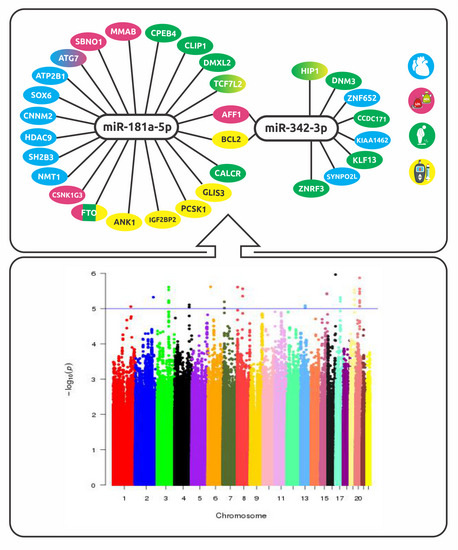
An Enrichment Analysis for Cardiometabolic Traits Suggests Non-Random Assignment of Genes to microRNAs
Abstract
:1. Introduction
2. Results
2.1. Retrieval of Cardiometabolic Genes
2.2. miRNA Target Prediction Databases
2.3. Pleiotropy
2.4. Enrichment Analysis
2.5. Biological Interpretation
3. Discussion
4. Methods
4.1. Retrieval of Cardiometabolic Genes
4.2. miRNA Target Prediction Databases
4.3. Pleiotropy
4.4. Enrichment Analysis
4.5. miRNA Expression and Association with Cardiometabolic Diseases
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CardiogramplusC4D | Coronary ARtery DIsease Genome-wide Replication and Meta-analysis (CARDIoGRAM) plus The Coronary Artery Disease (C4D) Genetics Consortium |
| DIAGRAM | DIAbetes Genetics Replication and Meta-analysis Consortium |
| FDR | false discovery rate |
| GIANT | Genetic Investigation of ANthropometric Traits Consortium |
| GLGC | Global Lipid Genetics Consortium |
| GWAS | genome-wide association studies |
| HOMA-B | homeostasis model assessment for beta cell function |
| HOMA-IR | homeostasis model assessment for insulin resistance |
| ICBP | International Consortium for Blood Pressure |
| MAGIC | the Meta-Analyses of Glucose and Insulin-related traits Consortium |
| mRNA | messenger RNA |
| miRNA | microRNA |
| NCBI | the National Centre for Biotechnology Information |
| RNA | ribonucleic acid |
| SNP | single-nucleotide polymorphism |
References
- Engels, B.M.; Hutvagner, G. Principles and Effects of microRNA-Mediated Post-Transcriptional Gene Regulation. Oncogene 2006, 25, 6163–6169. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Chou, C.; Chang, N.; Shrestha, S.; Hsu, S.; Lin, Y.; Lee, W.; Yang, C.; Hong, H.; Wei, T.; Tu, S. miRTarBase 2016: Updates to the Experimentally Validated miRNA-Target Interactions Database. Nucleic Acids Res. 2015, 44, D239–D247. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.A.; Li, Y.I.; Pritchard, J.K. An Expanded View of Complex Traits: From Polygenic to Omnigenic. Cell 2017, 169, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Deiuliis, J. MicroRNAs as Regulators of Metabolic Disease: Pathophysiologic Significance and Emerging Role as Biomarkers and Therapeutics. Int. J. Obes. 2015, 40, 88–101. [Google Scholar] [CrossRef] [PubMed]
- Evangelou, M.; Rendon, A.; Ouwehand, W.H.; Wernisch, L.; Dudbridge, F. Comparison of Methods for Competitive Tests of Pathway Analysis. PLoS ONE 2012, 7, e41018. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene Set Enrichment Analysis: A Knowledge-Based Approach for Interpreting Genome-Wide Expression Profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Glaab, E.; Baudot, A.; Krasnogor, N.; Schneider, R.; Valencia, A. EnrichNet: Network-Based Gene Set Enrichment Analysis. Bioinformatics 2012, 28, i451–i457. [Google Scholar] [CrossRef] [PubMed]
- Irizarry, R.A.; Wang, C.; Zhou, Y.; Speed, T.P. Gene Set Enrichment Analysis made Simple. Stat. Methods Med. Res. 2009, 18, 565–575. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Shi, B.; Wang, J.; Cao, Q.; Cui, Q. TAM: A Method for Enrichment and Depletion Analysis of a microRNA Category in a List of microRNAs. BMC Bioinform. 2010, 11, 419. [Google Scholar] [CrossRef] [PubMed]
