhibitors of protein kinase CK2: An update. Curr. Top. Med. Chem. 2011, 11, 1340–1351. [Google Scholar] [CrossRef]
- Wang, J.; Wu, Y.; Ma, C.; Fiorin, G.; Wang, J.; Pinto, L.H.; Lamb, R.A.; Klein, M.L.; Degrado, W.F. Structure and inhibition of the drug-resistant S31N mutant of the M2 ion channel of influenza A virus. Proc. Natl. Acad. Sci. USA 2013, 110, 1315–1320. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Canturk, B.; Jo, H.; Ma, C.; Gianti, E.; Klein, M.L.; Pinto, L.H.; Lamb, R.A.; Fiorin, G.; Wang, J.; et al. Flipping in the Pore: Discovery of Dual Inhibitors That Bind in Different Orientations to the Wild-Type versus the Amantadine-Resistant S31N Mutant of the Influenza A Virus M2 Proton Channel. J. Am. Chem. Soc. 2014, 136, 17987–17995. [Google Scholar] [CrossRef] [Green Version]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [Green Version]
- Buchanan, C.J.; Gaunt, B.; Harrison, P.J.; Yang, Y.; Liu, J.; Khan, A.; Giltrap, A.M.; Le Bas, A.; Ward, P.N.; Gupta, K.; et al. Pathogen-sugar interactions revealed by universal saturation transfer analysis. Science 2022, 377, eabm3125. [Google Scholar] [CrossRef]
- Sheldrick, G. A short history of SHELX. Acta Cryst. Sect. A 2008, 64, 112–122. [Google Scholar] [CrossRef]
- Kohn, W.; Sham, L.J. Self-Consistent Equations Including Exchange and Correlation Effects. Phys. Rev. 1965, 140, A1133–A1138. [Google Scholar] [CrossRef] [Green Version]
- Levy, M. Universal variational functionals of electron densities, first-order density matrices, and natural spin-orbitals and solution of the v-representability problem. Proc. Natl. Acad. Sci. USA 1979, 76, 6062–6065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Becke, A.D. A new mixing of Hartree–Fock and local density-functional theories. J. Chem. Phys. 1993, 98, 1372. [Google Scholar] [CrossRef]
- Lee, C.; Yang, W.; Parr, R.G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 1988, 37, 785–789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimme, S. Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J. Comp. Chem. 2006, 27, 1787–1799. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. Gaussian 09, revision D.01; Gaussian, Inc.: Wallingford, CT, USA, 2009. [Google Scholar]
- Espinosa, E.; Souhassou, M.; Lachekar, H.; Lecomte, C. Topological analysis of the electron density in hydrogen bonds. Acta Crystallogr. B Struct. Sci. 1999, 55, 563–572. [Google Scholar] [CrossRef]
- Abramov, Y. On the Possibility of Kinetic Energy Density Evaluation from the Experimental Electron-Density Distribution. Acta Crystallogr. Sect. A 1997, 53, 264–272. [Google Scholar] [CrossRef]
