A Comparative Analysis of Edwardsiella tarda-Induced Transcriptome Profiles in RAW264.7 Cells Reveals New Insights into the Strategy of Bacterial Immune Evasion
Abstract
:1. Introduction
2. Results
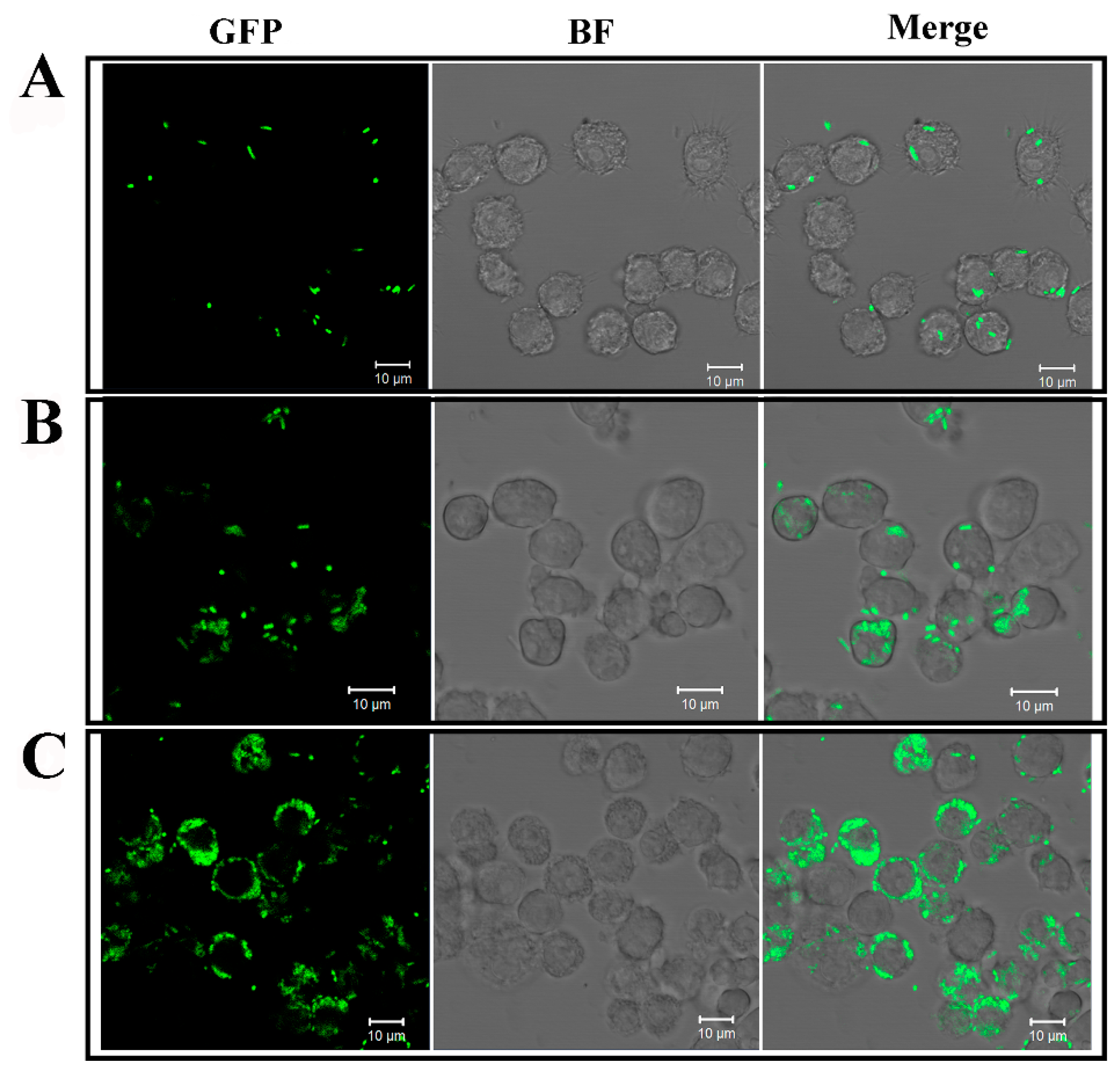
2.1. Infection of E. tarda in RAW264.7 Cells
2.2. RNA Sequencing and Read Mapping
2.3. Differential Expression Analysis
2.4. Validation of DEGs
2.5. GO and KEGG Enrichment Analysis of the DEGs
2.6. DEGs Involved in the Immune Response Induced by Live E. tarda
2.7. The Interaction Networks of Immune-Related DEGs
2.8. Phosphorylation Status of NF-κB in RAW264.7 Cells Infected with Live and Dead E. tarda
3. Discussion
3.1. RIG-I-Like Receptors
3.2. Cytokines
3.3. Interferon-Related Genes
3.4. Other Immune-Related Genes
4. Materials and Methods
4.1. Bacterial Strains and Cell Culture
4.2. Cellular Infection and Confocal Microscopy
4.3. RNA Preparation, Library Construction, and Sequencing
4.4. Sequence Quality Control and Data Processing
4.5. Differentially Expressed Genes (DEGs) Analysis
4.6. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analysis
4.7. Validation of DEGs by Quantitative Real-Time Reverse Transcription-PCR (qRT-PCR)
4.8. Determination of NOS2 Expression and Nitric Oxide (NO) Production
4.9. Construction of Protein-Protein Interaction Networks
4.10. Western Blot to Detect Phospho-NF-κB p65
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Leung, K.Y.; Siame, B.A.; Tenkink, B.J.; Noort, R.J.; Mok, Y.K. Edwardsiella tarda—Virulence mechanisms of an emerging gastroenteritis pathogen. Microbes Infect. 2012, 14, 26–34. [Google Scholar] [CrossRef]
- Park, S.B.; Aoki, T.; Jung, T.S. Pathogenesis of and strategies for preventing Edwardsiella tarda infection in fish. Vet. Res. 2012, 43, 67. [Google Scholar] [CrossRef] [PubMed]
- Golub, V.; Kim, A.C.; Krol, V. Surgical wound infection, tuboovarian abscess, and sepsis caused by Edwardsiella tarda: Case reports and literature review. Infection 2010, 38, 487–489. [Google Scholar] [CrossRef] [PubMed]
- Hirai, Y.; Asahata-Tago, S.; Ainoda, Y.; Fujita, T.; Kikuchi, K. Edwardsiella tarda bacteremia. A rare but fatal water- and foodborne infection: Review of the literature and clinical cases from a single centre. Can. J. Infect. Dis. Med. Microbiol. 2015, 26, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Leung, K.Y.; Wang, Q.; Yang, Z.; Siame, B.A. Edwardsiella piscicida: A versatile emerging pathogen of fish. Virulence 2019, 10, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Kumar, G.; Rathore, G.; El-Matbouli, M. Outer membrane protein assembly factor YaeT (omp85) and GroEL proteins of Edwardsiella tarda are immunogenic antigens for Labeo rohita (Hamilton). J. Fish. Dis. 2014, 37, 1055–1059. [Google Scholar] [CrossRef]
- Das, B.K.; Chakraborty, H.J.; Rout, A.K.; Behera, B.K. De novo whole transcriptome profiling of Edwardsiella tarda isolated from infected fish (Labeo catla). Gene 2019, 701, 152–160. [Google Scholar] [CrossRef]
