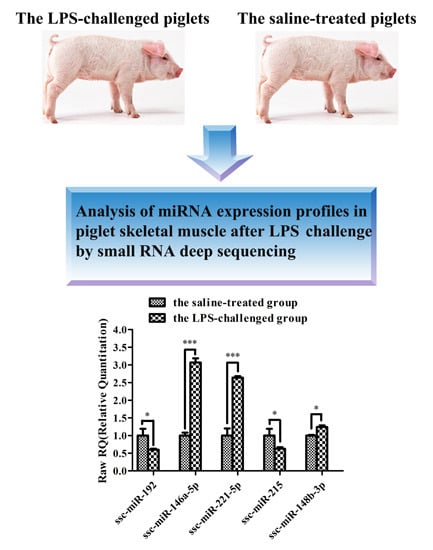
Analysis of MicroRNA Expression Profiles in Weaned Pig Skeletal Muscle after Lipopolysaccharide Challenge
Abstract
:1. Introduction
2. Results and Discussion
2.1. General Analysis of Small RNAs



| Classes of Reads | The LPS-Challenged Group | The Saline-Treated Group | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| L1 Library | L2 Library | L3 Library | L4 Library | L5 Library | L6 Library | |||||||
| Total | Unique | Total | Unique | Total | Unique | Total | Unique | Total | Unique | Total | Unique | |
| Clean date | 8,166,036 | 424,320 | 8,578,151 | 393,231 | 7,974,032 | 426,707 | 8,261,735 | 532,894 | 7,740,201 | 456,559 | 9,095,380 | 536,545 |
| Genome | 7,392,469 | 306,807 | 7,875,613 | 288,075 | 7,280,396 | 316,514 | 7,339,331 | 383,868 | 6,784,012 | 329,527 | 7,966,211 | 395,908 |
| miRBase (mature) | 3,979,370 | 7498 | 4,339,317 | 7753 | 3,940,011 | 7770 | 3,609,976 | 7597 | 3,995,522 | 8251 | 5,332,846 | 8842 |
| Rfam | 1,972,825 | 71,199 | 2,119,590 | 65,099 | 1,847,538 | 74,226 | 2,269,168 | 81,568 | 1,466,801 | 85,054 | 1,304,709 | 86,567 |
| RepBase | 166,565 | 28,787 | 132,226 | 25,094 | 157,402 | 30,124 | 259,354 | 31,758 | 180,018 | 30,089 | 158,490 | 35,920 |
| mRNA/EST | 501,200 | 128,735 | 437,848 | 116,045 | 516,196 | 129,355 | 608,415 | 173,253 | 594,752 | 126,662 | 586,024 | 183,958 |
2.2. Identification of Known miRNAs

2.3. Identification of Novel miRNAs by miRDeep2

| miRDeep2 Score 1 | Novel miRNAs | Known miRNAs | - | ||||
|---|---|---|---|---|---|---|---|
| Predicted 2 | False Positives 3 | True Positives 4 | In Species 5 | In Data 6 | Detected 7 | Signal-to-Noise 8 | |
| 10 | 65 | 3 ± 1 | 62 ± 1 (96% ± 2%) | 411 | 304 | 222 (72%) | 43.7 |
| 9 | 68 | 3 ± 1 | 65 ± 1 (96% ± 2%) | 411 | 304 | 223 (73%) | 43.5 |
| 8 | 69 | 3 ± 2 | 66 ± 3 (96% ± 2%) | 411 | 304 | 223 (73%) | 42.7 |
| 7 | 73 | 3 ± 2 | 70 ± 2 (96% ± 2%) | 411 | 304 | 223 (73%) | 42 |
| 6 | 78 | 3 ± 2 | 75 ± 3 (96% ± 2%) | 411 | 304 | 223 (73%) | 39.7 |
| 5 | 126 | 6 ± 2 | 120 ± 3 (95% ± 2%) | 411 | 304 | 256 (83%) | 30.7 |
| 4 | 156 | 15 ± 4 | 141 ± 4 (90% ± 3%) | 411 | 304 | 259 (84%) | 17.3 |
| 3 | 170 | 42 ± 6 | 128 ± 5 (75% ± 4%) | 411 | 304 | 261 (85%) | 7.5 |
| 2 | 206 | 57 ± 7 | 149 ± 6 (72% ± 3%) | 411 | 304 | 265 (86%) | 6.2 |
| 1 | 300 | 81 ± 8 | 219 ± 7 (73% ± 3%) | 411 | 304 | 278 (91%) | 5.4 |
2.4. Differential Expression Analysis
| miRNAs ID | The LPS-Challenged Group | The Saline-Treated Group | FoldChange_Log2 | p-Value | ||||
|---|---|---|---|---|---|---|---|---|
| L1_TPM | L2_TPM | L3_TPM | L4_TPM | L5_TPM | L6_TPM | |||
| ssc-miR-221-5p | 33.69 | 64.15 | 41.10 | 10.94 | 6.95 | 6.59 | 2.50 | 1.65 × 10−11 |
| ssc-miR-146a-5p | 2.73 | 14.64 | 50.54 | 2.98 | 13.26 | 4.71 | 1.70 | 1.97 × 10−5 |
| ssc-miR-9860-5p | 8.50 | 9.58 | 12.18 | 3.65 | 4.10 | 3.53 | 1.42 | 1.22 × 10−3 |
| ssc-miR-148b-3p | 216.10 | 294.41 | 242.05 | 106.78 | 119.34 | 146.86 | 1.01 | 1.73 × 10−5 |
| ssc-miR-192 | 191.82 | 104.61 | 123.61 | 545.85 | 142.38 | 183.34 | −1.05 | 1.00 × 10−2 |
| 4_16129 | 9.41 | 3.46 | 7.92 | 27.19 | 13.58 | 9.41 | −1.27 | 4.33 × 10−2 |
| 13_5595 | 5.77 | 5.86 | 7.92 | 18.57 | 26.20 | 8.94 | −1.46 | 1.87 × 10−2 |
| ssc-miR-215 | 16.09 | 1.06 | 1.83 | 43.11 | 4.74 | 18.12 | −1.80 | 9.43 × 10−4 |
