Abstract
Increasing agricultural productivity via modern breeding strategies is of prime interest to attain global food security. An array of biotic and abiotic stressors affect productivity as well as the quality of crop plants, and it is a primary need to develop crops with improved adaptability, high productivity, and resilience against these biotic/abiotic stressors. Conventional approaches to genetic engineering involve tedious procedures. State-of-the-art OMICS approaches reinforced with next-generation sequencing and the latest developments in genome editing tools have paved the way for targeted mutagenesis, opening new horizons for precise genome engineering. Various genome editing tools such as transcription activator-like effector nucleases (TALENs), zinc-finger nucleases (ZFNs), and meganucleases (MNs) have enabled plant scientists to manipulate desired genes in crop plants. However, these approaches are expensive and laborious involving complex procedures for successful editing. Conversely, CRISPR/Cas9 is an entrancing, easy-to-design, cost-effective, and versatile tool for precise and efficient plant genome editing. In recent years, the CRISPR/Cas9 system has emerged as a powerful tool for targeted mutagenesis, including single base substitution, multiplex gene editing, gene knockouts, and regulation of gene transcription in plants. Thus, CRISPR/Cas9-based genome editing has demonstrated great potential for crop improvement but regulation of genome-edited crops is still in its infancy. Here, we extensively reviewed the availability of CRISPR/Cas9 genome editing tools for plant biotechnologists to target desired genes and its vast applications in crop breeding research.
1. Introduction
Food security is the most crucial challenge in the current scenario of a rapidly growing global population. According to cautious estimates, the global population will escalate to ten billion by the end of 2050 and a 60–100% rise in global food production will be necessary [1]. Besides extreme weather, increasing biotic and abiotic stressors, a growing population, and shrinking availability of agricultural land and water resources are important constraints for food production and farming. Over the past few decades, improvements in crop plants have contributed by deciphering numerous biological mechanisms and elucidating the role of genetic and epigenetics factors [2]. Crop breeders and plant scientist are striving hard to understand mainly the genetic mechanism underlying unique plant responses towards environmental stressors. Recently, numerous novel genes and their regulatory pathways have been identified in plants [3,4]. For crop improvement and development of elite cultivars with increased productivity, a breeding strategy of “cross the elite with the elite and wait for the best” has been applied, concentrating on the genes linked with vital agronomic traits [5]. Classical plant breeding strategies for crop improvement are more challenging which take a long time for germplasm selection. On the other hand, modern tools for genome editing (GE) exhibit the capability of integrating a foreign gene into a predetermined site of the genome precisely, allowing for accurate substitution of an existing allele with an alternative one [6,7]. Genome editing has emerged as a tremendous strategy for efficient and targeted genome manipulations, especially for crops which have complex genomes and which are difficult to improve through conventional breeding approaches [6].
For basic as well as applied plant biology, the unstable and non-specific transgene incorporation in the host genome has been a matter of concern for edible crop species [8]. The discovery of programmed sequence-specific nucleases (SSNs) has facilitated precise gene editing. In both plant and animal systems, application of SSNs for accurate GE has been recognized as a breakthrough in genome engineering. The SSNs can be applied to produce several kinds of mutations, such as insertions, deletions, replacement, substitutions, integration of specific sequence of DNA at a desired locus, and site-directed substitutions across many organisms and cell types. Though all types of SSNs have unique features, the mechanism for producing double-strand breaks (DSBs) in the target DNA is similar for all. The DSBs created by SSNs are reconstructed via non-homologous end joining (NHEJ) or homology-directed recombination (HDR). Non-homologous end joining is an error-prone DNA repair mechanism that facilitates direct end-joining of DSBs without involving a homologous template and can generate insertions or deletions at target sites to develop gene knockouts. Additionally, NHEJ can also be applied to introduce insertions at the point of the DSB during operation of the repair mechanism. On the other hand, the HDR repair pathway is a highly accurate mechanism that needs a homologous template to mediate repair and can be used to attain precise changes like gene insertion and gene replacement [6,9,10]. As compared to transgenic strategies, which result in inadvertent gene insertions and sometimes random phenotypical characters, GE approaches produce well-defined mutants, proving GE as a powerful technique for plant breeding and functional genomics. In contrast to transgenic plants, genome-edited plants have the added benefit of site specificity [11]. In breeding programs, these improved plants can be proven useful and subsequent species can be employed reliably with less concerns and comparatively minor monitoring methods are needed in contrast to traditional genetically engineered plants [12].
2. Modern Trends in Plant Genome Editing
In recent years, many fascinating GE approaches have been established because of the advancements in molecular biology, which have permitted site-specific and accurate editing in many genomes [8]. In GE, engineered nucleases are composed of a sequence-specific DNA binding domain merged with a non-specific nucleases domain. Targeted genes can be precisely cleaved by such fused nucleases and nicks can be repaired with the help of HDR or NHEJ [13,14]. A vital strategy to execute targeted GE through SSNs is to generate DSBs at targeted sites; these nicks prompt the activation of the DNA repair mechanism through the HDR or NHEJ pathway [15]. The DNA repair system of the HDR pathways requires a homologous template to repair the DSB, whereas the two ends of DSBs are directly ligated in the NHEJ pathway [15]. Though NHEJ is more common, there are some flaws which make it undesirable in many studies. The major disadvantage of this process is that it produces insertions or deletions of different sizes during the repair mechanism, which may produce off-targets. In contrast to NHEJ, the repair mechanism via HDR is more accurate and reliable, which depends on homologous DNA to repair the DSB [16]. Thus, SSNs can be applied to manipulate the genomic sequences by targeted addition or deletion of specific nucleotides in the targeted locus [10].
Recently, great achievements have been made in the era of genome engineering with the development of meganulceases (MNs), zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), and clustered regularly interspaced short palindromic repeats/ CRISPR-associated protein 9 (CRISPR/Cas9). Experimental proofs gradually showed that these SSNs were not only used for gene insertion or inactivation, but also significantly enhanced the effectiveness of homologous recombination and, thus, allowed more precise gene replacement events. In 1993, Puchta and co-workers [17] provided the first evidence of homologous recombination in plant cells by using SSNs. After the discovery of ZFNs in 1996 by Kim and colleagues [18], extensive efforts have been made for progressive advancement with this tool, which offered a significant breakthrough in plant GE. In 2003, scientists were able to inactivate genes using ZFNs for the first time [19]. In 2005, the first SSN-based mutagenesis via ZFNs was carried out successfully in plants [20]. Therefore, ZFNs have the ability to produce site-specific DSBs and have many applications in genome engineering [21]. Later TALENs were included to the toolbox of SSNs for programmed genome engineering [22]. Genome manipulation through engineered nucleases had gained much importance by the end of 2011, and Nature Methods crowned it as the “Method of the Year”. In 2012, Science chose it as the “Breakthrough of the Year” due to the significant progress achieved in GE using TALENs. Recently, an emerging GE nuclease, “CRISPR/Cas9”, was added to the toolbox for editing nucleases. In 2013, the first CRISPR/Cas9-based GE event was reported in eukaryotes [23]. In 2015, Ma’s group [24] developed the multiplex genome editing mechanism in monocots and dicots. In 2013 and 2015, it was selected by Science as the “Breakthrough of the Year”. Furthermore, advancements in the CRISPR/Cas system introduced a more precise technique of base editing, which was heralded again by Science in 2017 as the “Breakthrough of the Year”. CRISPR/Cas9-based GE has tremendously revolutionized genome engineering since the initial few research articles were published in Nature Biotechnology [25,26]. All the above mentioned approaches have been extensively employed for GE and caused mutations via site-directed substitutions, replacement, deletions, and insertions at specific sites in the genome [10].
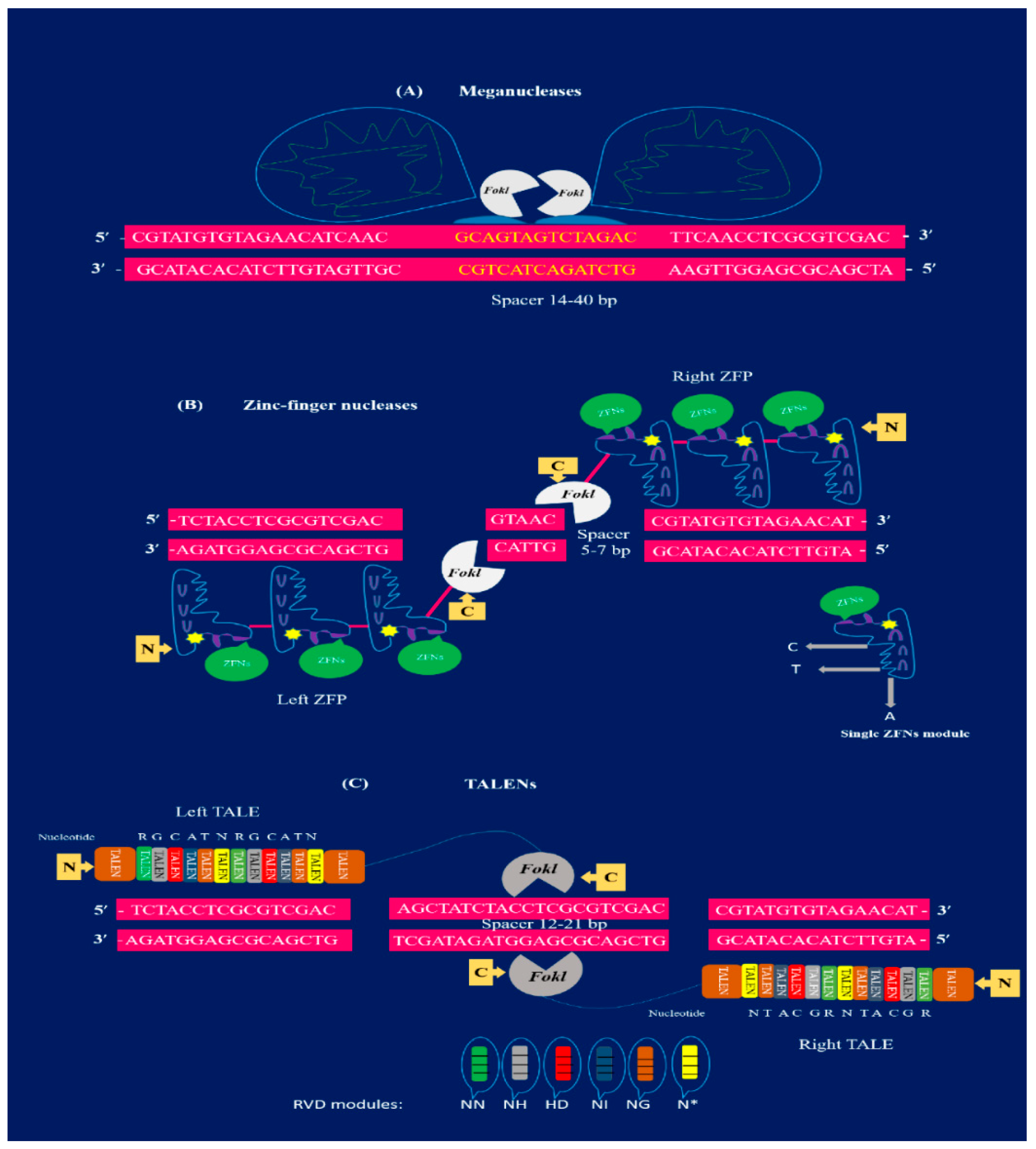
TALENs, ZFNs, and MNs are the first-generation editing tools for genome manipulation as, illustrated in Figure 1. However, they are time consuming and require lengthy protocols to attain target specificity. As compared to first-generation GE approaches, second-generation GE tools such as the CRISPR-Cas9 technique are easier to design, cost effective, and robust [26,27,28]. The CRISPR/Cas9 toolkit is very simple to design, as it involves only single-guided RNA (sgRNA) and the Cas9 protein in contrast to TALENs and ZFNs. Additionally, the procedure involved in TALENs and ZFNs are complex because they require protein engineering for their construction. Due to the presence of these constraints, applications of TALENs and ZFNs in plants have been limited [9]. Continuous innovation for efficient GE has expanded the applications of the CRISPR/Cas9 system in several fields of plant science and is quickly becoming a highly promising GE tool [6,11,23,24,25,26]. The plant GE tools and their corresponding applications are depicted in Figure 2, while successive steps involving the GE strategies are shown in Figure 3.
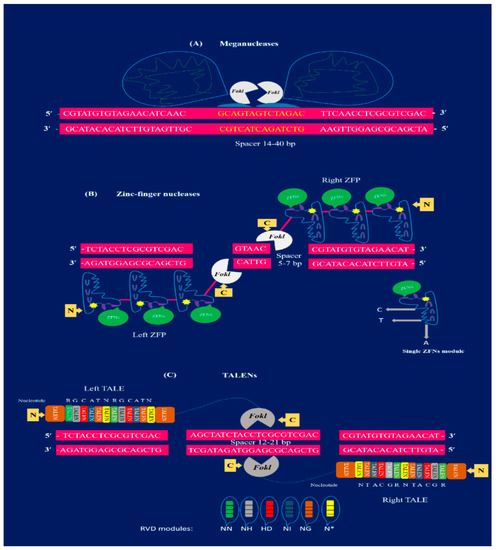
Figure 1.
Structural illustration of first-generation genome editing tools: (A) meganucleases (MNs) have multifunctional domains with the ability to bind double-stranded target DNA and generate DSBs. The meganuclease is demonstrated to bind a target spacer sequence of 14–40 bp (yellow). The FokI nuclease cuts the target sequence (color). (B) Representation of ZFN bound to the target sequence of 18–36 bp long. Each monomer of ZFN (blue) is made by ZFP. There are two basic domains: the DNA binding domain at N-terminus and the catalytic domain with FokI nuclease (white) present at the C-terminus. The connection among these domains is indicated with a pink line. The ZFN modules are merged with Fok1 (white) and dimerized to cut the target sequence at a spacer 5–7 bp (pink) to produce DSBs. (C) Two TALEN dimers bound to the target sequence (pink) site. Each module of TALENs are composed of TALE that contain 33–35 amino acid repeats. The pair of TALENs are separated by a spacer region of 12–21 bp (pink). There are specific RVD modules (green NN, grey NH, red HD, dark blue NI, orange NG, and yellow N) that can recognize only one single nucleotide. TALE modules are dimerized to fuse with FokI (at C-terminus) to produce DSBs in the spacer region.
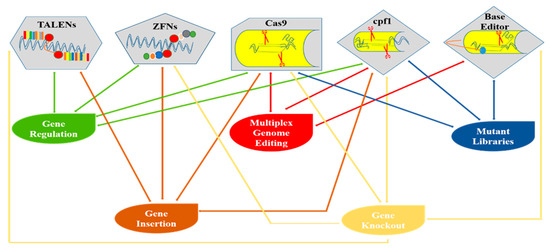
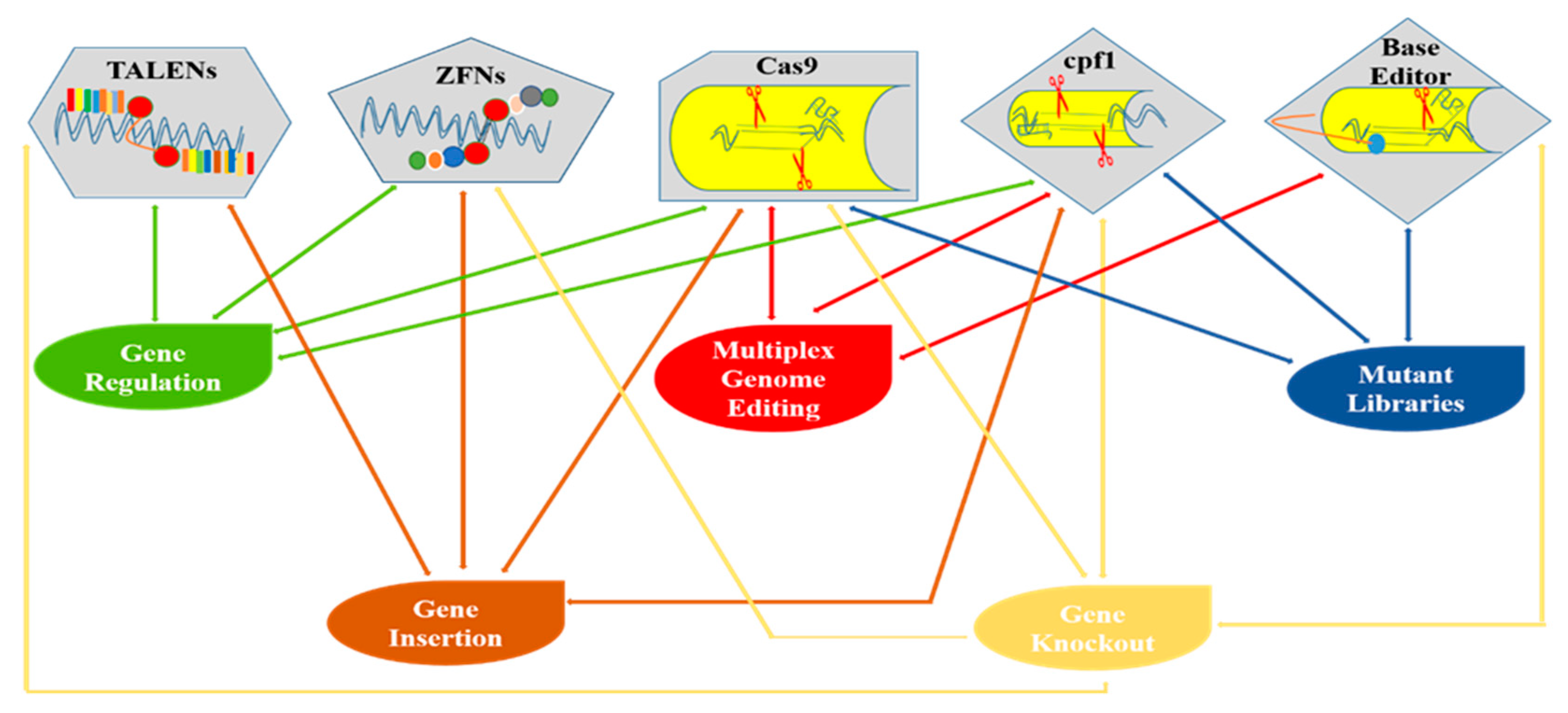
Figure 2.
Diagrammatical representation of various genome editing tools and their applications in plants.
