Abstract
We herein review available computational and experimental data pointing to the abundance of structural disorder within the nucleoprotein (N) and phosphoprotein (P) from three paramyxoviruses, namely the measles (MeV), Nipah (NiV) and Hendra (HeV) viruses. We provide a detailed molecular description of the mechanisms governing the disorder-to-order transition that the intrinsically disordered C-terminal domain (NTAIL) of their N proteins undergoes upon binding to the C-terminal X domain (PXD) of the homologous P proteins. We also show that NTAIL–PXD complexes are “fuzzy”, i.e., they possess a significant residual disorder, and discuss the possible functional significance of this fuzziness. Finally, we emphasize the relevance of N–P interactions involving intrinsically disordered proteins as promising targets for new antiviral approaches, and end up summarizing the general functional advantages of disorder for viruses.
1. Overview of the Replicative Complex of Paramyxoviruses
Negative-stranded RNA viruses (NSRVs) are causative agents of a large number of human and animal diseases with some of them being identified as potential agents of bioterrorism, and several being included in the NIAID (National Institute of Allergy and Infectious Diseases) and CDC (Center for Disease Control and Prevention) priority pathogen lists. Research on these viruses, as well as efforts aimed at the development of vaccines and antiviral drugs, have been paid an increased attention for many years. NSRVs can be divided into viruses with segmented RNA genomes and those with non-segmented RNA genomes. The latter are grouped within the Mononegavirales order. Among the viral families belonging to this order is the Paramyxoviridae family. The Nipah (NiV), Hendra (HeV) and measles (MeV) viruses belong to the Paramyxovirinae sub-family within the Paramyxoviridae family, where the latter also embraces the Pneumovirinae subfamily. Based on distinguishing nucleotide sequence features, the MeV and the NiV and HeV and have been classified into two distinct genera, the Morbillivirus genus and the Henipavirus genus, respectively [1,2,3].
Paramyxoviruses display a pleomorphic structure. The viral particule contains the non-segmented, negative-stranded RNA genome, which encodes for at least six proteins (Figure 1A). The genome of paramyxoviruses indeed encodes the fusion (F) and the attachment (H) glycoproteins, which are responsible for virus entry, the matrix (M), which is required for virus assembly and budding, and the proteins of the replicative complex (Figure 1B). In paramyxoviruses, RNA transcription and replication require an intricate interplay between three components: the RNA-dependent RNA polymerase (L), the phosphoprotein (P), and the nucleoprotein (N). As in all Mononegavirales members, the genome is encapsidated by N within a helical nucleocapsid. The N:RNA complex, rather than naked RNA, is the template for both transcription and replication. During RNA synthesis, P tethers L onto the N–RNA template through the N-P interaction. The complex formed by the N, P and L proteins constitutes the viral replicative unit, and these proteins are necessary and sufficient to sustain replication of viral RNA in Paramyxovirinae (Figure 1C) [4,5].
The N protein is the most abundant viral protein (Figure 1A). Within infected cells, the N protein from Paramyxoviridae members is found in a soluble, monomeric form (referred to as N°) and in a nucleocapsid assembled form (referred to as NNUC) [6,7]. Following synthesis of the N protein, a chaperone is required to maintain this latter protein in the unassembled form in the cytoplasm. This role is played by the P protein, whose association prevents illegitimate self-assembly of N and retains N in the cytoplasm [8,9]. This soluble N°-P complex is used as the substrate for the encapsidation of the nascent genomic RNA chain during replication. The assembled form of N also forms complexes with P, either isolated (NNUC-P) or bound to L (NNUC-P-L), which are essential to RNA synthesis by the viral polymerase. Hence, the components of the viral replication machinery, namely P, N and L, engage in a complex macromolecular ballet (Figure 1C) (see [2,5,10,11,12,13,14] for reviews on transcription and replication).
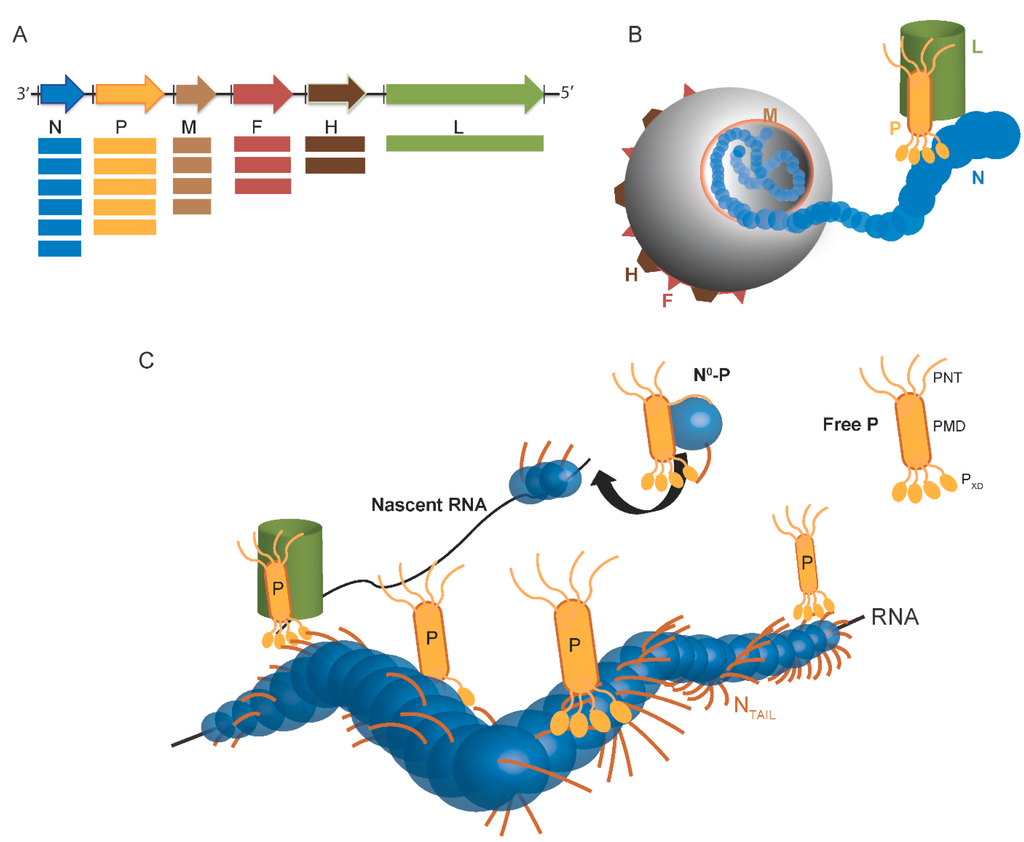
Figure 1.
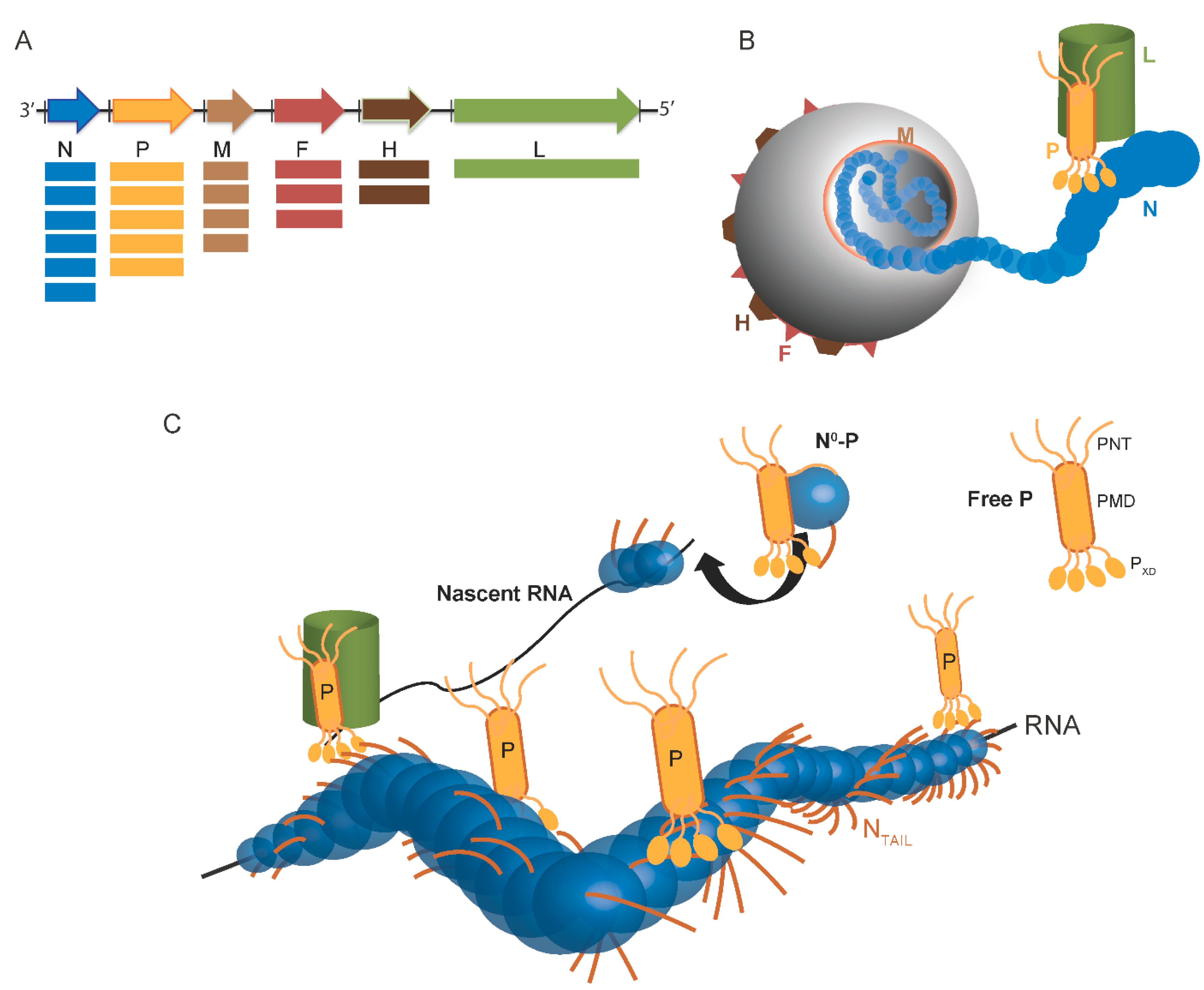
Scheme of Paramyxovirinae genome and viral particule. (A) Schematic representation of the genome of Paramyxovirinae. The negative-sense genomic RNA is presented in the 3′ to 5′ orientation. The open reading frames are represented by colored arrows and encode the nucleocapsid protein (N), phosphoprotein (P), matrix protein (M), fusion protein (F), attachment protein (H) and large polymerase protein (L). Vertical lines represent gene start and stop signals. Below the genome, shown is a schematic representation of the expression gradient of the encoded proteins as a result of the stop and re-initiation mechanism of the polymerase during transcription [2]; (B) Schematic illustration of the virion. The viral membrane is decorated by the F and H glycoproteins and M is located beneath the membrane. N is bound to genomic RNA and together with P and L forms the viral replication unit; (C) Schematic illustration of the Paramyxoviridae replicative complex. The RNA is represented as a solid black line. The neo-synthetized RNA is shown already partially encapsidated by N. The N and P intrinsically disordered regions are symbolized by lines. The extended conformation of the disordered regions is thought to allow the formation of a tripartite complex between N°, P and L required for nucleocapsid assembly. The P/L complex forms the RNA-dependent RNA polymerase (RdRp) complex that cartwheels onto the nucleocapsid complex via the X domain of P (PXD). P is shown as a tetramer to reflect the prevalence of this oligomeric state in paramyxoviral P proteins.