| ssc-miR-429 | 0.61 | 0.00 | 0.61 | 3.32 | 2.53 | 0.94 | −2.48 | 1.20 × 10−2 |

2.5. LPS Induces miR-146a-5p and miR-221-5p Expression in C2C12 Myotubes

3. Experimental Section
3.1. Animal and Tissue Collection
3.2. TNF-α Concentration in Plasma Measurements
3.3. Reverse Transcription qPCR
3.4. Small-RNA Library Construction and Deep Sequencing
3.5. Basic Data Processing
3.6. Differential Expression Analysis of miRNAs
3.7. Cell Culture
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lecker, S.H.; Goldberg, A.L.; Mitch, W.E. Protein degradation by the ubiquitin-proteasome pathway in normal and disease states. J. Am. Soc. Nephrol. 2006, 17, 1807–1819. [Google Scholar]
- Palus, S.; von Haehling, S.; Springer, J. Muscle wasting: An overview of recent developments in basic research. Int. J. Cardiol. 2014, 176, 640–644. [Google Scholar] [CrossRef]
- Frisard, M.I.; McMillan, R.P.; Marchand, J.; Wahlberg, K.A.; Wu, Y.; Voelker, K.A.; Heilbronn, L.; Haynie, K.; Muoio, B.; Li, L.; et al. Toll-like receptor 4 modulates skeletal muscle substrate metabolism. Am. J. Physiol. Endocrinol. Metab. 2010, 298, E988–E998. [Google Scholar] [CrossRef] [PubMed]
- Frost, R.A.; Lang, C.H. Regulation of muscle growth by pathogen-associated molecules. J. Anim. Sci. 2008, 86, E84–E93. [Google Scholar] [PubMed]
- Orellana, R.A.; O’Connor, P.M.; Nguyen, H.V.; Bush, J.A.; Suryawan, A.; Thivierge, M.C.; Fiorotto, M.L.; Davis, T.A. Endotoxemia reduces skeletal muscle protein synthesis in neonates. Am. J. Physiol. Endocrinol. Metab. 2002, 283, E909–E916. [Google Scholar] [CrossRef] [PubMed]
- Tarabees, R.; Hill, D.; Rauch, C.; Barrow, P.A.; Loughna, P.T. Endotoxin transiently inhibits protein synthesis through Akt and MAPK mediating pathways in C2C12 myotubes. Am. J. Physiol. Cell Physiol. 2011, 301, C895–C902. [Google Scholar] [CrossRef] [PubMed]
- Philippou, A.; Maridaki, M.; Theos, A.; Koutsilieris, M. Cytokines in muscle damage. Adv. Clin. Chem. 2012, 58, 49–87. [Google Scholar] [PubMed]
- Zoico, E.; Roubenoff, R. The role of cytokines in regulating protein metabolism and muscle function. Nutr. Rev. 2002, 60, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.P.; Chen, Y.; John, J.; Moylan, J.; Jin, B.; Mann, D.L.; Reid, M.B. TNF-α acts via p38 MAPK to stimulate expression of the ubiquitin ligase atrogin1/MAFbx in skeletal muscle. FASEB J. 2005, 19, 362–370. [Google Scholar] [CrossRef] [PubMed]
- Cai, D.; Frantz, J.D., Jr.; Tawa, N.E.; Melendez, P.A.; Oh, B.C.; Lidov, H.G.; Hasselgren, P.O.; Frontera, W.R.; Lee, J.; Glass, D.J.; et al. IKKβ/NF-κB activation causes severe muscle wasting in mice. Cell 2004, 119, 285–298. [Google Scholar] [PubMed]
- Doyle, A.; Zhang, G.; Abdel Fattah, E.A.; Eissa, N.T.; Li, Y.P. Toll-like receptor 4 mediates lipopolysaccharide-induced muscle catabolism via coordinate activation of ubiquitin-proteasome and autophagy-lysosome pathways. FASEB J. 2011, 25, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.H.; Liu, N.; van Rooij, E.; Olson, E.N. MicroRNA control of muscle development and disease. Curr. Opin. Cell Biol. 2009, 21, 461–469. [Google Scholar] [CrossRef] [PubMed]