- Li, M.F.; Sun, L.; Li, J. Edwardsiella tarda evades serum killing by preventing complement activation via the alternative pathway. Fish. Shellfish. Immunol 2015, 43, 325–329. [Google Scholar] [CrossRef]
- Ishibe, K.; Osatomi, K.; Hara, K.; Kanai, K.; Yamaguchi, K.; Oda, T. Comparison of the responses of peritoneal macrophages from Japanese flounder (Paralichthys olivaceus) against high virulent and low virulent strains of Edwardsiella tarda. Fish. Shellfish. Immunol. 2008, 24, 243–251. [Google Scholar] [CrossRef]
- Srinivasa Rao, P.S.; Lim, T.M.; Leung, K.Y. Opsonized virulent Edwardsiella tarda strains are able to adhere to and survive and replicate within fish phagocytes but fail to stimulate reactive oxygen intermediates. Infect. Immun. 2001, 69, 5689–5697. [Google Scholar]
- Qin, L.; Sun, Y.; Zhao, Y.; Xu, J.; Bi, K. In vitro model to estimate Edwardsiella tarda-macrophage interactions using RAW264.7 cells. Fish. Shellfish. Immunol. 2017, 60, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Ni, C.; Xu, W.; Dai, T.; Yang, D.; Wang, Q.; Zhang, Y.; Liu, Q. Intramacrophage infection reinforces the virulence of Edwardsiella tarda. J. Bacteriol. 2016, 198, 1534–1542. [Google Scholar] [CrossRef] [PubMed]
- Furci, L.; Schena, E.; Miotto, P.; Cirillo, D.M. Alteration of human macrophages microRNA expression profile upon infection with Mycobacterium tuberculosis. Int. J. Mycobacteriol. 2013, 2, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Schnappinger, D.; Ehrt, S.; Voskuil, M.I.; Liu, Y.; Mangan, J.A.; Monahan, I.M.; Dolganov, G.; Efron, B.; Butcher, P.D.; Nathan, C.; et al. Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: Insights into the phagosomal environment. J. Exp. Med. 2003, 198, 693–704. [Google Scholar] [CrossRef] [PubMed]
- Barbel, R.; Kaufmann, S.H.E. Immune responses to intracellular bacteria. Curr. Opin. Immunol. 2001, 13, 417–428. [Google Scholar]
- Flannagan, R.S.; Cosio, G.; Grinstein, S. Antimicrobial mechanisms of phagocytes and bacterial evasion strategies. Nat. Rev. Microbiol. 2009, 7, 355–366. [Google Scholar] [CrossRef]
- Shah, S.; Bohsali, A.; Ahlbrand, S.E.; Srinivasan, L.; Rathinam, V.A.; Vogel, S.N.; Fitzgerald, K.A.; Sutterwala, F.S.; Briken, V. Cutting edge: Mycobacterium tuberculosis but not nonvirulent mycobacteria inhibits IFN-beta and AIM2 inflammasome-dependent IL-1beta production via its ESX-1 secretion system. J. Immunol. 2013, 191, 3514–3518. [Google Scholar] [CrossRef]
- Myers, J.T.; Tsang, A.W.; Swanson, J.A. Localized reactive oxygen and nitrogen intermediates inhibit escape of Listeria monocytogenes from vacuoles in activated macrophages. J. Immunol. 2003, 171, 5447–5453. [Google Scholar] [CrossRef]
- Zhou, Z.J.; Sun, L. Edwardsiella tarda-induced inhibition of apoptosis: A strategy for intracellular survival. Front. Cell. Infect. Microbiol. 2016, 6, 76. [Google Scholar] [CrossRef]