- Espinosa, E.; Molins, E.; Lecomte, C. Hydrogen bond strengths revealed by topological analyses of experimentally observed electron densities. Chem. Phys. Lett. 1998, 285, 170–173. [Google Scholar] [CrossRef]
- Mata, I.; Alkorta, I.; Espinosa, E.; Molins, E. Relationships between interaction energy, intermolecular distance and electron density properties in hydrogen bonded complexes under external electric fields. Chem. Phys. Lett. 2011, 507, 185–189. [Google Scholar] [CrossRef]
- Emamian, S.; Lu, T.; Kruse, H.; Emamian, H. Exploring Nature and Predicting Strength of Hydrogen Bonds: A Correlation Analysis Between Atoms-in-Molecules Descriptors, Binding Energies, and Energy Components of Symmetry-Adapted Perturbation Theory. J. Comp. Chem. 2019, 40, 2868–2881. [Google Scholar] [CrossRef]
- Afonin, A.V.; Vashchenko, A.V.; Sigalov, M.V. Estimating the energy of intramolecular hydrogen bonds from 1H NMR and QTAIM calculations. Org. Biomol. Chem. 2016, 14, 11199. [Google Scholar] [CrossRef] [PubMed]
- Vener, M.V.; Egorova, A.N.; Churakov, A.V.; Tsirelson, V.G. Intermolecular hydrogen bond energies in crystals evaluated using electron density properties: DFT computations with periodic boundary conditions. J. Comput. Chem. 2012, 33, 2303–2309. [Google Scholar] [CrossRef] [PubMed]
- Jabłoński, M. A Critical Overview of Current Theoretical Methods of Estimating the Energy of Intramolecular Interactions. Molecules 2020, 25, 5512. [Google Scholar] [CrossRef] [PubMed]
- Cioslowski, J.; Mixon, S.T. Topological properties of electron density in search of steric interactions in molecules: Electronic structure calculations on ortho-substituted biphenyls. J. Am. Chem. Soc. 1992, 114, 4382–4387. [Google Scholar] [CrossRef]
- Jabłoński, M. Bond paths between distant atoms do not necessarily indicate dominant interactions. J. Comput. Chem. 2018, 39, 2183–2195. [Google Scholar] [CrossRef]
- Jabłoński, M. On the Uselessness of Bond Paths Linking Distant Atoms and on the Violation of the Concept of Privileged Exchange Channels. ChemistryOpen 2019, 8, 497–507. [Google Scholar] [CrossRef] [Green Version]
- Shahbazian, S. Why Bond Critical Points Are Not “Bond” Critical Points. Eur. J. Chem. 2017, 24, 5401–5405. [Google Scholar] [CrossRef]
- Kuznetsov, M.L. Relationships between Interaction Energy and Electron Density Properties for Homo Halogen Bonds of the [(A)nY–X⋯X–Z(B)m] Type (X = Cl, Br, I). Molecules 2019, 24, 2733. [Google Scholar] [CrossRef] [Green Version]