- Guller, I.; Russell, A.P. MicroRNAs in skeletal muscle: Their role and regulation in development, disease and function. J. Physiol. 2010, 588, 4075–4087. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, I.; Eran, A.; Nishino, I.; Moggio, M.; Lamperti, C.; Amato, A.A.; Lidov, H.G.; Kang, P.B.; North, K.N.; Mitrani-Rosenbaum, S.; et al. Distinctive patterns of microRNA expression in primary muscular disorders. Proc. Natl. Acad. Sci. USA 2007, 104, 17016–17021. [Google Scholar] [CrossRef] [PubMed]
- Greco, S.; de Simone, M.; Colussi, C.; Zaccagnini, G.; Fasanaro, P.; Pescatori, M.; Cardani, R.; Perbellini, R.; Isaia, E.; Sale, P.; et al. Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB J. 2009, 23, 3335–3346. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.H.; Valdez, G.; Moresi, V.; Qi, X.; McAnally, J.; Elliott, J.L.; Bassel-Duby, R.; Sanes, J.R.; Olson, E.N. MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science 2009, 326, 1549–1554. [Google Scholar] [CrossRef] [PubMed]
- Hudson, M.B.; Rahnert, J.A.; Zheng, B.; Woodworth-Hobbs, M.E.; Franch, H.A.; Price, S.R. miR-182 attenuates atrophy-related gene expression by targeting FoxO3 in skeletal muscle. Am. J. Physiol. Cell Physiol. 2014, 307, C314–C319. [Google Scholar] [CrossRef] [PubMed]
- Wada, S.; Kato, Y.; Okutsu, M.; Miyaki, S.; Suzuki, K.; Yan, Z.; Schiaffino, S.; Asahara, H.; Ushida, T.; Akimoto, T. Translational suppression of atrophic regulators by microRNA-23a integrates resistance to skeletal muscle atrophy. J. Biol. Chem. 2011, 286, 38456–38465. [Google Scholar] [CrossRef] [PubMed]
- Moschos, S.A.; Williams, A.E.; Perry, M.M.; Birrell, M.A.; Belvisi, M.G.; Lindsay, M.A. Expression profiling in vivo demonstrates rapid changes in lung microRNA levels following lipopolysaccharide-induced inflammation but not in the anti-inflammatory action of glucocorticoids. BMC Genom. 2007, 8. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Liu, Z.; Yang, Y. In vitro screening of LPS-induced miRNAs in leukocytes derived from cord blood and their possible roles in regulating TLR signals. Pediatr. Res. 2014, 75, 595–602. [Google Scholar] [PubMed]
- Taganov, K.D.; Boldin, M.P.; Chang, K.J.; Baltimore, D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.G.; Yang, J.; Zheng, Y.; Jin, Y. miR-15a/16 regulates macrophage phagocytosis after bacterial infection. J. Immunol. 2014, 193, 4558–4567. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Kuang, W.; Hao, Y.; Zhang, D.; Lei, M.; Du, L.; Jiao, H.; Zhang, X.; Wang, F. Downregulation of miR-27a* and miR-532-5p and upregulation of miR-146a and miR-155 in LPS-induced RAW264.7 macrophage cells. Inflammation 2012, 35, 1308–1313. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Jing, Z.; Cheng, G. MicroRNAs: New regulators of Toll-like receptor signalling pathways. Biomed. Res. Int. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Takeda, K.; Akira, S. TLR signaling pathways. Semin. Immunol. 2004, 16, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Bodine, S.C.; Latres, E.; Baumhueter, S.; Lai, V.K.; Nunez, L.; Clarke, B.A.; Poueymirou, W.T.; Panaro, F.J.; Na, E.; Dharmarajan, K.; et al. Identification of ubiquitin ligases required for skeletal muscle atrophy. Science 2001, 294, 1704–1708. [Google Scholar] [PubMed]
- Moylan, J.S.; Reid, M.B. Oxidative stress, chronic disease, and muscle wasting. Muscle Nerve 2007, 35, 411–429. [Google Scholar] [CrossRef] [PubMed]