- Sui, Z.H.; Xu, H.; Wang, H.; Jiang, S.; Chi, H.; Sun, L. Intracellular Trafficking Pathways of Edwardsiella tarda: From Clathrin- and Caveolin-Mediated Endocytosis to Endosome and Lysosome. Front. Cell. Infect. Microbiol. 2017, 7, 400. [Google Scholar] [CrossRef]
- Chen, H.; Yang, D.; Han, F.; Tan, J.; Zhang, L.; Xiao, J.; Zhang, Y.; Liu, Q. The Bacterial T6SS Effector EvpP Prevents NLRP3 Inflammasome Activation by Inhibiting the Ca(2+)-Dependent MAPK-Jnk Pathway. Cell Host Microbe 2017, 21, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Finlay, B.B.; Falkow, S. Common themes in microbial pathogenicity revisited. Microbiol. Mol. Biol. R. 1997, 61, 136–169. [Google Scholar]
- Loo, Y.M.; Gale, M., Jr. Immune signaling by RIG-I-like receptors. Immunity 2011, 34, 680–692. [Google Scholar] [CrossRef] [PubMed]
- Dixit, E.; Kagan, J.C. Intracellular pathogen detection by RIG-I-like receptors. Adv. Immunol. 2013, 117, 99–125. [Google Scholar] [PubMed]
- Sheikh, F.; Dickensheets, H.; Gamero, A.M.; Vogel, S.N.; Donnelly, R.P. An essential role for IFN-beta in the induction of IFN-stimulated gene expression by LPS in macrophages. J. Leukocyte. Biol. 2014, 96, 591–600. [Google Scholar] [CrossRef]
- Asadpour-Behzadi, A.; Kariminik, A. RIG-1 and MDA5 are the important intracellular sensors against bacteria in septicemia suffering patients. J. Appl. Biomed. 2018, 16, 358–361. [Google Scholar] [CrossRef]
- Monroe, K.M.; McWhirter, S.M.; Vance, R.E. Identification of host cytosolic sensors and bacterial factors regulating the type I interferon response to Legionella pneumophila. Plos Pathog 2009, 5, 11. [Google Scholar] [CrossRef]
- Satoh, T.; Kato, H.; Kumagai, Y.; Yoneyama, M.; Sato, S.; Matsushita, K.; Tsujimura, T.; Fujita, T.; Akira, S.; Takeuchi, O. LGP2 is a positive regulator of RIG-I- and MDA5-mediated antiviral responses. Proc. Natl. Acad. Sci. USA 2010, 107, 1512–1517. [Google Scholar] [CrossRef]
- Borish, L.C.; Steinke, J.W. 2. Cytokines and chemokines. J. Allergy. Clin. Immunol. 2003, 111, S460–S475. [Google Scholar] [CrossRef]
- Medzhitov, R. Origin and physiological roles of inflammation. Nature 2008, 454, 428–435. [Google Scholar] [CrossRef]
- Moreilhon, C.; Gras, D.; Hologne, C.; Bajolet, O.; Cottrez, F.; Magnone, V.; Merten, M.; Groux, H.; Puchelle, E.; Barbry, P. Live Staphylococcus aureus and bacterial soluble factors induce different transcriptional responses in human airway cells. Physiol. Genom. 2005, 20, 244–255. [Google Scholar] [CrossRef] [PubMed]
- Giacomini, E.; Iona, E.; Ferroni, L.; Miettinen, M.; Fattorini, L.; Orefici, G.; Julkunen, I.; Coccia, E.M. Infection of human macrophages and dendritic cells with Mycobacterium tuberculosis induces a differential cytokine gene expression that modulates T cell response. J. Immunol. 2001, 166, 7033–7704. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, A.M.; Savage, N.D.; van Zon, M.; Wilson, L.; Vandenbroucke-Grauls, C.M.; van der Wel, N.N.; Ottenhoff, T.H.; Bitter, W. The ESX-5 secretion system of Mycobacterium marinum modulates the macrophage response. J. Immunol. 2008, 181, 7166–7175. [Google Scholar] [CrossRef] [PubMed]