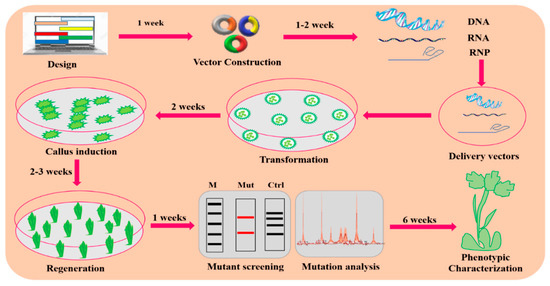
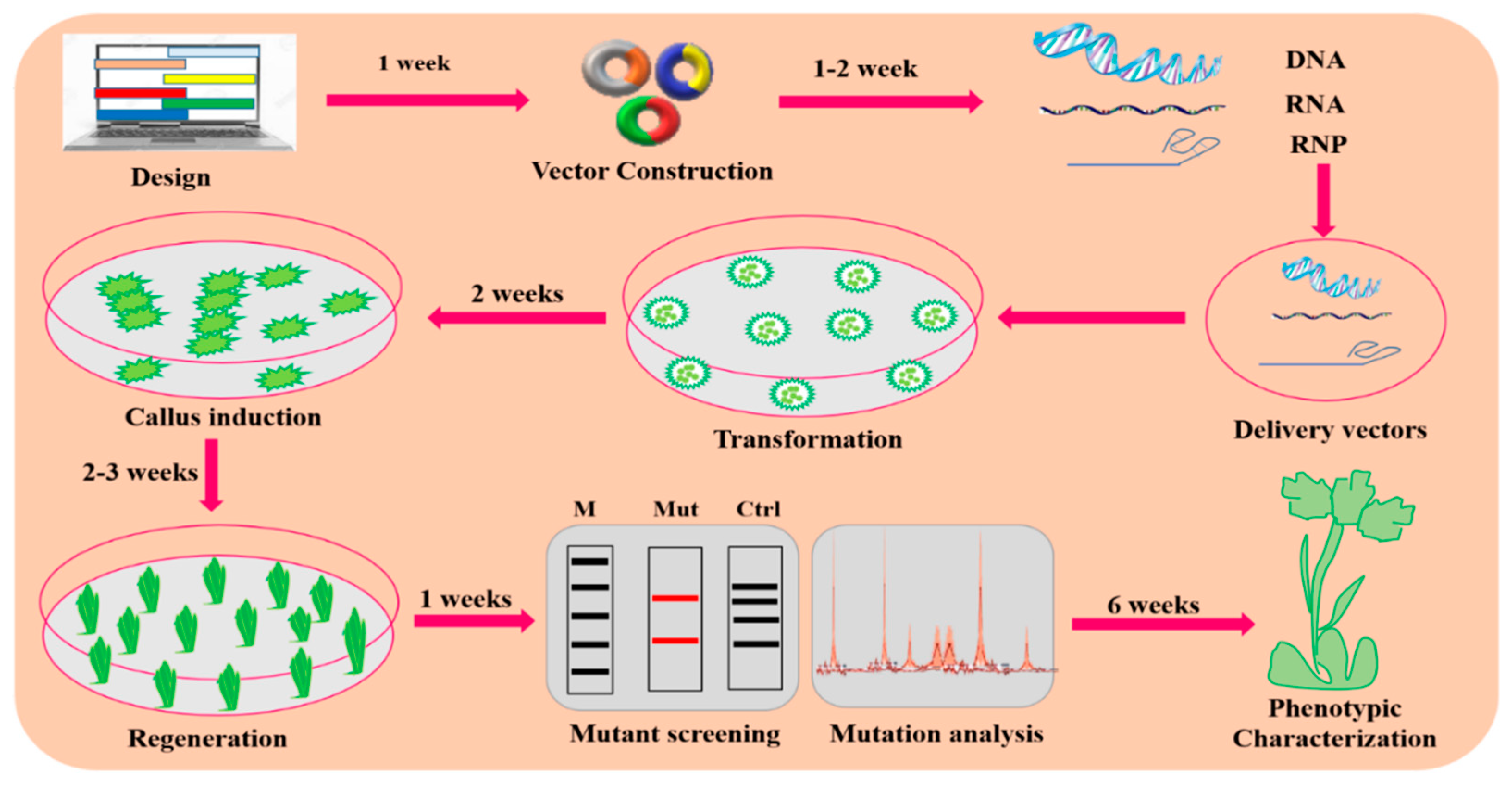
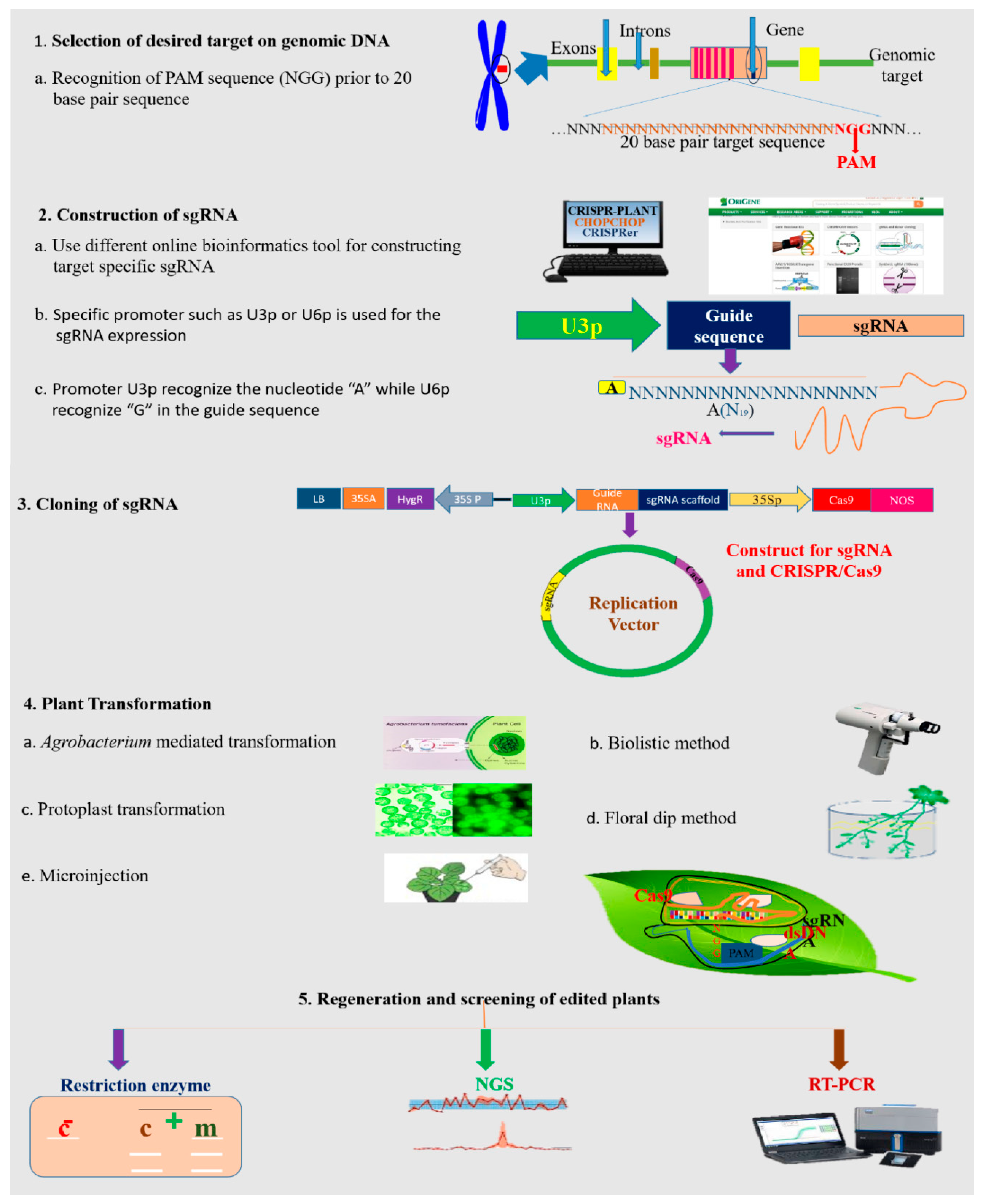
Figure 3.
A general description of the GE mechanism in plants. Plant GE typically consists of the following steps: designing and construction of vectors, targeted delivery of vectors via Agrobacterium-mediated transformation or biolistic for transformation, callus induction and regeneration, mutation screening and analysis, and phenotypic characterization for the desired trait.
In the present review, we discuss fascinating GE tools for crop improvement. We briefly describe first-generation genome editing tools such as TALENs, ZFNs, and MNs and comprehensively elaborate on second-generation genome editing strategies with special focus on the applications of the CRISPR/Cas9 system in plant breeding for crop improvement. We briefly outline historical background, structural organization, and mode of action of the CRISPR/Cas9 toolbox. We describe the workflow of CRISPR/Cas9 from vector design to mutant screening. We also highlight recent breakthrough events in technology improvement in the CRISPR/Cas9 system. Furthermore, we discuss the recent role of CRISPR/Cas9 technology in crop breeding to develop the best performing cultivars with biotic and abiotic stress resilience, improving yield-related traits and production of high-quality crops. Finally, we outline the future outlook of CRISPR/Cas9 and pinpoint the current challenges with respect to the regulation of edited crops and their safe use.
2.1. Meganulceases
In SSNs, MNs were the pioneering class of nucleases (Figure 1A), extensively applied for plant GE [29,30]. Meganulceases were also termed as homing endonucleases. Later, they were utilized for generating DSBs in several genomes [31]. Meganulceases have the ability to recognize target DNA sequences of about ~12–40 bp, that make MNs the most efficient delivery approach for all vectors including plant RNA viruses [32]. As compared to different SSNs, MNs are difficult to re-design for target sequences different than their natural ones. Non-modular properties of the specific proteins are the main reason for hindrance in re-designing MNs. So, for plants, the applications of MNs have been restricted to only naturally existing MNs such as I-SceI and I-CreI nucleases [29].
2.2. Zinc-Finger Nucleases
In plants, ZFNs are extensively being applied for plant GE [33]. Zinc-finger nucleases are one of the main techniques for genome manipulations which are very beneficial in various GE applications. Zinc-finger nucleases have been widely used for target specific mutagenesis to disrupt the gene function and produce several gene knockouts [34]. GE with ZFNs has demonstrated the production of herbicide-resistant plants, and various kinds of targeted and specific gene insertion have also been unveiled [35]. In plant biotechnology, zinc-finger proteins (ZFPs) can be exploited in two ways: ZFNs and ZFN-TFs. Due to the flexible nature of ZFPs, it provides a striking basis for modeling ZFNs with desired sequence-specific domains to produce DSBs and facilitate GE [36,37]. In 1996, ZFNs were reported for the first time and named as chimeric restriction enzymes. According to this research, chimeric restriction enzymes were developed by associating the non-specific FokI with the DNA binding domain of two dissimilar ZFPs. The ZFNs were constructed by the fusion of chimeric proteins that were composed of DNA cleavage and a DNA binding domain. A set of 3–6 Cys2His 2 ZFs constructed the DNA binding domain, while a Fok1 restriction enzyme was generated by the DNA cleavage domain [18]. FokI is homodimeric in nature and belongs to the type IIS class of restriction enzymes isolated from Flavobacterium okeanokoites [18]. The domain of FokI nuclease needs to dimerize in order to cut DNA [38]. Two ZFN monomers are required having FokI dimerization and C-terminal fusion for active cleavage when binding to DNA. Target DNA with 9–18 bp has been recognized by each monomer containing 3–6 ZFs (Figure 1B). Consequently, each monomer of ZFN targets the spacer region of 5–7 bp located in the adjacent half-site and dimerize to perform the cleavage activity for targeted DNA [10]. ZFNs, as compared to MNs, are small in size (about 300 aa in one monomer and 600 aa in a pair of nucleases), enabling them responsive to many delivery procedures. In the last two decades, ZFNs have been applied for site-specific mutations in plants such as Arabidopsis thaliana, soybean, maize, tobacco, and petunia [36,39,40]. However, modular association of ZFs has gained partial achievement [41]. Currently, ZFNs are not recommended in several cases due to the lower target specificity, limited amount of specific target domains, and large number of non-targeted editing [42].
2.3. TALENs
Another interesting tool for GE is termed as transcription activator-like effector nucleases (TALENs). These are modern inclusions to the SSNs resources and have been extensively applied for GE in plants [43]. In 1989, TALENs were first discovered when a pathogenic bacterium called Xanthomonas was studied for many plant varieties [44]. Xanthomonas is responsible for uncontrolled growth of plant cells due to the synthesis of a novel protein termed as transcription activator-like effectors (TALEs) that target specific DNA sequences and greatly influence gene expression [45]. For targeted GE, TALENs are manipulated by changing the TALE repeated domains required for specific target identification and are successively linked to Fok1 nuclease to obtain suitable TALEN. The TALENs that recognize 12–21 bp extend, likewise to ZFNs, and require a spacer region of 14–20 bp for Fok1 dimerization with a pair of TALENs (Figure 1C) [10].
TALENs have the advantage over other SSNs such as MNs and ZFNs because of their modular domain. The domain for TALENs contains 33–35 aa in direct sequence repeats and two amino acids are called repeat variable di-residues (RVDs) in these repeats. The RVDs are responsible for recognition of specific nucleotides which includes thymine; NI, HD, cytosine; NG and adenine; and NN. A single RVD associated with every single nucleotide in combined mechanisms was identified that has the ability to design specific DNA binding motifs and remove the remodeling issues faced in the case of ZFNs and MNs [22,46,47,48]. Target specificity is another advantage of TALENs over other nucleases. Typically, 15–20 RVDs are used in order to design TALEN monomers with more than a 30 bp target site. As compared to ZFNs, TALENs reduce the toxicity and are more specific having large target sites [49]. A large size of about ~950 aa to ~1900 aa is the only drawback of TALENs for use as an accurate tool for GE. TALENs are usually carried to cells via direct integration of DNA or by integration of a construct-harboring TALEN-encoding unit into the genome. TALENs have been successfully applied in plants such as rice [43], Arabidopsis [22], tobacco [42], and Brachypodium [26] for GE. TALENs are more extensively exploited for targeted GE as compared to ZFNs, but they still need an efficient way to assemble tandem repeats for binding to the targeted DNA region. Furthermore, the repetitive nature and large size pose as big hurdles for the successful delivery of TALENs [45]. Some reported events of MN-, TALEN-, and ZFN-mediated mutagenesis in plants are described in Table 1.

Table 1.
Applications of first-generation genome editing tools in crop plants.
3. CRISPR/Cas9 System
Recently, a fascinating GE tool CRISPR/Cas9 was identified for targeted genome manipulations and to express desired genes in numerous organisms [6,65]. The CRISPR/Cas9 system has emerged as the most powerful tool for GE in many species including plants [26]. The latest ground-breaking technology of CRISPR/Cas9 is basically present as an adaptive immune system of type II prokaryotes and protects them against invading organisms during phage infection by spacer acquisition, biogenesis, and target degradation [9]. The toolbox of CRISPR/Cas9 was adapted from bacteria as well as Archaea and in included in the toolbox of engineered nucleases [23,66]. There are two main components of the CRISPR/Cas9 system: a single guide RNA (sgRNA) that identifies a specific DNA sequence and the Cas9 protein which produces DSBs at a targeted site [9]. Therefore, when changing the design of sgRNA, numerous desired sites can be targeted, which makes it simpler to handle than TALENs and ZFNs [10].
3.1. Discovery of CRISPR/Cas9 Wonder
The discovery of the CRISPR/Cas9 system dates back to 1987 when Ishino and his colleagues first identified CRISPR while studying the iap gene in the genome of E. coli. During the cloning of the iap gene, they unexpectedly cloned a specific portion of CRISPR, and at the conclusion of their experiment, revealed that the bacterial genome consisted of a successive array of repeats [67]. After this discovery, an Archaea (Haloferax mediteranii) was also found to contain the CRISPR sequences [68]. Mojica et al. (2000) reported a similar type of regularly spaced repeats in Haloferax mediterranei and Haloferax volcanii, having interrelated functions [69]. Only prokaryotes were considered to have such repetitive sequences, which were named as CRISPR but were not present in eukaryotes and viruses [70]. These short repeats have an average length of 32 bp but are of different sizes from 21 to 47 bp in different organisms. Every repeat has a unique sequence of nucleotides that are extremely conserved in specific species [71]. It was unveiled that the short regular repeats are transcribed into small RNAs [72].
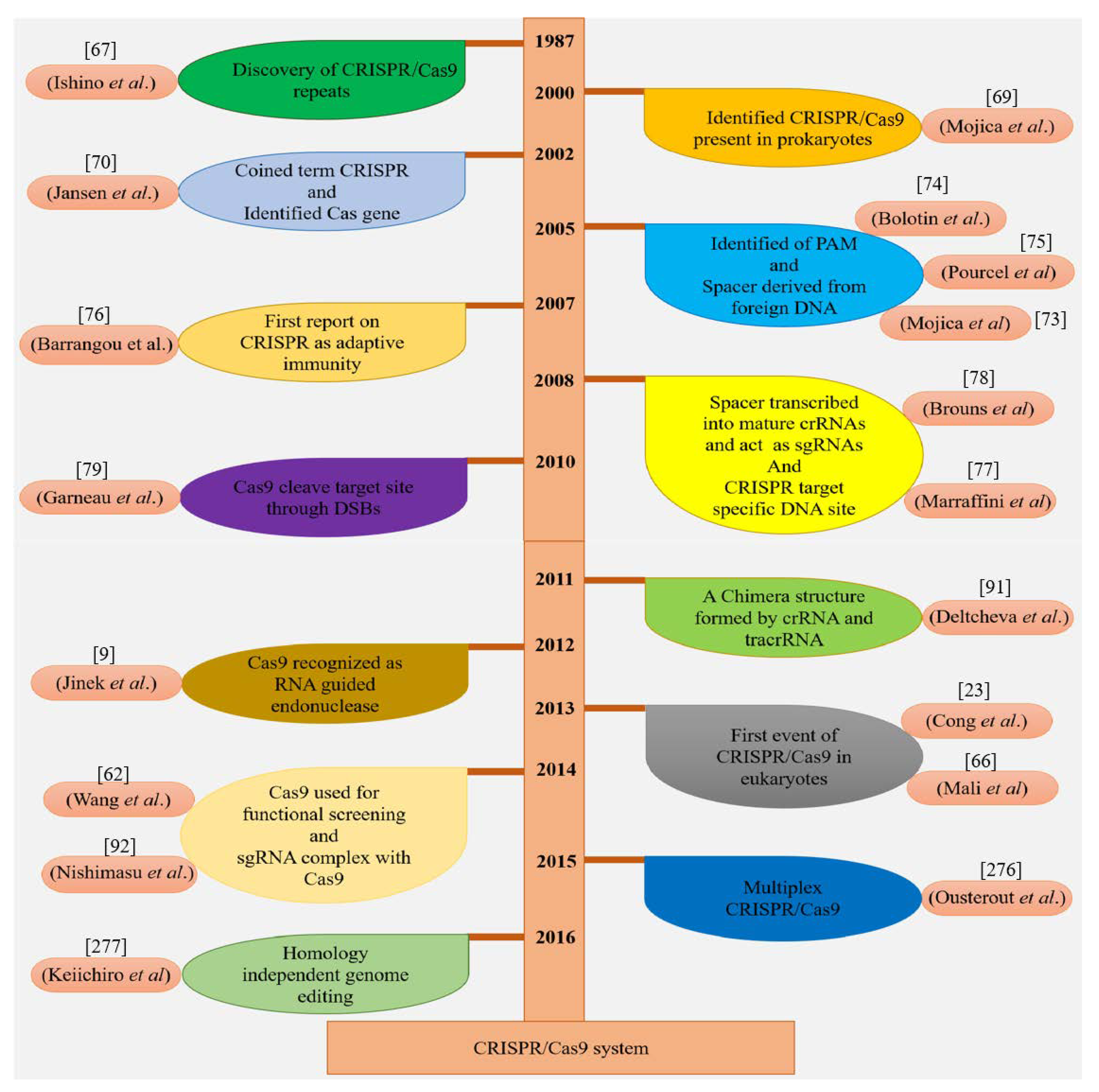
Four Cas genes (Cas1–4) were discovered in prokaryotes having CRISPR DNA sequences during that period [70]. From then on, many CRISPR/Cas sequences and multiple Cas proteins were identified [72]. In 2005, CRISPR spacers were discovered in plasmids and phages by three independent research groups by applying computational and sequencing technologies [73,74,75]. The function of CRISPR/Cas was still ambiguous before Barrangou et al. (2007) successfully demonstrated for the first time that CRISPR protected Streptococcus thermophilus from viral attack [76]. It was revealed that the CRISPR defense mechanism prevents the horizontal gene flow in Staphylococci [77]. In another study, it was observed that CRISPR RNAs regulate the CRISPR interference [78]. The presence of the CRISPR/Cas system in the bacterial genome was identified to cut specific sites in plasmid DNA and bacteriophages [79]. In 2011, the CRISPR/Cas machinery of S. thermophilus was exploited to confer immunity in E. coli [80]. Some of these important events are highlighted in Figure 4.
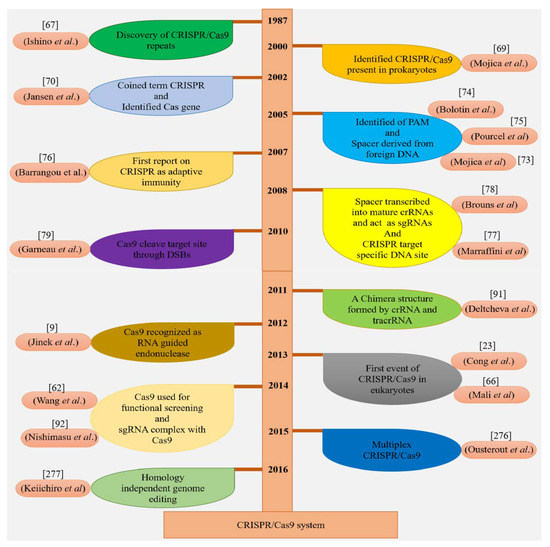
Figure 4.
Historical chart illuminating key developments in the CRISPR/Cas9 system.
3.2. Architectural Organization of CRISPR/Cas9 System and Its Functions
Prokaryotic organisms such as bacteria and Archaea have a special type of defense machinery called CRISPR/Cas [81] in their adaptive immune network to protect them against the attack of viruses and phages [82].
All the natural CRSIPR/Cas networks are composed of many Cas genes, which are encoded by homologous palindromic repeated units, RNA-mediated endonucleases, and novel short RNAs, termed as “spacers” produced by the introduction of short mobile sequences called protospacers. The protospacers are derived from the unique spacers which move among the homologous palindromic sequences repeats when the cell is attacked by invaders. These spacers work as identifying units for the invaded cell and allow the CRISPR-Cas system to cut foreign DNA sequences. There are three steps to the CRISPR/Cas-mediated immune system, including adaptation, expression, and interference. The first step in this mechanism is adaptation, which is associated with the sequential layout of the CRISPR-array via a new spacer’s procurement. Precursor CRISPR-RNA (pre-crRNA) and Cas genes are expressed in the expression stage. Mature cr-RNAs are produced from precursor CRISPR-RNA using RNase III and Cas proteins in the interference event-specific targeted portion memorized by the combinative properties of both Cas proteins and cr-RNA [83]. The CRISPR motif, termed as protospacer adjacent motif (PAM), is connected with each protospacer and closely situated in the target portion of the sequence. The PAM was found to be a highly specific part of the foreign phage or virus genome but is not present on the CRISPR locus in bacterial genome [78].
The PAM sequence consisting of conserved dinucleotides is required upstream of the binding sites of crRNA for Cas proteins to recognize the target sequence [9]. The Cas proteins are unable to detect target DNA for effective cleavage during PAM site recognition. The PAM is also exceptionally crucial to distinguish between the bacteria’s own DNA and invader DNA sequences. Such features enable bacteria to defend their own DNA from nucleases [84]. In several kinds of CRISPR networks, PAM sequences are essential for Cas proteins functions, such as PAM sequence 5′-NNNNGATT which is targeted by Cas proteins in Neissseria meningiditis [85]. Similarly, Cas9 proteins target the PAM sequence 5′-NGGNG or 5′-NNAGAA in S. thermophiles [79,86] and 5′-NGG in S. pyogenes [9].
Two independent groups of scientists discovered the CRISPR/Cas9 machinery with three major kinds (type I–II–III) and two classes in host cells [87]. In 2015, Markarova and coworkers executed the comparative genomic analysis of existing data and found two further reputed types and five subtypes [88]. During the defense mechanism of CRISPR/Cas9 against invader DNA, these two classes behave differently, such as class 1 which consists of subtypes (I, III, IV) and uses many Cas proteins, while only a large Cas protein is used by the class 2 system, which has the subtypes (II, V) [89]. In the adaptation phase of the CRISPR/Cas mechanism, spacers are added by Cas1 and Cas2 proteins and pre-crRNA develop involving Cas5 or Cas6 in the type I system. The Cas6 protein is also used in the type III system for a similar process but stimulation of 3′ end is accomplished by an uncertain element. For crRNA maturation, trans-activating crRNA (tracrRNA) and RNase are utilized in the type II mechanism [90], as shown in (Figure 5A). Currently, the immune system of S. pyogenes operates as a type II system, which is a well-established GE technique known as CRISPR. This CRISPR/Cas9 system is modified by two major units: a non-coding chimeric RNA and Cas9 endonucleases for double-stranded (dsDNA) breaks in DNA (Figure 5B) [9]. The Cas9 protein is directed by the guide RNA (gRNA) and Cas9 proteins recognize targeted DNA in the presence of the “seed” sequence, which is produced by spacers derived from crRNA and the S. pyogenses Cas9 (SpCas9) 5′ NGG ′3 sequence lying closely to the target region [9]. The crRNA and tracrRNA are complementary to each other and it directs pre-crRNA to mature crRNA by means of RNase III. After the maturation of crRNA, it guides Cas9 proteins to break specific DNA sequences [91]. In 2014, Nishimasu et al. (2014) demonstrated that the SpCas9 and gRNA DNA endonuclease has unique lobes, such as an assembly composed of a target detection lobe which is attached to the heteroduplex of sgRNA: a DNA molecule and a nuclease lobe which nicked the target DNA sequence [92], as illustrated in (Figure 5C).