- Small, E.M.; O’Rourke, J.R.; Moresi, V.; Sutherland, L.B.; McAnally, J.; Gerard, R.D.; Richardson, J.A.; Olson, E.N. Regulation of PI3-kinase/Akt signaling by muscle-enriched microRNA-486. Proc. Natl. Acad. Sci. USA 2010, 107, 4218–4223. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, J.J.; Esser, K.A. MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J. Appl. Physiol. 2007, 102, 306–313. [Google Scholar] [CrossRef]
- Callis, T.E.; Deng, Z.; Chen, J.F.; Wang, D.Z. Muscling through the microRNA world. Exp. Biol. Med. 2008, 233, 131–138. [Google Scholar] [CrossRef] [PubMed]
- Zuo, J.; Wu, F.; Liu, Y.; Xiao, J.; Xu, M.; Yu, Q.; Xia, M.; He, X.; Zou, S.; Tan, H.; et al. MicroRNA transcriptome profile analysis in porcine muscle and the effect of miR-143 on the MYH7 gene and protein. PLoS ONE 2015, 10, e0124873. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Chen, Y.; Liu, X.; Ye, S.; Yu, K.; Huang, Z.; Yu, J.; Zhou, X.; Chen, H.; Mo, D. Integrative analysis of porcine microRNAome during skeletal muscle development. PLoS ONE 2013, 8, e72418. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Tang, Z.; Liu, H.; Wang, N.; Ju, H.; Li, K. Discovery of microRNAs associated with myogenesis by deep sequencing of serial developmental skeletal muscles in pigs. PLoS ONE 2012, 7, e52123. [Google Scholar] [CrossRef] [PubMed]
- Nagpal, N.; Kulshreshtha, R. miR-191: An emerging player in disease biology. Front. Genet. 2014, 5. [Google Scholar] [CrossRef]
- Zhou, W.; Yin, H.; Wang, T.; Liu, T.; Li, Z.; Yan, W.; Song, D.; Chen, H.; Chen, J.; Xu, W.; et al. miR-126-5p regulates osteolysis formation and stromal cell proliferation in giant cell tumor through inhibition of PTHrP. Bone 2014, 66, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, M.R.; Chen, W.; Adamidi, C.; Maaskola, J.; Einspanier, R.; Knespel, S.; Rajewsky, N. Discovering microRNAs from deep sequencing data using miRDeep. Nat. Biotechnol. 2008, 26, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Shiota, C.; Abe, T.; Kawai, N.; Ohno, A.; Teshima-Kondo, S.; Mori, H.; Terao, J.; Tanaka, E.; Nikawa, T. Flavones inhibit LPS-induced atrogin-1/MAFbx expression in mouse C2C12 skeletal myotubes. J. Nutr. Sci. Vitaminol. 2015, 61, 188–194. [Google Scholar] [PubMed]
- Saba, R.; Sorensen, D.L.; Booth, S.A. MicroRNA-146a: A dominant, negative regulator of the innate immune response. Front. Immunol. 2014, 5. [Google Scholar] [CrossRef]
- Lu, Y.; Cao, L.; Jiang, B.C.; Yang, T.; Gao, Y.J. MicroRNA-146a-5p attenuates neuropathic pain via suppressing TRAF6 signaling in the spinal cord. Brain Behav. Immun. 2015, 49, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.Y.; Xue, D.; Xie, Y.F.; Zhu, D.W.; Dong, Y.Y.; Wei, C.C.; Deng, J.Y. The negative feedback regulation of microRNA-146a in human periodontal ligament cells after Porphyromonas gingivalis lipopolysaccharide stimulation. Inflamm. Res. 2015, 64, 441–451. [Google Scholar] [CrossRef]
- Hou, J.; Wang, P.; Lin, L.; Liu, X.; Ma, F.; An, H.; Wang, Z.; Cao, X. MicroRNA-146a feedback inhibits RIG-I-dependent Type I IFN production in macrophages by targeting TRAF6, IRAK1, and IRAK2. J. Immunol. 2009, 183, 2150–2158. [Google Scholar] [PubMed]