Figure 1.
Scheme of Paramyxovirinae genome and viral particule. (A) Schematic representation of the genome of Paramyxovirinae. The negative-sense genomic RNA is presented in the 3′ to 5′ orientation. The open reading frames are represented by colored arrows and encode the nucleocapsid protein (N), phosphoprotein (P), matrix protein (M), fusion protein (F), attachment protein (H) and large polymerase protein (L). Vertical lines represent gene start and stop signals. Below the genome, shown is a schematic representation of the expression gradient of the encoded proteins as a result of the stop and re-initiation mechanism of the polymerase during transcription [2]; (B) Schematic illustration of the virion. The viral membrane is decorated by the F and H glycoproteins and M is located beneath the membrane. N is bound to genomic RNA and together with P and L forms the viral replication unit; (C) Schematic illustration of the Paramyxoviridae replicative complex. The RNA is represented as a solid black line. The neo-synthetized RNA is shown already partially encapsidated by N. The N and P intrinsically disordered regions are symbolized by lines. The extended conformation of the disordered regions is thought to allow the formation of a tripartite complex between N°, P and L required for nucleocapsid assembly. The P/L complex forms the RNA-dependent RNA polymerase (RdRp) complex that cartwheels onto the nucleocapsid complex via the X domain of P (PXD). P is shown as a tetramer to reflect the prevalence of this oligomeric state in paramyxoviral P proteins.
Although in vitro L can synthesize short RNA transcripts using naked RNA as substrate in the absence of P [15], in infected cells P is required to stabilize L and to allow recognition of the N:RNA template. L is thought to carry out most (if not all) enzymatic activities required for transcription and replication, including nucleotide polymerization, mRNA capping and polyadenylation. It is found in low amounts in infected cells and is unstable unless bound to the P protein, thus making its full characterization challenging [16]. Since no functional paramyxoviral polymerase has been biochemically characterized so far, most of our present knowledge arises from bioinformatics studies. Among Paramyxovirinae members, the only exceptions are represented by the L/P complex from two Paramyxovirinae members, namely Rinderpest virus (RDV) whose polymerase has been partially purified [17], and Sendai virus (SeV) whose polymerase was shown to possess a methyltransefrase activity in its C-terminal region [18]. Among Pneumovirinae, the only exception is the L protein from respiratory syncytial virus (RSV) that could be partially purified and whose RNA polymerase activity was documented in vitro [19]. In addition, minireplicon studies allowed the identification within RSV L of a flexible hinge region tolerating insertion and demonstrated the crucial role of the GDNQ motif [20].
Accordingly, most of our present knowledge of the replicative complex of paramyxoviruses concerns the N and P proteins. In the last decades, many efforts have been devoted to the molecular characterization of paramyxoviral N and P proteins. The N–P interaction has attracted much interest not only from a fundamental point of view, but also from a more applied perspective: in fact, since abolishing the N–P interaction prevents the recruitment of L onto the nucleocapsid template, this interaction is regarded as a potential target for antiviral approaches.
In the course of a thorough structural and functional characterization of paramyxoviral N and P proteins that made use of a wide range of bioinformatics and experimental approaches, we showed that these proteins are enriched in intrinsically disordered regions (IDRs) and that these IDRs play key roles in the formation of the tripartite N–P–L complex and in the establishment of a broad molecular partnership (for reviews see [11,21,22,23,24,25,26,27]). Our seminal observations on MeV P and N proteins fostered subsequent studies that brought awareness of the prevalence and conservation of structural disorder within paramyxoviruses N and P proteins thereby pointing to its functional significance.
Intrinsically disordered proteins (IDPs) and IDRs are widespread functional proteins/regions that lack stable structures under physiological conditions. Behind their inability to fold resides an important raison d’être, which is tightly coupled to specific features of their amino acid sequence. Indeed, a specific imbalance in the content of hydrophobic vs. polar residues in IDP/Rs, confers them the ability to populate a wide conformational space with conformations ranging from completely extended (i.e., random coils, RC) to more compact (i.e., pre-molten globules, PMGs, and molten globules, MGs). The nature of the conformational ensemble that IDP/Rs could sample is coupled to their function and, hence, to the interactions they establish with their partners. Indeed, in many cases, IDP/Rs fold upon binding leading to either stable complexes amenable to crystallization, or, more often, to fuzzy complexes [28], i.e., complexes with significant residual disorder. These peculiar characteristics award IDP/Rs a number of advantages over folded proteins that promote their frequent involvement in particular functions, such as for instance, hubs in protein interaction networks and cell signaling (for a recent review on IDP/Rs see [29]).
In the current review, we provide a detailed description of the molecular information that exist to date on the N and P proteins from paramyxoviruses while highlighting the unique role of structural disorder in ensuring an efficient replication and transcription of the paramyxoviral genome. We accordingly discuss through the manuscript the functional role of induced folding and residual flexibility in terms of transcription, replication and molecular partnership. We also underscore how targeting the N–P interaction holds promises for new antiviral approaches. We finally conclude by highlighting the functional implications and general advantages of structural disorder within viruses.
6. Conclusions
When we analyzed the modular organization of the P proteins within the Paramyxovirinae subfamily, we noticed that a larger PNT domain in Henipavirus P proteins accounts for the extra length of their P protein with respect to other paramyxoviruses [30]. This finding is consistent with the higher tolerance of disordered regions to insertions or major rearrangements as compared to ordered ones. Moreover, since the P-encoded proteins are believed to possess anti-interferon functions, the extension in Henipavirus P proteins might have evolved to better equip these viruses so as to enhance their capacity to overcome the cellular interferon response. Furthermore, the disordered nature of PNT and of the “spacer” region connecting PNT to PMD likely reflects a way of alleviating evolutionary constraints within overlapping reading frames. Indeed, PNT partially overlaps with the C protein (being encoded by the same RNA region), and the “spacer” region partially overlaps with the C-terminal domain of the V protein [30,31]. This observation is in agreement with previous reports pointing out a relationship between overlapping genes and structural disorder [31,170,171,172,173]. We thus reasoned that structural disorder, which is encoded by a much wider portion of sequence space as compared to order, can indeed represent a strategy by which genes encoding overlapping reading frames can lessen evolutionary constraints imposed on their sequence by the overlap, allowing the encoded overlapping protein products to sample a wider sequence space without losing function.
Following our seminal studies that pointed out the abundance of disorder in Paramyxovirinae N and P proteins [31,32,57,174], several subsequent studies have documented the prevalence of disorder in viral proteins using both computational and experimental approaches (see [175] and references therein cited). Bioinformatics studies showed that viral proteins, and in particular proteins from RNA viruses, have a high disorder content [176,177]. In those studies, the authors propose that beyond affording a broad partnership, the wide occurrence of disordered regions in viral proteins could also be related to the typical high mutation rates of RNA viruses, i.e., it could represent a strategy for buffering the deleterious effects of mutations.