- Jelsch, C.; Ejsmont, K.; Huder, L. The enrichment ratio of atomic contacts in crystals, an indicator derived from the Hirshfeld surfaces analysis. IUCrJ 2014, 1, 119–128. [Google Scholar] [CrossRef]
- Politzer, P.; Murray, J.S.; Peralta-Inga, Z. Molecular surface electrostatic potentials in relation to noncovalent interactions in biological systems. Int. J. Quant. Chem. 2001, 85, 676–684. [Google Scholar] [CrossRef]
- Politzer, P.; Murray, J.S. Quantitative Analyses of Molecular Surface Electrostatic Potentials in Relation to Hydrogen Bonding and Co-Crystallization. Cryst. Growth Des. 2015, 15, 3767–3774. [Google Scholar] [CrossRef]
- Politzer, P.; Murray, J.S.; Clark, T. Mathematical modeling and physical reality in noncovalent interactions. J. Mol. Model. 2015, 21, 52. [Google Scholar] [CrossRef] [PubMed]
- Parr, R.G.; Pearson, R.G. Absolute hardness: Companion parameter to absolute electronegativity. J. Am. Chem. Soc. 1983, 105, 7512–7516. [Google Scholar] [CrossRef]
- Brooijmans, N.; Kuntz, I.D. Molecular Recognition and Docking Algorithms. Annu. Rev. Biophys. Biomol. Struct. 2003, 32, 332–373. [Google Scholar] [CrossRef]
- Gázquez, J.L.; Cedillo, A.; Vela, A. Electrodonating and Electroaccepting Powers. J. Phys. Chem. A 2007, 111, 1966–1970. [Google Scholar] [CrossRef]
- Vavra, O.; Filipovic, J.; Plhak, J.; Bednar, D.; Marques, S.M.; Brezovsky, J.; Stourac, J.; Matyska, L.; Damborsky, J. CaverDock: A Molecular Docking-Based Tool to Analyse Ligand Transport through Protein Tunnels and Channels. Bioinformatics 2019, 35, 4986–4993. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. Software news and update AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comp. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [Green Version]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A.F.; Forli, S. AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings. J. Am. Chem. Model. 2021, 61, 3891–3898. [Google Scholar] [CrossRef]
- BIOVIA; Dassault Systèmes. Discovery Studio Visualizer, version v21.1.0.20298; Dassault Systèmes: San Diego, CA, USA, 2021. [Google Scholar]












| AB | tClAB | tBrAB | tIAB | |
|---|---|---|---|---|
| No of deposit | CCDC 1478829 | CCDC 1478831 | CCDC 1478832 | CCDC 1478834 |
| Empirical formula | C19H25N2 | C19H20Cl4N2 | C19H20Br4N2 | C19H20I4N2 |
| Formula weight | 281.41 | 418.17 | 596.01 | 783.97 |
| Temperature | 293(2) K | 293(2) K | 293(2) K | 293(2) K |
| Wavelength | 1.54184 Å | 1.54184 Å | 1.54184 Å | 1.54184 Å |
| Crystal system | Triclinic | Monoclinic | Monoclinic | Triclinic |
