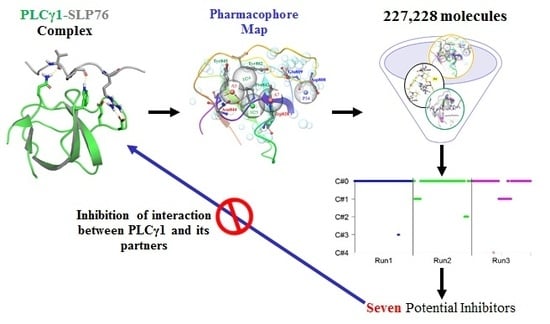
Investigation of Phospholipase Cγ1 Interaction with SLP76 Using Molecular Modeling Methods for Identifying Novel Inhibitors
Abstract
1. Introduction
2. Results and Discussion
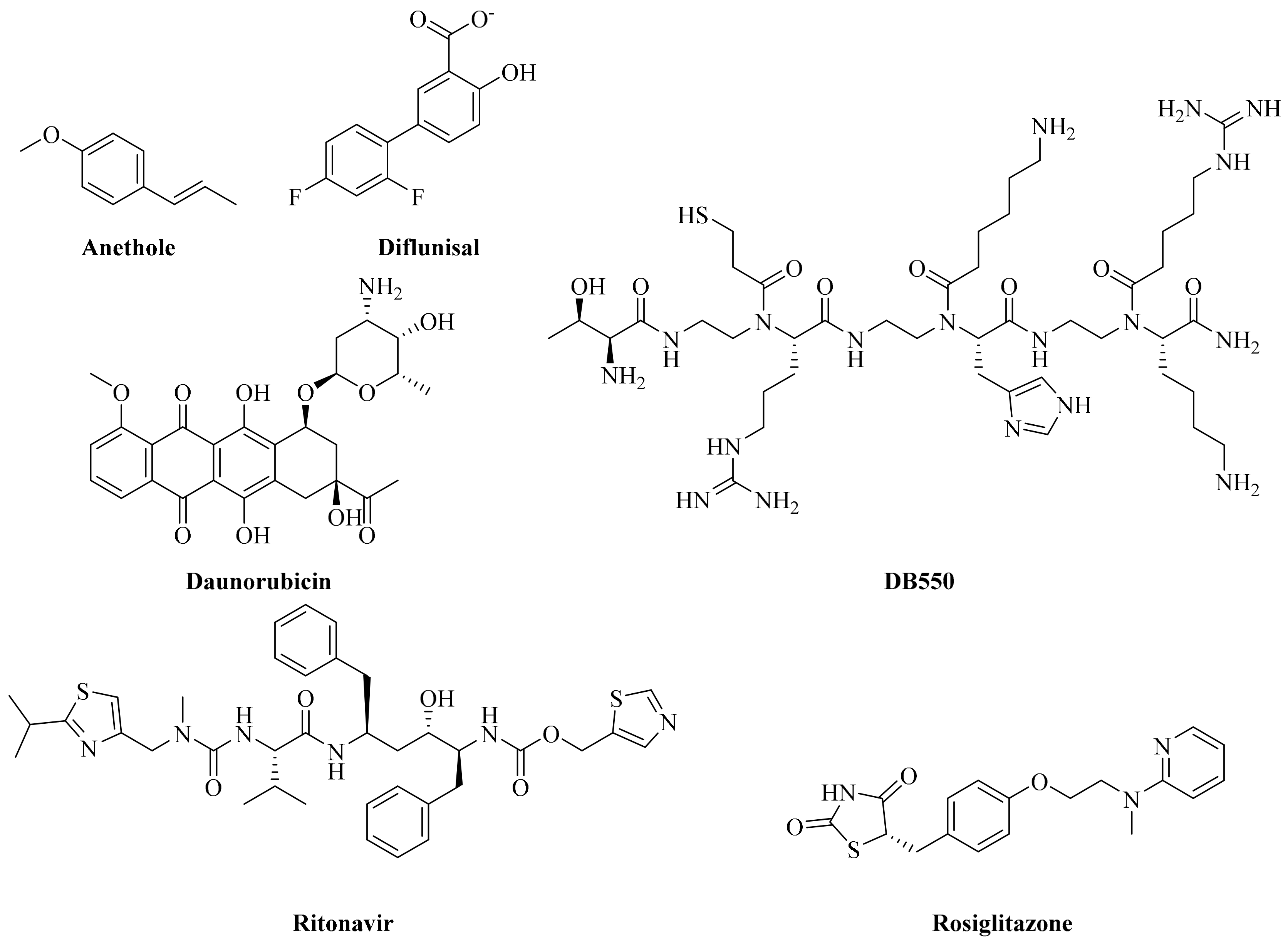
2.1. Molecular Recognition of SLP76 and Known Inhibitors by PLCγ1
2.2. Pharmacophore Mapping and Molecular Docking Based Identification of Promising Hits
2.3. MD Simulations
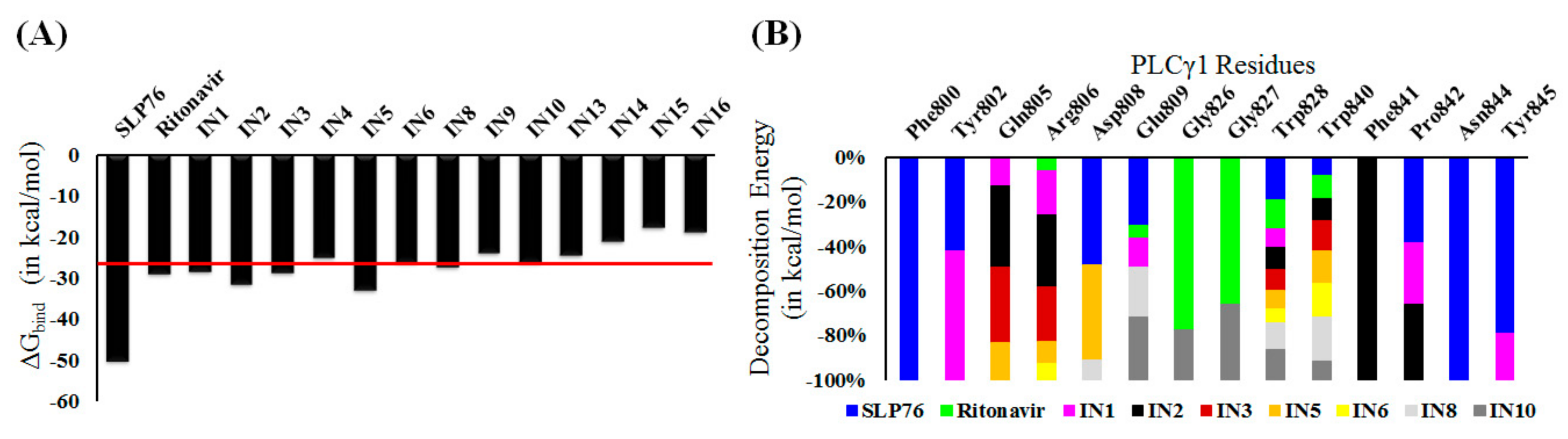
2.4. Stable Binding of Identified Molecules to PLCγ1
2.5. Molecular Recognition of the Selected Molecules to PLCγ1 Considering MD Simulations
3. Materials and Methods
3.1. Pharmacophore Modeling
3.2. Molecular Docking-Based Virtual Screening
3.3. Molecular Dynamics Simulations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 3D | Three Dimensional |
| ∆Gbind | Binding energy |
| AMBER | Assisted Model Building with Energy Refinement |
| CADD | Computer Aided Drug Design |
| CD95 | Cluster of differentiation 95 |
| CID | Calcium inducing domain |
| COM_dist | Center of Mass of the Bound Ligand and AsnA844 of PLCγ1 |
| GAFF | General Amber Force Field |
| H-bond | Hydrogen-bond |
| HTVS: | High-throughput virtual screening |
| IFD | Induced fit docking |
| IP3 | Inositol 1,4,5-triphosphate |
| MD | Molecular dynamics |
| MM/GBSA | Molecular Mechanics-Generalized Born Surface Area |
| PDB | Protein Data bank |
| PIP2 | Phosphatidylinositol 4,5-bisphosphate |
| PLCγ1 | Phospholipase C gamma 1 |
| PLCγ1-SH3 | SH3 domain of PLCγ1 |
| PME | Particle-Mesh Ewald |
| RMSD | Root mean square deviation |
| SP | Standard Precision |
| TIP3P | Three-site Transferrable Intermolecular Potential |
| VMD | Visual Molecular Dynamics software |
| XP | eXtra Precision |
References
- Perl, A.; Gergely, P.J.; Puskas, F.; Banki, K. Metabolic switches of T-cell activation and apoptosis. Antioxidants Redox Signal. 2002, 4, 427–443. [Google Scholar] [CrossRef] [PubMed]
- Solary, E.; Dubrez, L.; Eymin, B. The role of apoptosis in the pathogenesis and treatment of diseases. Eur. Respir. J. 1996, 9, 1293–1305. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.B. Apoptosis in the pathogenesis and treatment of disease. Science 1995, 267, 1456–1462. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y. E3 ubiquitin ligases as cancer targets and biomarkers. Neoplasia 2006, 8, 645–654. [Google Scholar] [CrossRef] [PubMed]