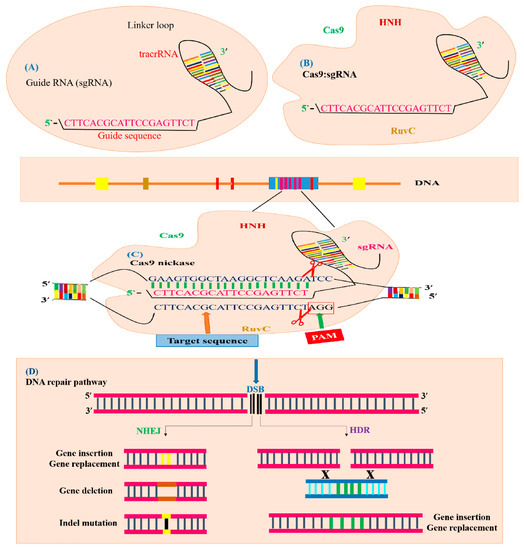
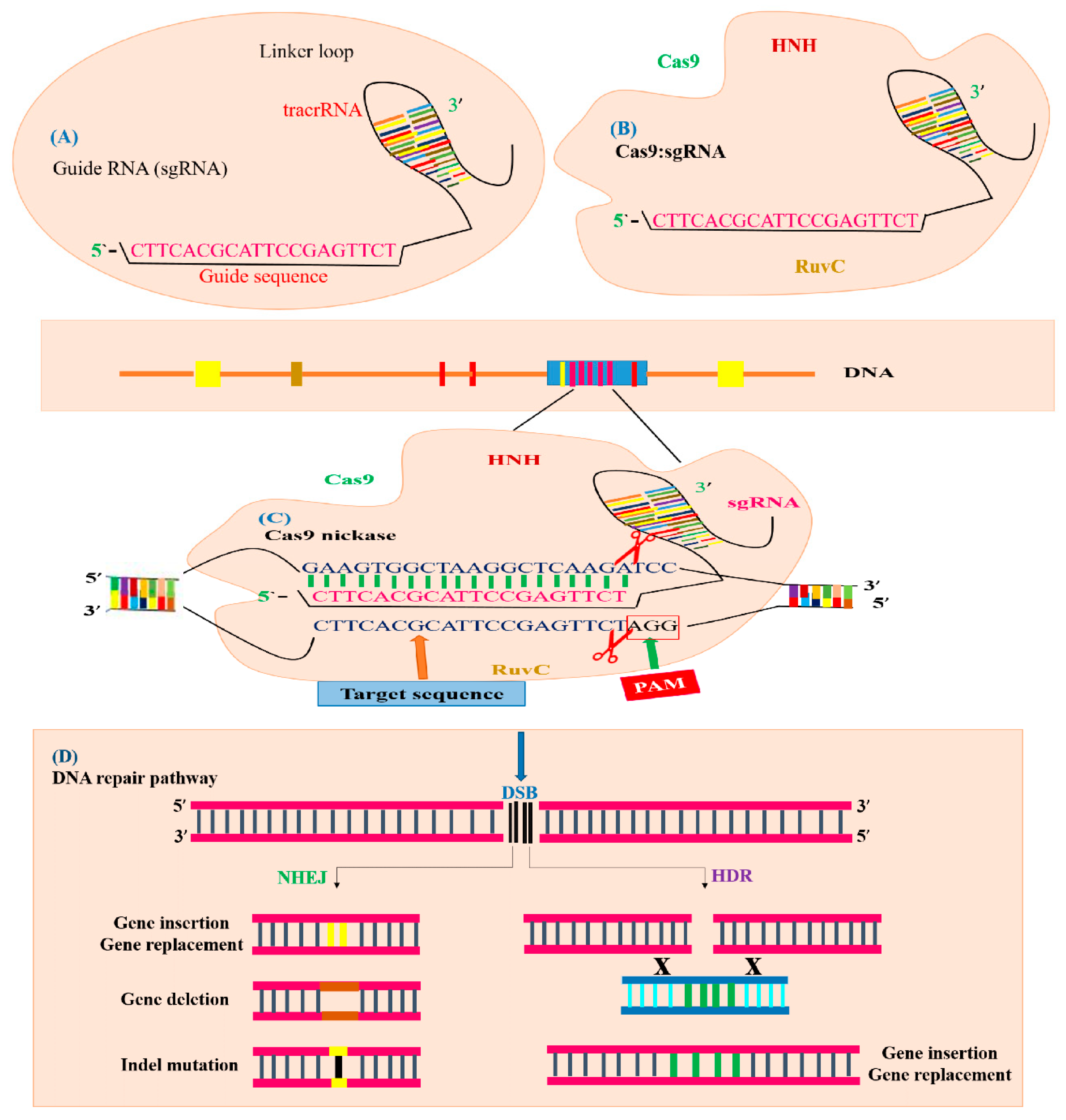
Figure 5.
Illustration of CRISPR/Cas9-mediated GE. The CRISPR/Cas9 system is composed of sgRNA and Cas9. (A) sgRNA with a guide sequence (colored pink) is developed by the combination of protospacer with crRNA and tracrRNA. (B) Cas9 machinery combines with sgRNA to form a complex to trigger CRISPR/Cas9 editing. The Cas9 nuclease consists of two parts, depending on its function and structure. The recognition site identifies the target DNA and interacts with sgRNA. The nuclease site contains two domains RuvC-like and HNH which cleave the target DNA site non-complementary by the RuvC domain and complementary by the HNH domain to the gRNA. (C) The Cas9 nuclease detects the genomic target site (indicated with blue color) having a 20 bp target sequence that is homologous to seed or guide sequence (indicated with pink color), which is crucial for Cas9 activity and specificity. The specific PAM sequence (indicated with red color) is detected by Cas9: sgRNA complex and DSBs created by the Cas9 endonuclease three base pairs upstream of the PAM sequence. (D) Targeted mutagenesis of a desired gene is achieved by filling the DSB (indicated with black color) by means of the HDR or NHEJ mechanism. The NHEJ repair mechanism generally produces insertion (indicated with yellow color), deletion (indicated with brown color) or indels (indicated with black line) at the break point, generating targeted mutants. The HDR repair mechanism uses a template DNA sequence for homologous recombination to produce gene replacement or gene insertion (indicated with green color).
3.3. Genome Editing Mechanism of CRISPR/Cas9 System
The mode of GE is established by the healing process of the genome. After the identification of the target site, Cas9 allows sgRNA to pair with the target DNA sequence. The Cas9 endonuclease is composed of the HNH and RucV-like domain, which cuts the target DNA strands three to four bases upstream of the PAM site. The HNH domain cuts the complementary DNA strands while the RuvC domain cleaves the non-complementary to gRNA. The blunt-ended DSBs can be repaired by the HDR and NHEJ repair pathways (Figure 5D). The NHEJ is error prone and causes DNA insertion or deletion at the target sequence [23]. The expression of sgRNA as pair, NHEJ mechanism came up with large deletions. The large deletions in chromosomes were attained by the NHEJ mechanism utilizing co-expressed sgRNAs. The HDR repair mechanism is only operational when a specific homologous target site is available with respect to the DSB site. In plants, through GE, many outstanding repairs were achieved via HDR, such as gene replacement, DNA correction, and targeted knock-in [93,94].
Biolistic and Agrobacterium-mediated transformation can be applied to transfer the sgRNA and Cas9 protein into desired cells [95]. GE by CRISPR/Cas9 is heavily dependent on the choice of sgRNA promoters and ubiquitous expression of the Cas9 enzyme. Universal CaMV35S RNA polymerase II promoters have been extensively used for Cas9 expression in plants. Similarly, for sgRNA expression, U3 or U6 RNA pol III promoters are applied [96]. The expression level of sgRNAs is significantly greater in endogenous promoters as compared to exogenous promoters [97]. Moreover, sgRNA expression is guided by U6 promoters which were derived from monocotyledonous or dicotyledonous varieties and can only be used in monocot or dicot plants [98]. For successful integration of CRISPR/Cas9 machinery in plant nuclei, Cas9 proteins must join with nuclear localization signals [96].
To bind the target DNA by synthetically developed short gRNA sequences of approximately 20 nucleotides, the mechanism of CRISPR/Cas editing demands the PAM 5′ NGG motif for the Cas9 enzyme to cleave 3–4 bases in the target DNA sequence after the generation of the protospacer [9]. There are two domains of Cas nucleases which have the ability to cut one strand of DNA like the HNH domain and RuvC-like domain. Simple steps involving the execution of the CRISPR mechanism are recognition of the PAM sequence; sgRNA development; cloning of sgRNA; transformation into the host cell; selection of transformed individual organisms; and edited lines confirmation, as described in Figure 6.
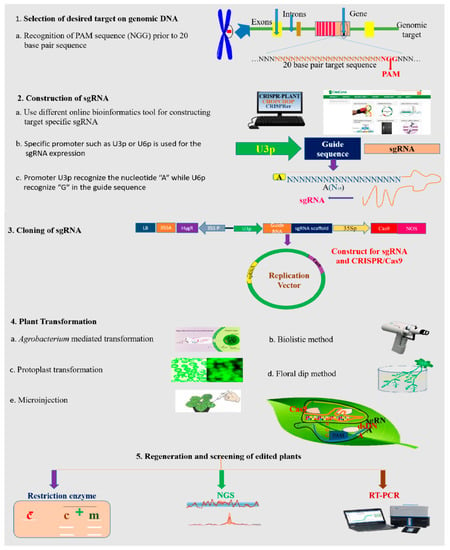
Figure 6.
Basic steps in the workflow of CRISPR/Cas9-based genome editing.
4. CRISPR/Cas9-Mediated GE in Plants
To date, numerous efforts have been successfully employed for targeted gene editing in model as well as in major crop plants via the CRISPR/Cas9-based GE toolbox. Many factors have been reported that affect the editing ability of the CRISPR/Cas9 system including targeted DNA, GC contents, Cas9 codons, sgRNA structure, and expression of cas9 and sgRNA. All these factors must be highly optimized to achieve greater efficiency of CRISPR/Cas9 system [24].
4.1. Designing the CRISPR/Cas9 Delivery System
In the past, numerous attempts have been made at gene editing in plants, but the efficiency of CRIPSR/Cas9 was low [25,26]. With the improvements in technology, many highly efficient vector delivery systems for CRISPR/Cas9 have been designed for plant GE, such as supersession of viral infection, gene disruption of cis-elements, genomic deletion, gene knockout, and multiplex genome editing. Continuous progress in this editing toolkit has allowed more precise, accurate, and targeted delivery of the Cas9 system into plant cells, which include discovery of new Cas9 variants, efficient screening methods for knockout mutants, vector selection, and construction and employment of the most appropriate delivery system for the Cas9 expression cassette. In this section, we describe the construction, screening, and delivery of the CRISPR/Cas9 system into plant cells.
4.2. Cargo-Vectors for the CRISPR/Cas9 System
Two kinds of vector systems are used in CRISPR/Cas9-mediated GE, such as a single-vector system and a binary-vector system. A binary-vector system has been utilized for many years because of its ability for fast primary testing. Any specific vector having several gRNAs and a Cas9 protein expression cassette already constructed in it can be applied for plant transformation. Different structural construct of gRNAs can be utilized for numerous Cas9 proteins to design a unique gRNA: Cas9 nuclease, which allows more accuracy and easiness in experimental design. A single vector harboring both expression cassettes of gRNA and Cas9 protein is becoming more promising. Generally, in a single-vector system, RNA polymerase III-driven promoters (U6/U3) are designed for gRNA expression, whereas ubiquitin and CaMV35S promoters based on RNA polymerase II are exploited for Cas9 gene expression. Continuous advancements in CRISPR/Cas9 permit researchers to develop smarter vector systems to regulate expression of gRNA and the Cas9 gene. Recently, some new adjustments were made in the single-vector system, such as single polymerase II and dual polymerase II promoters. Single polymerase II vectors are applied to govern the expression of gRNA and the Cas9 gene at the same time, while the dual polymerase II vectors exploit two different promoters to drive the expression of gRNA and the Cas9 gene. The addition of all these latest technologies in the CRISPR/Cas9 delivery system assists to decrease the vector length, which eventually, improves the transformation efficiencies [99].
4.3. Bioinformatics Tools for Designing the CRISPR/Cas9 Construct
One of the most crucial steps for highly precise GE is to design an sgRNA construct for CRISPR/Cas9. To date, numerous bioinformatics tools have been developed and are available online for sgRNA designing. There are many accessible online tools with plant databases and which permit the design of sgRNA for identification of new target sites [100], as shown in Table 2. For example CRISPR Design (http://www.genome-engineering.org) was developed by Zhang and colleagues for designing sgRNA and it also assists in assessing off-target mutation [101]. In 2014, Xie and coworkers successfully developed a web tool named CRISPR-PLANT (http://www.genome.arizona.edu/CRISPR) in order to design efficient sgRNA constructs for CRISPR/Cas9-based GE [102]. For example, a novel web tool was developed by Michano and colleagues for rapid detection of target loci in soybean for CRISPR/Cas9-mediated GE [103]. Similarly, CRISPR-P (http://cbi.hzau.edu.cn/crispr/) eases the designing of sgRNA for every plant having an available sequenced genome and also helps to evaluate off-targets [104].

Table 2.
List of various single guide RNA (sgRNA) designing bioinformatics tools for the CRISPR/Cas9 system.
4.4. Construction of the sgRNA Expression Cassette
Construction of a unique sgRNA expression cassette is the most important step in CRISPR/Cas9-mediated gene editing, and it works as a guide system for the Cas9/sgRNA complex consisting of 98 nucleotides with a 20 nucleotide target sequence [92]. In plants, RNA polymerase III is used to transcribe sgRNA and its expression is mostly governed by U3 or U6 promoters [93]. As the expression cassettes of sgRNA:U3/U6 promoters are very small in length, approximately ~300–600 bp, overlapping PCR or adaptor ligation can be applied to construct these expression cassettes [123]. In 2015, Ma and colleagues developed a robust cloning-free approach for sgRNA expression cassette development based on the PCR technique. Gibson assembly or the Golden Gate cloning strategy was used for direct cloning of sgRNA expression cassette into binary vectors for the CRISPR/Cas9 system [24]. In another approach, Gao and Zhao utilized a ribozyme mechanism to generate sgRNA by transcription of pre-RNA through RNA polymerase II, whereby inducible or constitutive promoters can be ligated to obtain the desired function of sgRNA [124].
4.5. Construction of Cas9 Expression Cassettes
Cas9 is composed of 4107 bp of coding sequence. For Cas9 nuclear localization in eukaryotes, the Cas9 coding sequence must be fused with the nuclear localization signal. Plant usage–bias codons have been used to design highly efficient and optimized Cas9 expression cassettes for improved GE in plants [125]. For example, utilization of codon-optimized Cas9p in rice and Gramineae family has been improved by enhancing GC contents [24], which imitate the genes from the Gramineae family [126]. Commonly, constitutive promoters like 35S Cauliflower mosaic virus (CaMV) and ubiquitin from A. thaliana, rice, and maize can govern the expression of Cas9 in dicots and monocots for highly targeted gene editing using callus-based transformation approaches.
4.6. Transformation Approaches for CRISPR/Cas9-Based Vector Delivery into Plants
For CRISPR/Cas9-mediated GE, cargo–vector harboring the expression cassettes of both sgRNA and the Cas9 gene must be carried to targeted sites in plant cells. For cargo–vector transformation, floral dip and biolistic approaches are generally executed. Nowadays, advanced strategies like ribonucleo-protein complex, plasmid delivery, and virus-mediated delivery systems are applied for plant transformation. There are certain limitations in using the virus-mediated delivery system, but several studies have been carried out in plants using the virus delivery system [127,128]. A transient expression system is commonly used by researchers to transfer the vector into protoplast for analyzing the efficiency and feasibility of the CRISPR/Cas9 toolkit [129]. Biolistic and PEG-mediated transformation techniques can be used for direct delivery of Cas9 gene expression cassettes [130]. However, it is difficult to regenerate plants from protoplast due to the heritable targeted mutations, which poses a major drawback associated with this approach in many plant species. Agrobacterium-mediated transformation is a highly efficient approach for stable transformation of the CRISPR/Cas9 system in dicot and monocots [131,132].
4.7. Strategies for Mutant Screening
The CRISPR/Cas9 system is a groundbreaking innovation in GE technology for developing desired mutants and numerous mutant libraries have been created by the CRISPR/Cas9 system so far, such as the genomic-scale mutant library for tomato [133] and rice [134,135]. As the applications for GE approaches are increasing day by day, scientists are required to screen huge numbers of mutants, using a strategy that includes the detection of off-target and on-target edits and which later removes the transgenes in edited plant off-springs.
To overcome the limitations associated with mutant screening, different techniques have been developed, including annealing at critical temperature polymerase chain reaction (ACT-PCR) [136], high-resolution melting analysis (HRMA) [137], polyacrylamide gel electrophoresis (PAGE)-mediated genotyping [138], T7 endonuclease I (T7EI) approach [139], and restriction enzyme site loss technique [140]. There are certain pros and cons for each technique and they are centered on genotyping differences. A mutant can be detected rapidly when it has a clear evident phenotype. For example, a visible albino phenotype was observed when a gene phytoen desaturase mutated via the CRISPR/Cas9 system. It was applied as a phenotypic marker to detect rice- and tobacco-edited plants [141,142]. Additionally, transgenic plants can also be screened using some herbicide/antibiotic selectable markers [141,143]. But making a connection among visual phenotypes and targeted genes is the only challenge associated with phenotyping-mediated screening [135]. In other approaches, high-throughput sequencing is highly efficient and precise strategy to screen all the mutants generated by the CRISPR/Cas9 system [144]. For the detection of DNA-free plants edited by CRISPR/Cas9, whole genome sequencing is quite beneficial and helpful [145].
5. Recent Breakthroughs in CRISPR/Cas9-Mediated Genome Editing in Plants
The GE tool of the CRISPR/Cas9 system has gained remarkable importance in agriculture; however, there are some drawbacks which limit its application in plant GE. The major concerns related to this technology are non-specific off-targets; HR inefficiency; PAM sequences constriction; cargo-vector inefficiency; and many others. With the advancement of plant biotechnology many unique innovative steps are regularly incorporated into the GE network to tackle these challenges.
5.1. CRISPR/Cas DNA as Cargo-Delivery Vector
The conventional cargo-vectors for GE have certain limitations which hinder the accurate editing mechanism in plants. The CRISPR/Cas DNA-based cargo-vector is now extensively used to overcome the drawbacks in plant GE.
5.1.1. Stable Expression
The abovementioned approaches of vector delivery into plant cells, such as biolistic and Agrobacterium-mediated transformation, are used to deliver the CRISPR/Cas DNA, and by screening the mutants, the DNA is inserted into the target location in the plant genome and carry out the editing phenomena. Currently, there are numerous applications of this system in plant GE. In 2016, Gao and colleagues incorporated a fluorescent gene in the expression cassette of CRISPR/Cas9 [146]. Recently, an interesting strategy was developed in which BARNASE and CMS were used as suicide genes to kill embryos and pollens containing the transgene of T0 plant progeny [147].
5.1.2. Transient Expression
Another delivery strategy that can be employed to obtain transgene-free edited plant is the CRISPR transient expression system. In this technique, the screening of mutants using selectable markers (antibiotic/herbicide) is omitted to regenerate the edited progeny without the incorporation of foreign genes into the plant genome. In 2016, Zhang and co-workers successfully executed this strategy for the first time in wheat to obtain transgene-free plants [148]. Furthermore, DNA base editors such as adenine and cytidine base editors have also been transferred by the transient expression approach for attaining DNA-free base substitutions [149,150]. Agrobacterium-mediated transformation-based transient expression has also been successfully developed in tobacco [142], and protoplast transformation-based transgenes-free expression can also be achieved in tobacco and potato protoplasts [151,152].
5.2. DNA-Free Genome Editing Through Ribonucleoproteins (RNPs)
Foreign DNA-free GE is another approach in the CRSIPR/Cas9 toolkit to produced transgene-free plants [153]. In conventional approaches to genetic engineering, the foreign DNA with editing components is incorporated into the host organism. Due to the random incorporation, the genetic modifications can be unpredictable. Even if the expression cassettes are abolished, the fragments of foreign DNA can still be integrated into the host genome and cause mutations [154]. Furthermore, the introduction of genetically modified organisms has increased globally [155]. So, there is an increased demand to produce transgene-free plants. To develop DNA-free genome-edited plants, particle bombardment and protoplast transformation techniques have been employed.
Transgene integration may be reduced by using the CRISPR/Cas DNA transient expression system, but it does not entirely eliminate it; additionally, the discarded DNA portion may still get incorporated into different non-targeted locations within the plant genome. To escape the shortcomings of mRNA and plasmid-based expression system of sgRNA/Cas9, a most competent transgene-free editing strategy was established by designing the RNPs sgRNA/Cas9 system in plants [156,157,158]. Hence, sgRNA/Cas9 RNPs have a greater ability to produce DNA-free edited plants with low off-target frequency and is more efficient than a plasmid-mediated editing system. The RNP-based system does not need transcriptional and translational apparatuses for creating nicks in the target sites and, after cleaving, it is then disintegrates itself. In 2015, Woo and colleagues performed DNA-free GE for the first time in rice, tobacco, lettuce, and Arabidopsis using the RNPs system [156]. Potato, apple, and grape explants were subjected to targeted mutations carrying CRISPR/Cas9-mediated RNPs [159,160,161]. In addition, DNA-free GE in maize and wheat has been developed by particle bombardment-mediated transformation of RNPs and Cas9 proteins into cells [58,148,162]. Recently, a new CRISPR/Cas variant Cpf1 has been added to the RNP-based GE toolkit, carrying AsCpf1/crRNA and LbCpf1/crRNA RNPs into tobacco and soybean [163]. However, in several cereal crops protoplast regeneration is a bigger task, hence, the biolistic-mediated RNP editing system is the most appropriate technique for GE in plants. The delivery of RNP-mediated CRISPR/Cas9 machinery has been demonstrated by two different groups in wheat and maize [145,146]. The discovery of a DNA-free editing system will surely simplify the GE of plants and helps to commercialize the edited plants in the future.