- Çorapçıoğlu, M.E.; Oğul, H. miSEA: microRNA Set Enrichment Analysis. BioSystems 2015, 134, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Steinfeld, I.; Navon, R.; Ach, R.; Yakhini, Z. miRNA Target Enrichment Analysis Reveals Directly Active miRNAs in Health and Disease. Nucleic Acids Res. 2012, 41, e45. [Google Scholar] [CrossRef] [PubMed]
- Backes, C.; Khaleeq, Q.T.; Meese, E.; Keller, A. miEAA: microRNA Enrichment Analysis and Annotation. Nucleic Acids Res. 2016, 44, W110–W116. [Google Scholar] [CrossRef] [PubMed]
- Kirk, E.P.; Klein, S. Pathogenesis and Pathophysiology of the Cardiometabolic Syndrome. J. Clin. Hypertens. 2009, 11, 761–765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alberti, K.G.M.M.; Zimmet, P.; Shaw, J. Metabolic Syndrome—A New World-wide Definition. A Consensus Statement from the International Diabetes Federation. Diabet. Med. 2006, 23, 469–480. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Definition, Diagnosis and Classification of Diabetes Mellitus and its Complications: Report of A WHO Consultation. Part 1, Diagnosis and Classification of Diabetes Mellitus; World Health Organization: Geneva, Switzerland, 1999. [Google Scholar]
- Gami, A.S.; Witt, B.J.; Howard, D.E.; Erwin, P.J.; Gami, L.A.; Somers, V.K.; Montori, V.M. Metabolic Syndrome and Risk of Incident Cardiovascular Events and Death: A Systematic Review and Meta-Analysis of Longitudinal Studies. J. Am. Coll. Cardiol. 2007, 49, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Manolio, T.A.; Collins, F.S.; Cox, N.J.; Goldstein, D.B.; Hindorff, L.A.; Hunter, D.J.; McCarthy, M.I.; Ramos, E.M.; Cardon, L.R.; Chakravarti, A.; et al. Finding the Missing Heritability of Complex Diseases. Nature 2009, 461, 747–753. [Google Scholar] [CrossRef] [PubMed]
- CARDIoGRAMplusC4D Consortium. A Comprehensive 1000 Genomes-Based Genome-Wide Association Meta-Analysis of Coronary Artery Disease. Nat. Genet. 2015, 47, 1121–1130. [Google Scholar] [CrossRef] [PubMed]
- Ehret, G.B.; Ferreira, T.; Chasman, D.I.; Jackson, A.U.; Schmidt, E.M.; Johnson, T.; Thorleifsson, G.; Luan, J.; Donnelly, L.A.; Kanoni, S. The Genetics of Blood Pressure Regulation and its Target Organs from Association Studies in 342,415 Individuals. Nat. Genet. 2016, 48, 1171. [Google Scholar] [CrossRef] [PubMed]
- Global Lipids Genetics Consortium. Discovery and Refinement of Loci Associated with Lipid Levels. Nat. Genet. 2013, 45, 1274–1283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J. Genetic Studies of Body Mass Index Yield New Insights for Obesity Biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shungin, D.; Winkler, T.W.; Croteau-Chonka, D.C.; Ferreira, T.; Locke, A.E.; Mägi, R.; Strawbridge, R.J.; Pers, T.H.; Fischer, K.; Justice, A.E. New Genetic Loci Link Adipose and Insulin Biology to Body Fat Distribution. Nature 2015, 518, 187–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fuchsberger, C.; Flannick, J.; Teslovich, T.M.; Mahajan, A.; Agarwala, V.; Gaulton, K.J.; Ma, C.; Fontanillas, P.; Moutsianas, L.; McCarthy, D.J. The