- Hunter, A.M.; LaCasse, E.C.; Korneluk, R.G. The inhibitors of apoptosis (IAPs) as cancer targets. Apoptosis 2007, 12, 1543–1568. [Google Scholar] [CrossRef] [PubMed]
- Faustman, D.; Davis, M. TNF receptor 2 pathway: Drug target for autoimmune diseases. Nat. Rev. Drug Discov. 2010, 9, 482–493. [Google Scholar] [CrossRef] [PubMed]
- Paplomata, E.; O’regan, R. The PI3K/AKT/mTOR pathway in breast cancer: Targets, trials and biomarkers. Ther. Adv. Med. Oncol. 2014, 6, 154–166. [Google Scholar] [CrossRef]
- Isono, F.; Fujita-Sato, S.; Ito, S. Inhibiting RORγt/Th17 axis for autoimmune disorders. Drug Discov. Today 2014, 19, 1205–1211. [Google Scholar] [CrossRef]
- Nayak, A.; Satapathy, S.R.; Das, D.; Siddharth, S.; Tripathi, N.; Bharatam, P.V.; Kundu, C.N. Nanoquinacrine induced apoptosis in cervical cancer stem cells through the inhibition of hedgehog-GLI1 cascade: Role of GLI-1. Sci. Rep. 2016, 6, 20600. [Google Scholar] [CrossRef]
- Das, S.; Tripathi, N.; Preet, R.; Siddharth, S.; Nayak, A.; Bharatam, P.V.; Kundu, C.N. Quinacrine induces apoptosis in cancer cells by forming a functional bridge between TRAIL-DR5 complex and modulating the mitochondrial intrinsic cascade. Oncotarget 2017, 8, 248–267. [Google Scholar] [CrossRef]
- Das, S.; Tripathi, N.; Siddharth, S.; Nayak, A.; Nayak, D.; Sethy, C.; Bharatam, P.V.; Kundu, C.N. Etoposide and doxorubicin enhance the sensitivity of triple negative breast cancers through modulation of TRAIL-DR5 axis. Apoptosis 2017, 22, 1205–1224. [Google Scholar] [CrossRef]
- Fasching, P.; Stradner, M.; Graninger, W.; Dejaco, C.; Fessler, J. Therapeutic potential of targeting the Th17/Treg axis in autoimmune disorders. Molecules 2017, 22, E134. [Google Scholar] [CrossRef]
- Sala, G.; Dituri, F.; Raimondi, C.; Previdi, S.; Maffucci, T.; Mazzoletti, M.; Rossi, C.; Iezzi, M.; Lattanzio, R.; Piantelli, M.; et al. Phospholipase Cγ1 is required for metastasis development and progression. Cancer Res. 2008, 68, 10187–10196. [Google Scholar] [CrossRef]