5.3. CRISPR/Cas9 Toolbox: Ways Toward Precise Editing
5.3.1. Base Editing
As compared to DSB-governed GE, single-nucleotide modification at a specific site of the genome is called base editing, and is not based on donor DNA or an HDR mechanism and also does not require DSB generation, which provides a simple, highly accurate, and universal mechanism for editing a single base at a target site. Thus, base editing with the CRISPR/Cas9 tool is gaining interest for precise targeted gene editing in plants [164]. Currently, the use of the HDR repair mechanism with donor DNA for DSBs has been found to be less effective in contrast to NHEJ repair with a template-free system, posing a great hurdle in plants for base substitution. Genome-wide association studies (GWAS) have demonstrated that crop plants having a single nucleotide insertion/deletion are more significant for screening the elite germplasm [16]. Therefore, powerful tools are required immediately for generating accurate base editing in crop plants [165]. The CRISPR/Cas9 is an exceptional technique for precise substitution of a single base in target DNA [166]. The CRISPR/Cas9-directed base editing strategy has used the gRNA system, which is homologous to the natural CRISPR system. But in case of a cytosine base-editor (CBE) system, modified Cas9 endonucleases called nickase (nCas9) are used as compared to the natural CRISPR system. These nCas9 proteins, in addition to dead Cas9 proteins fused with an enzyme having base cleaving activity such as cytidine, are converted to uridine by the cytidine deaminase [149,167]. Recently, an effective base-editor 3 (BE3) platform was developed, which involves the merger of APOBECI known as rat cytidine deaminase and which has been extensively employed for GE in many organisms including plants [168]. In addition, several improvements in the BE3 system has allowed modifications in PAM sites to enhance its editing specificity and accuracy [168]. Likewise, three cytidine deaminase orthologs such as human APOBEC3A [150,169], human AID [170], and lamprey PmCDA1 [167] have been fused with nCas9 to attain highly precise C-to-T substitution. For example, a plant CBE system based on APOBEC3A has been widely applied for C-to-T substitution in potato, rice, and wheat [150,171]. In rice and Arabidopsis, CBE has been used to create point mutations. In addition, CBE can also be applied to generate non-specific mutations that manipulate the desired gene and disrupt its function. CBE was found to be precise and more accurate than SSN-mediated editors, producing rare if any indels [172].
Similarly, adenine converts to inosine by adenine deaminase [168]. In wheat and watermelon, this strategy has been adopted to develop herbicide-resistance plants [50,173]. Yan and colleagues identified a fluorescence-tracking mechanism in rice which converts the adenine to guanine by a single-base editing system [174]. An adenine base editor (ABE) was developed for multiplex base substitutions in rice [175]. Similarly, ABE was applied to study the germline transmission and preferred phenotypic changes in Arabidopsis [176]. Recently, Li et al. (2018) upgraded the ABE for generating base editing in wheat and rice plants. They have also successfully developed an herbicide-resistant rice by producing the point mutation [177]. So, GE has been provided novel dimensions by base-editing tools, widening its prospective applications by manipulating desired nucleotides in the plant genome.
5.3.2. Multiplex Genome Editing
In plants, cellular processes are fine-tuned by several redundant genes. Sometimes, mutating a single gene may not confer a desired phenotype because of the compensation effect produced by other genes in same gene family. Hence, an upgraded editing system with improved efficiency is needed for multiplex gene editing in plants. In CRISPR/Cas9-mediated multiplex GE, many sgRNA cassettes can be designed by using single or multiple promoters into a single-vector system [108,178]. In 2013, Mao and coworkers designed two sgRNAs for two homologous of magnesium-chelatase subunit I (CHLI) having function in the photosynthesis mechanism, and it successfully transformed the vector in Arabidopsis thaliana. The result showed the albino phenotype in plants in which both genes were disrupted [179]. In another study, four subunits of katanin p80 were mutated in A. thaliana using multiplex genome editing. For this, three sgRNA expression cassettes were designed for simultaneous gene editing and the results demonstrated the dwarf phenotype in quadruple-mutant plants [180].
The group of Xie reported the editing of eight genes simultaneously by designing multiple sgRNA expression cassettes. An endogenous t-RNA-processing platform was used for the expression of multiple sgRNAs. All the sgRNAs were released after the nick produced by endogenous t-RNA-processing-based RNase [181]. Similarly, this t-RNA-based strategy has also been efficiently demonstrated in Zea mays [182]. A multiplexing system was developed by Tang and colleagues in which hammerhead self-cleaving ribozyme was applied. Additionally, the same promoter Po1II was used for the expression of multiple sgRNAs that govern Cas9 activity. Ribozyme cleavages separated sgRNAs and Cas9 after transcription and released functional sgRNAs and Cas9 [183]. Furthermore, the ability of the CRISPR-Cpf1 system was harnessed for multiplex GE in rice. A single promoter was applied to produce a construct composed of numerous repeated units of crRNA attached with a target sequence. A target repeat sequence was recognized by Cpf1 and produced cleavage, which resulted in releasing of crRNAs [184]. Hence, CRISPR/Cas9-mediated multiplex GE is a convenient approach for knocking out multiple genes at once and helping to decipher the function of a desired gene family that regulates multiple biological networks. Moreover, it is also beneficial in finding out the epistatic association among genes in numerous genetic processes.
5.4. Beyond Cas9: New Cas Variants Broadening the CRISPR Toolbox
The CRISPR/Cas9 system which originated from Streptococcus pyogenes has some drawbacks which hinder its editing activity like multiple incompatible off-targets due to the gRNA mismatches. Thus, several changes have been made to enhance the editing efficiency and to minimize the off-target nicking of Cas9 enzymes including SpCas9n (Cas9n) [23], Dead cas9 (dcas9) [185], and FokI Cas9 (fCas9) [186,187]. Various bacterial species have been used for the extraction of Cas9 proteins having novel- and stretched-PAM sequences that can help in enhancing the non-target cleavages. Neisseria meningitides have unique a CRISPR/Cas machinery named Nmecas9 which is specific for 8-mer (5′-NNNNGATT) PAM sequence targets that can minimize the chance of off-target cleavage and enhance specificity [188]. Besides, of the other identified orthologs of Cas9, SpCas9 has been most commonly used for GE. SpCas9, derived from Staphylcoccus aureus, detects the 6-mer PAM sequence (5′-NNGRRT) [189]. The modification of SpCas9 has been carried out which targets the PAM sequence (5′ NGA) and edits the target gene efficiently [190]. Besides SpCas9, the shorter length of SaCas9 permits it to overcome the delivery challenges faced by SpCas9 in utilizing the multi-dimensional adeno-virus cargo vectors [189]. The CRISPR/SaCas9 system has been efficiently used to edit many plant genomes such as citrus, rice, tobacco, and A. thaliana [191]. Furthermore, Streptococcus thermophilus-derived St1Cas9 and St3Cas9 have also been employed for CRISPR-mediated GE [191]. Different types of tracrRNA and crRNA are used by these orthologs to identify different PAM sites [192].
CRISPR/Cpf1 System
Recently, Francisella novicida was studied to discover the Class II type CRISPR-Cpfl system [193], recently named Cas13 [194]. In comparison to Cas9 for cleavage and production of cohesive ends, Cpfl needs a single RNA-guided complex having 4–5 nucleotides 5′-overhangs. The CRISPR-Cpfl system has been used successfully with none or fewer off-targets in both animals and plants. In 2016, the CRISPR/Cpf1 tool was effectively applied for GE in plants [195]. Due to the exceptional properties of Cpf1, type V CRISPR/Cpf1 has been considered as another powerful technique for plant GE [196]. The CRISPR/Cpf1 machinery like the conventional CRISPR/Cas9 system is formed by the two major elements: one Cpfl nuclease for target specificity and the other one for target sequence identification called crRNA. Although, in contrast to the Cas9 network, which recognizes PAM sequences with G-rich contents (5′-NGG-3′), the Cpf1 recognizes a PAM sequence (5′-TTN-3′) having T-rich contents [194]. Furthermore, CrRNA and tracrRNA interaction is not needed in the Cpfl system, although it is necessary for the Cas9 technique. A size of about 42 to 44 crRNA is required in the CpfI system, which is smaller than that of gRNA [194]. In rice and tobacco, targeted mutagenesis has been carried out through the CRISPR/Cpf1 mechanism derived from Francisella novicida (FnCpf1) [195]. In rice, Lachnospiraceae-derived Cpf1 (LbCpf1)-mediated targeted mutations have been reported [197,198]. Similarly, LbCpf1 and FnCpf1 nucleases have great potential for precise GE for specific gene addition via HR mechanism [199].
6. Applications of CRISPR/Cas9 in Plant Breeding
Climate change and rapid increases in the world’s population are two major concerns that threaten agriculture production and food security globally [200]. Several biotic stressors (bacteria, viruses, fungi, insects, nematodes, etc.) and abiotic stresses (drought, salinity, heat, cold, waterlogging, etc.) hamper crop production and compromise food security around the world. Crop breeders are striving hard to develop climate-resilient, stress-tolerant crops with better quality and increased production [201]. Thus, the CRISPR/Cas9 system has numerous applications for the functional genomic research of plant genes that play a crucial role in genetic improvement of many significant agronomic traits. Especially, the knockout of some genes can encourage superior traits including disease resistance, adaptation to various abiotic stressors, nutrient usage, and yield improvements. Thus, CRISPR/Cas9-mediated GE has great potential in plant breeding for crop improvement.
6.1. CRISPR/Cas9 System for Plant Disease Resistance
Virus, bacteria, fungi, nematodes, and insects are the major causal agents inducing biotic stressors and crop yield reduction. Moreover, the persistent upsurge in several new strains of lethal pests make the battle very challenging against these pathogens [202]. Thus, to protect agriculture from the devastating impact of biotic stressors, it is very crucial to understand the plant–pathogen interaction [203]. GE strategies have been successfully applied to explore plant–pathogen interactions and mechanisms underlying plant responses against pathogen attack.
CRISPR/Cas9-mediated GE can be employed directly to disrupt disease-causing genes, known as “S-genes” and develop disease-resistant crops. For example, targeted knockout plants for the ethylene-responsive gene OsERF922 were generated via the CRISPR/Cas9 tool, which showed reduced blast lesions and increased resistance against rice blast caused by Magnaporthe oryzae [204]. Likewise, bacterial blight-resistant plants were produced by targeted mutagenesis of the SWEET13 gene [205]. CRISPR/Cas9-based mutagenesis was applied to the promoter region, and transcription factor (TF) of canker CsLOB1 in Citrus paradise was identified. Due to the presence of such mutations, two mutant lines DLOB10 and DLOB9 with high mutation rates have been produced. The frame-shift mutation and disruption of the CsLOB1 gene improved resistance against Xanthomonas citri [206]. Peng and colleagues reported the editing of effector binding elements (EBEs) by the CRISPR/Cas9 system in the CsLOB1 gene promoter region to increase disease resistance in Citrus sinensis against Xanthomonas citri [207].
In wheat protoplasts, the CRISPR/Cas9 technique was applied by Shan et al. to edit the TaMLO gene [141] and produce wheat lines resistant to powdery mildew caused by Blumeria graminis f. sp. Tritici [62]. In another study, CRISPR/Cas9-mediated multiplex GE was performed to mutate three homologs of the EDR1 gene to develop resistance against powdery mildew in wheat [208]. Similarly, CRISPR/Cas9-based mutants of MLO were produced in tomatoes which conferred resistance against powdery mildew [209].
It has been estimated that about half of plant diseases are caused by virulent viruses, which result in heavy crop losses globally [201]. Gene-targeting efficiencies were improved many folds by DNA virus amplicons. Geminiviral-based DNA replicons of wheat was utilized for transient expression of the CRISPR/Cas9 system against wheat dwarf virus (WDV), in hexaploid wheat and 12 fold upregulation was observed in ubiquitin gene expression [210]. Stable over-expression of sgRNAs and Cas9 that particularly target the genome of the Gemini-virus to prevent its growth has been applied for virus-resistant crop breeding programs [211,212,213]. Furthermore, the CRISPR/Cas9 system can also be used to mutate viral genomes in addition to tackling diseases caused by them [201]. The efficiency of CRISPR/Cas9-mediated viral GE can be increased by using virus promoters to govern sgRNA/Cas9 expression cassettes [211]. Recently, a new ortholog of Cas9 has been discovered in Francisella novicida (FnCas9) to edit RNA virus genomes. FnCas9 has successfully inhibited the replication of the tobacco mosaic virus as well as the cucumber mosaic virus and provides immunity against them [214]. Therefore, CRISPR/Cas9-mediated GE is an exceptional tool to improve genetic make-up and enables them to combat various pathogens. A list of recent studies indicating the significant success of the CRISPR/Cas9 system against various plant diseases is compiled in Table 3.

Table 3.
Summary of disease-resistant crops developed via CRISPR/Cas9.
6.2. CRISPR/Cas9 for the Production of Climate Smart Crops
The CRISPR/Cas9 technology has been extensively applied in major crop plants such as wheat, rice, maize, cotton, soybean, tomato, and potato to cope with various abiotic stressors. Development of climate smart abiotic stress-tolerant crops via the CRISPR/Cas9 tool has modernized plant breeding programs. Major events for crop improvement via CRISPR/Cas9 are described in Table 4.

Table 4.
Summary of CRISPR/Cas9 applications in major crops for abiotic stress tolerance.
For example, in wheat protoplast, two genes related to abiotic stress, TaDREB3 and TaDREB2, have been studied using the CRISPR/Cas9 technique. With a T7 endonuclease assay, the expression of mutated genes has been confirmed in approximately 70% of transfected protoplasts. The mutated plants showed increased tolerance against drought as compared to wild cultivars [221]. Three rice genes named mitogen-activated protein kinase (OsMPK2), phytoene desaturase (OsPDS), and betaine aldehyde dehydrogenase (OsBADH2) have been edited using the CRISPR/Cas9 technique. For transformation of CRISPR/Cas9 machinery, particle bombardment and protoplast transformation methods were used and revealed that these genes are responsible for regulating many abiotic stressors [26].
To protect plants from abiotic stressors, plant annexins play a major role. The annexin OsAnn3 gene in rice has been studied under cold stress and its function was determined in edited knockouts developed by the CRISPR/Cas9 system [222]. Similarly, the gene SAPK2 was mutated to study the stress tolerance mechanism in rice. The results revealed that the expression level of SAPK2 was enhanced under drought and salinity stress conditions [223]. Drought tolerance in transgenic maize was enhanced by the overexpression of AGROS genes and they are of great prominence for maize breeding. To identify new allelic variants, CRISPR/Cas9 was applied to mutate the ARGOS8 gene [224]. Curtin and colleagues carried out CRISPR/Cas9-based knockout mutagenesis of two genes Drb2a and Drb2b and found that these genes regulate salt and drought tolerance in soybean [225]. In tomato, important signaling molecules, i.e., mitogen-activated protein kinases (MAPKs) that respond against drought stress by protecting the membrane of cells from oxidative destruction and regulating genes transcription to tackle drought stress. The association of the SlMAPK3 gene in controlling the drought tolerance mechanism has been reported in tomato by creating knockout mutants of the SlMAPK3 gene under drought stress through the CRISPR/Cas9 system [226].
Many important traits like stress tolerance and crop yield are controlled by multiple genes. Many studies have been carried out to locate the quantitative trait loci (QTLs) that are controlling important traits in crop improvement programs. For the development of better performing varieties, such QTLs have been transferred into the elite lines. But this introgression is laborious for closely associated QTLs and if non-target regions are introduced into best performing varieties, it may produce many deleterious effects. Conversely, CRISPR/Cas technology can be a fascinating approach to generate and examine targeted mutagenesis. Using a CRISPR/Cas9-mediated QTL editing approach, the functions of grain number QTLs (Gn1a) and grain size (GS3) in rice varieties were examined [227]. Hence, the above studies revealed that CRISPR/Cas9-based GE has massive potential for the development of climate-resilient crops.
6.3. Crop Yield and Quality Improvements via CRISPR/Cas9
Two other important agricultural traits are crop yield and quality that need to be improved through the CRISPR/Cas9 system to ensure food security worldwide (Table 5). Crop yield is a complex, multi-genic, and quantitative trait that is influenced by several features. The CRISPR/Cas9 technology has demonstrated its worth for quick yield improvement in crops.

Table 5.
Summary of CRISPR/Cas9 applications in major crops for yield and quality improvement.
In many studies, the CRISPR/Cas9 technique has been used to knockout the genes that negatively regulate yield related-traits including tiller number (OsAAP3), panicle size (OsDEP1, TaDEP1), grain weight (TaGW2, TaGASR7), grain size (OsGS3, OsGRF4), and grain number (OsGn1a). The results demonstrated that CRISPR/Cas9 is an efficient technology for improving crop yield [236,237,238,239,240]. A multiplexing GE strategy has been employed to mutate three genes simultaneously including GS3, GW2, and GW5, and TGW6 which headed towards trait pyramiding and enhancing grain size and weight in rice [241]. Similarly, Li and coworkers applied the CRISPR/Cas9 system to knockout three yield-related genes, Hd2, Hd 4 and Hd5, which resulted in early heading in rice [242]. It was reported that the OsSWEET11 gene has a crucial role in grain filling and sucrose transportation. So, the CRISPR/Cas9 system was applied to disrupt the OsSWEET11 gene, which led to decreased sucrose concentration and reduced grain weight. This study suggested that the overexpression of this gene may be beneficial for maximizing rice yield [243]. In wheat, CRISPR/Cas9-mediated GE knockout of the GASR7 gene increased kernel weight [148]. Recently, a new approach has been established for gene identification on a large-scale that assists in examining the complex quantitative traits, including yield, by integrating the CRISPR/Cas9 tool, whole genome sequencing, and pedigree analysis. In a study, 30 varieties of “Green Revolution miracle rice” were subjected to genome sequencing and 57 genes controlling yield-related traits were screened. Knockout mutants of those 57 genes were created using the CRISPR/Cas9 technique. Phenotyping indicated that several genes are crucial for regulating yield-related traits in rice [244].
There are also many applications of CRISPR/Cas9 technology for quality improvement in crops such as storage quality, nutritional value, fragrance, and starch contents. For example, the cooking and eating quality of rice has been improved by mutating the Waxy gene using CRISPR/Cas9 [245]. The nutritional value of rice has also been improved by knocking out the SBEIIb gene which resulted in more amylose synthesis [246]. Similarly, the starch synthase gene GBSS was mutated via CRISPR/Cas9 in potato. The mutated lines showed decrease levels of amylose and enhanced the concentration of the amylose/amylopectin ratio [152]. In 2018, Sanchez and colleagues, carried out CRISPR/Cas9-mediated GE of the gluten-encoding gene family α–gliadin to produce low-gluten wheat [247]. To improve oil composition of soybean, the CRISPR/Cpf1 system was employed to disrupt the FAD2-1B and FAD2-1A. The results revealed high-yielding soybean plants with improved levels of oleic acid [163]. Moreover, it was reported that the zein protein has been reduced by 12.5% in kernels by disrupting the PPR and RPL genes in maize. The mutated plants indicate increased production levels of healthy tryptophan and lysine in maize [182]. Sorghum nourishment quality has been improved by targeting k1C genes which were responsible for poor digestibility and hindered production of important amino acids [248]. Recently, some other studies for quality improvement have been carried out via the CRISPR/Cas9 system, such as Brassica napus with high oleic acid concentration [249], long shelf life of tomatoes [250], and increased lycopene levels in tomato [251].
In summary, the above described studies reveal that CRISPR/Cas9-mediated modern breeding techniques can be utilized to attain valuable mutations for improving crop yield and quality.
7. Regulatory Affairs of Genome-Edited Crops
Since the development of the first genetically modified organism (GMO) in 1995, firm rules and sanctions have been imposed to regulate GM crops worldwide. In most European countries, GM crops are still banned for commercial production and release of GM crops in the field and consumer market is prohibited. Similarly, GE non-transgenic crops may also be banned if regulatory bodies consider them as GMOs. GM crops continue to provoke extensive public misunderstanding and mistrust despite 22 years of commercialization and cultivation on 189.8 million hectares in 2017 with approximately US$18.2 billion economic gains in 2016. Globally, 82% of the total crop area for soybeans, 68% for cotton, 30% for maize, and 25% for oilseed rape were planted with GM varieties in 2014. Despite high adoption rates by farmers, the cumbersome regulatory processes and delayed cultivation approval procedures have reduced the value of innovation “the GM crops”.
Quick action may be needed for strict legislation to distinguish between GE and transgenic crop. Modern breeding technologies, particularly GE, are highly feasible alternative options to GM crops and involve a reduced degree of regulatory oversight. New discoveries in GE technology and continuous progress in delivery systems that do not need to insert any specific foreign DNA in host cells for crop improvement may strongly challenge the legislative laws regulating the transgenic crops [130].
Traditional plant breeding approaches mainly depend on chromosomal modification via homologous recombination. Selection and crossbreeding have been applied for many years to screen the best performing varieties. Conventional plant breeding techniques have been used for several decades to detect novel traits and introduce them into individual plants for desired results. Traditional breeding has successfully developed many new cultivars, but these approaches need rigorous and continuous selection for many generations [256]. Genetic variability has significantly decreased due to the progressive evolution of many major crops through traditional breeding [257]. Therefore, modern plant breeding approaches have become essential to overcome the certain limitations of traditional breeding such as self-incompatibility, long generation time, heterozygosity, polyploidy, and time consuming.