| Space group | P-1 | P 1 21/c 1 | P 1 21/c 1 | P-1 |
| Unit cell dimensions | a = 6.9937(3) Å, γ = 70.000(3)° | a = 14.9004(4) Å | a = 15.1138(6) Å | a = 6.5333(2) Å, γ = 98.886(2)° |
| b = 9.9348(4) Å, γ = 85.448(3)° | b = 12.0172(3) Å, γ = 98.071(3)° | b = 12.3261(7) Å, γ = 96.753(5)° | b = 10.9704(3) Å, γ = 98.514(2)° | |
| c = 12.0737(4) Å, γ = 85.551(3)° | c = 10.6991(3) Å | c = 10.7864(6) Å | c = 15.3814(3) Å, γ = 94.170(2)° | |
| Volume | 784.67(5) Å3 | 1896.81(9) Å3 | 1995.50(18) Å3 | 1072.03(5) Å3 |
| Z | 2 | 4 | 4 | 2 |
| Density (calculated) | 1.191 Mg/m3 | 1.464 Mg/m3 | 1.984 Mg/m3 | 2.429 Mg/m3 |
| Absorption coefficient | 0.528 mm−1 | 5.700 mm−1 | 9.876 mm−1 | 45.683 mm−1 |
| F(000) | 306 | 864 | 1152 | 720 |
| Crystal size | 0.3758 × 0.2469 × 0.1231 mm3 | 0.4219 × 0.3013 × 0.1671 mm3 | 0.2644 × 0.1388 × 0.0971 mm3 | 1.0284 × 0.0934 × 0.0185 mm3 |
| Theta range for data collection | 3.90 to 71.20° | 3.00 to 71.09° | 2.94 to 71.62° | 2.95 to 71.19° |
| Index ranges | −8 ≤ h ≤ 8, −12 ≤ k ≤ 12, −14 ≤ l ≤ 14 | −18 ≤ h ≤ 17, −14 ≤ k ≤ 14, −12 ≤ l ≤ 13 | −18 ≤ h ≤ 18, −14 ≤ k ≤ 14, −13 ≤ l ≤ 12 | −7 ≤ h ≤ 6, −13 ≤ k ≤ 13, −18 ≤ l ≤ 18 |
| Reflections collected | 22,919 | 25,427 | 16,160 | 30,943 |
| Independent reflections | 3029 [R(int) = 0.0380] | 3654 [R(int) = 0.0323] | 3802 [R(int) = 0.0806] | 4136 [R(int) = 0.0464] |
| Completeness to theta = 71.20° | 99.2% | 99.6% | 97.5% | 99.6% |
| Absorption correction | Analytical | Analytical | Analytical | Analytical |
| Max. and min. transmission | 0.941 and 0.853 | 0.505 and 0.197 | 0.471 and 0.200 | 0.469 and 0.008 |
| Refinement method | Full-matrix least-squares on F2 | Full-matrix least-squares on F2 | Full-matrix least-squares on F2 | Full-matrix least-squares on F2 |
| Data/restraints/parameters | 3029/0/193 | 3654/0/226 | 3802/0/226 | 4136/0/226 |
| Goodness-of-fit on F2 | 1.040 | 1.046 | 1.791 | 1.101 |
| Final R indices [I > 2sigma(I)] | R1 = 0.0434, wR2 = 0.1186 | R1 = 0.0632, wR2 = 0.1876 | R1 = 0.1300, wR2 = 0.4177 | R1 = 0.0570, wR2 = 0.1746 |
| R indices (all data) | R1 = 0.0524, wR2 = 0.1265 | R1 = 0.0683, wR2 = 0.1934 | R1 = 0.1397, wR2 = 0.4323 | R1 = 0.0609, wR2 = 0.1780 |
| Largest diff. peak and hole | 0.174 and −0.226 e·Å−3 | 0.847 and −0.318 e·Å−3 | 3.881 and −1.041 e·Å−3 | 2.763 and −2.355 e·Å−3 |
| Melting point | 137 K | 183 K | 205 K | 225 K (decomposition) |
| Molecular System | X | Ecoulombic [kJ/mol] | Epolarization [kJ/mol] | Edispersion [kJ/mol] | Erepulsion [kJ/mol] | Etotal lattice [kJ/mol] |
|---|---|---|---|---|---|---|