- Yang, Y.R.; Choi, J.H.; Chang, J.-S.; Kwon, H.M.; Jang, H.-J.; Ryu, S.H.; Suh, P.-G. Diverse cellular and physiological roles of phospholipase C-γ1. Adv. Enzyme Regul. 2012, 52, 138–151. [Google Scholar] [CrossRef] [PubMed]
- Lattanzio, R.; Piantelli, M.; Falasca, M. Role of phospholipase C in cell invasion and metastasis. Adv. Biol. Regul. 2013, 53, 309–318. [Google Scholar] [CrossRef]
- Poissonnier, A.; Guégan, J.P.; Nguyen, H.T.; Best, D.; Levoin, N.; Kozlov, G.; Gehring, K.; Pineau, R.; Jouan, F.; Morere, L.; et al. Disrupting the CD95–PLCγ1 interaction prevents Th17-driven inflammation. Nat. Chem. Biol. 2018, 14, 1079–1089. [Google Scholar] [CrossRef]
- Kadamur, G.; Ross, E.M. Mammalian Phospholipase, C. Annu. Rev. Physiol. 2012, 75, 127–154. [Google Scholar] [CrossRef]
- Deng, L.; Velikovsky, C.A.; Swaminathan, C.P.; Cho, S.; Mariuzza, R.A.; Huber, R. Structural basis for recognition of the T Cell adaptor protein SLP-76 by the SH3 domain of phospholipase Cγ1. J. Mol. Biol. 2005, 352, 1–10. [Google Scholar] [CrossRef]
- Gierschik, P.; Buehler, A.; Walliser, C. Activated PLCγ breaking loose. Structure 2012, 20, 1989–1990. [Google Scholar] [CrossRef][Green Version]
- Bunney, T.D.; Esposito, D.; Mas-Droux, C.; Lamber, E.; Baxendale, R.W.; Martins, M.; Cole, A.; Svergun, D.; Driscoll, P.C.; Katan, M. Structural and functional integration of the PLCγ interaction domains critical for regulatory mechanisms and signaling deregulation. Structure 2012, 20, 2062–2075. [Google Scholar] [CrossRef]
- Koss, H.; Bunney, T.D.; Behjati, S.; Katan, M. Dysfunction of phospholipase Cγ in immune disorders and cancer. Trends Biochem. Sci. 2014, 39, 603–611. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Zhou, Y.; Sun, D.; Dong, L.; Xia, J.; Yang, B. Oncogenic role of PLCG1 in progression of hepatocellular carcinoma. Hepatol. Res. 2019, 49, 559–569. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Ma, M.H.; Brown, K.R.; Geisler, M.; Li, L.; Tzeng, E.; Jia, C.Y.H.; Jurisica, I.; Li, S.S.C. Systematic identification of SH3 domain-mediated human protein-protein interactions by peptide array target screening. Proteomics 2007, 7, 1775–1785. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.S.; Davis, L.; Huber, H.; Goodhart, P.J.; Wegrzyn, R.E.; Oliff, A.; Heimbrook, D.C. An SH3 domain is required for the mitogenic activity of microinjected phospholipase C-γ1. FEBS Lett. 1995, 358, 287–292. [Google Scholar] [CrossRef]