A detailed comparative examination of viral and non-viral proteins showed that, amongst several distinguishing characteristics, viral proteins possess (i) a larger fraction of residues that are not organized into regular secondary structural elements; (ii) conformational stabilities that are less affected by mutations; (iii) a high rate of mutations; (iv) enrichment in proteins encoded by overlapping reading frames; and (v) a higher content of polar residues. These features indicate that they have been shaped by evolution to be endowed with better adaptation to their hostile habitats and to rapid changes in their biological and physical environment. Indeed, one of the many noteworthy features of viruses is their ability to adapt to very harsh and hostile environments and to adjust themselves according to the biological and genetic features of the hosts, which in turn are often adapted to exist at extreme conditions (see [175] and references therein cited). The above-mentioned features are interestingly in intimate relationship with structural disorder, which indeed provides several advantages.
In fact, because viruses are obligate intracellular parasites, they have to interact with various components of the host, including membranes, nucleic acids, and proteins. The lack of a rigid 3D structure imparts to IDP/Rs the necessary plasticity to establish various interactions with several partners at once. In the course of evolution, viruses have “learned” to hijack and manipulate host proteins for their benefit, and to evade the host defense mechanisms. A recent study by Davey and co-workers showed that viruses have achieved this ability through broad mimicry of host protein short linear motifs (SLiMs) [178], where the latter are embedded in disordered regions and play a variety of roles, including targeting host proteins for proteosomal degradation, cell signaling, directing proteins to the correct subcellular localization, deregulating cell cycle checkpoints, and altering transcription of host proteins [179]. Importantly, binding to cell proteins through sites that mimic SLiMs also helps viral proteins in eluding the host cell’s immune system, by rendering viral epitopes poorly recognizable by the host immune system (see [175] and references therein cited).
Based on all these considerations, we proposed that the main advantage of the abundance of disorder within viral proteins would reside in pleiotropy and genetic compaction [175]. Indeed, disorder provides a solution to reduce both genome size and molecular crowding, where a single gene would (i) encode a single (regulatory) protein product that can establish multiple interactions via its disordered regions and hence exert multiple concomitant biological effects including evasion of the host immune response; and/or (ii) would encode more than one product by means of overlapping reading frames.
Acknowledgments
Sonia Longhi wishes to thank all the members of her lab and her co-workers for their critical contribution to the studies herein summarized. Within her group, she thanks David Karlin, François Ferron, Jean-Marie Bourhis, Kenth Johansson, Antoine Gruet, David Blocquel, Jenny Erales, Lorenzo Baronti, and Jennifer Roche (previous members) and Marion Dosnon, Matilde Beltrandi and Christophe Bignon (present members). Among her numerous past and present co-workers, she thanks Bruno Canard (AFMB, Marseille, France), Denis Gerlier (CIRI, Lyon, France), Michael Oglesbee (Ohio State University, Columbus, OH, USA), Hélène Valentin (CIRI, Lyon, France), Valerie Belle and Bruno Guigliarelli (BIP, Marseille, France), Janez Strancar (Jozef Stefan Institute, Ljubljana, Slovenia), Gary Daughdrill (University of South Florida, Tampa, FL, USA), Martin Blackledge and Malene Ringkjobin-Jensen (Institut de Biologie Structurale, Grenoble, France), Jin Wang (Changchun Institute of Applied Chemistry, Chinese Academy of Sciences, Changchun, China), Roberta Pierattelli and Isabella Felli (CERM, Florence, Italy), Rita Grandori (Universita’ degli Studi Milano-Bicocca, Milan, Italy), Paolo Carloni (Institute for Advanced Simulation IAS-5 and Institute of Neuroscience and Medicine INM-9, Jülich, Düren, Germany) and Stefano Gianni (La Sapienza, Universita’ di Roma, Rome, Italy). She is particularly grateful to Denis Gerlier (CIRI, Lyon, France) and to Vladimir Uversky (University of South Florida) for the numerous stimulating discussions and for their useful advice on various issues. The studies herein reviewed were carried out with the financial support of the Agence Nationale de la Recherche, specific programs “Physico-Chimie du Vivant”, ANR-08-PCVI-0020-01, and “ASTRID”, ANR-11-ASTR-003-01. They also benefited from support from the CNRS, the Direction Générale de l’Armement (DGA) and the Fondation pour la Recherche Médicale (FRM). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author Contributions
Conflicts of Interest
The authors declare no conflict of interest.
References
- Eaton, B.T.; Mackenzie, J.S.; Wang, L.F. Henipaviruses. In Fields Virology, 5th ed.; Fields, B.N., Knipe, D.M., Howley, P.M., Eds.; Lippincott-Raven: Philadelphia, PA, USA, 2007; pp. 1587–1600. [Google Scholar]
- Lamb, R.A.; Parks, G.D. Paramyxoviridae: The viruses and their replication. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 1450–1497. [Google Scholar]
- Wang, L.F.; Yu, M.; Hansson, E.; Pritchard, L.I.; Shiell, B.; Michalski, W.P.; Eaton, B.T. The exceptionally large genome of Hendra virus: Support for creation of a new genus within the family Paramyxoviridae. J. Virol. 2000, 74, 9972–9979. [Google Scholar] [CrossRef] [PubMed]
- Halpin, K.; Bankamp, B.; Harcourt, B.H.; Bellini, W.J.; Rota, P.A. Nipah virus conforms to the rule of six in a minigenome replication assay. J. Gen. Virol. 2004, 85, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Lamb, R.A.; Kolakofsky, D. Paramyxoviridae: The viruses and their replication. In Fields Virology, 4th ed.; Fields, B.N., Knipe, D.M., Howley, P.M., Eds.; Lippincott-Raven: Philadelphia, PA, USA, 2001; pp. 1305–1340. [Google Scholar]
- Bhella, D.; Ralph, A.; Murphy, L.B.; Yeo, R.P. Significant differences in nucleocapsid morphology within the Paramyxoviridae. J. Gen. Virol. 2002, 83, 1831–1839. [Google Scholar] [PubMed]
- Tan, W.S.; Ong, S.T.; Eshaghi, M.; Foo, S.S.; Yusoff, K. Solubility, immunogenicity and physical properties of the nucleocapsid protein of nipah virus produced in Escherichia coli. J. Med. Virol. 2004, 73, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Huber, M.; Cattaneo, R.; Spielhofer, P.; Orvell, C.; Norrby, E.; Messerli, M.; Perriard, J.C.; Billeter, M.A. Measles virus phosphoprotein retains the nucleocapsid protein in the cytoplasm. Virology 1991, 185, 299–308. [Google Scholar] [CrossRef]
- Spehner, D.; Drillien, R.; Howley, P.M. The assembly of the measles virus nucleoprotein into nucleocapsid-like particles is modulated by the phosphoprotein. Virology 1997, 232, 260–268. [Google Scholar] [CrossRef] [PubMed]
- Albertini, A.A.V.; Schoehn, G.; Ruigrok, R.W. Structures impliquées dans la réplication et la transcription des virus à arn non segmentés de sens négatif. Virologie 2005, 9, 83–92. (In French) [Google Scholar]
- Blocquel, D.; Bourhis, J.M.; Eléouët, J.F.; Gerlier, D.; Habchi, J.; Jamin, M.; Longhi, S.; Yabukarski, F. Transcription et réplication des mononégavirales: Une machine moléculaire originale. Virologie 2012, 16, 225–257. (In French) [Google Scholar]
- Lamb, R.A.; Krug, R.M. Orthomyxoviridae: The viruses and their replication. In Fields Virology, 4th ed.; Fields, B.N., Knipe, D.M., Howley, P.M., Eds.; Lippincott-Raven: Philadelphia, PA, USA, 2001; pp. 1487–1531. [Google Scholar]
- Longhi, S.; Canard, B. Mécanismes de transcription et de réplication des Paramyxoviridae. Virologie 1999, 3, 227–240. (In French) [Google Scholar]
- Roux, L. Dans le génome des paramyxovirinae, les promoteurs et leurs activités sont façonnés par la «règle de six». Virologie 2005, 9, 19–34. (In French) [Google Scholar]