- Tecchio, C.; Cassatella, M.A. Neutrophil-derived chemokines on the road to immunity. Semin. Immunol. 2016, 28, 119–128. [Google Scholar] [CrossRef]
- Arango Duque, G.; Descoteaux, A. Macrophage cytokines: Involvement in immunity and infectious diseases. Front. Immunol. 2014, 5, 491. [Google Scholar] [CrossRef] [Green Version]
- Audrey, L.K.; Mario, O.; Delia, G.; Alessnadra, O.; Drew, W.; Kira, G.; Elena, H.; Robert, G.; Linda, E.; Anthony, S.F. HIV replication in CD4+ T cells of HIV-infected individuals is regulated by a balance between the viral suppressive effects of endogenous β-chemokines and the viral inductive effects of other endogenous cytokines. P. Natl. Acad. Sci. USA 1996, 93, 14076–14081. [Google Scholar]
- Roach, D.R.; Bean, A.G.; Demangel, C.; France, M.P.; Briscoe, H.; Britton, W.J. TNF regulates chemokine induction essential for cell recruitment, granuloma formation, and clearance of mycobacterial infection. J. Immunol. 2002, 168, 4620–4627. [Google Scholar] [CrossRef] [Green Version]
- Vallerskog, T.; Martens, G.W.; Kornfeld, H. Diabetic mice display a delayed adaptive immune response to Mycobacterium tuberculosis. J. Immunol. 2010, 184, 6275–6282. [Google Scholar] [CrossRef] [Green Version]
- Auerbuch, V.; Brockstedt, D.G.; Meyer-Morse, N.; O’Riordan, M.; Portnoy, D.A. Mice lacking the type I interferon receptor are resistant to Listeria monocytogenes. J. Exp. Med. 2004, 200, 527–533. [Google Scholar] [CrossRef] [Green Version]
- Devitt, A.; Lund, P.A.; Morris, A.G.; Pearce, J.H. Induction of alpha/beta interferon and dependent nitric oxide synthesis during Chlamydia trachomatis infection of McCoy cells in the absence of exogenous cytokine. Infect. Immun. 1996, 64, 3951–3956. [Google Scholar]
- Henry, T.; Brotcke, A.; Weiss, D.S.; Thompson, L.J.; Monack, D.M. Type I interferon signaling is required for activation of the inflammasome during Francisella infection. J. Exp. Med. 2007, 204, 987–994. [Google Scholar] [CrossRef] [PubMed]
- Stetson, D.B.; Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 2006, 24, 93–103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meunier, E.; Wallet, P.; Dreier, R.F.; Costanzo, S.; Anton, L.; Ruhl, S.; Dussurgey, S.; Dick, M.S.; Kistner, A.; Rigard, M.; et al. Guanylate-binding proteins promote activation of the AIM2 inflammasome during infection with Francisella novicida. Nat. Immunol. 2015, 16, 476–484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shenoy, A.R.; Wellington, D.A.; Kumar, P.; Kassa, H.; Booth, C.J.; Cresswell, P.; MacMicking, J.D. GBP5 promotes NLRP3 inflammasome assembly and immunity in mammals. Science 2012, 336, 481–485. [Google Scholar] [CrossRef]