- Smith, M.R.; Liu, Y.L.; Kim, S.R.; Bae, Y.S.; Kim, C.G.; Kwon, K.S.; Rhee, S.G.; Kung, H.F. PLCy1 Src homology domain induces mitogenesis in quiescent NIH 3T3 fibroblasts. Biochem. Biophys. Res. Commun. 1996, 222, 186–193. [Google Scholar] [CrossRef]
- Seedorf, K.; Kostka, G.; Lammers, R.; Bashkin, P.; Daly, R.; Burgess, W.H.; Van der Bliek, A.M.; Schlessinger, J.; Ullrich, A. Dynamin binds to SH3 domains of phospholipase Cγ and GRB-2. J. Biol. Chem. 1994, 269, 16009–16014. [Google Scholar] [PubMed]
- Kitchen, D.B.; Decornez, H.; Furr, J.R.; Bajorath, J. Docking and scoring in virtual screening for drug discovery: Methods and applications. Nat. Rev. Drug Discov. 2004, 3, 935–949. [Google Scholar] [CrossRef]
- Kwofie, S.K.; Dankwa, B.; Enninful, K.S.; Adobor, C.; Broni, E.; Ntiamoah, A.; Wilson, M.D. Molecular docking and dynamics simulation studies predict munc18b as a target of mycolactone: A plausible mechanism for granule exocytosis impairment in Buruli Ulcer Pathogenesis. Toxins (Basel) 2019, 11, 181. [Google Scholar] [CrossRef]
- Wade, R.C.; Salo-Ahen, O.M.H. Molecular Modeling in Drug Design. Molecules 2019, 24, 321. [Google Scholar] [CrossRef]
- Wang, S.Q.; Du, Q.S.; Huang, R.B.; Zhang, D.W.; Chou, K.C. Insights from investigating the interaction of oseltamivir (Tamiflu) with neuraminidase of the 2009 H1N1 swine flu virus. Biochem. Biophys. Res. Commun. 2009, 386, 432–436. [Google Scholar] [CrossRef]
- Li, X.B.; Wang, S.Q.; Xu, W.R.; Wang, R.L.; Chou, K.C. Novel inhibitor design for hemagglutinin against H1N1 influenza virus by core hopping method. PLoS ONE 2011, 6, e28111. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Wang, S.Q.; Xu, W.R.; Wang, R.L.; Chou, K.C. Design novel dual agonists for treating type-2 diabetes by targeting peroxisome proliferator-activated receptors with core hopping approach. PLoS ONE 2012, 7, e38546. [Google Scholar] [CrossRef] [PubMed]