Genetic Architecture of Type 2 Diabetes. Nature 2016, 536, 41–47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walford, G.A.; Gustafsson, S.; Rybin, D.; Stancakova, A.; Chen, H.; Liu, C.T.; Hong, J.; Jensen, R.A.; Rice, K.; Morris, A.P.; et al. Genome-Wide Association Study of the Modified Stumvoll Insulin Sensitivity Index Identifies BCL2 and FAM19A2 as Novel Insulin Sensitivity Loci. Diabetes 2016, 65, 3200–3211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prokopenko, I.; Poon, W.; Mägi, R.; Prasad, R.; Salehi, S.A.; Almgren, P.; Osmark, P.; Bouatia-Naji, N.; Wierup, N.; Fall, T. A Central Role for GRB10 in Regulation of Islet Function in Man. PLoS Genet. 2014, 10, e1004235. [Google Scholar] [CrossRef] [PubMed]
- Scott, R.A.; Lagou, V.; Welch, R.P.; Wheeler, E.; Montasser, M.E.; Luan, J.; Mägi, R.; Strawbridge, R.J.; Rehnberg, E.; Gustafsson, S. Large-Scale Association Analyses Identify New Loci Influencing Glycemic Traits and Provide Insight into the Underlying Biological Pathways. Nat. Genet. 2012, 44, 991–1005. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soranzo, N.; Sanna, S.; Wheeler, E.; Gieger, C.; Radke, D.; Dupuis, J.; Bouatia-Naji, N.; Langenberg, C.; Prokopenko, I.; Stolerman, E.; et al. Common Variants at 10 Genomic Loci Influence Hemoglobin A(1)(C) Levels Via Glycemic and Nonglycemic Pathways. Diabetes 2010, 59, 3229–3239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strawbridge, R.J.; Dupuis, J.; Prokopenko, I.; Barker, A.; Ahlqvist, E.; Rybin, D.; Petrie, J.R.; Travers, M.E.; Bouatia-Naji, N.; Dimas, A.S.; et al. Genome-Wide Association Identifies Nine Common Variants Associated with Fasting Proinsulin Levels and Provides New Insights into the Pathophysiology of Type 2 Diabetes. Diabetes 2011, 60, 2624–2634. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dupuis, J.; Langenberg, C.; Prokopenko, I.; Saxena, R.; Soranzo, N.; Jackson, A.U.; Wheeler, E.; Glazer, N.L.; Bouatia-Naji, N.; Gloyn, A.L. New Genetic Loci Implicated in Fasting Glucose Homeostasis and their Impact on Type 2 Diabetes Risk. Nat. Genet. 2010, 42, 105–116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bush, W.S.; Moore, J.H. Genome-Wide Association Studies. PLoS Comput. Biol. 2012, 8, e1002822. [Google Scholar] [CrossRef] [PubMed]
- Aken, B.L.; Achuthan, P.; Akanni, W.; Amode, M.R.; Bernsdorff, F.; Bhai, J.; Billis, K.; Carvalho-Silva, D.; Cummins, C.; Clapham, P. Ensembl 2017. Nucleic Acids Res. 2016, 45, D635–D642. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Qiu, C.; Tu, J.; Geng, B.; Yang, J.; Jiang, T.; Cui, Q. HMDD V2. 0: A Database for Experimentally Supported Human microRNA and Disease Associations. Nucleic Acids Res. 2013, 42, D1070–D1074. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, N.; Leidinger, P.; Becker, K.; Backes, C.; Fehlmann, T.; Pallasch, C.; Rheinheimer, S.; Meder, B.; Stähler, C.; Meese, E. Distribution of miRNA Expression Across Human Tissues. Nucleic Acids Res. 2016, 44, 3865–3877. [Google Scholar] [CrossRef] [PubMed]
- Panwar, B.; Omenn, G.S.; Guan, Y. miRmine: A Database of Human miRNA Expression Profiles. Bioinformatics 2017, 33, 1554–1560. [Google Scholar] [CrossRef] [PubMed]