- Morin, B.; Rahmeh, A.A.; Whelan, S.P. Mechanism of RNA synthesis initiation by the vesicular stomatitis virus polymerase. EMBO J. 2012, 31, 1320–1329. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, S.; Banerjee, A.K. Phosphoprotein, P of human parainfluenza virus type 3 prevents self-association of RNA-dependent RNA polymerase, L. Virology 2009, 383, 226–236. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, M.; Shaila, M.S. Recombinant L and P protein complex of Rinderpest virus catalyses mRNA synthesis in vitro. Virus Res. 2008, 135, 150–154. [Google Scholar] [CrossRef] [PubMed]
- Ogino, T.; Kobayashi, M.; Iwama, M.; Mizumoto, K. Sendai virus RNA-dependent RNA polymerase L protein catalyzes cap methylation of virus-specific mRNA. J. Biol. Chem. 2005, 280, 4429–4435. [Google Scholar] [CrossRef] [PubMed]
- Noton, S.L.; Deflube, L.R.; Tremaglio, C.Z.; Fearns, R. The respiratory syncytial virus polymerase has multiple RNA synthesis activities at the promoter. PLoS Pathog. 2012, 8, e1002980. [Google Scholar] [CrossRef] [PubMed]
- Fix, J.; Galloux, M.; Blondot, M.L.; Eleouet, J.F. The insertion of fluorescent proteins in a variable region of respiratory syncytial virus L polymerase results in fluorescent and functional enzymes but with reduced activities. Open Virol. J. 2011, 5, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Bourhis, J.M.; Longhi, S. Measles virus nucleoprotein: Structural organization and functional role of the intrinsically disordered C-terminal domain. In Measles Virus Nucleoprotein; Longhi, S., Ed.; Nova Publishers Inc.: Hauppage, NY, USA, 2007; pp. 1–35. [Google Scholar]
- Longhi, S. Measles Virus Nucleoprotein; Nova Publishers Inc.: Hauppage, NY, USA, 2007. [Google Scholar]
- Longhi, S. Nucleocapsid structure and function. Curr. Top. Microbiol. Immunol. 2009, 329, 103–128. [Google Scholar] [PubMed]
- Longhi, S.; Oglesbee, M. Structural disorder within the measles virus nucleoprotein and phosphoprotein. Protein Pept. Lett. 2010, 17, 961–978. [Google Scholar] [CrossRef] [PubMed]
- Longhi, S. Structural disorder within the measles virus nucleoprotein and phosphoprotein: Functional implications for transcription and replication. In Negative Strand RNA Virus; Luo, M., Ed.; World Scientific Publishing: Singapore, Singapore, 2011; pp. 95–125. [Google Scholar]
- Habchi, J.; Longhi, S. Structural disorder within paramyxovirus nucleoproteins and phosphoproteins. Mol. Biosyst. 2012, 8, 69–81. [Google Scholar] [CrossRef] [PubMed]
- Habchi, J.; Mamelli, L.; Longhi, S. Structural disorder within the nucleoprotein and phosphoprotein from measles, nipah and hendra viruses. In Flexible Viruses: Structural Disorder in Viral Proteins; Uversky, V.N., Longhi, S., Eds.; John Wiley and Sons: Hoboken, NJ, USA, 2012; pp. 47–94. [Google Scholar]
- Tompa, P.; Fuxreiter, M. Fuzzy complexes: Polymorphism and structural disorder in protein–protein interactions. Trends Biochem. Sci. 2008, 33, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Habchi, J.; Tompa, P.; Longhi, S.; Uversky, V.N. Introducing protein intrinsic disorder. Chem. Rev. 2014, 114, 6561–6588. [Google Scholar] [CrossRef] [PubMed]
- Habchi, J.; Mamelli, L.; Darbon, H.; Longhi, S. Structural disorder within Henipavirus nucleoprotein and phosphoprotein: From predictions to experimental assessment. PLoS ONE 2010, 5, e11684. [Google Scholar] [CrossRef] [PubMed]
- Karlin, D.; Ferron, F.; Canard, B.; Longhi, S. Structural disorder and modular organization in Paramyxovirinae N and P. J. Gen. Virol. 2003, 84, 3239–3252. [Google Scholar] [CrossRef] [PubMed]
- Karlin, D.; Longhi, S.; Receveur, V.; Canard, B. The N-terminal domain of the phosphoprotein of morbilliviruses belongs to the natively unfolded class of proteins. Virology 2002, 296, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Bourhis, J.; Johansson, K.; Receveur-Bréchot, V.; Oldfield, C.J.; Dunker, A.K.; Canard, B.; Longhi, S. The C-terminal domain of measles virus nucleoprotein belongs to the class of intrinsically disordered proteins that fold upon binding to their physiological partner. Virus Res. 2004, 99, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Houben, K.; Marion, D.; Tarbouriech, N.; Ruigrok, R.W.; Blanchard, L. Interaction of the C-terminal domains of sendai virus N and P proteins: Comparison of polymerase-nucleocapsid interactions within the Paramyxovirus family. J. Virol. 2007, 81, 6807–6816. [Google Scholar] [CrossRef] [PubMed]
- Johansson, K.; Bourhis, J.M.; Campanacci, V.; Cambillau, C.; Canard, B.; Longhi, S. Crystal structure of the measles virus phosphoprotein domain responsible for the induced folding of the C-terminal domain of the nucleoprotein. J. Biol. Chem. 2003, 278, 44567–44573. [Google Scholar] [CrossRef] [PubMed]
- Kingston, R.L.; Hamel, D.J.; Gay, L.S.; Dahlquist, F.W.; Matthews, B.W. Structural basis for the attachment of a paramyxoviral polymerase to its template. Proc. Natl. Acad. Sci. USA 2004, 101, 8301–8306. [Google Scholar] [CrossRef] [PubMed]
- Kingston, R.L.; Walter, A.B.; Gay, L.S. Characterization of nucleocapsid binding by the measles and the mumps virus phosphoprotein. J. Virol. 2004, 78, 8630–8640. [Google Scholar] [CrossRef] [PubMed]
- Bernado, P.; Blanchard, L.; Timmins, P.; Marion, D.; Ruigrok, R.W.; Blackledge, M. A structural model for unfolded proteins from residual dipolar couplings and small-angle X-ray scattering. Proc. Natl. Acad. Sci. USA 2005, 102, 17002–17007. [Google Scholar] [CrossRef] [PubMed]
- Blanchard, L.; Tarbouriech, N.; Blackledge, M.; Timmins, P.; Burmeister, W.P.; Ruigrok, R.W.; Marion, D. Structure and dynamics of the nucleocapsid-binding domain of the sendai virus phosphoprotein in solution. Virology 2004, 319, 201–211. [Google Scholar] [CrossRef] [PubMed]
- Houben, K.; Blanchard, L.; Blackledge, M.; Marion, D. Intrinsic dynamics of the partly unstructured PX domain from the sendai virus RNA polymerase cofactor P. Biophys. J. 2007, 93, 2830–2844. [Google Scholar] [CrossRef] [PubMed]
- Communie, G.; Crepin, T.; Maurin, D.; Jensen, M.R.; Blackledge, M.; Ruigrok, R.W. Structure of the tetramerization domain of measles virus phosphoprotein. J. Virol. 2013, 87, 7166–7169. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, S.M.; Sun, D.; Schmitt, A.P.; He, B. Phosphorylation of paramyxovirus phosphoprotein and its role in viral gene expression. Future Microbiol. 2010, 5, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Gerard, F.C.; Ribeiro Ede, A., Jr.; Leyrat, C.; Ivanov, I.; Blondel, D.; Longhi, S.; Ruigrok, R.W.; Jamin, M. Modular organization of rabies virus phosphoprotein. J. Mol. Biol. 2009, 388, 978–996. [Google Scholar] [CrossRef] [PubMed]
- Leyrat, C.; Gerard, F.C.; de Almeida Ribeiro, E., Jr.; Ivanov, I.; Ruigrok, R.W.; Jamin, M. Structural disorder in proteins of the rhabdoviridae replication complex. Protein Pept. Lett. 2010, 17, 979–987. [Google Scholar] [CrossRef] [PubMed]
- Iakoucheva, L.M.; Radivojac, P.; Brown, C.J.; O’Connor, T.R.; Sikes, J.G.; Obradovic, Z.; Dunker, A.K. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res. 2004, 32, 1037–1049. [Google Scholar] [CrossRef] [PubMed]
- Ringkjøbing Jensen, M.; Communie, G.; Ribeiro, E.D., Jr.; Martinez, N.; Desfosses, A.; Salmon, L.; Mollica, L.; Gabel, F.; Jamin, M.; Longhi, S.; et al. Intrinsic disorder in measles virus nucleocapsids. Proc. Natl. Acad. Sci. USA 2011, 108, 9839–9844. [Google Scholar]
- Ruigrok, R.W.; Crepin, T.; Kolakofsky, D. Nucleoproteins and nucleocapsids of negative-strand RNA viruses. Curr. Opin. Microbiol. 2011, 14, 504–510. [Google Scholar] [CrossRef] [PubMed]
- Warnes, A.; Fooks, A.R.; Dowsett, A.B.; Wilkinson, G.W.; Stephenson, J.R. Expression of the measles virus nucleoprotein gene in Escherichia coli and assembly of nucleocapsid-like structures. Gene 1995, 160, 173–178. [Google Scholar] [CrossRef]