- Franchini, S.; Battisti, U.M.; Prandi, A.; Tait, A.; Borsari, C.; Cichero, E.; Fossa, P.; Cilia, A.; Prezzavento, O.; Ronsisvalle, S.; et al. Scouting new sigma receptor ligands: Synthesis, pharmacological evaluation and molecular modeling of 1,3-dioxolane-based structures and derivatives. Eur. J. Med. Chem. 2016, 112, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Franchini, S.; Manasieva, L.; Sorbi, C.; Battisti, U.; Fossa, P.; Cichero, E.; Denora, N.; Iacobazzi, R.; Cilia, A.; Pirona, L.; et al. Synthesis, biological evaluation and molecular modeling of 1-oxa-4-thiaspiro- and 1,4-dithiaspiro[4.5]decane derivatives as potent and selective 5-HT1A receptor agonists. Eur. J. Med. Chem. 2017, 125, 435–452. [Google Scholar] [CrossRef] [PubMed]
- Ghamari, N.; Zarei, O.; Reiner, D.; Dastmalchi, S.; Stark, H.; Hamzeh-Mivehroud, M. Histamine H3 receptor ligands by hybrid virtual screening, docking, molecular dynamics simulations, and investigation of their biological effects. Chem. Biol. Drug Des. 2019, 93, 832–843. [Google Scholar] [CrossRef] [PubMed]
- Selvakumar, J.N.; Chandrasekaran, S.D.; Doss, G.P.C.; Kumar, T.D. Inhibition of the ATPase Domain of Human Topoisomerase IIa on HepG2 Cells by 1, 2-benzenedicarboxylic Acid, Mono (2-ethylhexyl) Ester: Molecular Docking and Dynamics Simulations. Curr. Cancer Drug Targets 2019, 19, 495–503. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Bellis, L.J.; Bento, A.P.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; et al. ChEMBL: A large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012, 40, D1100–D1107. [Google Scholar] [CrossRef] [PubMed]
- Dixon, S.L.; Smondyrev, A.M.; Knoll, E.H.; Rao, S.N.; Shaw, D.E.; Friesner, R.A. PHASE: A new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J. Comput. Aided. Mol. Des. 2006, 20, 647–671. [Google Scholar] [CrossRef] [PubMed]
- Dixon, S.L.; Smondyrev, A.M.; Rao, S.N. PHASE: A novel approach to pharmacophore modeling and 3D database searching. Chem. Biol. Drug Des. 2006, 67, 370–372. [Google Scholar] [CrossRef]
- Schrödinger Release 2018-2: Phase; Schrödinger, LLC: New York, NY, USA, 2018.
- Madhavi Sastry, G.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided. Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef]
- Schrödinger Release 2018-2: Protein Preparation Wizard; Schrödinger, LLC: New York, NY, USA, 2016.