| AB | H | −14.3 | −29.1 | −138.3 | 44.0 | −137.6 |
| tClAB | Cl | 1.6 | −23.5 | −165.7 | 40.3 | −147.3 |
| tBrAB | Br | −5.9 | −16.5 | −191.5 | 55.0 | −159.0 |
| tIAB | I | −9.5 | −15.1 | −225.4 | 78.0 | −172.0 |
| Molecular System | X | Ecoulombic [kJ/mol] | Epolarization [kJ/mol] | Edispersion [kJ/mol] | Erepulsion [kJ/mol] | Etotal lattice [kJ/mol] | Dipole Moment |
|---|---|---|---|---|---|---|---|
| AB | H | −38.5 | −16.9 | −156.4 | 90.8 | −121.0 | 4.522 |
| tClAB | Cl | −30.4 | −14.4 | −189.5 | 76.0 | −158.2 | 6.661 |
| tBrAB | Br | −39.3 | −16.4 | −182.7 | 83.8 | −154.7 | 7.724 |
| tIAB | I | −75.4 | −29.5 | −224.8 | 162.2 | −167.6 | 6.673 |
| Molecular System | X | Ecoulombic [kJ/mol] | Epolarization [kJ/mol] | Edispersion [kJ/mol] | Erepulsion [kJ/mol] | Etotal lattice [kJ/mol] | Level |
|---|---|---|---|---|---|---|---|
| AB | H | −29.1 | −9.3 | −130.0 | 63.3 | −111.6 | B3LYP |
| −29.9 | −15.6 | −171.9 | 54.1 | −152.7 | HF | ||
| tClAB | Cl | −26.5 | −9.1 | −170.5 | 81.6 | −133.3 | B3LYP |
| −20.8 | −14.1 | −182.4 | 62.9 | −144.9 | HF | ||
| tBrAB | Br | −42.1 | −11.6 | −157.6 | 86.4 | −138.9 | B3LYP |
| −19.8 | −21.2 | −211.1 | 73.6 | −175.4 | HF | ||
| tIAB | I | - | - | - | - | - | B3LYP * |
| −51.65 | −15.8 | −254.2 | 112.7 | −201.7 | HF |
| Compound | X | X⋯X [%] | X⋯H [%] | H⋯X [%] | X⋯C [%] | C⋯X [%] | X⋯N [%] | N⋯X [%] | C⋯H [%] | H⋯C [%] | N⋯H [%] | H⋯N [%] | H⋯H [%] | N⋯N [%] | C⋯N [%] | N⋯C [%] | C⋯C [%] | Surface [Å2] | Volume [Å3] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AB | H | - | - | - | - | - | - | - | 9.3 | 7.8 | 5.4 | 4.2 | 72.7 | 0.1 | 0.1 | 0.0 | 0.0 | 331.4 | 385.9 |
| tClAB | Cl | 1.0 | 25.3 | 17.4 | 3.8 | 3.9 | 1.2 | 1.1 | 5.1 | 4.0 | 3.8 | 3.3 | 29.1 | 0.0 | 0.2 | 0.2 | 0.6 | 383.2 | 467.2 |
| tBrAB | Br | 0.9 | 27.0 | 17.3 | 4.2 | 4.0 | 1.6 | 1.3 | 4.9 | 3.9 | 3.8 | 3.1 | 26.8 | 0.0 | 0.2 | 0.2 | 0.7 | 397.2 | 491.3 |
| tIAB | I | 8.6 | 22.4 | 12.6 | 5.4 | 5.1 | 0.5 | 0.4 | 3.5 | 2.8 | 3.3 | 2.9 | 29.9 | 0.3 | 0.5 | 0.4 | 0.8 | 423.9 | 526.9 |
| Molecular System | Atom | C | H | X | N |
|---|---|---|---|---|---|
| AB | Surface% | 8.7 | 86.3 | - | 4.8 |
| C | 0 | - | - | - | |
| H | 1.15 | 0.98 | - | - | |
| N | 0.12 | 1.12 | - | 0.42 | |
| tClAB | Surface% | 9.2 | 58.6 | 27.3 | 4.8 |
| C | 0.71 | - | - | - | |
| H | 0.85 | 0.85 | - | - | |
| X | 1.53 | 1.33 | 0.13 | - | |
| N | 0.34 | 1.26 | 0.84 | 0 | |
| tBrAB | Surface% | 9.4 | 56.8 | 28.6 | 5.1 |
| C | 0.79 | - | - | - | |
| H | 0.82 | 0.98 | - | - | |
| X | 1.53 | 1.36 | 0.11 | - | |
| N | 0.42 | 1.19 | 0.99 | 0 | |