- Backes, C.; Kehl, T.; Stöckel, D.; Fehlmann, T.; Schneider, L.; Meese, E.; Lenhof, H.; Keller, A. miRPathDB: A New Dictionary on microRNAs and Target Pathways. Nucleic Acids Res. 2016, 45, D90–D96. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.; Leidinger, P.; Bauer, A.; ElSharawy, A.; Haas, J.; Backes, C.; Wendschlag, A.; Giese, N.; Tjaden, C.; Ott, K. Toward the Blood-Borne miRNome of Human Diseases. Nat. Methods 2011, 8, 841. [Google Scholar] [CrossRef] [PubMed]
- Riffo-Campos, Á.L.; Riquelme, I.; Brebi-Mieville, P. Tools for Sequence-Based miRNA Target Prediction: What to Choose? Int. J. Mol. Sci. 2016, 17, 1987. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting Effective microRNA Target Sites in Mammalian mRNAs. Elife 2015, 4, e05005. [Google Scholar] [CrossRef] [PubMed]
- Ghanbari, M. The Role of MicroRNAs in Age-Related Disorders: From Population-Based Genetic Studies to Experimental Validation; Erasmus University Rotterdam: Rotterdam, The Netherlands, 5 July 2017. [Google Scholar]
- Manolio, T.A. Genomewide Association Studies and Assessment of the Risk of Disease. N. Engl. J. Med. 2010, 363, 166–176. [Google Scholar] [CrossRef] [PubMed]
- 1000 Genomes Project Consortium; Abecasis, G.R.; Altshuler, D.; Auton, A.; Brooks, L.D.; Durbin, R.M.; Gibbs, R.A.; Hurles, M.E.; McVean, G.A. A Map of Human Genome Variation from Population-Scale Sequencing. Nature 2010, 467, 1061–1073. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hammond, S.M. An Overview of microRNAs. Adv. Drug Deliv. Rev. 2015, 87, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Khatri, P.; Sirota, M.; Butte, A.J. Ten Years of Pathway Analysis: Current Approaches and Outstanding Challenges. PLoS Comput. Biol. 2012, 8, e1002375. [Google Scholar] [CrossRef] [PubMed]
- Sudlow, C.; Gallacher, J.; Allen, N.; Beral, V.; Burton, P.; Danesh, J.; Downey, P.; Elliott, P.; Green, J.; Landray, M. UK Biobank: An Open Access Resource for Identifying the Causes of a Wide Range of Complex Diseases of Middle and Old Age. PLoS Med. 2015, 12, e1001779. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Zuo, Z.; Cai, G.; Kang, S.; Gao, X.; Li, T. miRecords: An Integrated Resource for microRNA–target Interactions. Nucleic Acids Res. 2008, 37, D105–D110. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V.; Bartel, B.; Bartel, D.P.; Burge, C.B.; Carrington, J.C.; Chen, X.; Dreyfuss, G.; Eddy, S.R.; Griffiths-Jones, S.; Marshall, M.; et al. A Uniform System for microRNA Annotation. RNA 2003, 9, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S.; Grocock, R.J.; Van Dongen, S.; Bateman, A.; Enright, A.J. miRBase: microRNA Sequences, Targets and Gene Nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.A. Statistical Methods for Research Workers; Genesis Publishing Pvt Ltd.: New Delhi, India, 1925. [Google Scholar]
- Holmans, P.; Green, E.K.; Pahwa, J.S.; Ferreira, M.A.; Purcell, S.M.; Sklar, P.; Owen, M.J.; O’Donovan, M.C.; Craddock, N.; Wellcome Trust Case-Control Consortium. Gene Ontology Analysis of GWA Study Data Sets Provides Insights into the Biology of Bipolar Disorder. Am. J. Hum. Genet. 2009, 85, 13–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B-Stat. Methodol. 1995, 57, 289–300. [Google Scholar]