- Schrödinger Release 2018-2: LigPrep; Schrödinger, LLC: New York, NY, USA, 2018.
- Shelley, J.C.; Cholleti, A.; Frye, L.L.; Greenwood, J.R.; Timlin, M.R.; Uchimaya, M. Epik: A software program for pKa prediction and protonation state generation for drug-like molecules. J. Comput. Aided. Mol. Des. 2007, 21, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Greenwood, J.R.; Calkins, D.; Sullivan, A.P.; Shelley, J.C. Towards the comprehensive, rapid, and accurate prediction of the favorable tautomeric states of drug-like molecules in aqueous solution. J. Comput. Aided. Mol. Des. 2010, 24, 591–604. [Google Scholar] [CrossRef] [PubMed]
- Schrödinger Release 2018-2: Epik; Schrödinger, LLC: New York, NY, USA, 2018.
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Halgren, T.A.; Murphy, R.B.; Friesner, R.A.; Beard, H.S.; Frye, L.L.; Pollard, W.T.; Banks, J.L. Glide: A new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 2004, 47, 1750–1759. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Murphy, R.B.; Repasky, M.P.; Frye, L.L.; Greenwood, J.R.; Halgren, T.A.; Sanschagrin, P.C.; Mainz, D.T. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef] [PubMed]
- Schrödinger Release 2018-2: Glide; Schrödinger, LLC: New York, NY, USA, 2018.
- Sherman, W.; Day, T.; Jacobson, M.P.; Friesner, R.A.; Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J. Med. Chem. 2006, 49, 534–553. [Google Scholar] [CrossRef]
- Sherman, W.; Beard, H.S.; Farid, R. Use of an induced fit receptor structure in virtual screening. Chem. Biol. Drug Des. 2006, 67, 83–84. [Google Scholar] [CrossRef]
- Schrödinger Release 2018-2: Induced Fit Docking protocol; Glide; Schrödinger, LLC: New York, NY, USA, 2016.
- Jorgensen, W.L.; Maxwell, D.S.; Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 1996, 118, 11225–11236. [Google Scholar] [CrossRef]
- Schrödinger Release 2018-2: Prime; Schrödinger, LLC: New York, NY, USA, 2018.
- Case, D.A.; Ben-Shalom, I.Y.; Brozell, S.R.; Cerutti, D.S.; Cheatham, T.E.I.; Cruzeiro, V.W.D.; Darden, T.A.; Duke, R.E.; Ghoreishi, D.; Gilson, M.K.; et al. Amber 2018; University of California: San Francisco, CA, USA, 2018. [Google Scholar]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general Amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins Struct. Funct. Genet. 2006, 65, 712–725. [Google Scholar] [CrossRef]