- Fisch, D.; Bando, H.; Clough, B.; Hornung, V.; Yamamoto, M.; Shenoy, A.R.; Frickel, E.M. Human GBP 1 is a microbe-specific gatekeeper of macrophage apoptosis and pyroptosis. Embo. J. 2019, 38, 1–19. [Google Scholar] [CrossRef]
- Fensterl, V.; Sen, G.C. Interferon-induced Ifit proteins: Their role in viral pathogenesis. J. Virol. 2015, 89, 2462–2468. [Google Scholar] [CrossRef] [Green Version]
- MacMicking, J.D. Immune control of phagosomal bacteria by p47 GTPases. Curr. Opin. Microbiol. 2005, 8, 74–82. [Google Scholar] [CrossRef]
- Taylor, G.A. IRG proteins: Key mediators of interferon-regulated host resistance to intracellular pathogens. Cell Microbiol. 2007, 9, 1099–1107. [Google Scholar] [CrossRef]
- Dao, C.T.; Zhang, D.E. ISG15: A ubiquitin-like enigma. Front. Biosci. 2005, 10, 2701–2722. [Google Scholar] [CrossRef]
- Pitha-Rowe, I.F.; Pitha, P.M. Viral defense, carcinogenesis and ISG15: Novel roles for an old ISG. Cytokine. Growth F. R. 2007, 18, 409–417. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.S.; Sun, Y.; Zhang, M.; Sun, L. Identification and analysis of a Sciaenops ocellatus ISG15 homologue that is involved in host immune defense against bacterial infection. Fish. Shellfish. Immunol. 2010, 29, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Chakravortty, D.; Hensel, M. Inducible nitric oxide synthase and control of intracellular bacterial pathogens. Microbes Infect. 2003, 5, 621–627. [Google Scholar] [CrossRef]
- Eriksson, S.; Bjorkman, J.; Borg, S.; Syk, A.; Pettersson, S.; Andresson, D.I.; Rhen, M. Salmonella typhimurium mutants that downregulate phagocyte nitric oxide production. Cell Microbiol. 2010, 2, 239–250. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, L.; Carmichael, J.; Lowrie, D.B.; Andrew, P.W. Strains of Mycobacterium tuberculosis differ in susceptibility to reactive nitrogen intermediates in vitro. Infect. Immun. 1994, 62, 5187–5190. [Google Scholar]
- Ishibe, K.; Yamanishi, T.; Wang, Y.; Osatomi, K.; Hara, K.; Kanai, K.; Yamaguchi, K.; Oda, T. Comparative analysis of the production of nitric oxide (NO) and tumor necrosis factor-alpha (TNF-alpha) from macrophages exposed to high virulent and low virulent strains of Edwardsiella tarda. Fish. Shellfish. Immunol. 2009, 27, 386–389. [Google Scholar] [CrossRef] [Green Version]
- Grayfer, L.; Hodgkinson, J.W.; Belosevic, M. Antimicrobial responses of teleost phagocytes and innate immune evasion strategies of intracellular bacteria. Dev. Comp. Immunol. 2014, 43, 223–242. [Google Scholar] [CrossRef]
- Fast, M.D.; Tse, B.; Boyd, J.M.; Johnson, S.C. Mutations in the Aeromonas salmonicida subsp. salmonicida type III secretion system affect Atlantic salmon leucocyte activation and downstream immune responses. Fish. Shellfish. Immunol. 2009, 27, 721–728. [Google Scholar]
- Teder, P.; Noble, P.W. Endothelin-1 in pulmonary inflammation and fibrosis. Am. J. Resp. Cell Mol. 2000, 23, 7–10. [Google Scholar] [CrossRef]