| Cardiometabolic Trait | Consortium | Number of SNPs Associated * | Ref |
|---|---|---|---|
| trait group 1: coronary artery disease & blood pressure traits | |||
| coronary artery disease | CardiogramplusC4D | 51 | [19] |
| systolic blood pressure | ICBP | 54 | [20] |
| diastolic blood pressure | ICBP | 53 | [20] |
| hypertension | ICBP | 11 | [20] |
| trait group 2: lipid traits | |||
| total cholesterol | GLGC | 39 | [21] |
| triglyceride | GLGC | 35 | [21] |
| high-density lipoprotein cholesterol | GLGC | 60 | [21] |
| low-density lipoprotein cholesterol | GLGC | 30 | [21] |
| trait group 3: anthropometric traits | |||
| body mass index | GIANT | 108 | [22] |
| waist-hip ratio | GIANT | 39 | [23] |
| trait group 4: type 2 diabetes and glycaemic traits | |||
| type 2 diabetes | DIAGRAM | 18 | [24] |
| insulin sensitivity index (Stumvoll index) | MAGIC | 4 | [25] |
| insulin secretion during oral glucose tolerance test | MAGIC | 8 | [26] |
| fasting glucose | MAGIC | 36 | [27] |
| 2 h-glucose | MAGIC | 9 | [27] |
| fasting insulin | MAGIC | 26 | [27] |
| HbA1c | MAGIC | 11 | [28] |
| fasting pro-insulin | MAGIC | 9 | [29] |
| HOMA-B | MAGIC | 2 | [30] |
| HOMA-IR | MAGIC | 7 | [30] |
| Total | 610 | ||
| TargetScan (Predicted) | miRTarBase (Validated) | miRecords (Validated) | miRTarBase & miRecords * | |
|---|---|---|---|---|
| H0(a): the miRNA is not enriched within cardiometabolic genes | ||||
| Number of miRNAs # | 306 | 54 | 7 | 61 |
| Nominal p-value < 0.05 | 102 | 13 | 3 | 19 |
| FDR-adjusted p-value < 0.05 + | 29 | 0 | 3 | 0 |
| H0(b): the miRNA is not enriched within pleiotropic cardiometabolic genes | ||||
| Number of miRNAs # | 129 | - $ | - $ | - $ |
| Nominal p-value < 0.05 | 19 | - | - | - |
| FDR-adjusted p-value < 0.05 + | 0 | - | - | - |
| Cardiometabolic-Associated Diseases | Cardiometabolic Tissues with Evidence of miRNA Expression | |
|---|---|---|
| TargetScan | ||
| miR-1251-5p | - | pancreas |
| miR-125a-5p | heart failure, obesity, acute coronary syndrome, coronary atherosclerosis | liver, pancreas, adipocyte, myocardium |
| miR-125b-5p | myocardial infarction, atherosclerosis, cardiomyopathy, coronary artery disease, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-1271-5p | type 2 diabetes | liver, pancreas, adipocyte, myocardium |
| miR-141-3p | ischemic heart disease | liver, pancreas, adipocyte |
| miR-142-3p.2 | type 2 diabetes, heart failure, ischemic heart disease, atherosclerosis | liver, pancreas, adipocyte, myocardium |
| miR-150-5p | type 2 diabetes, myocardial infarction, obesity, hypertension, heart failure, atherosclerosis, cardiomyopathy | liver, pancreas, adipocyte, myocardium |
| miR-155-5p | type 2 diabetes, myocardial infarction, cardiomyopathy, coronary artery disease, heart failure, hypertension, acute coronary syndrome, atherosclerosis | liver, pancreas, adipocyte, myocardium |
| miR-181a-5p | myocardial infarction, atherosclerosis, cardiomegaly, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-181b-5p | atherosclerosis, diabetic cardiomyopathy, obesity, type 2 diabetes cardiovascular disease (unspecific) | liver, pancreas, adipocyte, myocardium |