| tIAB | Surface% | 9.65 | 54 | 32.15 | 4.3 |
| C | 0.86 | - | - | - | |
| H | 0.60 | 1.03 | - | - | |
| X | 1.69 | 1.03 | 0.83 | - | |
| N | 1.08 | 1.34 | 0.33 | 1.62 |
| Molecular System | X | VSmax [kJ/mol] | VSmin [kJ/mol] | VS+ [kJ/mol] | σ+2 [kJ/mol] | VS− [kJ/mol] | σ−2 [kJ/mol] | π [kJ/mol] | ηS [-] | σT2 [kJ/mol] | ηSσT2 [kJ/mol] |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AB | H | 50.14 | −97.09 | 18.89 | 0.06 | −31.25 | 0.40 | 19.45 | 0.12 | 0.47 | 0.06 |
| tClAB | Cl | 67.09 | −87.23 | 23.89 | 0.11 | −23.06 | 0.21 | 21.95 | 0.23 | 0.32 | 0.07 |
| tBrAB | Br | 85.01 | −103.20 | 28.06 | 0.15 | −28.06 | 0.31 | 26.81 | 0.22 | 0.46 | 0.10 |
| tIAB | I | 63.76 | −82.23 | 22.09 | 0.10 | −20.84 | 0.16 | 20.56 | 0.24 | 0.46 | 0.11 |
| Molecular System | Type of Interaction | Interaction | ρBCP(r) [a.u.] | ΔρBCP(r) [a.u.] | VBCP [a.u.] | GBCP [a.u.] | HBCP [a.u.] | EHBE [kJ/mol] | EHBM [kJ/mol] | EHBEM [kJ/mol] | EHBA [kJ/mol] | EXXK [kJ/mol] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| tAB | intra | H9 ⋯ H25 | 0.0128 | 0.0459 | −0.0074 | 0.0095 | 0.0020 | −9.79 | −10.67 | −8.84 | −5.88 | - |
| inter | N3 ⋯ H67, N24 ⋯ H43 | 0.0068 | 0.0209 | −0.0035 | 0.0044 | 0.0009 | −4.59 | −4.96 | −3.24 | −3.00 | - | |
| tClAB | intra | Cl4 ⋯ H56, Cl29 ⋯ H76 | 0.0028 | 0.0094 | −0.0013 | 0.0018 | 0.0005 | −1.71 | −2.03 | 0.49 | −1.40 | - |
| intra | Cl4 ⋯ H55, Cl29 ⋯ H75 | 0.0070 | 0.0246 | −0.0038 | 0.0050 | 0.0012 | −4.99 | −5.63 | −3.43 | −3.21 | - | |
| intra | Cl4 ⋯ H52, Cl29 ⋯ H72 | 0.0109 | 0.0394 | −0.0063 | 0.0081 | 0.0018 | −8.27 | −9.12 | −7.07 | −5.03 | - | |
| inter | Cl29 ⋯ H64, Cl29 ⋯ H54 | 0.0032 | 0.0121 | −0.0016 | 0.0023 | 0.0007 | −2.10 | −2.59 | 0.12 | −1.61 | - | |
| inter | H69 ⋯ H87 | 0.0032 | 0.0097 | −0.0016 | 0.0020 | 0.0004 | −2.10 | −2.25 | 0.12 | −1.61 | - | |
| inter | H64 ⋯ H76 | 0.0035 | 0.0107 | −0.0018 | 0.0022 | 0.0004 | −2.36 | −2.48 | −0.16 | −1.76 | - | |
| inter | H66 ⋯ H78 | 0.0045 | 0.0130 | −0.0024 | 0.0028 | 0.0004 | −3.15 | −3.15 | −1.09 | −2.20 | - | |
| inter | Cl1 ⋯ Cl27 | 0.0011 | 0.0033 | −0.0004 | 0.0006 | 0.0002 | −0.53 | −0.68 | - | - | −3.51 | |
| tBrAB | intra | Br4 ⋯ H56, Br32 ⋯ H84 | 0.0040 | 0.0125 | −0.0018 | 0.0025 | 0.0007 | −2.36 | −2.82 | −0.63 | −1.76 | - |
| intra | Br4 ⋯ H55, Br32 ⋯ H83 | 0.0077 | 0.0251 | −0.0040 | 0.0051 | 0.0011 | −5.25 | −5.74 | −4.08 | −3.36 | - | |
| intra | Br4 ⋯ H52, Br32 ⋯ H81 | 0.0111 | 0.0368 | −0.0061 | 0.0077 | 0.0015 | −8.01 | −8.67 | −7.25 | −4.89 | - | |
| inter | Br4 ⋯ H74, Br4 ⋯ H71 | 0.0032 | 0.0115 | −0.0015 | 0.0022 | 0.0007 | −1.97 | −2.48 | 0.12 | −1.54 | - | |