Mutation breeding and engineering of transgenic plants are other crucial strategies used for crop improvement [258]. Conventional mutagenesis has helped to produced genetic variations that ultimately help in improving food quality and crop yield. For genetic analysis, natural or artificial mutagenesis has been induced using chemical and physical mutagens like ethyl methanesulfonate (EMS) and gamma rays [259]. Screening of large numbers of mutants is the biggest challenge. Such laborious, time-consuming, and untargeted breeding platforms cannot maintain pace with global food demands [258]. Foreign genes have been transferred into the elite crop lines to obtain desired traits through transgenic breeding. As compared to traditional breeding, transgenic techniques eliminate all crossing barriers and have increased genetic variability. Transgenic technology has provided enormous opportunities for crop improvement but, at the same time, provoked public concerns about its potential effect on human health and environment. Hence, commercialization of genetically modified crops (GMOs) is under strict control and limited by lengthy and expensive regulatory assessment procedures [260].
The advanced GE technology assists to produce precise and targeted mutations without the integration of any DNA sequence in plant genomes. This permits the development of non-transgenic crops with increased yield and improved quality and stress tolerance [261]. These approaches are speedy in contrast to traditional breeding techniques and allow development of transgene-free plants [156]. Plants produced via GE approaches are very similar to plants developed by conventional breeding. Additionally, GE technology takes less time to incorporate desired traits into the plant genome as compared to transgenic breeding, mutation breeding, and traditional breeding. It will take approximately 4–6 years to develop a GE plant with desired trait as compared to the transgenic breeding, which require 8–12 years for the creation of transgenic plant. On the other hand, mutation breeding and conventional breeding need 8–10 years approximately to obtain a desired phenotype [258]. The emergence of advanced GE tools not only revolutionized the world of science, but its economics are also very spectacular as compared to genetically engineered plants. Generally, the total expenditure required to execute a single transformation event would have been approximately a quarter of a million US dollars [262], while the budget needed for GE could be $30 [263], which is astonishingly economical and reliable. Consequently, large amounts of money can be saved on developing and approving genome-edited crops, avoiding laborious and time-consuming field experiments which normally demand many years to regulate a GM crop. In addition, it will eliminate the uncertainty and fear regarding the use of GM crops [212]
There is a primary need to review the current rules and regulations regarding GMOs. In addition, genome manipulation done via GE tools are quite different from a transgenic approach. For example, mutations produced by CIRSPR/Cas9 are small indels as compared to large gene sequence insertion or deletion [264]. Such small indels are most often produced in plants under normal growth environments and can also be generated through conventional mutagens. Additionally, in contrast to GMOs which need stable integration of foreign DNA in the genome, CRISPR/Cas9-mediated GE can be used to develop DNA-free non-transgenic plants with improved traits. As far as the regulatory affairs of these gene-edited plants are concerned, there is no international regulatory framework present at the moment. Two major stakeholders, the USA and European Union (EU), have opposite policies for the regulation of genome-edited plants. The United States Department of Agriculture (USDA) has exempted GE crops from its strict rules and regulations [265], while the EU holds the position to treat genome-edited plants as GMOs. Recently, the European Court of Justice has ordered the verdict that GE plants should be subjected to similar regulatory procedures as in the case of GMOs [266]. This judicial ruling may impede investment in GE techniques and limit their use in modern plant breeding platforms in European countries. In Germany, CIBUSTM canola cases are undecided and no legitimate information has been issued by the European Commission (EC) [267]. In 2011, independent legal experts at the EU suggested some legal categorization of modern plant breeding approaches, including GE technology. A committee was formed by the EC, called the “New Techniques Working Group”, to evaluate plants developed using different breeding techniques that fall under the category of GMOs legislation. By the end of 2011, the assessment was completed and the report finalized but it was never published [268]. Although, some important regulatory entities of the EU (including the French High Council for Biotechnology, the European Plant Science Organization, German Academy of Sciences, and the British Biotechnology and Biological Sciences Research Council) have already proposed that the assessment of GE plants should be based on the specific trait improvement instead of technology executed to develop them. However, a study was carried out by the German Federal Agency for Nature Conservation about GE organisms and they decided that GE organisms must be treated under the same regulations as GMOs, arguing that since GE is a strategy to manipulate the genome to produce targeted modification linked with unfamiliar risks, regardless of genome alterations that happen in nature [269].
In several countries, emerging crop editing tools like meganulcease, TALENs, ZFNs, and CRISPR/Cas9 have been applied for the past decade and they do not come under the category of GMO regulatory laws. The USA and Canada regard gene editing as equivalent to traditional breeding. The United States Department of Agriculture (USDA) granted permission to regularize and develop the CRISPR/Cas9-mediated genome-edited crops. Besides the USA, many other countries such as Brazil, Chile, and Argentina have established advanced regulatory principles for genome-edited crops. Every new discovery in biotechnology has been flawlessly approved due to the trait-based scheme in Canada. A strong and authentic regulatory policy is required to distinguish between GMOs and GE plants. Unfortunately, many countries have not established a clear regulatory policy for GE plants. The extensive use of GE strategies brings many challenges for regulatory bodies, as it requires great technical expertise and reliable evaluating procedures for the regulation of GE crops. Evidently, science-based guidelines that judge genome edited plants in a similar way as plants developed by conventional breeding programs are required to boost the applications of GE for crop improvement. For this, many countries such as the United States, Argentina, Australia, Brazil, Canada, and Chile have issued legal interpretations of various omissions in regulatory rules and exempted GE crops from the strict regulations of GMOs. However, this exemption may be dependent on some strict requirements like absence of foreign gene (Australia), no signs of pest characteristics (USA), and type of trait modification (Canada and other countries).
The USA is the main stakeholder in the world and several regulatory authorities govern the regulation of GE crops including the Food and Drug Administration (FDA), Environmental Protection Agency (EPA), Animal and Plant Health Inspection Service (APHIS), and the and United States Department of Agriculture (USDA). The current policy of the USA regarding GE crops was developed by the USDA and depends on the “Plant Protection Act”. Any GE plant that poses pest characteristics and food safety issues is closely assessed and monitored by the regulatory bodies (USDA, EPA, and FDA). The USDA does not treat GE plants under GMOs regulations (https://www.aphis.usda.gov/aphis/ourfocus/biotechnology/brs-news-and-information/pbi-details). The APHIS has proposed many verdicts regarding the risk assessment of DNA-free GE crops and suggested modifications to the rules in order to eradicate the legislation application about pest and GE crops in 2017 [270]. The United States Department of Agriculture (USDA) acknowledged gene editing as a much faster form of traditional breeding. The USDA has allowed more than ten case-by-case studies of genome editing for cultivation without regulatory permits. These included the development of a high level of amylopectin producing Wax corn by applying the CRISPR/Cas9 tool which has been mutated for the Wx1 gene. Similarly, a CRISPR/Cas9 strategy was applied to produce browning resistance to white button mushrooms by mutating the polyphenol oxidase gene at the Pennsylvania State University [265]. Herbicide-resistant rape seed was produced by the RTDS mechanism. In soybean, drought-resistant genes such as Drb2b and Drb2a were knocked out using the CRIPSR/Cas9 system. Setaria viridis was subjected to the CRISPR/Cas9 technique for delayed flowering by disrupting the ID1 gene and the Camelina genome was edited by Yield10 Bioscience for enhanced oil production. In addition, low phytate level corn has been established using Dow’s ZFN, and resistant wheat against powdery mildew, soybean with a mutated FAD3 gene, and potato with PPO knockout using a TALENs approach were also approved.
Similarly, there is no difference in Canada’s approach towards the GE techniques from the techniques that have foreshadowed it. Canadian plants with novel traits (PNTs) regulations are activated only if the technique produces any specific trait, causing toxicity, allergenicity, and effects on any other organism. All the plant cultivars with specific traits are subjected to PNT regulations, irrespective of how they were produced, suggesting that the plant cultivar could be produced via conventional breeding, conventional mutagenesis, genetic engineering or gene editing. It is anticipated that some of GE techniques may produce novel cultivars that are PNTs, while many of them may not be treated under the PNTs regulations. Thus, in Canada, plant cultivars that are carried through the PNT regulations need open release approval from Health Canada and the Canadian Food Inspection Agency (CFIA) in order to register as approved cultivars for commercial use by industry [271].
Argentina established a functional regulatory framework for regularization of modern plant breeding products (Whelan 2015) [272]. Policy-makers and regulatory bodies have made flexible assessment protocols that depend on case-by-case evaluations. Fundamentally, the regulatory framework of Argentina determines the overall process of developing a GE plant. The plant developed without any transgene integration has been designated as non-GMO. Moreover, if any transgene strategy was applied but the final product is DNA-free, then this is also treated as non-GMO. A regulating body in Argentina, CONABIA, assesses the genome-edited material before giving approval. Under the rule No. 763/11, a simple deletion in genome is not regarded as a GM crop.
Most of the GE tools are introduced by industries, but the CRISPR/Cas technology was discovered by academic research groups. These academic institutions and different companies are contesting to establish intellectual property (IP) sets for speedy commercialization of CRISPR/Cas-based products. Since 2005, a 15 fold increase has been reported in the number of patent applications and 42 patent applications were registered in the USA in 2014. Over the last couple of years, investment in GE bio-enterprise has increased fivefold [273]. It was estimated that the market value of GE technology was about $1.84 billion at the end of 2014, and it is predicted to grow with a 13.75% compound annual growth rate of about $3.51 billion by 2019 [274]. Additionally, the private companies that use CRISPR/Cas in the sector of agriculture, health, and industry have equally played a significant part in the current growth of the GE market. Over $600 million has been received by major companies which use CRISPR/Cas technology over the last decade [275].
To conclude, regulatory authorities need to develop comprehensive regulatory frameworks which direct the utilization of GE tools without constraining research. Furthermore, issues of IP rights and licensing policies need to be scrutinized for GE plants which can be used for commercial purposes.
8. Conclusion and Outlook
The production of safe, low-cost, and nutritive food by adopting sustainable agricultural practices will be a huge task. In this regard, the availability of modern technologies to improve cultivars will be a vital aspect. GE is a powerful tool which is expected to play a crucial role in meeting the increasing demands of crop production to fulfill the needs of an exploding population under a climate change scenario. As compared to conventional breeding methodologies, the molecular breeding strategies aided by GE tools allow scientists to precisely target and edit for desired traits. GE can be used to enhance crop productivity, nutritional value, and develop resistance against biotic as well as abiotic stressors by improving the crop genome. The advanced tools in plant GE have been extensively employed to edit crops for a specific agronomic trait and have been utilized in several breeding platforms for carrying the desired trait for the development of an elite local variety. Thus, modern plant breeding approaches will increase the performance of plant breeding, and gene-edited elite cultivars can be approved for cultivation in specified locations without strict regulatory laws. For the last two decades SSNs such as meganucleases, ZFNs, and TALENs have revolutionized plant GE. These SSNs have many applications in plant GE and can be used for gene insertion, gene deletion, and increasing the efficiency of homologous recombination which allows for more precise and accurate events of gene replacement.
Beside other GE techniques, CRISPR/Cas9 is the most powerful tool for crop improvement. In many plant systems it has been vigorously applied over the last five years for combating abiotic and biotic stressors and to improve other agronomic traits. CRISPR/Cas9 as a GE technology for site-direct mutagenesis has many excellent characteristics including great target specificity, easiness to execute, and low cost, which are unachievable through conventional mutagenic strategies. Further, CRISPR/Cas9 is superior to other first-generation SSNs because RNAs guide the Cas9 nuclease instead of proteins. Several Cas9-mediated techniques are being employed in different plant varieties, and these techniques will offer exceptional knowledge about plant biology and facilitate us to develop improved cultivars with great accuracy and speed via modern plant breeding. Recently some striking developments have been achieved in the CRISPR/Cas9 toolbox to increase the targeted mutagenesis with increased efficiency via base editing, multiplex GE [276], and generation of DNA-free plants. The CRSIPR/Cas9 is a versatile tool for plant GE, due to the fact of its sophisticated toolbox of Cas9 variants such as the CRISPR/Cpf1 system and online accessible bioinformatics tools for designing highly precise delivery systems. The CRISPR/Cas9-based precise GE produces gene replacement, gene insertion, and knockout mutations that are rapidly being used to increase yield, improve quality, and enhance tolerance in crops to boost crop domestication and hybrid breeding. Moreover, CRISPR/Cas9 technology is gaining interest day by day and will be a fundamental GE approach to developing improved plants with desired traits that will aid in accomplishing the goal of zero hunger in the world.
Although CRISPR/Cas9-mediated GE has gained remarkable achievements in crop improvement, there are certain challenges that need to be addressed to develop a more efficient system for plant GE. This includes assembling pangenomes for crop improvement, programmed identification of candidate sites for gene editing via functional genomics, designing of highly efficient delivery systems for GE, and reducing the frequency of off-target editing, deciphering novel pathways for this reduction, and optimization of the Cas9 function. The major pitfalls of CRISPR/Cas9 is the inefficient delivery system for plant transformation because the current protocols are limited to certain tissues [277], genotypes, and crop varieties. The packaging of Cas proteins into delivery vectors poses large barriers for efficient delivery of CRISPR/Cas machinery. Recently, some novel cargo-vector systems have been introduced which show promising potential for efficient delivery systems. For example, carbon nanotubes have been utilized to transfer CRISPR/Cas9 editing constructs into plant leaves. Some other nano-products such as mesoporous silica nanoparticles and layered double hydroxides also have great potential to broaden the accessibility of delivery systems, as they have high transformation efficiencies and little toxicity and cellular damage. Developing improved delivery systems will be vital for efficient targeted and more precise GE for crop improvement. On the other hand, the frequency of off-target effects needs to be addressed more comprehensively, as there are many safety issues linked with CRISRP/Cas9-based bio-products. Luckily, off-target mutations are mostly bearable in plants and mutants, and off-target effects can be detected and eliminated through segregation over successive crosses. Selection of Cas9 requiring long PAMs and designing of sgRNA with close affinity for target sequence, may help to reduce the off-target effects in the future. Therefore, continuous efforts are required to overcome these hurdles in order to increase the experimental versatility and applied skills of the CRISPR/Cas9 toolbox in the future.
The advancement in modern breeding approaches has been greatly acknowledged as an innovation in our capacity to manipulate genomes and has subsequently challenged our understanding and assessment of current regulatory policies. As GE tools are extensively employed in plants, the safety of GE plants is the matter of debate around the globe. Development of regulatory policies for novel crop innovations should be multidimensional, transparent, and be able to distinguish between GMOs and GE events. Hence, to explore the large prospective of modern plant breeding approaches for improved yield and food security, it is necessary to illuminate the clear status of these approaches, including GE, and to fix current regulatory uncertainties. Harnessing the innovative ideas of system biology, synthetic biology, next-generation sequencing, and the latest developments in functional genomic approaches integrated with the advanced tools of CRISPR/Cas9 will permit the development of smart crops with higher yields and improved qualities. In the near future, CRISPR/Cas9 technology can be integrated with speed breeding programs to revolutionize the global agriculture and promise of food security.
Author Contributions
A.R. conceived the idea. A.R. and M.K. wrote the manuscript. F.S. helped in original draft preparation. F.A.J. and A.R. developed figures and organized tables. M.K.H. helped in the literature review. F.A.J., S.Y., F.S., G.M., and H.M.I.A. read, reviewed, and edited the manuscript. M.S.K. provided expert opinion, technical assistance, and finalized the manuscript. All authors listed have made substantial, direct, and intellectual contributions to the work and approved the manuscript.
Funding
This research received no external funding.