- Jakalian, A.; Jack, D.B.; Bayly, C.I. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J. Comput. Chem. 2002, 21, 132–146. [Google Scholar] [CrossRef]
- Mark, P.; Nilsson, L. Structure and dynamics of the TIP3P, SPC, and SPC/E water models at 298 K. J. Phys. Chem. A 2001, 105, 9954–9960. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Forester, T.R.; Smith, W. SHAKE, rattle, and roll: Efficient constraint algorithms for linked rigid bodies. J. Comput. Chem. 1998, 19, 102–111. [Google Scholar] [CrossRef]
- Roe, D.R.; Cheatham, T.E. PTRAJ and CPPTRAJ: Software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 2013, 9, 3084–3095. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Genheden, S.; Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 2015, 10, 449–461. [Google Scholar] [CrossRef]

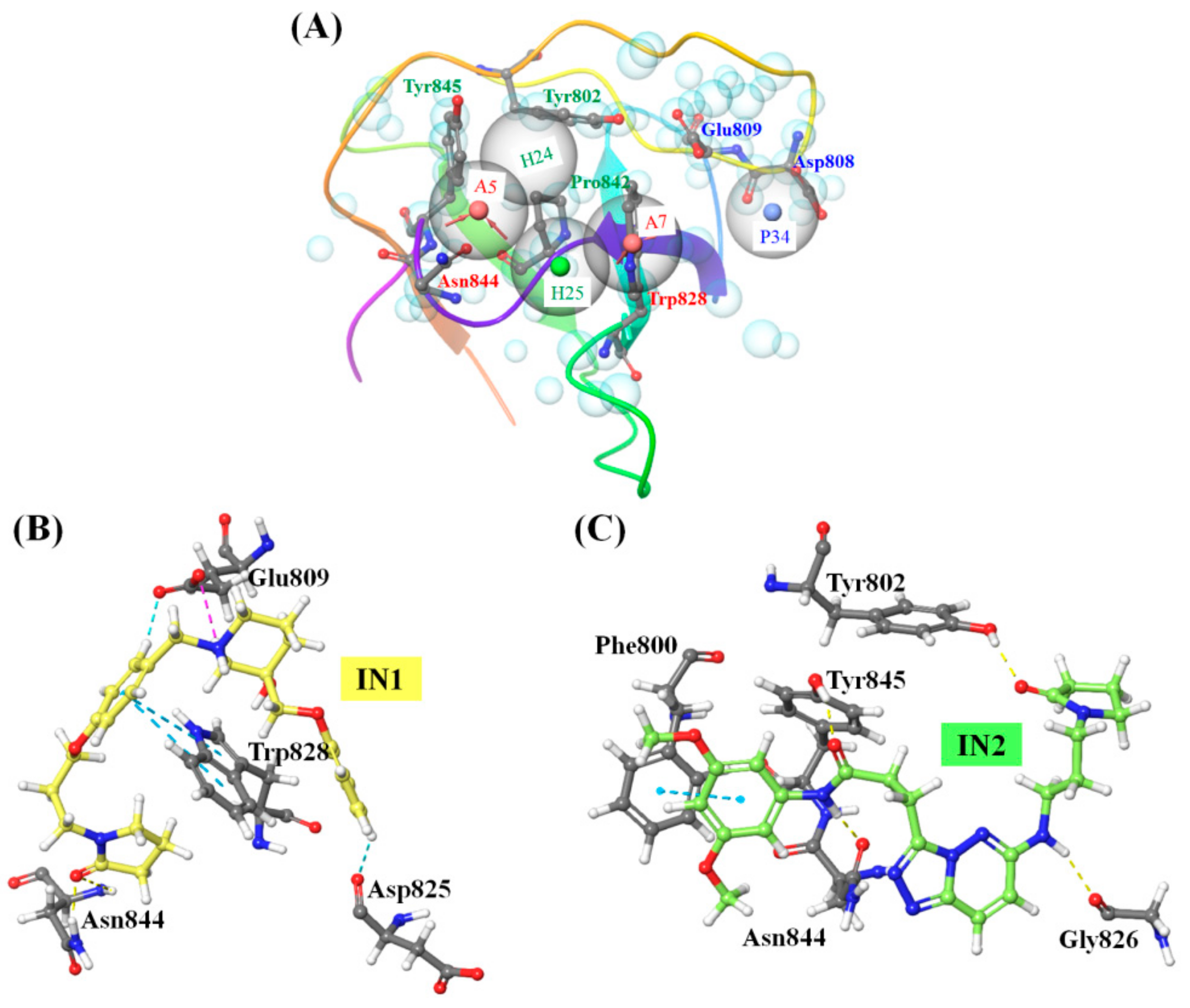


| Title | H-Bond | NH···π/π···π Stacking Interactions | Hydrophobic Interactions | Other Residues within 5Å |
|---|---|---|---|---|
| IN1 | Glu809, Asn844 | Trp828 | Tyr802, Gly826, Gly827, Trp840, Phe841, Pro842, Tyr845 | Gln805, Arg806, Asp808, Gln824, Asp825, Ser843 |
| IN2 | Phe800, Tyr802, Gly826, Asn844 | Tyr845 | Leu799, Gly827, Trp828, Trp840, Phe841, Pro842 | Gln805, Arg806, Asp808, Glu809 |
| IN3 | Arg806, Asp808, Gly826, Asn844 | Trp828, Trp840 | Tyr802, Gly825, Phe841, Pro842, Tyr845 | Gln805, Glu809, Ser843 |
| IN4 | Gln805, Arg806, Asp808, Glu809, Tyr845 | Tyr802, Trp828, Trp840, Pro842, Tyr845, | Lys803, Glu807, Gln824, Asp825, Asn844 | |
| IN5 | Asp808, Glu809 Trp828 | Trp840 | Tyr802, Gly827, Gly826, Trp829, Phe841, Pro842, Tyr845 | Gln805, Arg806, Gln824, Asp825, Arg830, Ser843, Asn844 |
| IN6 | Gln805, Arg806, Asp808, Trp840 | Tyr802, Trp828, Gly826, Gly827 | Gln824, Asp825, Arg830 | |
| IN7 | Glu809, Gly826 | Trp828, Trp840 | Pro842, Tyr845, Tyr802, Gly827 | Gln805, Arg806, Asp808, Asp825, Ser843, Asn844 |
| IN8 | Arg806, Asp808, Asn844 | Trp828, Trp840 | Tyr802, Gly826, Gly827, Phe841, Pro842, Tyr845 | Gln805, Glu809, Gln824, Asp825, Ser843 |