| miR-181c-5p | heart failure | liver, pancreas, adipocyte, myocardium |
| miR-181d-5p | - | liver, pancreas, adipocyte, myocardium |
| miR-182-5p | atherosclerosis, type 2 diabetes, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-186-5p | myocardial infarction, type 2 diabetes, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-188-5p | cardiovascular disease (unspecific) | liver, pancreas, adipocyte, myocardium |
| miR-200a-3p | ischemic cardiomyopathy, type 2 diabetes, obesity | liver, pancreas, adipocyte |
| miR-200b-3p | type 2 diabetes, obesity | liver, pancreas, adipocyte, myocardium |
| miR-200c-3p | cardiovascular disease (unspecific), type 2 diabetes | liver, pancreas, adipocyte |
| miR-204-5p | type 2 diabetes, hypertension, myocardial infarction, cardiomyopathy, obesity, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-211-5p | heart failure | liver, pancreas, adipocyte, myocardium |
| miR-23a-3p | type 2 diabetes, cardiovascular disease (unspecific), cardiomyopathy, coronary artery disease, heart failure, myocardial infarction | liver, pancreas, adipocyte, myocardium |
| miR-23b-3p | atherosclerosis, type 2 diabetes, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-23c | - | liver, pancreas, adipocyte, myocardium |
| miR-342-3p | heart failure, atherosclerosis, obesity, type 2 diabetes, | liver, pancreas, adipocyte, myocardium |
| miR-371a-5p | cardiomyopathy | liver, pancreas, adipocyte, myocardium |
| miR-429 | type 2 diabetes, heart failure | liver, pancreas, adipocyte, myocardium |
| miR-485-5p | - | liver, pancreas, adipocyte, myocardium |
| miR-493-5p | - | liver, pancreas, adipocyte, myocardium |
| miR-96-5p | type 2 diabetes, cardiomyopathy | liver, pancreas, adipocyte, myocardium |
| miRecords | ||
| miR-181a | myocardial infarction, atherosclerosis, cardiomegaly, heart failure | liver, pancreas, adipocyte, myocardium (miR-181a-3p and miR-181a-5p) |
| miR-302d | - | - |
| miR-372 | heart failure | liver (miR-372-3p) |
| Target Gene for miRNA | Not a Target Gene for miRNA | Total | |
|---|---|---|---|
| number of cardiometabolic genes | a | b | a + b |
| number of non-cardiometabolic genes | c | d | c + d |
| a + c | b + d | N |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mustafa, R.; Ghanbari, M.; Evangelou, M.; Dehghan, A. An Enrichment Analysis for Cardiometabolic Traits Suggests Non-Random Assignment of Genes to microRNAs. Int. J. Mol. Sci. 2018, 19, 3666. https://doi.org/10.3390/ijms19113666
Mustafa R, Ghanbari M, Evangelou M, Dehghan A. An Enrichment Analysis for Cardiometabolic Traits Suggests Non-Random Assignment of Genes to microRNAs. International Journal of Molecular Sciences. 2018; 19(11):3666. https://doi.org/10.3390/ijms19113666
Chicago/Turabian StyleMustafa, Rima, Mohsen Ghanbari, Marina Evangelou, and Abbas Dehghan. 2018. "An Enrichment Analysis for Cardiometabolic Traits Suggests Non-Random Assignment of Genes to microRNAs" International Journal of Molecular Sciences 19, no. 11: 3666. https://doi.org/10.3390/ijms19113666