- Bhella, D. Measles virus nucleocapsid structure, conformational flexibility and the rule of six. In Measles Virus Nucleoprotein; Longhi, S., Ed.; Nova Publishers Inc.: Hauppage, NY, USA, 2007. [Google Scholar]
- Eshaghi, M.; Tan, W.S.; Ong, S.T.; Yusoff, K. Purification and characterization of Nipah virus nucleocapsid protein produced in insect cells. J. Clin. Microbiol. 2005, 43, 3172–3177. [Google Scholar] [CrossRef] [PubMed]
- Jensen, M.R.; Bernado, P.; Houben, K.; Blanchard, L.; Marion, D.; Ruigrok, R.W.; Blackledge, M. Structural disorder within sendai virus nucleoprotein and phosphoprotein: Insight into the structural basis of molecular recognition. Protein Pept. Lett. 2010, 17, 952–960. [Google Scholar] [CrossRef] [PubMed]
- Heggeness, M.H.; Scheid, A.; Choppin, P.W. Conformation of the helical nucleocapsids of paramyxoviruses and vesicular stomatitis virus: Reversible coiling and uncoiling induced by changes in salt concentration. Proc. Natl. Acad. Sci. USA 1980, 77, 2631–2635. [Google Scholar] [CrossRef] [PubMed]
- Heggeness, M.H.; Scheid, A.; Choppin, P.W. The relationship of conformational changes in the sendai virus nucleocapsid to proteolytic cleavage of the NP polypeptide. Virology 1981, 114, 555–562. [Google Scholar] [CrossRef]
- Karlin, D.; Longhi, S.; Canard, B. Substitution of two residues in the measles virus nucleoprotein results in an impaired self-association. Virology 2002, 302, 420–432. [Google Scholar] [CrossRef] [PubMed]
- Bankamp, B.; Horikami, S.M.; Thompson, P.D.; Huber, M.; Billeter, M.; Moyer, S.A. Domains of the measles virus N protein required for binding to P protein and self-assembly. Virology 1996, 216, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Liston, P.; Batal, R.; DiFlumeri, C.; Briedis, D.J. Protein interaction domains of the measles virus nucleocapsid protein (NP). Arch. Virol. 1997, 142, 305–321. [Google Scholar] [CrossRef] [PubMed]
- Longhi, S.; Receveur-Brechot, V.; Karlin, D.; Johansson, K.; Darbon, H.; Bhella, D.; Yeo, R.; Finet, S.; Canard, B. The C-terminal domain of the measles virus nucleoprotein is intrinsically disordered and folds upon binding to the C-terminal moiety of the phosphoprotein. J. Biol. Chem. 2003, 278, 18638–18648. [Google Scholar] [CrossRef] [PubMed]
- Bhella, D.; Ralph, A.; Yeo, R.P. Conformational flexibility in recombinant measles virus nucleocapsids visualised by cryo-negative stain electron microscopy and real-space helical reconstruction. J. Mol. Biol. 2004, 340, 319–331. [Google Scholar] [CrossRef] [PubMed]
- Schoehn, G.; Mavrakis, M.; Albertini, A.; Wade, R.; Hoenger, A.; Ruigrok, R.W. The 12Å structure of trypsin-treated measles virus N–RNA. J. Mol. Biol. 2004, 339, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Tawar, R.G.; Duquerroy, S.; Vonrhein, C.; Varela, P.F.; Damier-Piolle, L.; Castagné, N.; MacLellan, K.; Bedouelle, H.; Bricogne, G.; Bhella, D.; et al. 3D structure of a nucleocapsid-like nucleoprotein-RNA complex of respiratory syncytial virus. Science 2009, 326, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Alayyoubi, M.; Leser, G.P.; Kors, C.A.; Lamb, R.A. Structure of the paramyxovirus parainfluenza virus 5 nucleoprotein-RNA complex. Proc. Natl. Acad. Sci. USA 2015, 112, E1792–E1799. [Google Scholar] [CrossRef] [PubMed]
- Desfosses, A.; Goret, G.; Farias Estrozi, L.; Ruigrok, R.W.; Gutsche, I. Nucleoprotein-RNA orientation in the measles virus nucleocapsid by three-dimensional electron microscopy. J. Virol. 2011, 85, 1391–1395. [Google Scholar] [CrossRef] [PubMed]
- Communie, G.; Habchi, J.; Yabukarski, F.; Blocquel, D.; Schneider, R.; Tarbouriech, N.; Papageorgiou, N.; Ruigrok, R.W.; Jamin, M.; Ringkjøbing-Jensen, M.; et al. Atomic resolution description of the interaction between the nucleoprotein and phosphoprotein of hendra virus. PLoS Pathog. 2013, 9, e1003631. [Google Scholar] [CrossRef] [PubMed]
- Baronti, L.; Erales, J.; Habchi, J.; Felli, I.C.; Pierattelli, R.; Longhi, S. Dynamics of the intrinsically disordered C-terminal domain of the nipah virus nucleoprotein and interaction with the X domain of the phosphoprotein as unveiled by NMR spectroscopy. ChemBioChem 2015, 16, 268–276. [Google Scholar] [CrossRef] [PubMed]
- Gutsche, I.; Desfosses, A.; Effantin, G.; Ling, W.L.; Haupt, M.; Ruigrok, R.W.; Sachse, C.; Schoehn, G. Near-atomic cryo-EM structure of the helical measles virus nucleocapsid. Science 2015, 348, 704–707. [Google Scholar] [CrossRef] [PubMed]
- Yabukarski, F.; Lawrence, P.; Tarbouriech, N.; Bourhis, J.M.; Delaforge, E.; Jensen, M.R.; Ruigrok, R.W.; Blackledge, M.; Volchkov, V.; Jamin, M. Structure of Nipah virus unassembled nucleoprotein in complex with its viral chaperone. Nat. Struct. Mol. Biol. 2014, 21, 754–759. [Google Scholar] [CrossRef] [PubMed]
- Garner, E.; Romero, P.; Dunker, A.K.; Brown, C.; Obradovic, Z. Predicting binding regions within disordered proteins. Genome Inform. 1999, 10, 41–50. [Google Scholar]
- Oldfield, C.J.; Cheng, Y.; Cortese, M.S.; Romero, P.; Uversky, V.N.; Dunker, A.K. Coupled folding and binding with α-helix-forming molecular recognition elements. Biochemistry 2005, 44, 12454–12470. [Google Scholar] [CrossRef] [PubMed]
- Mohan, A.; Oldfield, C.J.; Radivojac, P.; Vacic, V.; Cortese, M.S.; Dunker, A.K.; Uversky, V.N. Analysis of molecular recognition features (MoRFs). J. Mol. Biol. 2006, 362, 1043–1059. [Google Scholar] [CrossRef] [PubMed]
- Vacic, V.; Oldfield, C.J.; Mohan, A.; Radivojac, P.; Cortese, M.S.; Uversky, V.N.; Dunker, A.K. Characterization of molecular recognition features, morfs, and their binding partners. J. Proteome Res. 2007, 6, 2351–2366. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, N.; Kawano, M.; Tsurudome, M.; Nishio, M.; Ito, M.; Ohgimoto, S.; Suga, S.; Komada, H.; Ito, Y. Binding of the V proteins to the nucleocapsid proteins of human parainfluenza type 2 virus. Med. Microbiol. Immunol. 1996, 185, 89–94. [Google Scholar] [CrossRef] [PubMed]
- Karlin, D.; Belshaw, R. Detecting remote sequence homology in disordered proteins: Discovery of conserved motifs in the N-termini of Mononegavirales phosphoproteins. PLoS ONE 2012, 7, e31719. [Google Scholar] [CrossRef] [PubMed]
- Leyrat, C.; Jensen, M.R.; Ribeiro, E.A., Jr.; Gerard, F.C.; Ruigrok, R.W.; Blackledge, M.; Jamin, M. The N0-binding region of the vesicular stomatitis virus phosphoprotein is globally disordered but contains transient α-helices. Protein Sci. 2011, 20, 542–556. [Google Scholar] [CrossRef] [PubMed]
- Leyrat, C.; Yabukarski, F.; Tarbouriech, N.; Ribeiro, E.A., Jr.; Jensen, M.R.; Blackledge, M.; Ruigrok, R.W.; Jamin, M. Structure of the vesicular stomatitis virus N0-P complex. PLoS Pathog. 2011, 7, e1002248. [Google Scholar] [CrossRef] [PubMed]
- Sweetman, D.A.; Miskin, J.; Baron, M.D. Rinderpest virus C and V proteins interact with the major (L) component of the viral polymerase. Virology 2001, 281, 193–204. [Google Scholar] [CrossRef] [PubMed]
- Ding, B.; Zhang, G.; Yang, X.; Zhang, S.; Chen, L.; Yan, Q.; Xu, M.; Banerjee, A.K.; Chen, M. Phosphoprotein of human parainfluenza virus type 3 blocks autophagosome-lysosome fusion to increase virus production. Cell Host Microbe 2014, 15, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Blocquel, D.; Beltrandi, M.; Erales, J.; Barbier, P.; Longhi, S. Biochemical and structural studies of the oligomerization domainof the Nipah virus phosphoprotein: evidence for an elongated coiled-coil homotrimer. Virology 2013, 446, 162–172. [Google Scholar] [CrossRef] [PubMed]
- Beltrandi, M.; Blocquel, D.; Erales, J.; Barbier, P.; Cavalli, A.; Longhi, S. Insights into the coiled-coil organization of the Hendra virus phosphoprotein from combined biochemical and SAXS studies. Virology 2015, 477, 42–55. [Google Scholar] [CrossRef] [PubMed]
- Blocquel, D.; Habchi, J.; Durand, E.; Sevajol, M.; Ferron, F.; Erales, J.; Papageorgiou, N.; Longhi, S. Coiled-coil deformations in crystal structures: The measles virus phosphoprotein multimerization domain as an illustrative example. Acta Cryst. D 2014, 70, 1589–1603. [Google Scholar] [CrossRef] [PubMed]