Acknowledgments
We are grateful to all the researchers whose contributions have been cited in this review paper, which have helped us to prepare this review paper. Furthermore, we apologize to those authors whose excellent work could not be cited due to space limitations.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| CRISPR | Clustered regularly interspaced short palindromic repeats |
| Cas9 | CRISPR-associated protein 9 |
| GE | Genome Editing |
| SSNs | Site-specific nucleases |
| DSB | Double-stranded breaks |
| NHEJ | Non-homologous end joining |
| HDR | Homology-directed recombination |
| MNs | Meganulceases |
| ZFNs | Zinc-finger nucleases |
| TALENs | Transcription activator-like effector nucleases |
| TALEs | Transcription activator like effectors |
| RVD | Repeat variable di-residues |
| sgRNA | single guide RNA |
| Pre-crRNA | Precursor CRISPR-RNA |
| PAM | Protospacer adjacent motif |
| RNPs | Ribonucleoproteins |
| GMO | Genetically modified organism |
| CBE | Cytosine base editor |
| ABE | Adenine base editor |
References
- Dhankher, O.P.; Foyer, C.H. Climate resilient crops for improving global food security and safety. Plant Cell Environ. 2018, 41, 877–884. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhu, Q.; Chen, Y.; Liu, Y.G. CRISPR/Cas9 platforms for genome editing in plants: Developments and applications. Mol. Plant 2016, 9, 961–974. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.; Li, Y.; Liu, B.; Kong, F.; Tran, L.S.P. Adaptive Mechanisms of Soybean Grown on Salt-Affected Soils. Land Degrad. Dev. 2018, 29, 1054–1064. [Google Scholar] [CrossRef]
- Esfahani, M.N.; Inoue, K.; Chu, H.D.; Nguyen, K.H.; Van Ha, C.; Watanabe, Y.; Burritt, D.J.; Herrera-Estrella, L.; Mochida, K.; Phan Tran, L.S. Comparative transcriptome analysis of nodules of two Mesorhizobium-chickpea associations with differential symbiotic efficiency under phosphate deficiency. Plant J. 2017, 91, 911–926. [Google Scholar] [CrossRef] [PubMed]
- Abdelrahman, M.; Sawada, Y.; Nakabayashi, R.; Sato, S.; Hirakawa, H.; El-Sayed, M.; Hirai, M.Y.; Saito, K.; Yamauchi, N.; Shigyo, M. Integrating transcriptome and target metabolome variability in doubled haploids of Allium cepa for abiotic stress protection. Mol. Breed. 2015, 35, 195. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, B.; Ding, W.; Liu, X.; Yang, D.L.; Wei, P.; Cao, F.; Zhu, S.; Zhang, F.; Mao, Y.; et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229–1232. [Google Scholar] [CrossRef]
- Abdelrahman, M.; Jogaiah, S.; Burritt, D.J.; Tran, L.S.P. Legume genetic resources and transcriptome dynamics under abiotic stress conditions. Plant Cell Environ. 2018, 41, 1972–1983. [Google Scholar] [CrossRef]
- Brooks, C.; Nekrasov, V.; Lippman, Z.B.; Van Eck, J. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 2014, 166, 1292–1297. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A Programmable Dual-RNA —Guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Kim, H.; Kim, J.S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 2014, 15, 321–334. [Google Scholar] [CrossRef]
- Zhang, Z.; Hua, L.; Gupta, A.; Tricoli, D.; Edwards, K.J.; Yang, B.; Li, W. Development of an Agrobacterium-delivered CRISPR/Cas9 system for wheat genome editing. Plant Biotechnol. J. 2019, 17, 1623–1635. [Google Scholar] [CrossRef]
- Waltz, E. With a free pass, CRISPR-edited plants reach market in record time. Nat. Biotechnol. 2018, 36, 6–7. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Chen, K.; Liang, Z.; Li, J.; Zhang, Y.; Zhang, K.; Liu, J.; Voytas, D.F.; Zheng, X.; et al. Rapid and efficient gene modification in rice and Brachypodium using TALENs. Mol. Plant 2013, 6, 1365–1368. [Google Scholar] [CrossRef]
- Puchta, H.; Dujon, B.; Hohn, B. Two different but related mechanisms are used in plants for the repair of genomic double-strand breaks by homologous recombination. Proc. Natl. Acad. Sci. USA 1996, 93, 5055–5060. [Google Scholar] [CrossRef]
- Puchta, H. The repair of double-strand breaks in plants: Mechanisms and consequences for genome evolution. J. Exp. Bot. 2005, 56, 1–14. [Google Scholar] [CrossRef]
- Li, J.; Meng, X.; Zong, Y.; Chen, K.; Zhang, H.; Liu, J.; Li, J.; Gao, C. Gene replacements and insertions in rice by intron targeting using CRISPR-Cas9. Nat. Plants 2016, 2, 1–6. [Google Scholar] [CrossRef]
- Puchta, H.; Dujon, B.; Hohn, B. Homologous recombination in plant cells is enhanced by in vivo induction of double strand breaks into DNA by a site-specific endonuclease. Nucleic Acids Res. 1993, 21, 5034–5040. [Google Scholar] [CrossRef]
- Kim, Y.G.; Cha, J.; Chandrasegaran, S. Hybrid restriction enzymes: Zinc finger fusions to FokI cleavage domain. Proc. Natl. Acad. Sci. USA 1996, 93, 1156–1160. [Google Scholar] [CrossRef]
- Bibikova, M.; Beumer, K.; Trautman, J.K.; Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 2003, 300, 764. [Google Scholar] [CrossRef]
- Lloyd, A.; Plaisier, C.L.; Carroll, D.; Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 2005, 102, 2232–2237. [Google Scholar] [CrossRef]
- Gupta, M.; Dekelver, R.C.; Palta, A.; Clifford, C.; Gopalan, S.; Miller, J.C.; Novak, S.; Desloover, D.; Gachotte, D.; Connell, J.; et al. Transcriptional activation of Brassica napus β-ketoacyl-ACP synthase II with an engineered zinc finger protein transcription factor. Plant Biotechnol. J. 2012, 10, 783–791. [Google Scholar] [CrossRef]
- Cermak, T.; Doyle, E.L.; Christian, M.; Wang, L.; Zhang, Y.; Schmidt, C.; Baller, J.A.; Somia, N.V.; Bogdanove, A.J.; Voytas, D.F. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 2011, 39, 7879. [Google Scholar] [CrossRef]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas Systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef]
- Ma, X.; Zhang, Q.; Zhu, Q.; Liu, W.; Chen, Y.; Qiu, R.; Wang, B.; Yang, Z.; Li, H.; Lin, Y.; et al. A Robust CRISPR/Cas9 System for Convenient, High-Efficiency Multiplex Genome Editing in Monocot and Dicot Plants. Mol. Plant 2015, 8, 1274–1284. [Google Scholar] [CrossRef]
- Nekrasov, V.; Staskawicz, B.; Weigel, D.; Jones, J.D.G.; Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 691–693. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qiu, J.L.; et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 686–688. [Google Scholar] [CrossRef]
- Moscou, M.J.; Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 2009, 326, 1501. [Google Scholar] [CrossRef]
- Boch, J.; Scholze, H.; Schornack, S.; Landgraf, A.; Hahn, S.; Kay, S.; Lahaye, T.; Nickstadt, A.; Bonas, U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009, 326, 1509–1512. [Google Scholar] [CrossRef]
- Prieto, J.; Redondo, P.; Padró, D.; Arnould, S.; Epinat, J.C.; Pâques, F.; Blanco, F.J.; Montoya, G. The Cterminal loop of the homing endonuclease I-CreI is essential for site recognition, DNA binding and cleavage. Nucleic Acids Res. 2007, 35, 3262–3271. [Google Scholar] [CrossRef][Green Version]
- D’Halluin, K.; Vanderstraeten, C.; Van Hulle, J.; Rosolowska, J.; Van Den Brande, I.; Pennewaert, A.; D’Hont, K.; Bossut, M.; Jantz, D.; Ruiter, R.; et al. Targeted molecular trait stacking in cotton through targeted double-strand break induction. Plant Biotechnol. J. 2013, 11, 933–941. [Google Scholar] [CrossRef]
- Stoddard, B.L. Homing endonucleases: From microbial genetic invaders to reagents for targeted DNA modification. Structure 2011, 19, 7–15. [Google Scholar] [CrossRef]
- Marton, I.; Zuker, A.; Shklarman, E.; Zeevi, V.; Tovkach, A.; Roffe, S.; Ovadis, M.; Tzfira, T.; Vainstein, A. Nontransgenic genome modification in plant cells. Plant Physiol. 2010, 154, 1079–1087. [Google Scholar] [CrossRef]
- Ran, Y.; Patron, N.; Kay, P.; Wong, D.; Buchanan, M.; Cao, Y.Y.; Sawbridge, T.; Davies, J.P.; Mason, J.; Webb, S.R.; et al. Zinc finger nuclease-mediated precision genome editing of an endogenous gene in hexaploid bread wheat (Triticum aestivum) using a DNA repair template. Plant Biotechnol. J. 2018, 16, 2088–2101. [Google Scholar] [CrossRef]
- Bonawitz, N.D.; Ainley, W.M.; Itaya, A.; Chennareddy, S.R.; Cicak, T.; Effinger, K.; Pareddy, D.R. Zinc finger nuclease-mediated targeting of multiple transgenes to an endogenous soybean genomic locus via non-homologous end joining. Plant Biotechnol. J. 2018, 17, 750–761. [Google Scholar] [CrossRef]
- Cai, C.Q.; Doyon, Y.; Ainley, W.M.; Miller, J.C.; DeKelver, R.C.; Moehle, E.A.; Rock, J.M.; Lee, Y.L.; Garrison, R.; Schulenberg, L.; et al. Targeted transgene integration in plant cells using designed zinc finger nucleases. Plant Mol. Biol. 2009, 69, 699–709. [Google Scholar] [CrossRef]
- Zhang, F.; Maeder, M.L.; Unger-Wallace, E.; Hoshaw, J.P.; Reyon, D.; Christian, M.; Li, X.; Pierick, C.J.; Dobbs, D.; Peterson, T.; et al. High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. Proc. Natl. Acad. Sci. USA 2010, 107, 12028–12033. [Google Scholar] [CrossRef]
- Cantos, C.; Francisco, P.; Trijatmiko, K.R.; Slamet-Loedin, I.; Chadha-Mohanty, P.K. Identification of “safe harbor” loci in indica rice genome by harnessing the property of zinc-finger nucleases to induce DNA damage and repair. Front. Plant Sci. 2014, 5, 302. [Google Scholar] [CrossRef]
- Vanamee, É.S.; Santagata, S.; Aggarwal, A.K. FokI requires two specific DNA sites for cleavage. J. Mol. Biol. 2001, 309, 69–78. [Google Scholar] [CrossRef]
- Shukla, V.K.; Doyon, Y.; Miller, J.C.; Dekelver, R.C.; Moehle, E.A.; Worden, S.E.; Mitchell, J.C.; Arnold, N.L.; Gopalan, S.; Meng, X.; et al. Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 2009, 459, 437–441. [Google Scholar] [CrossRef]
- Curtin, S.J.; Zhang, F.; Sander, J.D.; Haun, W.J.; Starker, C.; Baltes, N.J.; Reyon, D.; Dahlborg, E.J.; Goodwin, M.J.; Coffman, A.P.; et al. Targeted mutagenesis of duplicated genes in soybean with Zinc-Finger Nucleases. Plant Physiol. 2011, 156, 466–473. [Google Scholar] [CrossRef]
- Ramirez, C.L.; Foley, J.E.; Wright, D.A.; Müller-Lerch, F.; Rahman, S.H.; Cornu, T.I.; Winfrey, R.J.; Sander, J.D.; Fu, F.; Townsend, J.A.; et al. Unexpected failure rates for modular assembly of engineered zinc fingers. Nat. Methods 2008, 5, 575. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, F.; Li, X.; Baller, J.A.; Qi, Y.; Starker, C.G.; Bogdanove, A.J.; Voytas, D.F. Transcription Activator-Like Effector Nucleases enable efficient plant genome engineering. Plant Physiol. 2013, 161, 20–27. [Google Scholar] [CrossRef]
- Li, T.; Liu, B.; Spalding, M.H.; Weeks, D.P.; Yang, B. High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat. Biotechnol. 2012, 30, 390–392. [Google Scholar] [CrossRef]
- Bonas, U.; Stall, R.E.; Staskawicz, B. Genetic and structural characterization of the avirulence gene avrBs3 from Xanthomonas campestris pv. vesicatoria. MGG Mol. Gen. Genet. 1989, 218, 127–136. [Google Scholar] [CrossRef]
- Pennisi, E. The tale of the TALEs. Science 2012, 338, 1408–1411. [Google Scholar] [CrossRef]
- Reyon, D.; Tsai, S.Q.; Khgayter, C.; Foden, J.A.; Sander, J.D.; Joung, J.K. FLASH assembly of TALENs for high-throughput genome editing. Nat. Biotechnol. 2012, 30, 460–465. [Google Scholar] [CrossRef]
- Briggs, A.W.; Rios, X.; Chari, R.; Yang, L.; Zhang, F.; Mali, P.; Church, G.M. Iterative capped assembly: Rapid and scalable synthesis of repeat-module DNA such as TAL effectors from individual monomers. Nucleic Acids Res. 2012, 40, e117. [Google Scholar] [CrossRef]
- Schmid-Burgk, J.L.; Schmidt, T.; Kaiser, V.; Höning, K.; Hornung, V. A ligation-independent cloning technique for high-throughput assembly of transcription activator-like effector genes. Nat. Biotechnol. 2013, 31, 76–81. [Google Scholar] [CrossRef]
- Mussolino, C.; Morbitzer, R.; Lütge, F.; Dannemann, N.; Lahaye, T.; Cathomen, T. A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res. 2011, 39, 9283–9293. [Google Scholar] [CrossRef]
- Djukanovic, V.; Smith, J.; Lowe, K.; Yang, M.; Gao, H.; Jones, S.; Nicholson, M.G.; West, A.; Lape, J.; Bidney, D.; et al. Male-sterile maize plants produced by targeted mutagenesis of the cytochrome P450-like gene (MS26) using a re-designed I-CreI homing endonuclease. Plant J. 2013, 76, 888–899. [Google Scholar] [CrossRef]
- Gao, H.; Smith, J.; Yang, M.; Jones, S.; Djukanovic, V.; Nicholson, M.G.; West, A.; Bidney, D.; Falco, S.C.; Jantz, D.; et al. Heritable targeted mutagenesis in maize using a designed endonuclease. Plant J. 2010, 61, 176–187. [Google Scholar] [CrossRef]
- Ainley, W.M.; Sastry-Dent, L.; Welter, M.E.; Murray, M.G.; Zeitler, B.; Amora, R.; Corbin, D.R.; Miles, R.R.; Arnold, N.L.; Strange, T.L.; et al. Trait stacking via targeted genome editing. Plant Biotechnol. J. 2013, 11, 1126–1134. [Google Scholar] [CrossRef]
- Kannan, B.; Jung, J.H.; Moxley, G.W.; Lee, S.; Altpeter, F. TALEN-mediated targeted mutagenesis of more than 100 COMT copies/alleles in highly polyploid sugarcane improves saccharification efficiency without compromising biomass yield. Plant Biotechnol. J. 2018, 16, 856–866. [Google Scholar] [CrossRef]
- Kelliher, T.; Starr, D.; Richbourg, L.; Chintamanani, S.; Delzer, B.; Nuccio, M.L.; Green, J.; Chen, Z.; McCuiston, J.; Wang, W.; et al. MATRILINEAL, a sperm-specific phospholipase, triggers maize haploid induction. Nature 2017, 542, 105–109. [Google Scholar] [CrossRef]
- Du, H.; Zeng, X.; Zhao, M.; Cui, X.; Wang, Q.; Yang, H.; Cheng, H.; Yu, D. Efficient targeted mutagenesis in soybean by TALENs and CRISPR/Cas9. J. Biotechnol. 2016, 217, 90–97. [Google Scholar] [CrossRef]
- Demorest, Z.L.; Coffman, A.; Baltes, N.J.; Stoddard, T.J.; Clasen, B.M.; Luo, S.; Retterath, A.; Yabandith, A.; Gamo, M.E.; Bissen, J.; et al. Direct stacking of sequence-specific nuclease-induced mutations to produce high oleic and low linolenic soybean oil. BMC Plant Biol. 2016, 16, 225. [Google Scholar] [CrossRef]
- Clasen, B.M.; Stoddard, T.J.; Luo, S.; Demorest, Z.L.; Li, J.; Cedrone, F.; Tibebu, R.; Davison, S.; Ray, E.E.; Daulhac, A.; et al. Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotechnol. J. 2016, 14, 169–176. [Google Scholar] [CrossRef]
- Jung, J.H.; Altpeter, F. TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol. Biol. 2016, 92, 131–142. [Google Scholar] [CrossRef]
- Nicolia, A.; Proux-Wéra, E.; Åhman, I.; Onkokesung, N.; Andersson, M.; Andreasson, E.; Zhu, L.H. Targeted gene mutation in tetraploid potato through transient TALEN expression in protoplasts. J. Biotechnol. 2015, 204, 17–24. [Google Scholar] [CrossRef]
- Char, S.N.; Unger-Wallace, E.; Frame, B.; Briggs, S.A.; Main, M.; Spalding, M.H.; Vollbrecht, E.; Wang, K.; Yang, B. Heritable site-specific mutagenesis using TALENs in maize. Plant Biotechnol. J. 2015, 13, 1002–1010. [Google Scholar] [CrossRef]
- Shan, Q.; Zhang, Y.; Chen, K.; Zhang, K.; Gao, C. Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology. Plant Biotechnol. J. 2015, 13, 791–800. [Google Scholar] [CrossRef]
- Wang, Y.; Cheng, X.; Shan, Q.; Zhang, Y.; Liu, J.; Gao, C.; Qiu, J. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 2014, 32, 947–952. [Google Scholar] [CrossRef]
- Gurushidze, M.; Hensel, G.; Hiekel, S.; Schedel, S.; Valkov, V.; Kumlehn, J. True-breeding targeted gene knock-out in barley using designer TALE-nuclease in haploid cells. PLoS ONE 2014, 9, e92046. [Google Scholar] [CrossRef]
- Haun, W.; Coffman, A.; Clasen, B.M.; Demorest, Z.L.; Lowy, A.; Ray, E.; Retterath, A.; Stoddard, T.; Juillerat, A.; Cedrone, F.; et al. Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnol. J. 2014, 12, 934–940. [Google Scholar] [CrossRef]
- Alagoz, Y.; Gurkok, T.; Zhang, B.; Unver, T. Manipulating the biosynthesis of bioactive compound alkaloids for next-generation metabolic engineering in opium poppy using CRISPR-Cas 9 genome editing technology. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Guell, M.; DiCarlo, J.E.; Norville, J.E.; Church, G.M. RNA-guided human genome engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef]
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakatura, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isoenzyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 1987, 169, 5429–5433. [Google Scholar] [CrossRef]
- Mojica, F.J.M.; Ferrer, C.; Juez, G.; Rodríguez-Valera, F. Long stretches of short tandem repeats are present in the largest replicons of the Archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol. Microbiol. 1995, 17, 85–93. [Google Scholar] [CrossRef]
- Mojica, F.J.M.; Díez-Villaseñor, C.; Soria, E.; Juez, G. Biological significance of a family of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria. Mol. Microbiol. 2000, 36, 244–246. [Google Scholar] [CrossRef]
- Jansen, R.; Van Embden, J.D.A.; Gaastra, W.; Schouls, L.M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 2002, 43, 1565–1575. [Google Scholar] [CrossRef]
- Karginov, F.V.; Hannon, G.J. The CRISPR system: Small RNA-guided defense in bacteria and archaea. Mol. Cell 2010, 37, 7–19. [Google Scholar] [CrossRef]
- Haft, D.H.; Selengut, J.; Mongodin, E.F.; Nelson, K.E. A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/cas subtypes exist in prokaryotic genomes. PLoS Comput. Biol. 2005, 1, e60. [Google Scholar] [CrossRef]
- Mojica, F.J.M.; Díez-Villaseñor, C.; García-Martínez, J.; Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 2005, 60, 174–182. [Google Scholar] [CrossRef]
- Bolotin, A.; Quinquis, B.; Sorokin, A.; Dusko Ehrlich, S. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 2005, 151, 2551–2561. [Google Scholar] [CrossRef]
- Pourcel, C.; Salvignol, G.; Vergnaud, G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 2005, 151, 653–663. [Google Scholar] [CrossRef]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D.A.; Horvath, P. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef]
- Marraffini, L.A.; Sontheimer, E.J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 2008, 322, 1843–1845. [Google Scholar] [CrossRef]
- Brouns, S.J.J.; Jore, M.M.; Lundgren, M.; Westra, E.R.; Slijkhuis, R.J.H.; Snijders, A.P.L.; Dickman, M.J.; Makarova, K.S.; Koonin, E.V.; Oost, J. Antiviral defense in prokaryotes. Science 2008, 321, 960–964. [Google Scholar] [CrossRef]
- Garneau, J.E.; Dupuis, M.È.; Villion, M.; Romero, D.A.; Barrangou, R.; Boyaval, P.; Fremaux, C.; Horvath, P.; Magadán, A.H.; Moineau, S. He CRISPR/cas bacterial immune system cleaves bacteriophage and plasmid DNAT. Nature 2010, 468, 67–71. [Google Scholar] [CrossRef]
- Sapranauskas, R.; Gasiunas, G.; Fremaux, C.; Barrangou, R.; Horvath, P.; Siksnys, V. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 2011, 39, 9275–9282. [Google Scholar] [CrossRef]
- Pawluk, A.; Davidson, A.R.; Maxwell, K.L. Anti-CRISPR: Discovery, mechanism and function. Nat. Rev. Microbiol. 2018, 16, 12–17. [Google Scholar] [CrossRef]
- Bhaya, D.; Davison, M.; Barrangou, R. CRISPR-Cas systems in bacteria and archaea: Versatile small RNAs for adaptive defense and regulation. Annu. Rev. Genet. 2011, 45, 273–297. [Google Scholar] [CrossRef]
- Bortesi, L.; Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 2015, 33, 41–52. [Google Scholar] [CrossRef]
- Mojica, F.J.M.; Díez-Villaseñor, C.; García-Martínez, J.; Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 2009, 155, 733–740. [Google Scholar] [CrossRef]
- Zhang, Y.; Heidrich, N.; Ampattu, B.J.; Gunderson, C.W.; Seifert, H.S.; Schoen, C.; Vogel, J.; Sontherimer, E.J. Processing-Independent CRISPR RNAs limit natural transformation in Neisseria meningitidis. Mol. Cell 2013, 50, 488–503. [Google Scholar] [CrossRef]
- Gasiunas, G.; Barrangou, R.; Horvath, P.; Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. USA 2012, 109, E2579–E2586. [Google Scholar] [CrossRef]
- Makarova, K.S.; Aravind, L.; Wolf, Y.I.; Koonin, E.V. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol. Direct 2011, 6, 38. [Google Scholar] [CrossRef]