| inter | H67 ⋯ H79 | 0.0022 | 0.0073 | −0.0011 | 0.0015 | 0.0004 | −1.44 | −1.69 | 1.05 | −1.25 | - | |
| inter | H56 ⋯ H74 | 0.0034 | 0.0102 | −0.0017 | 0.0021 | 0.0004 | −2.23 | −2.37 | −0.07 | −1.69 | - | |
| inter | H58 ⋯ H75 | 0.0050 | 0.0140 | −0.0026 | 0.0031 | 0.0004 | −3.41 | −3.49 | −1.56 | −2.34 | - | |
| inter | Br2 ⋯ Br29 | 0.0014 | 0.0037 | −0.0005 | 0.0007 | 0.0002 | −0.66 | −0.79 | - | - | −6.27 | |
| tIAB | intra | I29 ⋯ H77, I4 ⋯ H57 | 0.0033 | 0.0099 | −0.0013 | 0.0019 | 0.0006 | −1.71 | −2.14 | 0.03 | −1.40 | - |
| intra | I29 ⋯ H72, I4 ⋯ H52 | 0.0147 | 0.0446 | −0.0085 | 0.0098 | 0.0013 | −11.16 | −11.04 | −10.61 | −6.63 | - | |
| intra | H30 ⋯ H17, H58 ⋯ H75 | 0.0125 | 0.0467 | −0.0076 | 0.0096 | 0.0021 | −9.8 | −10.78 | −8.57 | −8.87 | ||
| intra | I1 ⋯ I2, I26 ⋯ I27 | 0.0139 | 0.0387 | −0.0083 | 0.0090 | 0.0007 | −10.90 | −10.14 | - | - | −9.38 | |
| intra | I2 ⋯ I3, I27 ⋯ I28 | 0.0164 | 0.0420 | −0.0097 | 0.0101 | 0.0004 | −12.73 | −11.38 | - | - | −9.37 | |
| intra | I3 ⋯ I4, I28 ⋯ I29 | 0.0170 | 0.0433 | −0.0102 | 0.0105 | 0.0003 | −10.76 | −11.83 | - | - | −9.36 | |
| inter | I26 ⋯ H62, I1 ⋯ H82 | 0.0040 | 0.0119 | −0.0016 | 0.0023 | 0.0007 | −2.10 | −2.59 | −0.63 | −1.61 | - | |
| inter | N32 ⋯ H51, N7 ⋯ H71 | 0.0128 | 0.0428 | −0.0082 | 0.0095 | 0.0013 | −10.76 | −10.70 | −8.84 | −6.41 | - |
| Molecular System | HOMO [eV] | LUMO [eV] | ΔEHOMO-LUMO GAP [eV] | μ [eV] | χ [eV] | η [eV] | S [1/eV] | ω [eV] | ω+ [eV] | ω− [eV] | Δω [eV] | ΔNmax [-] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AB | −8.069 | 3.646 | 11.715 | −2.212 | 2.212 | 5.858 | 0.171 | 0.835 | 2.779 | 0.567 | 3346 | 0.378 |
| tClAB | −8.699 | 2.466 | 11.165 | −3.116 | 3.116 | 5.582 | 0.179 | 0.870 | 3.389 | 0.272 | 3661 | 0.558 |
| tBrAB | −7.751 | 2.234 | 9.985 | −2.758 | 2.758 | 4.993 | 0.200 | 0.762 | 3.008 | 0.250 | 3258 | 0.552 |
| tIAB | −8.497 | 1.420 | 9.917 | −3.538 | 3.538 | 4.959 | 0.202 | 1.262 | 3.640 | 0.102 | 3742 | 0.714 |
| Protein | Ligand | Score [kJ/mol] | Total Energy [kJ/mol] | Steric Protein-Ligand [kJ/mol] | Van der Waals [kJ/mol] |
|---|---|---|---|---|---|
| 3KXN | AB | −49.55 | −51.60 | −71.55 | −25.56 |
| tClAB | −60.33 | −70.14 | −93.91 | −25.66 | |
| tBrAB | −59.73 | −72.18 | −98.33 | −26.16 | |
| tIAB | −62.24 | −77.21 | −95.00 | −26.24 | |
| Actual ligand—TIBI | −35.98 | −51.18 | −55.34 | −12.92 | |
| 2MUV | AB | −44.99 | −45.55 | −70.06 | −28.51 |
| tClAB | −54.16 | −60.78 | −84.42 | −29.73 | |
| tBrAB | −52.48 | −59.71 | −83.28 | −28.70 | |