- Denisenko, E.; Guler, R.; Mhlanga, M.; Suzuki, H.; Brombacher, F.; Schmeier, S. Transcriptionally induced enhancers in the macrophage immune response to Mycobacterium tuberculosis infection. BMC Genom. 2019, 20, 71. [Google Scholar] [CrossRef] [Green Version]
- Allen, L.H.; Aderem, A. A role for MARCKS, the alpha isozyme of protein kinase C and myosin I in zymosan phagocytosis by macrophages. J. Exp. Med. 1995, 182, 829–840. [Google Scholar] [CrossRef]
- Aderem, A. Phagocytosis and the inflammatory response. J. Infect. Dis. 2003, 187, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Samanta, M.; Iwakiri, D.; Kanda, T.; Imaizumi, T.; Takada, K. EB virus-encoded RNAs are recognized by RIG-I and activate signaling to induce type I IFN. EMBO J. 2006, 25, 4207–4214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, Y.S.; Kim, J.H.; Woo, M.; Kim, T.S.; Sohn, K.M.; Lee, Y.H.; Jo, E.K.; Yuk, J.M. Innate signaling mechanisms controlling Mycobacterium chelonae-mediated CCL2 and CCL5 expression in macrophages. J. Microbiol. 2015, 53, 864–874. [Google Scholar] [CrossRef] [PubMed]
- Cao, L.; Li, R.; Chen, X.; Xue, Y.; Liu, D. Neougonin A Inhibits Lipopolysaccharide-Induced Inflammatory Responses via Downregulation of the NF-κB Signaling Pathway in RAW 264.7 Macrophages. Inflammation 2016, 39, 1939–1948. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Sun, K.; Sun, L. Regulation of autoinducer 2 production and luxS expression in a pathogenic Edwardsiella tarda strain. Microbiology 2008, 154, 2060–2069. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [Green Version]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [Green Version]
- Bo, L.; Colin, N.D. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T.; et al. KEGG for linking genomes to life and the environment. Nucleic. Acids. Res. 2008, 36, 480–484. [Google Scholar] [CrossRef]
- Li, Z.; Liu, X.; Liu, J.; Zhang, K.; Yu, H.; He, Y.; Wang, X.; Qi, J.; Wang, Z.; Zhang, Q. Transcriptome profiling based on protein-protein interaction networks provides a core set of genes for understanding blood immune response mechanisms against Edwardsiella tarda infection in Japanese flounder (Paralichthys olivaceus). Dev. Comp. Immunol. 2018, 78, 100–113. [Google Scholar] [CrossRef] [PubMed]







| Samples | Read Length (bp) | Raw Reads | Clean Reads | Clean Reads Ratio (%) | Mapped Reads Ratio (%) | Detected Gene Number |
|---|---|---|---|---|---|---|
| C4h-1 | 150 | 62058308 | 61216228 | 98.64 | 82.79 | 12605 |
| C4h-2 | 150 | 74317910 | 73087158 | 98.34 | 81.68 | 12765 |
| C4h-3 | 150 | 65623654 | 64806176 | 98.75 | 83.67 | 12676 |
| D4h-1 | 150 | 72355478 | 71475480 | 98.78 | 81.79 | 12727 |
| D4h-2 | 150 | 70127974 | 69159932 | 98.62 | 81.45 | 12700 |
| D4h-3 | 150 | 67394630 | 66417284 | 98.55 | 82.45 | 12748 |
| L4h-1 | 150 | 75659098 | 74755562 | 98.81 | 80.57 | 12801 |
| L4h-2 | 150 | 69452342 | 68558994 | 98.71 | 80.3 | 12785 |
| L4h-3 | 150 | 65264048 | 64465772 | 98.78 | 79.78 | 12694 |