- Makarova, K.S.; Wolf, Y.I.; Alkhnbashi, O.S.; Costa, F.; Shah, S.A.; Saunders, S.J.; Barrangou, R.; Brouns, S.J.J.; Charpentier, E.; Haft, D.H.; et al. An updated evolutionary classification of CRISPR-Cas systems. Nat. Rev. Microbiol. 2015, 13, 722–736. [Google Scholar] [CrossRef]
- Makarova, K.S.; Haft, D.H.; Barrangou, R.; Brouns, S.J.J.; Charpentier, E.; Horvath, P.; Moineau, S.; Mojica, F.J.M.; Wolf, Y.I.; Yakunin, A.F.; et al. Evolution and classification of the CRISPR-Cas systems. Nat. Rev. Microbiol. 2011, 9, 467–477. [Google Scholar] [CrossRef]
- Rath, D.; Amlinger, L.; Rath, A.; Lundgren, M. The CRISPR-Cas immune system: Biology, mechanisms and applications. Biochimie 2015, 117, 119–128. [Google Scholar] [CrossRef]
- Deltcheva, E.; Chylinski, K.; Sharma, C.M.; Gonzales, K.; Chao, Y.; Pirzada, Z.A.; Eckert, M.R.; Vogel, J.; Charpentier, E. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 2011, 471, 602–607. [Google Scholar] [CrossRef]
- Nishimasu, H.; Ran, F.A.; Hsu, P.D.; Konermann, S.; Shehata, S.I.; Dohmae, N.; Ishitani, R.; Zhang, F.; Nureki, O. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 2014, 156, 935–949. [Google Scholar] [CrossRef]
- Li, J.F.; Norville, J.E.; Aach, J.; McCormack, M.; Zhang, D.; Bush, J.; Church, G.M.; Sheen, J. Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 2013, 31, 688–691. [Google Scholar] [CrossRef]
- Schiml, S.; Fauser, F.; Puchta, H. The CRISPR/Cas system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J. 2014, 80, 1139–1150. [Google Scholar] [CrossRef]
- Miao, J.; Guo, D.; Zhang, J.; Huang, Q.; Qin, G.; Zhang, X.; Wan, J.; Gu, H.; Qu, L.J. Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res. 2013, 23, 1233–1236. [Google Scholar] [CrossRef]
- Belhaj, K.; Chaparro-Garcia, A.; Kamoun, S.; Nekrasov, V. Plant genome editing made easy: Targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 2013, 9, 39. [Google Scholar] [CrossRef]
- Sun, X.; Hu, Z.; Chen, R.; Jiang, Q.; Song, G.; Zhang, H.; Xi, Y. Targeted mutagenesis in soybean using the CRISPR-Cas9 system. Sci. Rep. 2015, 5, 10342. [Google Scholar] [CrossRef]
- Mao, Y.; Botella, J.R.; Zhu, J.K. Heritability of targeted gene modifications induced by plant-optimized CRISPR systems. Cell. Mol. Life Sci. 2017, 74, 1075–1093. [Google Scholar] [CrossRef]
- Lowder, L.G.; Zhang, D.; Baltes, N.J.; Paul, J.W.; Tang, X.; Zheng, X.; Voytas, D.F.; Hsieh, T.F.; Zhang, Y.; Qi, Y. A CRISPR/Cas9 Toolbox for Multiplexed Plant Genome Editing and Transcriptional Regulation. Plant Physiol. 2015, 169, 971–985. [Google Scholar] [CrossRef]
- Stemmer, M.; Thumberger, T.; Del Sol Keyer, M.; Wittbrodt, J.; Mateo, J.L. CCTop: An intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS ONE 2015, 10, e0124633. [Google Scholar] [CrossRef]
- Hsu, P.D.; Scott, D.A.; Weinstein, J.A.; Ran, F.A.; Konermann, S.; Agarwala, V.; Li, Y.; Fine, E.J.; Wu, X.; Shalem, O.; et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 2013, 31, 827–832. [Google Scholar] [CrossRef]
- Xie, K.; Zhang, J.; Yang, Y. Genome-wide prediction of highly specific guide RNA spacers for CRISPR–Cas9-Mediated genome editing in model plants and major crops. Mol. Plant 2014, 7, 923–926. [Google Scholar] [CrossRef]
- Michno, J.M.; Wang, X.; Liu, J.; Curtin, S.J.; Kono, T.J.; Stupar, R.M. CRISPR/Cas mutagenesis of soybean and Medicago truncatula using a new web-tool and a modified Cas9 enzyme. GM Crop. Food 2015, 6, 243–252. [Google Scholar] [CrossRef]
- Lei, Y.; Lu, L.; Liu, H.Y.; Li, S.; Xing, F.; Chen, L.L. CRISPR-P: A web tool for synthetic single-guide RNA design of CRISPR-system in Plants. Mol. Plant 2014, 7, 1494–1496. [Google Scholar] [CrossRef]
- Chen, W.; Zhang, G.; Li, J.; Zhang, X.; Huang, S.; Xiang, S.; Hu, X.; Liu, C. CRISPRlnc: A manually curated database of validated sgRNAs for lncRNAs. Nucleic Acids Res. 2019, 47, D63–D68. [Google Scholar] [CrossRef]
- Sun, J.; Liu, H.; Liu, J.; Cheng, S.; Peng, Y.; Zhang, Q.; Yan, J.; Liu, H.J.; Chen, L.L. CRISPR-Local: A local single-guide RNA (sgRNA) design tool for non-reference plant genomes. Bioinformatics 2018. [Google Scholar] [CrossRef]
- Chari, R.; Yeo, N.C.; Chavez, A.; Church, G. Sgrna Scorer 2.0—A species independent model to predict CRISPR/Cas9 activity. ACS Synth. Biol. 2017, 6, 902–904. [Google Scholar] [CrossRef]
- Liu, H.; Ding, Y.; Zhou, Y.; Jin, W.; Xie, K.; Chen, L.L. CRISPR-P 2.0: An Improved CRISPR-Cas9 Tool for Genome Editing in Plants. Mol. Plant 2017, 10, 530–532. [Google Scholar] [CrossRef]
- Rahman, M.K.; Rahman, M.S. CRISPRpred: A flexible and efficient tool for sgRNAs on-target activity prediction in CRISPR/Cas9 systems. PLoS ONE 2017, 12, e0181943. [Google Scholar] [CrossRef]
- Ma, J.; Köster, J.; Qin, Q.; Hu, S.; Li, W.; Chen, C.; Cao, Q.; Wang, J.; Mei, S.; Liu, Q.; et al. CRISPR-DO for genome-wide CRISPR design and optimization. Bioinformatics 2016, 32, 3336–3338. [Google Scholar] [CrossRef]
- Rastogi, A.; Murik, O.; Bowler, C.; Tirichine, L. PhytoCRISP-Ex: A web-based and stand-alone application to find specific target sequences for CRISPR/CAS editing. BMC Bioinform. 2016, 17, 261. [Google Scholar] [CrossRef]
- Blin, K.; Pedersen, L.E.; Weber, T.; Lee, S.Y. CRISPy-web: An online resource to design sgRNAs for CRISPR applications. Synth. Syst. Biotechnol. 2016, 1, 118–121. [Google Scholar] [CrossRef]
- Park, J.; Bae, S.; Kim, J.S. Cas-Designer: A web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 2015, 31, 4014–4016. [Google Scholar] [CrossRef]
- Fusi, N.; Smith, I.; Doench, J.; Listgarten, J. In Silico Predictive Modeling of CRISPR/Cas9 guide efficiency. BioRxiv 2015, 021568. [Google Scholar] [CrossRef]
- Naito, Y.; Hino, K.; Bono, H.; Ui-Tei, K. CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 2015, 31, 1120–1123. [Google Scholar] [CrossRef]
- Zhu, L.J.; Holmes, B.R.; Aronin, N.; Brodsky, M.H. CRISPRseek: A Bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS ONE 2014, 9, e108424. [Google Scholar] [CrossRef]
- Bae, S.; Park, J.; Kim, J.S. Cas-OFFinder: A fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 2014, 30, 1473–1475. [Google Scholar] [CrossRef]
- Heigwer, F.; Kerr, G.; Boutros, M. E-CRISP: Fast CRISPR target site identification. Nat. Methods 2014, 11, 122–123. [Google Scholar] [CrossRef]
- Upadhyay, S.K.; Sharma, S. SSFinder: High Throughput CRISPR-Cas Target Sites Prediction Tool. Biomed Res. Int. 2014, 2014, 4. [Google Scholar] [CrossRef]
- Doench, J.G.; Hartenian, E.; Graham, D.B.; Tothova, Z.; Hegde, M.; Smith, I.; Sullender, M.; Ebert, B.L.; Xavier, R.J.; Root, D.E. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat. Biotechnol. 2014, 32, 1262–1267. [Google Scholar] [CrossRef]
- Montague, T.G.; Cruz, J.M.; Gagnon, J.A.; Church, G.M.; Valen, E. CHOPCHOP: A CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 2014, 42, 401–407. [Google Scholar] [CrossRef]
- Xie, S.; Shen, B.; Zhang, C.; Huang, X.; Zhang, Y. SgRNAcas9: A software package for designing CRISPR sgRNA and evaluating potential off-target cleavage sites. PLoS ONE 2014, 9, e0100448. [Google Scholar] [CrossRef]
- Xie, K.; Yang, Y. RNA-Guided genome editing in plants using a CRISPR-Cas system. Mol. Plant 2013, 6, 1975–1983. [Google Scholar] [CrossRef]
- Gao, Y.; Zhao, Y. Self-processing of ribozyme-flanked RNAs into guide RNAs in vitro and in vivo for CRISPR-mediated genome editing. J. Integr. Plant Biol. 2014, 56, 343–349. [Google Scholar] [CrossRef]
- Fauser, F.; Schiml, S.; Puchta, H. Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J. 2014, 79, 348–359. [Google Scholar] [CrossRef]
- Wong, G.K.S.; Wang, J.; Tao, L.; Tan, J.; Zhang, J.; Passey, D.A.; Yu, J. Compositional gradients in Gramineae genes. Genome Res. 2002, 12, 851–856. [Google Scholar] [CrossRef]
- Baltes, N.J.; Gil-Humanes, J.; Cermak, T.; Atkins, P.A.; Voytas, D.F. DNA Replicons for plant genome engineering. Plant Cell 2014, 26, 151–163. [Google Scholar] [CrossRef]
- Ali, Z.; Eid, A.; Ali, S.; Mahfouz, M.M. Pea early-browning virus-mediated genome editing via the CRISPR/Cas9 system in Nicotiana benthamiana and Arabidopsis. Virus Res. 2018, 244, 333–337. [Google Scholar] [CrossRef]
- Jiang, W.; Zhou, H.; Bi, H.; Fromm, M.; Yang, B.; Weeks, D.P. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 2013, 41, e188. [Google Scholar] [CrossRef]
- Wolter, F.; Puchta, H. Knocking out consumer concerns and regulator’s rules: Efficient use of CRISPR/Cas ribonucleoprotein complexes for genome editing in cereals. Genome Biol. 2017, 18, 682–698. [Google Scholar] [CrossRef]
- Char, S.N.; Neelakandan, A.K.; Nahampun, H.; Frame, B.; Main, M.; Spalding, M.H.; Becraft, P.W.; Meyers, B.C.; Walbot, V.; Wang, K.; et al. An Agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol. J. 2017, 15, 257–268. [Google Scholar] [CrossRef]
- Zhang, S.; Zhang, R.; Song, G.; Gao, J.; Li, W.; Han, X.; Chen, M.; Li, Y.; Li, G. Targeted mutagenesis using the Agrobacterium tumefaciens-mediated CRISPR-Cas9 system in common wheat. BMC Plant Biol. 2018, 18, 302. [Google Scholar] [CrossRef]
- Jacobs, T.B.; Zhang, N.; Patel, D.; Martin, G.B. Generation of a collection of mutant tomato lines using pooled CRISPR libraries. Plant Physiol. 2017, 174, 2023–2037. [Google Scholar] [CrossRef]
- Meng, X.; Yu, H.; Zhang, Y.; Zhuang, F.; Song, X.; Gao, S.; Gao, C.; Li, J. Construction of a genome-wide mutant library in rice using CRISPR/Cas9. Mol. Plant 2017, 10, 1238–1241. [Google Scholar] [CrossRef]
- Lu, Y.; Ye, X.; Guo, R.; Huang, J.; Wang, W.; Tang, J.; Tan, L.; Zhu, J.; Chu, C.; Qian, Y. genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system. Mol. Plant 2017, 10, 1242–1245. [Google Scholar] [CrossRef]
- Hua, Y.; Wang, C.; Huang, J.; Wang, K. A simple and efficient method for CRISPR/Cas9-induced mutant screening. J. Genet. Genomics 2017, 44, 207–213. [Google Scholar] [CrossRef]
- Thomas, H.R.; Percival, S.M.; Yoder, B.K.; Parant, J.M. High-throughput genome editing and phenotyping facilitated by high resolution melting curve analysis. PLoS ONE 2014, 9, e114632. [Google Scholar] [CrossRef]
- Zhu, X.; Xu, Y.; Yu, S.; Lu, L.; Ding, M.; Cheng, J.; Song, G.; Gao, X.; Yao, L.; Fan, D.; et al. An efficient genotyping method for genome-modified animals and human cells generated with CRISPR/Cas9 system. Sci. Rep. 2014, 4, 6420. [Google Scholar] [CrossRef]
- Vouillot, L.; Thélie, A.; Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3 Genes Genomes Genet. 2015, 5, 407–415. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Li, J.; Gao, C. Genome editing in rice and wheat using the CRISPR/Cas system. Nat. Protoc. 2014, 9, 2395–2410. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, J.; Wei, P.; Zhang, B.; Gou, F.; Feng, Z.; Mao, Y.; Yang, L.; Zhang, H.; Xu, N.; et al. The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol. J. 2014, 12, 797–807. [Google Scholar] [CrossRef]
- Chen, L.; Li, W.; Katin-Grazzini, L.; Ding, J.; Gu, X.; Li, Y.; Gu, T.; Wang, R.; Lin, X.; Deng, Z.; et al. A method for the production and expedient screening of CRISPR/Cas9-mediated non-transgenic mutant plants. Hortic. Res. 2018, 5, 13. [Google Scholar] [CrossRef]
- Lu, H.P.; Liu, S.M.; Xu, S.L.; Chen, W.Y.; Zhou, X.; Tan, Y.Y.; Huang, J.Z.; Shu, Q.Y. CRISPR-S: An active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants. Plant Biotechnol. J. 2017, 15, 1371–1373. [Google Scholar] [CrossRef]
- Bell, C.C.; Magor, G.W.; Gillinder, K.R.; Perkins, A.C. A high-throughput screening strategy for detecting CRISPR-Cas9 induced mutations using next-generation sequencing. BMC Genomics 2014, 15, 1002. [Google Scholar] [CrossRef]
- Braatz, J.; Harloff, H.J.; Mascher, M.; Stein, N.; Himmelbach, A.; Jung, C. CRISPR-Cas9 targeted mutagenesis leads to simultaneous modification of different homoeologous gene copies in polyploid oilseed rape (Brassica napus). Plant Physiol. 2017, 174, 935–942. [Google Scholar] [CrossRef]
- Gao, X.; Chen, J.; Dai, X.; Zhang, D.; Zhao, Y. An effective strategy for reliably isolating heritable and Cas9 -free Arabidopsis mutants generated by CRISPR/Cas9-mediated genome editing. Plant Physiol. 2016, 171, 1794–1800. [Google Scholar] [CrossRef]
- He, Y.; Zhu, M.; Wang, L.; Wu, J.; Wang, Q.; Wang, R.; Zhao, Y. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 2018, 11, 1210–1213. [Google Scholar] [CrossRef]
- Zhang, Y.; Liang, Z.; Zong, Y.; Wang, Y.; Liu, J.; Chen, K.; Qiu, J.L.; Gao, C. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 2016, 7, 12617. [Google Scholar] [CrossRef]
- Zong, Y.; Wang, Y.; Li, C.; Zhang, R.; Chen, K.; Ran, Y.; Qiu, J.L.; Wang, D.; Gao, C. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 438–440. [Google Scholar] [CrossRef]
- Zong, Y.; Song, Q.; Li, C.; Jin, S.; Zhang, D.; Wang, Y.; Qiu, J.L.; Gao, C. Efficient c-to-t base editing in plants using a fusion of ncas9 and human apobec3a. Nat. Biotechnol. 2018, 36, 950. [Google Scholar] [CrossRef]
- Lin, C.S.; Hsu, C.T.; Yang, L.H.; Lee, L.Y.; Fu, J.Y.; Cheng, Q.W.; Wu, F.H.; Hsiao, H.C.W.; Zhang, Y.; Zhang, R.; et al. Application of protoplast technology to CRISPR/Cas9 mutagenesis: From single-cell mutation detection to mutant plant regeneration. Plant Biotechnol. J. 2018, 16, 1295–1310. [Google Scholar] [CrossRef]
- Andersson, M.; Turesson, H.; Nicolia, A.; Fält, A.S.; Samuelsson, M.; Hofvander, P. Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep. 2017, 36, 117–128. [Google Scholar] [CrossRef]
- Veillet, F.; Perrot, L.; Chauvin, L.; Kermarrec, M.P.; Guyon-Debast, A.; Chauvin, J.E.; Nogué, F.; Mazier, M. Transgene-free genome editing in tomato and potato plants using Agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int. J. Mol. Sci. 2019, 20, 402. [Google Scholar] [CrossRef]
- Kim, S.; Kim, D.; Cho, S.W.; Kim, J.; Kim, J.S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 2014, 24, 1012–1019. [Google Scholar] [CrossRef]
- Jones, H.D. Regulatory uncertainty over genome editing. Nat. Plants 2015, 1, 10–38. [Google Scholar] [CrossRef]
- Woo, J.W.; Kim, J.; Kwon, S.; Corvalán, C.; Cho, S.W.; Kim, H.; Kim, S.G.; Kim, S.T.; Choe, S.; Kim, J.S. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 2015, 33, 1162–1164. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Zhang, Y.; Liu, J.; Yin, K.; Qiu, J.L.; Gao, C. Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat. Protoc. 2018, 13, 413–430. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Li, T.; Zhang, Y.; Wang, Y.; Zhao, Q.; Liu, J.; Zhang, H.; Liu, C.; Ran, Y.; et al. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 2017, 8, 14261. [Google Scholar] [CrossRef]
- Andersson, M.; Turesson, H.; Olsson, N.; Fält, A.S.; Ohlsson, P.; Gonzalez, M.N.; Samuelsson, M.; Hofvander, P. Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol. Plant. 2018, 164, 378–384. [Google Scholar] [CrossRef]
- Mcvey, M.; Khodaverdian, V.Y.; Meyer, D.; Heyer, W.; Genetics, M.; Biology, C.; Cordova, R. Eukaryotic DNA polymerases in homologous recombination. Annu. Rev. Genet. 2017, 393–421. [Google Scholar] [CrossRef]
- Malnoy, M.; Viola, R.; Jung, M.H.; Koo, O.J.; Kim, S.; Kim, J.S.; Velasco, R.; Nagamangala Kanchiswamy, C. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 2016, 7, 1904. [Google Scholar] [CrossRef]
- Svitashev, S.; Schwartz, C.; Lenderts, B.; Young, J.K.; Mark Cigan, A. Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes. Nat. Commun. 2016, 7, 13274. [Google Scholar] [CrossRef]
- Kim, H.; Kim, S.T.; Ryu, J.; Kang, B.C.; Kim, J.S.; Kim, S.G. CRISPR/Cpf1-mediated DNA-free plant genome editing. Nat. Commun. 2017, 8, 14406. [Google Scholar] [CrossRef]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K.; et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef]
- Shimatani, Z.; Ariizumi, T.; Fujikura, U.; Kondo, A.; Ezura, H.; Nishida, K. Targeted base editing with CRISPR-deaminase in tomato. Methods Mol. Biol. 2019, 1917, 297–307. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef]
- Nishida, K.; Arazoe, T.; Yachie, N.; Banno, S.; Kakimoto, M.; Tabata, M.; Mochizuki, M.; Miyabe, A.; Araki, M.; Hara, K.Y.; et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 2016, 353. [Google Scholar] [CrossRef]
- Hess, G.T.; Tycko, J.; Yao, D.; Bassik, M.C. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 2017, 68, 26–43. [Google Scholar] [CrossRef]
- Gehrke, J.M.; Cervantes, O.; Clement, M.K.; Wu, Y.; Zeng, J.; Bauer, D.E.; Pinello, L.; Joung, J.K. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 2019, 36, 977–982. [Google Scholar] [CrossRef]
- Ma, Y.; Zhang, J.; Yin, W.; Zhang, Z.; Song, Y.; Chang, X. Targeted AID-mediated mutagenesis (TAM) enables efficient genomic diversification in mammalian cells. Nat. Methods 2016, 13, 1029–1035. [Google Scholar] [CrossRef]
- Li, Z.; Xiong, X.; Li, J.F. New cytosine base editor for plant genome editing. Sci. China Life Sci. 2018, 61, 1602–1603. [Google Scholar] [CrossRef]
- Billon, P.; Bryant, E.E.; Joseph, S.A.; Nambiar, T.S.; Hayward, S.B.; Rothstein, R.; Ciccia, A. CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of stop codons. Mol. Cell 2017, 67, 1068–1079. [Google Scholar] [CrossRef]
- Tian, S.; Jiang, L.; Cui, X.; Zhang, J.; Guo, S.; Li, M.; Zhang, H.; Ren, Y.; Gong, G.; Zong, M.; et al. Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep. 2018, 37, 1353–1356. [Google Scholar] [CrossRef]
- Yan, F.; Kuang, Y.; Ren, B.; Wang, J.; Zhang, D.; Lin, H.; Yang, B.; Zhou, X.; Zhou, H. Highly Efficient A·T to G·C Base Editing by Cas9n-Guided tRNA Adenosine Deaminase in Rice. Mol. Plant 2018, 11, 631–634. [Google Scholar] [CrossRef]
- Hua, K.; Tao, X.; Yuan, F.; Wang, D.; Zhu, J.K. Precise A·T to G·C base editing in the rice genome. Mol. Plant 2018, 11, 627–630. [Google Scholar] [CrossRef]
- Kang, B.C.; Yun, J.Y.; Kim, S.T.; Shin, Y.J.; Ryu, J.; Choi, M.; Woo, J.W.; Kim, J.S. Precision genome engineering through adenine base editing in plants. Nat. Plants 2018, 4, 427–431. [Google Scholar] [CrossRef]
- Li, C.; Zong, Y.; Wang, Y.; Jin, S.; Zhang, D.; Song, Q.; Zhang, R.; Gao, C. Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 2018, 19, 59. [Google Scholar] [CrossRef]
- Xing, H.L.; Wang, Z.P.; Zhang, H.Y.; Han, C.Y.; Liu, B.; Wang, X.C.; Chen, Q.J.; Dong, L. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 2014, 14, 327. [Google Scholar] [CrossRef]
- Mao, Y.; Zhang, H.; Xu, N.; Zhang, B.; Gou, F.; Zhu, J.K. Application of the CRISPR–Cas system for efficient genome engineering in plants. Mol. Plant 2013, 6, 2008–2011. [Google Scholar] [CrossRef]
- Wang, C.; Liu, W.; Wang, G.; Li, J.; Dong, L.; Han, L.; Wang, Q.; Tian, J.; Yu, Y.; Gao, C.; et al. KTN80 confers precision to microtubule severing by specific targeting of katanin complexes in plant cells. EMBO J. 2017, 36, 3435–3447. [Google Scholar] [CrossRef]
- Xie, K.; Minkenberg, B.; Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl. Acad. Sci. USA 2015, 112, 3570–3575. [Google Scholar] [CrossRef]
- Qi, W.; Zhu, T.; Tian, Z.; Li, C.; Zhang, W.; Song, R. High-efficiency CRISPR/Cas9 multiplex gene editing using the glycine tRNA-processing system-based strategy in maize. BMC Biotechnol. 2016, 16, 58. [Google Scholar] [CrossRef]
- Tang, X.; Zheng, X.; Qi, Y.; Zhang, D.; Cheng, Y.; Tang, A.; Voytas, D.F.; Zhang, Y. A single transcript CRISPR-Cas9 system for efficient genome editing in plants. Mol. Plant 2016, 9, 1088–1091. [Google Scholar] [CrossRef]
- Wang, M.; Mao, Y.; Lu, Y.; Tao, X.; Zhu, J. Multiplex gene editing in rice using the CRISPR-Cpf1 System. Mol. Plant 2017, 10, 1011–1013. [Google Scholar] [CrossRef]
- Mali, P.; Aach, J.; Stranges, P.B.; Esvelt, K.M.; Moosburner, M.; Kosuri, S.; Yang, L.; Church, G.M. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 2013, 31, 833–838. [Google Scholar] [CrossRef]