- Blocquel, D.; Habchi, J.; Gruet, A.; Blangy, S.; Longhi, S. Compaction and binding properties of the intrinsically disordered C-terminal domain of henipavirus nucleoprotein as unveiled by deletion studies. Mol. Biosyst. 2012, 8, 392–410. [Google Scholar] [CrossRef] [PubMed]
- Tompa, P. Structure and Function of Intrinsically Disordered Proteins; CRC Press, Taylor & Francis Group: Boca Raton, FL, USA, 2010. [Google Scholar]
- Hazy, E.; Tompa, P. Limitations of induced folding in molecular recognition by intrinsically disordered proteins. Chemphyschem 2009, 10, 1415–1419. [Google Scholar] [CrossRef] [PubMed]
- Martinho, M.; Habchi, J.; El Habre, Z.; Nesme, L.; Guigliarelli, B.; Belle, V.; Longhi, S. Assessing induced folding within the intrinsically disordered C-terminal domain of the Henipavirus nucleoproteins by site directed spin labeling EPR spectroscopy. J. Biomol. Struct. Dyn. 2013, 31, 453–471. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chu, X.; Longhi, S.; Roche, P.; Han, W.; Wang, E.; Wang, J. Multiscaled exploration of coupled folding and binding of an intrinsically disordered molecular recognition element in measles virus nucleoprotein. Proc. Natl. Acad. Sci. USA 2013, 110, E3743–E3752. [Google Scholar] [CrossRef] [PubMed]
- Morin, B.; Bourhis, J.M.; Belle, V.; Woudstra, M.; Carrière, F.; Guigliarelli, B.; Fournel, A.; Longhi, S. Assessing induced folding of an intrinsically disordered protein by site-directed spin-labeling epr spectroscopy. J. Phys. Chem. B 2006, 110, 20596–20608. [Google Scholar] [CrossRef] [PubMed]
- Belle, V.; Rouger, S.; Costanzo, S.; Liquiere, E.; Strancar, J.; Guigliarelli, B.; Fournel, A.; Longhi, S. Mapping α-helical induced folding within the intrinsically disordered C-terminal domain of the measles virus nucleoprotein by site-directed spin-labeling EPR spectroscopy. Proteins Struct. Funct. Bioinform. 2008, 73, 973–988. [Google Scholar] [CrossRef] [PubMed]
- Gely, S.; Lowry, D.F.; Bernard, C.; Ringkjobing-Jensen, M.; Blackledge, M.; Costanzo, S.; Darbon, H.; Daughdrill, G.W.; Longhi, S. Solution structure of the C-terminal X domain of the measles virus phosphoprotein and interaction with the intrinsically disordered C-terminal domain of the nucleoprotein. J. Mol. Recognit. 2010, 23, 435–447. [Google Scholar] [CrossRef] [PubMed]
- Jensen, M.R.; Houben, K.; Lescop, E.; Blanchard, L.; Ruigrok, R.W.; Blackledge, M. Quantitative conformational analysis of partially folded proteins from residual dipolar couplings: Application to the molecular recognition element of sendai virus nucleoprotein. J. Am. Chem. Soc. 2008, 130, 8055–8061. [Google Scholar] [CrossRef] [PubMed]
- Serrano, L.; Fersht, A.R. Capping and α-helix stability. Nature 1989, 342, 296–299. [Google Scholar] [CrossRef] [PubMed]
- Habchi, J.; Blangy, S.; Mamelli, L.; Ringkjobing Jensen, M.; Blackledge, M.; Darbon, H.; Oglesbee, M.; Shu, Y.; Longhi, S. Characterization of the interactions between the nucleoprotein and the phosphoprotein of henipaviruses. J. Biol. Chem. 2011, 286, 13583–13602. [Google Scholar] [CrossRef] [PubMed]
- Bourhis, J.M.; Receveur-Bréchot, V.; Oglesbee, M.; Zhang, X.; Buccellato, M.; Darbon, H.; Canard, B.; Finet, S.; Longhi, S. The intrinsically disordered C-terminal domain of the measles virus nucleoprotein interacts with the C-terminal domain of the phosphoprotein via two distinct sites and remains predominantly unfolded. Protein Sci. 2005, 14, 1975–1992. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Glendening, C.; Linke, H.; Parks, C.L.; Brooks, C.; Udem, S.A.; Oglesbee, M. Identification and characterization of a regulatory domain on the carboxyl terminus of the measles virus nucleocapsid protein. J. Virol. 2002, 76, 8737–8746. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Bourhis, J.M.; Longhi, S.; Carsillo, T.; Buccellato, M.; Morin, B.; Canard, B.; Oglesbee, M. Hsp72 recognizes a P binding motif in the measles virus N protein C-terminus. Virology 2005, 337, 162–174. [Google Scholar] [CrossRef] [PubMed]
- Couturier, M.; Buccellato, M.; Costanzo, S.; Bourhis, J.M.; Shu, Y.; Nicaise, M.; Desmadril, M.; Flaudrops, C.; Longhi, S.; Oglesbee, M. High affinity binding between Hsp70 and the C-terminal domain of the measles virus nucleoprotein requires an Hsp40 co-chaperone. J. Mol. Recognit. 2010, 23, 301–315. [Google Scholar] [CrossRef] [PubMed]
- Tenoever, B.R.; Servant, M.J.; Grandvaux, N.; Lin, R.; Hiscott, J. Recognition of the measles virus nucleocapsid as a mechanism of IRF-3 activation. J. Virol. 2002, 76, 3659–3669. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Bourhis, J.M.; Chamontin, C.; Soriano, C.; Villet, S.; Costanzo, S.; Couturier, M.; Belle, V.; Fournel, A.; Darbon, H.; et al. The interaction between the measles virus nucleoprotein and the interferon regulator factor 3 relies on a specific cellular environment. Virol. J. 2009, 6, 59. [Google Scholar] [CrossRef] [PubMed]
- Laine, D.; Bourhis, J.; Longhi, S.; Flacher, M.; Cassard, L.; Canard, B.; Sautès-Fridman, C.; Rabourdin-Combe, C.; Valentin, H. Measles virus nucleoprotein induces cell proliferation arrest and apoptosis through NTAIL/NR and NCORE/FCγRIIB1 interactions, respectively. J. Gen. Virol. 2005, 86, 1771–1784. [Google Scholar] [CrossRef] [PubMed]
- Laine, D.; Trescol-Biémont, M.; Longhi, S.; Libeau, G.; Marie, J.; Vidalain, P.; Azocar, O.; Diallo, A.; Canard, B.; Rabourdin-Combe, C.; et al. Measles virus (MV) nucleoprotein binds to a novel cell surface receptor distinct from FcγRII via its C-terminal domain: Role in MV-induced immunosuppression. J. Virol. 2003, 77, 11332–11346. [Google Scholar] [CrossRef] [PubMed]
- Sato, H.; Masuda, M.; Miura, R.; Yoneda, M.; Kai, C. Morbillivirus nucleoprotein possesses a novel nuclear localization signal and a CRM1-independent nuclear export signal. Virology 2006, 352, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, M.; Takeda, M.; Shirogane, Y.; Nakatsu, Y.; Nakamura, T.; Yanagi, Y. The matrix protein of measles virus regulates viral RNA synthesis and assembly by interacting with the nucleocapsid protein. J. Virol. 2009, 83, 10374–10383. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, A.; Yoneda, M.; Ikeda, F.; Sugai, A.; Sato, H.; Kai, C. Peroxiredoxin 1 is required for efficient transcription and replication of measles virus. J. Virol. 2011, 85, 2247–2253. [Google Scholar] [CrossRef] [PubMed]
- De, B.P.; Banerjee, A.K. Involvement of actin microfilaments in the transcription/replication of human parainfluenza virus type 3: Possible role of actin in other viruses. Microsc. Res. Tech. 1999, 47, 114–123. [Google Scholar] [CrossRef]
- Moyer, S.A.; Baker, S.C.; Horikami, S.M. Host cell proteins required for measles virus reproduction. J. Gen. Virol. 1990, 71, 775–783. [Google Scholar] [CrossRef] [PubMed]
- Krumm, S.A.; Takeda, M.; Plemper, R.K. The measles virus nucleocapsid protein tail domain is dispensable for viral polymerase recruitment and activity. J. Biol. Chem. 2013, 288, 29943–29953. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Cortay, J.C.; Gerlier, D. Measles virus protein interactions in yeast: New findings and caveats. Virus Res. 2003, 98, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Liston, P.; DiFlumeri, C.; Briedis, D.J. Protein interactions entered into by the measles virus P, V, and C proteins. Virus Res. 1995, 38, 241–259. [Google Scholar] [CrossRef]
- Curran, J.; Pelet, T.; Kolakofsky, D. An acidic activation-like domain of the sendai virus P protein is required for RNA synthesis and encapsidation. Virology 1994, 202, 875–884. [Google Scholar] [CrossRef] [PubMed]
- Curran, J.; Marq, J.B.; Kolakofsky, D. An N-terminal domain of the sendai paramyxovirus P protein acts as a chaperone for the NP protein during the nascent chain assembly step of genome replication. J. Virol. 1995, 69, 849–855. [Google Scholar] [PubMed]