| IN9 | Asp808, Glu809, Trp828, Trp840 | Arg806, Trp840 | Tyr802, Pro842, Tyr845 | Gln805 |
| IN10 | Asp808, Asn844 | Trp840, Trp828 | Tyr845, Tyr802, Gly826, Gly827, Pro842 | Arg806, Glu809, Gln824, Asp825, Ser843 |
| IN11 | Gln805, Trp828, Asn844, | Tyr802, Gly826, Gly827, Trp840, Pro842, Tyr845 | Lys803, Arg806, Asp808, Glu809, Arg830, Ser843, | |
| IN12 | Gln805, Arg806, Glu809 | Trp828 | Tyr802, Gly826, Trp840, Pro842, Tyr845 | Asp801, Lys803, Asp808, Gln824, Asp825, Asn844 |
| IN13 | Asp808, Gly826, Trp828 | Trp828, Trp840 | Tyr802, Gly827, Pro842, Tyr845, | Arg806, Glu809, Asp825, Ser843, Asn844 |
| IN14 | Asp808, Glu809, Gly826 | Trp828 | Tyr802, Gly827, Trp840 | Arg806, Gln824, Asp825 |
| IN15 | Asp808 | Trp828, Trp840 | Tyr802, Gly826, Phe841, Pro842 | Gln805, Arg806, Glu809, Asp825 |
| IN16 | Asp808, Gly826, Asn844 | Trp828 | Gly827, Trp840, Pro842, Tyr845 | Gln805, Arg806, Glu809, Asp825, Ser843 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tripathi, N.; Vetrivel, I.; Téletchéa, S.; Jean, M.; Legembre, P.; Laurent, A.D. Investigation of Phospholipase Cγ1 Interaction with SLP76 Using Molecular Modeling Methods for Identifying Novel Inhibitors. Int. J. Mol. Sci. 2019, 20, 4721. https://doi.org/10.3390/ijms20194721
Tripathi N, Vetrivel I, Téletchéa S, Jean M, Legembre P, Laurent AD. Investigation of Phospholipase Cγ1 Interaction with SLP76 Using Molecular Modeling Methods for Identifying Novel Inhibitors. International Journal of Molecular Sciences. 2019; 20(19):4721. https://doi.org/10.3390/ijms20194721
Chicago/Turabian StyleTripathi, Neha, Iyanar Vetrivel, Stéphane Téletchéa, Mickaël Jean, Patrick Legembre, and Adèle D. Laurent. 2019. "Investigation of Phospholipase Cγ1 Interaction with SLP76 Using Molecular Modeling Methods for Identifying Novel Inhibitors" International Journal of Molecular Sciences 20, no. 19: 4721. https://doi.org/10.3390/ijms20194721
APA StyleTripathi, N., Vetrivel, I., Téletchéa, S., Jean, M., Legembre, P., & Laurent, A. D. (2019). Investigation of Phospholipase Cγ1 Interaction with SLP76 Using Molecular Modeling Methods for Identifying Novel Inhibitors. International Journal of Molecular Sciences, 20(19), 4721. https://doi.org/10.3390/ijms20194721