| C8h-1 | 150 | 69453594 | 68637212 | 98.82 | 82.06 | 12734 |
| C8h-2 | 150 | 76623908 | 75674390 | 98.76 | 82.32 | 12891 |
| C8h-3 | 150 | 71397364 | 70552920 | 98.82 | 82.08 | 12833 |
| D8h-1 | 150 | 73407730 | 72504312 | 98.77 | 82.45 | 12850 |
| D8h-2 | 150 | 67833338 | 67023630 | 98.81 | 83.21 | 12713 |
| D8h-3 | 150 | 70832694 | 69984558 | 98.8 | 83.15 | 12837 |
| L8h-1 | 150 | 72237008 | 71372390 | 98.8 | 80.15 | 12885 |
| L8h-2 | 150 | 73850386 | 72797386 | 98.57 | 79.58 | 12863 |
| L8h-3 | 150 | 69607688 | 68648300 | 98.62 | 79.86 | 12858 |
| L4h-vs-C4h | D4h-vs-C4h | L4h-vs-D4h | L8h-vs-C8h | D8h-vs-C8h | L8h-vs-D8h | |
|---|---|---|---|---|---|---|
| Upregulated DEGs | 81 | 268 | 57 | 121 | 313 | 63 |
| Downregulated DEGs | 64 | 119 | 151 | 59 | 175 | 169 |
| Total DEGs | 145 | 387 | 208 | 180 | 488 | 232 |
| Category and Gene Name | Fold Change | |
|---|---|---|
| L4h-vs-D4h | L8h-vs-D8h | |
| RIG-I-like receptors | ||
| Interferon induced with helicase C domain 1 (IFIH1) | −2.69 | −2.91 |
| DEAD box polypeptide 58 (DDX58) | ns | −2.49 |
| DEXH box polypeptide 58 (DHX58) | ns | −2.17 |
| Cytokines | ||
| Chemokine (C-C motif) ligand 22 (CCL22) | −3.89 | ns |
| Chemokine (C-C motif) ligand 3 (CCL3) | −2.30 | ns |
| Chemokine (C-C motif) ligand 5 (CCL5) | −7.06 | −5.08 |
| Chemokine (C-X-C motif) ligand 10 (CXCL10) | −3.51 | ns |
| Chemokine (C-C motif) ligand 22 (CCL2) | −3.01 | –2.34 |
| Chemokine (C-X-C motif) ligand 2 (CXCL2) | −2.54 | ns |
| Chemokine (C-C motif) ligand 4 (CCL4) | −3.50 | ns |
| Chemokine (C-C motif) ligand 7 (CCL7) | −2.91 | −2.79 |
| Interleukin 6 (IL6) | −11.96 | −14.96 |
| Interleukin 27 (IL27) | −3.25 | −3.25 |
| Interleukin 1 family, member 9 (IL1F9) | −2.20 | −2.91 |
| Leukemia inhibitory factor (LIF) | −4.18 | ns |
| Colony stimulating factor 3 (CSF3) | −3.69 | −2.59 |
| Interferon-related genes | ||
| ISG15 ubiquitin-like modifier (ISG15) | −3.81 | −4.34 |
| Guanylate binding protein 5 (GBP5) | −4.30 | −5.75 |
| Guanylate binding protein 2 (GBP2) | −4.51 | −5.71 |
| Guanylate binding protein 2b (GBP2b) | −3.02 | −5.63 |
| Guanylate binding protein 7 (GBP7) | ns | −3.23 |
| Guanylate binding protein 9 (GBP9) | ns | −2.74 |
| Guanylate binding protein 3 (GBP3) | ns | −3.75 |
| Interferon-induced protein with tetratricopeptide repeats 3 (IFIT3) | −3.06 | −4.76 |
| Interferon-induced protein with tetratricopeptide repeats 3b (IFIT3b) | −3.30 | −4.01 |
| Interferon-induced protein with tetratricopeptide repeats 1 (IFIT1) | −4.07 | −3.88 |
| Interferon-induced protein with tetratricopeptide repeats 2 (IFIT2) | ns | −2.89 |
| Immunity-related GTPase family M member 1 (IRGM1) | −2.03 | −3.18 |
| Immunity-related GTPase family M member 2 (IRGM2) | −2.76 | −3.06 |
| Interferon gamma inducible protein 47 (IFI47) | ns | −2.92 |
| Interferon regulatory factor 9 (IRF9) | −2.13 | ns |
| Other immune-related genes | ||
| Myristoylated alanine rich protein kinase C substrate (MARCKS) | −6.32 | −4.07 |