- Nahid, M.A.; Pauley, K.M.; Satoh, M.; Chan, E.K. miR-146a is critical for endotoxin-induced tolerance: Implication in innate immunity. J. Biol. Chem. 2009, 284, 34590–34599. [Google Scholar] [CrossRef] [PubMed]
- Nahid, M.A.; Satoh, M.; Chan, E.K. Mechanistic role of microRNA-146a in endotoxin-induced differential cross-regulation of TLR signaling. J. Immunol. 2011, 186, 1723–1734. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Xu, C.; Kong, X.; Li, X.; Kong, X.; Wang, Y.; Ding, X.; Yang, Q. Trail resistance induces epithelial-mesenchymal transition and enhances invasiveness by suppressing PTEN via miR-221 in breast cancer. PLoS ONE 2014, 9, e99067. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, R.; Tomosugi, M.; Horinaka, M.; Sowa, Y.; Sakai, T. Metformin causes G1-phase arrest via down-regulation of miR-221 and enhances TRAIL sensitivity through DR5 up-pegulation in pancreatic cancer cells. PLoS ONE 2015, 10, e0125779. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.F.; Huang, J.; Li, H.; Zhang, C.; Huang, X.; Tong, G.; Xu, Y.Z. MicroRNA-221 regulates endothelial nitric oxide production and inflammatory response by targeting adiponectin receptor 1. Gene 2015, 565, 246–251. [Google Scholar] [PubMed]
- Ortega, F.J.; Moreno, M.; Mercader, J.M.; Moreno-Navarrete, J.M.; Fuentes-Batllevell, N.; Sabater, M.; Ricart, W.; Fernández-Real, J.M. Inflammation triggers specific microRNA profiles in human adipocytes and macrophages and in their supernatants. Clin. Epigenet. 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lu, J.; Shi, J.; Hou, Y.; Zhu, H.; Zhao, S.; Liu, H.; Ding, B.; Yin, Y.; Yi, G. Increased expression of the peroxisome proliferator-activated receptor gamma in the immune system of weaned pigs after Escherichia coli lipopolysaccharide injection. Vet. Immunol. Immunopathol. 2008, 124, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmitten, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆Ct method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Dhahbi, J.M.; Atamna, H.; Boffelli, D.; Magis, W.; Spindler, S.R.; Martin, D.I. Deep sequencing reveals novel microRNAs and regulation of microRNA expression during cell senescence. PLoS ONE 2011, 6, e20509. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.M.; Yi, S.K.; Zhong, J.; Nie, C.H.; Guan, N.N.; Chen, B.X.; Gao, Z.X. Identification of microRNA for intermuscular bone development in blunt snout bream (Megalobrama amblycephala). Int. J. Mol. Sci. 2015, 16, 10686–10703. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Fu, S.-L.; Liu, Y.; Liu, Y.-L.; Wang, W.-J. Analysis of MicroRNA Expression Profiles in Weaned Pig Skeletal Muscle after Lipopolysaccharide Challenge. Int. J. Mol. Sci. 2015, 16, 22438-22455. https://doi.org/10.3390/ijms160922438
Zhang J, Fu S-L, Liu Y, Liu Y-L, Wang W-J. Analysis of MicroRNA Expression Profiles in Weaned Pig Skeletal Muscle after Lipopolysaccharide Challenge. International Journal of Molecular Sciences. 2015; 16(9):22438-22455. https://doi.org/10.3390/ijms160922438
Chicago/Turabian StyleZhang, Jing, Shu-Lin Fu, Yan Liu, Yu-Lan Liu, and Wen-Jun Wang. 2015. "Analysis of MicroRNA Expression Profiles in Weaned Pig Skeletal Muscle after Lipopolysaccharide Challenge" International Journal of Molecular Sciences 16, no. 9: 22438-22455. https://doi.org/10.3390/ijms160922438