- Tarbouriech, N.; Curran, J.; Ruigrok, R.W.; Burmeister, W.P. Tetrameric coiled coil domain of sendai virus phosphoprotein. Nat. Struct. Biol. 2000, 7, 777–781. [Google Scholar] [PubMed]
- Rahaman, A.; Srinivasan, N.; Shamala, N.; Shaila, M.S. Phosphoprotein of the rinderpest virus forms a tetramer through a coiled coil region important for biological function. A structural insight. J. Biol. Chem. 2004, 279, 23606–23614. [Google Scholar] [CrossRef] [PubMed]
- Llorente, M.T.; Barreno-Garcia, B.; Calero, M.; Camafeita, E.; Lopez, J.A.; Longhi, S.; Ferron, F.; Varela, P.F.; Melero, J.A. Structural analysis of the human respiratory syncitial virus phosphoprotein: Characterization of an α-helical domain involved in oligomerization. J. Gen. Virol. 2006, 87, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Cox, R.; Green, T.J.; Purushotham, S.; Deivanayagam, C.; Bedwell, G.J.; Prevelige, P.E.; Luo, M. Structural and functional characterization of the mumps virus phosphoprotein. J. Virol. 2013, 87, 7558–7568. [Google Scholar] [CrossRef] [PubMed]
- Leyrat, C.; Renner, M.; Harlos, K.; Grimes, J.M. Solution and crystallographic structures of the central region of the phosphoprotein from human metapneumovirus. PLoS ONE 2013, 8, e80371. [Google Scholar] [CrossRef] [PubMed]
- Bruhn-Johannsen, J.F.; Barnett, K.; Bibby, J.; Thomas, J.; Keegan, R.; Rigden, D.; Bornholdt, Z.A.; Saphire, E.O. Crystal structure of the Nipah virus phosphoprotein tetramerization domain. J. Virol. 2014, 88, 758–762. [Google Scholar] [CrossRef] [PubMed]
- Dutta, K.; Alexandrov, A.; Huang, H.; Pascal, S.M. pH-induced folding of an apoptotic coiled coil. Protein Sci. 2001, 10, 2531–2540. [Google Scholar] [CrossRef] [PubMed]
- Lupas, A.N.; Gruber, M. The structure of α-helical coiled coils. Adv. Protein Chem. 2005, 70, 37–78. [Google Scholar] [PubMed]
- Oshaben, K.M.; Salari, R.; McCaslin, D.R.; Chong, L.T.; Horne, W.S. The native GCN4 leucine-zipper domain does not uniquely specify a dimeric oligomerization state. Biochemistry 2012, 51, 9581–9591. [Google Scholar] [CrossRef] [PubMed]
- Curran, J. A role for the sendai virus p protein trimer in RNA synthesis. J. Virol. 1998, 72, 4274–4280. [Google Scholar] [PubMed]
- Curran, J.; Boeck, R.; Lin-Marq, N.; Lupas, A.; Kolakofsky, D. Paramyxovirus phosphoproteins form homotrimers as determined by an epitope dilution assay, via predicted coiled coils. Virology 1995, 214, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Kingston, R.L.; Gay, L.S.; Baase, W.S.; Matthews, B.W. Structure of the nucleocapsid-binding domain from the mumps virus polymerase; an example of protein folding induced by crystallization. J. Mol. Biol. 2008, 379, 719–731. [Google Scholar] [CrossRef] [PubMed]
- Yegambaram, K.; Bulloch, E.M.; Kingston, R.L. Protein domain definition should allow for conditional disorder. Protein Sci. 2013, 22, 1502–1518. [Google Scholar] [CrossRef] [PubMed]
- D’Urzo, A.; Konijnenberg, A.; Rossetti, G.; Habchi, J.; Li, J.; Carloni, P.; Sobott, F.; Longhi, S.; Grandori, R. Molecular basis for structural heterogeneity of an intrinsically disordered protein bound to a partner by combined ESI-IM-MS and modeling. J. Am. Soc. Mass Spectrom. 2015, 26, 472–481. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Delano, W.L. The pymol molecular graphics system. Proteins Struct. Funct. Bioinform. 2002, 30, 442–454. [Google Scholar]
- Erales, J.; Beltrandi, M.; Roche, J.; Maté, M.; Longhi, S. Insights into the hendra virus NTAIL-XD complex: Evidence for a parallel organization of the helical more at the XD surface stabilized by a combination of hydrophobic and polar interactions. Biochim. Biophys. Acta 2015, 1854, 1038–1053. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, G.; Haran, G.; Zhou, H.X. Fundamental aspects of protein-protein association kinetics. Chem. Rev. 2009, 109, 839–860. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Yuwen, T.; Zhu, F.; Skrynnikov, N.R. Role of electrostatic interactions in binding of peptides and intrinsically disordered proteins to their folded targets. 1. NMR and MD characterization of the complex between the c-Crk N-SH3 domain and the peptide sos. Biochemistry 2014, 53, 6473–6495. [Google Scholar] [CrossRef] [PubMed]
- Dosnon, M.; Bonetti, D.; Morrone, A.; Erales, J.; di Silvio, E.; Longhi, S.; Gianni, S. Demonstration of a folding after binding mechanism in the recognition between the measles virus NTAIL and X domains. ACS Chem. Biol. 2015, 10, 795–802. [Google Scholar] [CrossRef] [PubMed]
- Kavalenka, A.; Urbancic, I.; Belle, V.; Rouger, S.; Costanzo, S.; Kure, S.; Fournel, A.; Longhi, S.; Guigliarelli, B.; Strancar, J. Conformational analysis of the partially disordered measles virus NTAIL-XD complex by SDSL EPR spectroscopy. Biophys. J. 2010, 98, 1055–1064. [Google Scholar] [CrossRef] [PubMed]
- Belle, V.; Rouger, S.; Costanzo, S.; Longhi, S.; Fournel, A. Site-directed spin labeling EPR spectroscopy. In Instrumental Analysis of Intrinsically Disordered Proteins: Assessing Structure and Conformation; Uversky, V.N., Longhi, S., Eds.; John Wiley and Sons: Hoboken, NJ, USA, 2010. [Google Scholar]
- Meszaros, B.; Tompa, P.; Simon, I.; Dosztanyi, Z. Molecular principles of the interactions of disordered proteins. J. Mol. Biol. 2007, 372, 549–561. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.D.; Ma, B.; Kumar, S.; Wolfson, H.; Nussinov, R. Protein folding: Binding of conformationally fluctuating building blocks via population selection. Crit. Rev. Biochem. Mol. Biol. 2001, 36, 399–433. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.J.; Ma, B.; Sham, Y.Y.; Kumar, S.; Nussinov, R. Structured disorder and conformational selection. Proteins Struct. Funct. Bioinform. 2001, 44, 418–427. [Google Scholar] [CrossRef] [PubMed]
- Espinoza-Fonseca, L.M. Reconciling binding mechanisms of intrinsically disordered proteins. Biochem. Biophys. Res. Commun. 2009, 382, 479–482. [Google Scholar] [CrossRef] [PubMed]
- Shoemaker, B.A.; Portman, J.J.; Wolynes, P.G. Speeding molecular recognition by using the folding funnel: The fly-casting mechanism. Proc. Natl. Acad. Sci. USA 2000, 97, 8868–8873. [Google Scholar] [CrossRef] [PubMed]
- Cox, R.; Pickar, A.; Qiu, S.; Tsao, J.; Rodenburg, C.; Dokland, T.; Elson, A.; He, B.; Luo, M. Structural studies on the authentic mumps virus nucleocapsid showing uncoiling by the phosphoprotein. Proc. Natl. Acad. Sci. USA 2014, 111, 15208–15213. [Google Scholar] [CrossRef] [PubMed]
- Oglesbee, M.; Ringler, S.; Krakowka, S. Interaction of canine distemper virus nucleocapsid variants with 70K heat-shock proteins. J. Gen. Virol. 1990, 71, 1585–1590. [Google Scholar] [CrossRef] [PubMed]
- Oglesbee, M.; Tatalick, L.; Rice, J.; Krakowka, S. Isolation and characterization of canine distemper virus nucleocapsid variants. J. Gen. Virol. 1989, 70, 2409–2419. [Google Scholar] [CrossRef] [PubMed]
- Brunel, J.; Chopy, D.; Dosnon, M.; Bloyet, L.M.; Devaux, P.; Urzua, E.; Cattaneo, R.; Longhi, S.; Gerlier, D. Sequence of events in measles virus replication: Role of phosphoprotein-nucleocapsid interactions. J. Virol. 2014, 88, 10851–10863. [Google Scholar] [CrossRef] [PubMed]
- Shu, Y.; Habchi, J.; Costanzo, S.; Padilla, A.; Brunel, J.; Gerlier, D.; Oglesbee, M.; Longhi, S. Plasticity in structural and functional interactions between the phosphoprotein and nucleoprotein of measles virus. J. Biol. Chem. 2012, 287, 11951–11967. [Google Scholar] [CrossRef] [PubMed]
- Fuxreiter, M.; Tompa, P. Fuzzy interactome: The limitations of models in molecular biology. Trends Biochem. Sci. 2009, 34, 3. [Google Scholar] [CrossRef]
- Fuxreiter, M. Fuzziness: Linking regulation to protein dynamics. Mol. Biosyst. 2012, 8, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Longhi, S. The measles virus NTAIL-XD complex: An illustrative example of fuzziness. Adv. Exp. Med. Biol. 2012, 725, 126–141. [Google Scholar] [PubMed]