| Complement component 5a receptor 1 (C5AR1) | −2.07 | ns |
| Programmed cell death 1 (PDCD1) | −2.01 | ns |
| Heat shock protein 1B (HSPA1B) | 2.46 | 2.06 |
| Endothelin 1 (EDN1) | −5.53 | −3.22 |
| Signal transducer and activator of transcription 2 (STAT2) | ns | −2.04 |
| Nitric oxide synthase 2, inducible (NOS2) | ns | −2.06 |
| Gene Name | Description | Number of Protein-Protein Interaction |
|---|---|---|
| CXCL10 | Chemokine (C-X-C motif) ligand 10 | 29 |
| IL6 | Interleukin 6 | 28 |
| IFIT2 | interferon-induced protein with tetratricopeptide repeats 2 | 23 |
| CCL5 | Chemokine (C-C motif) ligand 5 | 23 |
| ISG15 | ISG15 ubiquitin-like modifier | 21 |
| IFIH1 | Interferon induced with helicase C domain 1 | 21 |
| IFIT1 | Interferon-induced protein with tetratricopeptide repeats 1 | 21 |
| CCL2 | Chemokine (C-C motif) ligand 2 | 19 |
| DDX58 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 58 | 19 |
| IRGM1 | immunity-related GTPase family M member 1 | 18 |
| Gene Name | Forward Primer (5′-3′) | Reverse Primer (5′-3′) | Amplicon Length (bp) |
|---|---|---|---|
| CCL2 | TGCTGACCCCAAGAAGGAAT | TGAGGTGGTTGTGGAAAAGGTA | 184 |
| CCL5 | GACACCACTCCCTGCTGCTT | ACACTTGGCGGTTCCTTCG | 133 |
| CXCL10 | CATCCTGCTGGGTCTGAGTG | ACATTCTTTTTCATCGTGGCA | 177 |
| CSF3 | CCAGAGGCGCATGAAGCTAA | GCTCCAGGGACTTAAGCAGG | 233 |
| HSPA1B | AGAAGGTGCTGGACAAGTGC | AGGCTCCTTTCGGCGG | 192 |
| GBP5 | AGGTCAACGGACCTCGTCTA | CCGGGCCAAGGTTACTACTG | 104 |
| EDN1 | ACCGTATGGACTGGGAGGTT | GGTGAGCGCACTGACATCTA | 101 |
| IFIH1 | CCCAGAAGACAACACAGAATCA | TGGCTCGGGGGATACTCTTT | 163 |
| IFIT1 | AAGGCTGTCCGGTTAAATCC | GAGCTTTGTCTACGCGATGT | 190 |
| ISG15 | GTGCTCCAGGACGGTCTTAC | GACCTCATAGATGTTGCTGTGG | 138 |
| IL27 | CTTCCCAATGTTTCCCTGAC | CGAAGTGTGGTAGCGAGGA | 83 |
| IL6 | GGGAAATCGTGGAAATGAGA | AGGACTCTGGCTTTGTCTTTC | 247 |
| NOS2 | GAGCAACTACTGCTGGTGGT | CGATGTCATGAGCAAAGGCG | 178 |
| GAPDH | ATTCAACGGCACAGTCAAGG | GATGTTAGTGGGGTCTCGCTC | 91 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, H.; Sun, B.; Ning, X.; Jiang, S.; Sun, L. A Comparative Analysis of Edwardsiella tarda-Induced Transcriptome Profiles in RAW264.7 Cells Reveals New Insights into the Strategy of Bacterial Immune Evasion. Int. J. Mol. Sci. 2019, 20, 5724. https://doi.org/10.3390/ijms20225724
Li H, Sun B, Ning X, Jiang S, Sun L. A Comparative Analysis of Edwardsiella tarda-Induced Transcriptome Profiles in RAW264.7 Cells Reveals New Insights into the Strategy of Bacterial Immune Evasion. International Journal of Molecular Sciences. 2019; 20(22):5724. https://doi.org/10.3390/ijms20225724
Chicago/Turabian StyleLi, Huili, Boguang Sun, Xianhui Ning, Shuai Jiang, and Li Sun. 2019. "A Comparative Analysis of Edwardsiella tarda-Induced Transcriptome Profiles in RAW264.7 Cells Reveals New Insights into the Strategy of Bacterial Immune Evasion" International Journal of Molecular Sciences 20, no. 22: 5724. https://doi.org/10.3390/ijms20225724