| tIAB | −51.59 | −58.54 | −78.64 | −30.08 | |
| Actual ligand | −38.95 | −43.83 | −55.02 | −24.44 | |
| 2MUW | AB | −38.53 | −34.96 | −59.18 | −27.07 |
| tClAB | −46.77 | −53.77 | −71.91 | −27.16 | |
| tBrAB | −45.59 | −50.71 | −70.07 | −27.10 | |
| tIAB | −46.76 | −51.94 | −68.63 | −26.87 | |
| Actual ligand | −27.94 | −21.63 | −39.30 | −21.16 | |
| 2LY01 | AB | −48.85 | −48.40 | −72.40 | −29.20 |
| tClAB | −53.23 | −59.74 | −79.70 | −29.88 | |
| tBrAB | −51.13 | −58.19 | −76.96 | −28.75 | |
| tIAB | −52.93 | −58.81 | −76.71 | −29.29 | |
| Actual ligand | −54.39 | −54.53 | −70.94 | −29.55 | |
| 6M17 small cave | AB | −31.33 | −102.85 | −143.43 | 65.76 |
| tClAB | −3.312 | −62.39 | −99.57 | 55.85 | |
| tBrAB | −16.41 | −60.90 | −94.51 | 28.07 | |
| tIAB | −33.41 | −62.85 | −88.02 | 4.98 | |
| Actual ligand—Amantadine | −57.52 | −57.64 | −79.48 | −12.92 | |
| Actual ligand—Rimantadine | −75.22 | −73.04 | −100.15 | −30.28 | |
| 6M17 large cave | AB | −69.71 | −79.54 | −104.80 | −28.93 |
| tClAB | −78.58 | −95.92 | −120.56 | −27.69 | |
| tBrAB | −78.11 | −97.58 | −117.05 | −27.72 | |
| tIAB | −75.25 | −95.84 | −110.26 | −24.87 | |
| 7QUR | AB | −64.99 | −70.66 | −99.70 | −30.70 |
| tClAB | −75.84 | −87.30 | −119.69 | −27.31 | |
| tBrAB | −76.26 | −89.58 | −117.38 | −28.66 | |
| tIAB | −75.52 | −89.45 | −114.43 | −28.28 | |
| Actual ligand | −45.73 | −53.78 | −61.68 | −15.63 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Latosińska, J.N.; Latosińska, M.; Orzeszko, A.; Maurin, J.K. Synthesis and Crystal Structure of Adamantylated 4,5,6,7-Tetrahalogeno-1H-benzimidazoles Novel Multi-Target Ligands (Potential CK2, M2 and SARS-CoV-2 Inhibitors); X-ray/DFT/QTAIM/Hirshfeld Surfaces/Molecular Docking Study. Molecules 2023, 28, 147. https://doi.org/10.3390/molecules28010147
Latosińska JN, Latosińska M, Orzeszko A, Maurin JK. Synthesis and Crystal Structure of Adamantylated 4,5,6,7-Tetrahalogeno-1H-benzimidazoles Novel Multi-Target Ligands (Potential CK2, M2 and SARS-CoV-2 Inhibitors); X-ray/DFT/QTAIM/Hirshfeld Surfaces/Molecular Docking Study. Molecules. 2023; 28(1):147. https://doi.org/10.3390/molecules28010147
Chicago/Turabian StyleLatosińska, Jolanta Natalia, Magdalena Latosińska, Andrzej Orzeszko, and Jan Krzysztof Maurin. 2023. "Synthesis and Crystal Structure of Adamantylated 4,5,6,7-Tetrahalogeno-1H-benzimidazoles Novel Multi-Target Ligands (Potential CK2, M2 and SARS-CoV-2 Inhibitors); X-ray/DFT/QTAIM/Hirshfeld Surfaces/Molecular Docking Study" Molecules 28, no. 1: 147. https://doi.org/10.3390/molecules28010147