- Carsillo, T.; Zhang, X.; Vasconcelos, D.; Niewiesk, S.; Oglesbee, M. A single codon in the nucleocapsid protein C terminus contributes to in vitro and in vivo fitness of edmonston measles virus. J. Virol. 2006, 80, 2904–2912. [Google Scholar] [CrossRef] [PubMed]
- Oglesbee, M. Nucleocapsid protein interactions with the major inducible heat shock protein. In Measles Virus Nucleoprotein; Longhi, S., Ed.; Nova Publishers Inc.: Hauppage, NY, USA, 2007; pp. 53–98. [Google Scholar]
- Carsillo, T.; Traylor, Z.; Choi, C.; Niewiesk, S.; Oglesbee, M. Hsp72, a host determinant of measles virus neurovirulence. J. Virol. 2006, 80, 11031–11039. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.; Vucetic, S.; Iakoucheva, L.M.; Oldfield, C.J.; Dunker, A.K.; Obradovic, Z.; Uversky, V.N. Functional anthology of intrinsic disorder. 3. Ligands, post-translational modifications, and diseases associated with intrinsically disordered proteins. J. Proteome Res. 2007, 6, 1917–1932. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, K.; Sato, H.; Inoue, Y.; Watanabe, A.; Yoneda, M.; Ikeda, F.; Fujita, K.; Fukuda, H.; Takamura, C.; Kozuka-Hata, H.; et al. Phosphorylation of measles virus nucleoprotein upregulates the transcriptional activity of minigenomic RNA. Proteomics 2008, 8, 1871–1879. [Google Scholar] [CrossRef] [PubMed]
- Sugai, A.; Sato, H.; Yoneda, M.; Kai, C. Phosphorylation of measles virus nucleoprotein affects viral growth by changing gene expression and genomic RNA stability. J. Virol. 2013, 87, 11684–11692. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Sato, H.; Hagiwara, K.; Watanabe, A.; Sugai, A.; Ikeda, F.; Kozuka-Hata, H.; Oyama, M.; Yoneda, M.; Kai, C. Determination of a phosphorylation site in Nipah virus nucleoprotein and its involvement in virus transcription. J. Gen. Virol. 2011, 92, 2133–2141. [Google Scholar] [CrossRef] [PubMed]
- Gruet, A.; Longhi, S.; Bignon, C. One-step generation of error-prone PCR libraries using Gateway® technology. Microb. Cell Fact. 2012, 11, 14. [Google Scholar] [CrossRef] [PubMed]
- Pancsa, R.; Fuxreiter, M. Interactions via intrinsically disordered regions: What kind of motifs? IUBMB Life 2012, 64, 513–520. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.G.; Magliery, T.J.; Regan, L. Detecting protein–protein interactions with GFP-fragment reassembly. Nat. Methods 2004, 1, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Gruet, A.; Dosnon, M.; Vassena, A.; Lombard, V.; Gerlier, D.; Bignon, C.; Longhi, S. Dissecting partner recognition by an intrinsically disordered protein using descriptive random mutagenesis. J. Mol. Biol. 2013, 425, 3495–3509. [Google Scholar] [CrossRef] [PubMed]
- Sebolt-Leopold, J.S.; English, J.M. Mechanisms of drug inhibition of signalling molecules. Nature 2006, 441, 457–462. [Google Scholar] [CrossRef] [PubMed]
- Lagerstrom, M.C.; Schioth, H.B. Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat. Rev. Drug Discov. 2008, 7, 339–357. [Google Scholar] [CrossRef] [PubMed]
- Betzi, S.; Restouin, A.; Opi, S.; Arold, S.T.; Parrot, I.; Guerlesquin, F.; Morelli, X.; Collette, Y. Protein protein interaction inhibition (2P2I) combining high throughput and virtual screening: Application to the HIV-1 Nef protein. Proc. Natl. Acad. Sci. USA 2007, 104, 19256–19261. [Google Scholar] [CrossRef] [PubMed]
- Galloux, M.; Gabiane, G.; Sourimant, J.; Richard, C.A.; England, P.; Moudjou, M.; Aumont-Nicaise, M.; Fix, J.; Rameix-Welti, M.A.; Eleouet, J.F. Identification and characterization of the binding site of the respiratory syncytial virus phosphoprotein to RNA-free nucleoprotein. J. Virol. 2015, 89, 3484–3496. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Legall, T.; Oldfield, C.J.; Mueller, J.P.; Van, Y.Y.; Romero, P.; Cortese, M.S.; Uversky, V.N.; Dunker, A.K. Rational drug design via intrinsically disordered protein. Trends Biotechnol. 2006, 24, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Targeting intrinsically disordered proteins in neurodegenerative and protein dysfunction diseases: Another illustration of the D2 concept. Expert Rev. Proteomics 2010, 7, 543–564. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Uversky, V.N. Drugs for “protein clouds”: Targeting intrinsically disordered transcription factors. Curr. Opin. Pharmacol. 2010, 10, 782–788. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Intrinsically disordered proteins and novel strategies for drug discovery. Expert Opin. Drug Discov. 2012, 7, 475–488. [Google Scholar] [CrossRef] [PubMed]
- Lo Conte, L.; Chothia, C.; Janin, J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 1999, 285, 2177–2198. [Google Scholar] [CrossRef] [PubMed]
- Gunasekaran, K.; Tsai, C.J.; Nussinov, R. Analysis of ordered and disordered protein complexes reveals structural features discriminating between stable and unstable monomers. J. Mol. Biol. 2004, 341, 1327–1341. [Google Scholar] [CrossRef] [PubMed]
- Klein, C.; Vassilev, L.T. Targeting the p53-MDM2 interaction to treat cancer. Br. J. Cancer 2004, 91, 1415–1419. [Google Scholar] [CrossRef] [PubMed]
- Vassilev, L.T. Small-molecule antagonists of p53-MDM2 binding: Research tools and potential therapeutics. Cell Cycle 2004, 3, 419–421. [Google Scholar] [CrossRef] [PubMed]
- Vassilev, L.T.; Vu, B.T.; Graves, B.; Carvajal, D.; Podlaski, F.; Filipovic, Z.; Kong, N.; Kammlott, U.; Lukacs, C.; Klein, C.; et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 2004, 303, 844–848. [Google Scholar] [PubMed]
- Dey, S.; Pal, A.; Chakrabarti, P.; Janin, J. The subunit interfaces of weakly associated homodimeric proteins. J. Mol. Biol. 2010, 398, 146–160. [Google Scholar] [CrossRef] [PubMed]
- Bourgeas, R.; Basse, M.J.; Morelli, X.; Roche, P. Atomic analysis of protein–protein interfaces with known inhibitors: The 2P2I database. PLoS ONE 2010, 5, e9598. [Google Scholar] [CrossRef] [PubMed]
- Jordan, I.K.; Sutter, B.A.; McClure, M.A. Molecular evolution of the Paramyxoviridae and Rhabdoviridae multiple-protein-encoding P gene. Mol. Biol. Evol. 2000, 17, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Narechania, A.; Terai, M.; Burk, R.D. Overlapping reading frames in closely related human papillomaviruses result in modular rates of selection within E2. J. Gen. Virol. 2005, 86, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Rancurel, C.; Khosravi, M.; Dunker, K.A.; Romero, P.R.; Karlin, D. Overlapping genes produce proteins with unusual sequence properties and offer insight into de novo protein creation. J. Virol. 2009, 83, 10719–10736. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, E.; Tompa, P.; Liliom, K.; Kalmar, L. Dual coding in alternative reading frames correlates with intrinsic protein disorder. Proc. Natl. Acad. Sci. USA 2010, 107, 5429–5434. [Google Scholar] [CrossRef] [PubMed]
- Bourhis, J.M.; Canard, B.; Longhi, S. Structural disorder within the replicative complex of measles virus: Functional implications. Virology 2006, 344, 94–110. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.; Blocquel, D.; Habchi, J.; Uversky, A.V.; Kurgan, L.; Uversky, V.N.; Longhi, S. Structural disorder in viral proteins. Chem. Rev. 2014, 114, 6880–6911. [Google Scholar] [CrossRef] [PubMed]
- Tokuriki, N.; Oldfield, C.J.; Uversky, V.N.; Berezovsky, I.N.; Tawfik, D.S. Do viral proteins possess unique biophysical features? Trends Biochem. Sci. 2009, 34, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.; Williams, R.W.; Oldfield, C.J.; Goh, G.K.; Dunker, A.K.; Uversky, V.N. Viral disorder or disordered viruses: Do viral proteins possess unique features? Protein Pept. Lett. 2010, 17, 932–951. [Google Scholar] [CrossRef] [PubMed]
- Davey, N.E.; van Roey, K.; Weatheritt, R.J.; Toedt, G.; Uyar, B.; Altenberg, B.; Budd, A.; Diella, F.; Dinkel, H.; Gibson, T.J. Attributes of short linear motifs. Mol. Biosyst. 2012, 8, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Davey, N.E.; Trave, G.; Gibson, T.J. How viruses hijack cell regulation. Trends Biochem. Sci. 2011, 36, 159–169. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).