- Guilinger, J.P.; Thompson, D.B.; Liu, D.R. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 2014, 32, 577–582. [Google Scholar] [CrossRef]
- Chylinski, K.; Le Rhun, A.; Charpentier, E. The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems. RNA Biol. 2013, 10, 726–737. [Google Scholar] [CrossRef]
- Lee, C.M.; Cradick, T.J.; Bao, G. The Neisseria meningitidis CRISPR-Cas9 system enables specific genome editing in mammalian cells. Mol. Ther. 2016, 24, 645–654. [Google Scholar] [CrossRef]
- Ran, F.A.; Cong, L.; Yan, W.X.; Scott, D.A.; Gootenberg, J.S.; Kriz, A.J.; Zetsche, B.; Shalem, O.; Wu, X.; Makarova, K.S.; et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 2015, 520, 186–191. [Google Scholar] [CrossRef]
- Hu, X.; Meng, X.; Liu, Q.; Li, J.; Wang, K. Increasing the efficiency of CRISPR-Cas9-VQR precise genome editing in rice. Plant Biotechnol. J. 2018, 16, 292–297. [Google Scholar] [CrossRef]
- Jia, H.; Xu, J.; Orbović, V.; Zhang, Y.; Wang, N. Editing citrus genome via SaCas9/sgRNA System. Front. Plant Sci. 2017, 8, 2135. [Google Scholar] [CrossRef]
- Steinert, J.; Schiml, S.; Fauser, F.; Puchta, H. Highly efficient heritable plant genome engineering using Cas9 orthologues from Streptococcus thermophilus and Staphylococcus aureus. Plant J. 2015, 84, 1295–1305. [Google Scholar] [CrossRef]
- Zetsche, B.; Heidenreich, M.; Mohanraju, P.; Fedorova, I.; Kneppers, J.; Degennaro, E.M.; Winblad, N.; Choudhury, S.R.; Abudayyeh, O.O.; Gootenberg, J.S.; et al. Multiplex gene editing by CRISPR-Cpf1 using a single crRNA array. Nat. Biotechnol. 2017, 35, 31–34. [Google Scholar] [CrossRef]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; Van Der Oost, J.; Regev, A.; et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef]
- Endo, A.; Masafumi, M.; Kaya, H.; Toki, S. Efficient targeted mutagenesis of rice and tobacco genomes using Cpf1 from Francisella novicida. Sci. Rep. 2016, 6, 38169. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, J.; Lang, Z.; Botella, J.R.; Zhu, J.K. Genome Editing—Principles and Applications for Functional Genomics Research and Crop Improvement. Crit. Rev. Plant Sci. 2017, 34, 291–309. [Google Scholar] [CrossRef]
- Yin, X.; Biswal, A.K.; Dionora, J.; Perdigon, K.M.; Balahadia, C.P.; Mazumdar, S.; Chater, C.; Lin, H.C.; Coe, R.A.; Kretzschmar, T.; et al. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental gene EPFL9 in rice. Plant Cell Rep. 2017, 36, 745–757. [Google Scholar] [CrossRef]
- Xu, R.; Qin, R.; Li, H.; Li, D.; Li, L.; Wei, P.; Yang, J. Generation of targeted mutant rice using a CRISPR-Cpf1 system. Plant Biotechnol. J. 2017, 15, 713–717. [Google Scholar] [CrossRef]
- Begemann, M.B.; Gray, B.N.; January, E.; Gordon, G.C.; He, Y.; Liu, H.; Wu, X.; Brutnell, T.P.; Mockler, T.C.; Oufattole, M. Precise insertion and guided editing of higher plant genomes using Cpf1 CRISPR nucleases. Sci. Rep. 2017, 7, 11606. [Google Scholar] [CrossRef]
- Scheben, A.; Edwards, D. Genome editors take on crops. Science 2017, 355, 1122–1123. [Google Scholar] [CrossRef]
- Zaidi, S.S.A.; Tashkandi, M.; Mansoor, S.; Mahfouz, M.M. Engineering plant immunity: Using crispr/cas9 to generate virus resistance. Front. Plant Sci. 2016, 7, 1673. [Google Scholar] [CrossRef]
- Al-Sadi, A.M.; Al-Moqbali, H.S.; Al-Yahyai, R.A.; Al-Said, F.A. AFLP data suggest a potential role for the low genetic diversity of acid lime (Citrus aurantifolia Swingle) in Oman in the outbreak of witches’ broom disease of lime. Euphytica 2012, 188, 285–297. [Google Scholar] [CrossRef]
- Kettles, G.J.; Kanyuka, K. Dissecting the molecular interactions between wheat and the fungal pathogen Zymoseptoria tritici. Front. Plant Sci. 2016, 7, 508. [Google Scholar] [CrossRef]
- Wang, F.; Wang, C.; Liu, P.; Lei, C.; Hao, W.; Gao, Y.; Liu, Y.G.; Zhao, K. Enhanced rice blast resistance by CRISPR/ Cas9-Targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 2016, 11, e0154027. [Google Scholar] [CrossRef]
- Zhou, J.; Peng, Z.; Long, J.; Sosso, D.; Liu, B.; Eom, J.S.; Huang, S.; Liu, S.; Vera Cruz, C.; Frommer, W.B.; et al. Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J. 2015, 82, 632–643. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, Y.; Orbović, V.; Xu, J.; White, F.F.; Jones, J.B.; Wang, N. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 2017, 15, 817–823. [Google Scholar] [CrossRef]
- Peng, A.; Chen, S.; Lei, T.; Xu, L.; He, Y.; Wu, L.; Yao, L.; Zou, X. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef]
- Zhang, Y.; Bai, Y.; Wu, G.; Zou, S.; Chen, Y.; Gao, C.; Tang, D. Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J. 2017, 91, 714–724. [Google Scholar] [CrossRef]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Gil-Humanes, J.; Wang, Y.; Liang, Z.; Shan, Q.; Ozuna, C.V.; Sánchez-León, S.; Baltes, N.J.; Starker, C.; Barro, F.; Gao, C.; et al. High-efficiency gene targeting in hexaploid wheat using DNA replicons and CRISPR/Cas9. Plant J. 2017, 89, 1251–1262. [Google Scholar] [CrossRef]
- Ji, X.; Si, X.; Zhang, Y.; Zhang, H.; Zhang, F.; Gao, C. Conferring DNA virus resistance with high specificity in plants using virus-inducible genome-editing system. Genome Biol. 2018, 19, 197. [Google Scholar] [CrossRef]
- Baltes, N.J.; Hummel, A.W.; Konecna, E.; Cegan, R.; Bruns, A.N.; Bisaro, D.M.; Voytas, D.F. Conferring resistance to geminiviruses with the CRISPR-Cas prokaryotic immune system. Nat. Plants 2015, 1, 15145. [Google Scholar] [CrossRef]
- Ali, Z.; Abulfaraj, A.; Idris, A.; Ali, S.; Tashkandi, M.; Mahfouz, M.M. CRISPR/Cas9-mediated viral interference in plants. Genome Biol. 2015, 16, 238. [Google Scholar] [CrossRef]
- Zhang, T.; Zheng, Q.; Yi, X.; An, H.; Zhao, Y.; Ma, S.; Zhou, G. Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol. J. 2018, 16, 1415–1423. [Google Scholar] [CrossRef]
- Macovei, A.; Sevilla, N.R.; Cantos, C.; Jonson, G.B.; Slamet-Loedin, I.; Čermák, T.; Voytas, D.F.; Choi, I.R.; Chadha-Mohanty, P. Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotechnol. J. 2018, 16, 1918–1927. [Google Scholar] [CrossRef]
- Wang, X.; Tu, M.; Wang, D.; Liu, J.; Li, Y.; Li, Z.; Wang, Y.; Wang, X. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol. J. 2018, 16, 844–855. [Google Scholar] [CrossRef]
- Zhang, Z.; Ge, X.; Luo, X.; Wang, P.; Fan, Q.; Hu, G.; Xiao, J.; Li, F.; Wu, J. Simultaneous Editing of Two Copies of Gh14-3-3d Confers Enhanced Transgene-Clean Plant Defense Against Verticillium dahliae in Allotetraploid Upland Cotton. Front. Plant Sci. 2018, 9, 842. [Google Scholar] [CrossRef]
- Ortigosa, A.; Gimenez-Ibanez, S.; Leonhardt, N.; Solano, R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol. J. 2018, 17, 665–673. [Google Scholar] [CrossRef]
- Tashkandi, M.; Ali, Z.; Aljedaani, F.; Shami, A.; Mahfouz, M.M. Engineering resistance against Tomato yellow leaf curl virus via the CRISPR/Cas9 system in tomato. Plant Signal. Behav. 2018, 13. [Google Scholar] [CrossRef]
- Chandrasekaran, J.; Brumin, M.; Wolf, D.; Leibman, D.; Klap, C.; Pearlsman, M.; Sherman, A.; Arazi, T.; Gal-On, A. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 2016, 7, 1140–1153. [Google Scholar] [CrossRef]
- Kim, D.; Kim, D.; Alptekin, B.; Budak, H. CRISPR/Cas9 genome editing in wheat. Funct. Integr. Genomics 2017, 18, 31–41. [Google Scholar] [CrossRef]
- Shen, C.; Que, Z.; Xia, Y.; Tang, N.; Li, D.; He, R.; Cao, M. Knock out of the annexin gene OsAnn3 via CRISPR/Cas9-mediated genome editing decreased cold tolerance in rice. J. Plant Biol. 2017, 60, 539–547. [Google Scholar] [CrossRef]
- Lou, D.; Wang, H.; Liang, G.; Yu, D. OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front. Plant Sci. 2017, 8, 993. [Google Scholar] [CrossRef]
- Shi, J.; Gao, H.; Wang, H.; Lafitte, H.R.; Archibald, R.L.; Yang, M.; Hakimi, S.M.; Mo, H.; Habben, J.E. ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol. J. 2017, 15, 207–216. [Google Scholar] [CrossRef]
- Curtin, S.J.; Xiong, Y.; Michno, J.M.; Campbell, B.W.; Stec, A.O.; Čermák, T.; Starker, C.; Voytas, D.F.; Eamens, A.L.; Stupar, R.M. CRISPR/Cas9 and TALENs generate heritable mutations for genes involved in small RNA processing of Glycine max and Medicago truncatula. Plant Biotechnol. J. 2018, 16, 1125–1137. [Google Scholar] [CrossRef]
- Wang, L.; Chen, L.; Li, R.; Zhao, R.; Yang, M.; Sheng, J.; Shen, L. Reduced drought tolerance by CRISPR/Cas9-mediated SlMAPK3 mutagenesis in tomato plants. J. Agric. Food Chem. 2017, 65, 8674–8682. [Google Scholar] [CrossRef]
- Shen, L.; Wang, C.; Fu, Y.; Wang, J.; Liu, Q.; Zhang, X.; Yan, C.; Qian, Q.; Wang, K. QTL editing confers opposing yield performance in different rice varieties. J. Integr. Plant Biol. 2018, 60, 89–93. [Google Scholar] [CrossRef]
- Bo, W.; Zhaohui, Z.; Huanhuan, Z.; Xia, W.; Binglin, L.; Lijia, Y.; Xiangyan, H.; Deshui, Y.; Xuelian, Z.; Chunguo, W.; et al. Targeted Mutagenesis of NAC Transcription Factor Gene, OsNAC041, Leading to Salt Sensitivity in Rice. Rice Sci. 2019, 26, 98–108. [Google Scholar] [CrossRef]
- Sadanandom, A.; Srivastava, A.K.; Zhang, C. Targeted mutagenesis of the SUMO protease, Overly Tolerant to Salt1 in rice through CRISPR/Cas9-mediated genome editing reveals a major role of this SUMO protease in salt tolerance. BioRxiv 2019, 555706. [Google Scholar] [CrossRef]
- Zhang, A.; Liu, Y.; Wang, F.; Li, T.; Chen, Z.; Kong, D.; Bi, J.; Zhang, F.; Luo, X.; Wang, J.; et al. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef]
- Li, R.; Liu, C.; Zhao, R.; Wang, L.; Chen, L.; Yu, W.; Zhang, S.; Sheng, J.; Shen, L. CRISPR/Cas9-Mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol. 2019, 19, 38. [Google Scholar] [CrossRef]
- Shim, J.S.; Oh, N.; Chung, P.J.; Kim, Y.S.; Choi, Y.D.; Kim, J.K. Overexpression of OsNAC14 Improves Drought Tolerance in Rice. Front. Plant Sci. 2018, 9, 310. [Google Scholar] [CrossRef]
- Lou, D.; Wang, H.; Yu, D. The sucrose non-fermenting-1-related protein kinases SAPK1 and SAPK2 function collaboratively as positive regulators of salt stress tolerance in rice. BMC Plant Biol. 2018, 18, 203. [Google Scholar] [CrossRef]
- Zhang, M.; Cao, Y.; Wang, Z.; Wang, Z.Q.; Shi, J.; Liang, X.; Song, W.; Chen, Q.; Lai, J.; Jiang, C. A retrotransposon in an HKT1 family sodium transporter causes variation of leaf Na+exclusion and salt tolerance in maize. New Phytol. 2018, 217, 1161–1176. [Google Scholar] [CrossRef]
- Li, R.; Zhang, L.; Wang, L.; Chen, L.; Zhao, R.; Sheng, J.; Shen, L. Reduction of Tomato-Plant Chilling Tolerance by CRISPR-Cas9-Mediated SlCBF1 Mutagenesis. J. Agric. Food Chem. 2018, 66, 9042–9051. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, D.; Zhang, D.; Zhao, X.; Cao, X.; Dong, L.; Liu, J.; Chen, K.; Zhang, H.; Gao, C.; et al. Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J. 2018, 94, 857–866. [Google Scholar] [CrossRef]
- Lu, K.; Wu, B.; Wang, J.; Zhu, W.; Nie, H.; Qian, J.; Huang, W.; Fang, Z. Blocking amino acid transporter OsAAP3 improves grain yield by promoting outgrowth buds and increasing tiller number in rice. Plant Biotechnol. J. 2018, 16, 1710–1722. [Google Scholar] [CrossRef]
- Liu, J.; Chen, J.; Zheng, X.; Wu, F.; Lin, Q.; Heng, Y.; Tian, P.; Cheng, Z.J.; Yu, X.; Zhou, K.; et al. GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat. Plants 2017, 3, 17043. [Google Scholar] [CrossRef]
- Li, S.; Gao, F.; Xie, K.; Zeng, X.; Cao, Y.; Zeng, J.; He, Z.; Ren, Y.; Li, W.; Deng, Q.; et al. The OsmiR396c-OsGRF4-OsGIF1 regulatory module determines grain size and yield in rice. Plant Biotechnol. J. 2016, 14, 2134–2146. [Google Scholar] [CrossRef]
- Li, M.; Li, X.; Zhou, Z.; Wu, P.; Fang, M.; Pan, X.; Lin, Q.; Luo, W.; Wu, G.; Li, H. Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front. Plant Sci. 2016, 7, 377. [Google Scholar] [CrossRef]
- Xu, R.; Yang, Y.; Qin, R.; Li, H.; Qiu, C.; Li, L.; Wei, P.; Yang, J. Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice. J. Genet. Genomics 2016, 43, 529–532. [Google Scholar] [CrossRef]
- Li, X.; Zhou, W.; Ren, Y.; Tian, X.; Lv, T.; Wang, Z.; Fang, J.; Chu, C.; Yang, J.; Bu, Q. High-efficiency breeding of early-maturing rice cultivars via CRISPR/Cas9-mediated genome editing. J. Genet. Genomics 2017, 44, 175–178. [Google Scholar] [CrossRef]
- Ma, L.; Zhang, D.; Miao, Q.; Yang, J.; Xuan, Y.; Hu, Y. Essential role of sugar transporter OsSWEET11 during the early stage of rice grain filling. Plant Cell Physiol. 2017, 58, 863–873. [Google Scholar] [CrossRef]
- Huang, J.; Li, J.; Zhou, J.; Wang, L.; Yang, S.; Hurst, L.D.; Li, W.H.; Tian, D. Identifying a large number of high-yield genes in rice by pedigree analysis, whole-genome sequencing, and CRISPR-Cas9 gene knockout. Proc. Natl. Acad. Sci. USA 2018, 115, E7559–E7567. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, H.; Botella, J.R.; Zhu, J. Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J. Integr. Plant Biol. 2018, 60, 369–375. [Google Scholar] [CrossRef]
- Sun, Y.; Jiao, G.; Liu, Z.; Zhang, X.; Li, J.; Guo, X.; Du, W.; Du, J.; Francis, F.; Zhao, Y.; et al. Generation of High-Amylose Rice through CRISPR/Cas9-Mediated Targeted Mutagenesis of Starch Branching Enzymes. Front. Plant Sci. 2017, 8, 298. [Google Scholar] [CrossRef]
- Sánchez-León, S.; Gil-Humanes, J.; Ozuna, C.V.; Giménez, M.J.; Sousa, C.; Voytas, D.F.; Barro, F. Low-gluten, nontransgenic wheat engineered with CRISPR/Cas9. Plant Biotechnol. J. 2018, 16, 902–910. [Google Scholar] [CrossRef]
- Li, A.; Jia, S.; Yobi, A.; Ge, Z.; Sato, S.; Zhang, C.; Angelovici, R.; Clemente, T.E.; Holding, D.R. Editing of an alpha-kafirin gene family increases digestibility and protein quality in sorghum. Plant Physiol. 2018, 177, 1425–1438. [Google Scholar] [CrossRef]
- Okuzaki, A.; Ogawa, T.; Koizuka, C.; Kaneko, K.; Inaba, M.; Imamura, J.; Koizuka, N. CRISPR/Cas9-mediated genome editing of the fatty acid desaturase 2 gene in Brassica napus. Plant Physiol. Biochem. 2018, 131, 63–69. [Google Scholar] [CrossRef]
- Li, R.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. CRISPR/Cas9-mediated mutagenesis of lncRNA1459 alters tomato fruit ripening. Plant J. 2018, 94, 513–524. [Google Scholar] [CrossRef]
- Li, X.; Wang, Y.; Chen, S.; Tian, H.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. Lycopene Is Enriched in Tomato Fruit by CRISPR/Cas9-Mediated Multiplex Genome Editing. Front. Plant Sci. 2018, 9, 559. [Google Scholar] [CrossRef]
- Butt, H.; Jamil, M.; Wang, J.Y.; Al-Babili, S.; Mahfouz, M. Engineering plant architecture via CRISPR/Cas9-mediated alteration of strigolactone biosynthesis. BMC Plant Biol. 2018, 18, 174. [Google Scholar] [CrossRef]
- Cai, Y.; Chen, L.; Liu, X.; Guo, C.; Sun, S.; Wu, C.; Jiang, B.; Han, T.; Hou, W. CRISPR/Cas9-mediated targeted mutagenesis of GmFT2a delays flowering time in soya bean. Plant Biotechnol. J. 2018, 16, 176–185. [Google Scholar] [CrossRef]
- Soyk, S.; Müller, N.A.; Park, S.J.; Schmalenbach, I.; Jiang, K.; Hayama, R.; Zhang, L.; Van Eck, J.; Jiménez-Gómez, J.M.; Lippman, Z.B. Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat. Genet. 2017, 49, 162–168. [Google Scholar] [CrossRef]
- Nonaka, S.; Arai, C.; Takayama, M.; Matsukura, C.; Ezura, H. Efficient increase of Γ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis. Sci. Rep. 2017, 7, 7057. [Google Scholar] [CrossRef]
- Chahal, G.S.; Gosal, S.S. Principles and Procedures of Plant Breeding: Biotechnological and Conventional Approaches; Alpha Science Int’l Ltd.: Oxford, UK, 2002. [Google Scholar]
- Pacher, M.; Puchta, H. From classical mutagenesis to nuclease-based breeding–directing natural DNA repair for a natural end-product. Plant J. 2017, 90, 819–833. [Google Scholar] [CrossRef]
- Scheben, A.; Wolter, F.; Batley, J.; Puchta, H.; Edwards, D. Towards CRISPR/Cas crops–bringing together and genome editing. New Phytol. 2017, 216, 682–698. [Google Scholar] [CrossRef]
- Sikora, P.; Chawade, A.; Larsson, M.; Olsson, J.; Olsson, O. Mutagenesis as a tool in plant genetics, functional genomics, and breeding. Int. J. Plant Genomics 2011, 2011, 1–3. [Google Scholar] [CrossRef]
- Hartung, F.; Schiemann, J. Precise plant breeding using new genome editing techniques: Opportunities, safety and regulation in the EU. Plant J. 2014, 78, 742–752. [Google Scholar] [CrossRef]
- Xu, R.F.; Li, H.; Qin, R.Y.; Li, J.; Qiu, C.H.; Yang, Y.C.; Ma, H.; Li, L.; Wei, P.C.; Yang, J.B. Generation of and “transgene clean” targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Sci. Rep. 2015, 5, 11491. [Google Scholar] [CrossRef]
- Visser, B.; Eaton, D.; Louwaars, N.; van der Meer, I.M.; Beekwilder, J.; van Tongeren, F. Potential of Genetic Use Restriction Technologies (GURTs) on Agrobiodiversity and Agricultural Production Systems; FAO: Rome, Italy, 2001. [Google Scholar]
- Ledford, H. CRISPR, the disruptor. Nat. News 2015, 522, 20–24. [Google Scholar] [CrossRef]
- Feng, Z.; Mao, Y.; Xu, N.; Zhang, B.; Wei, P.; Yang, D.L.; Wang, Z.; Zhang, Z.; Zheng, R.; Yang, L.; et al. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 4632–4637. [Google Scholar] [CrossRef]
- Waltz, E. Gene-edited CRISPR mushroom escapes US regulation. Nat. News 2016, 532, 293. [Google Scholar] [CrossRef]
- Callaway, E. CRISPR plants now subject to tough GM laws in European Union. Nature 2018, 560, 16. [Google Scholar] [CrossRef]
- Sprink, T.; Eriksson, D.; Schiemann, J.; Hartung, F. Regulatory hurdles for genome editing: Process-vs. Product-based approaches in different regulatory contexts. Plant Cell Rep. 2016, 35, 1493–1506. [Google Scholar] [CrossRef]
- Kahrmann, J.; Bömeke, O.; Leggewie, G. Aged GMO legislation meets new genome editing techniques. Z. Eur. Umw. Plan. 2017, 15, 176–182. [Google Scholar] [CrossRef]
- Fears, R.; Ter Meulen, V. Point of View: How should the applications of genome editing be assessed and regulated? Elife 2017, 6, e26295. [Google Scholar] [CrossRef]
- Ishii, T.; Araki, M. A future scenario of the global regulatory landscape regarding genome-edited crops. GM Crop. Food 2017, 8, 44–56. [Google Scholar] [CrossRef]
- Smyth, S.J. Canadian regulatory perspectives on genome engineered crops. GM Crop. Food 2017, 8, 35–43. [Google Scholar] [CrossRef]
- Whelan, A.I.; Lema, M.A. Regulatory framework for gene editing and other new breeding techniques (NBTs) in Argentina. GM Crop. Food 2015, 6, 253–265. [Google Scholar] [CrossRef]
- Brinegar, K.K.; Yetisen, A.; Choi, S.; Vallillo, E.; Ruiz-Esparza, G.U.; Prabhakar, A.M.; Khademhosseini, A.; Yun, S.H. The commercialization of genome-editing technologies. Crit. Rev. Biotechnol. 2017, 37, 924–932. [Google Scholar] [CrossRef]
- Genome Editing/Genome Engineering Market by Application (Cell Line Engineering, Animal & Plant Genetic Engineering), Technology (CRISPR, Antisense, TALEN, Zinc Finger Nuclease) & End User (Biotechnology & Pharmaceutical, Crop)-Global Forecast to 2019. Available online: http://www.marketsandmarkets.com/Market-Reports/genome-editing-engineerin market231037000.html (accessed on 12 April 2016).
- Van Erp, P.B.; Bloomer, G.; Wilkinson, R.; Wiedenheft, B. The history and market impact of CRISPR RNA-guided nucleases. Curr. Opin. Virol. 2015, 12, 85–90. [Google Scholar] [CrossRef]
- Ousterout, D.G.; Kabadi, A.M.; Thakore, P.I.; Majoros, W.H.; Reddy, T.E.; Gersbach, C.A. Multiplex CRISPR/Cas9-based genome editing for correction of dystrophin mutations that cause Duchenne muscular dystrophy. Nat. Commun. 2015, 6, 6244. [Google Scholar] [CrossRef]
- Suzuki, K.; Tsunekawa, Y.; Hernandez-Benitez, R.; Wu, J.; Zhu, J.; Kim, E.J.; Hatanaka, F.; Yamamoto, M.; Araoka, T.; Li, Z.; et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 2016, 540, 144. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).