The Implications of the Long Non-Coding RNA NEAT1 in Non-Cancerous Diseases
Abstract
:1. Introduction
2. NEAT1: Overview, Domain Architecture, Function
2.1. NEAT1 Overview
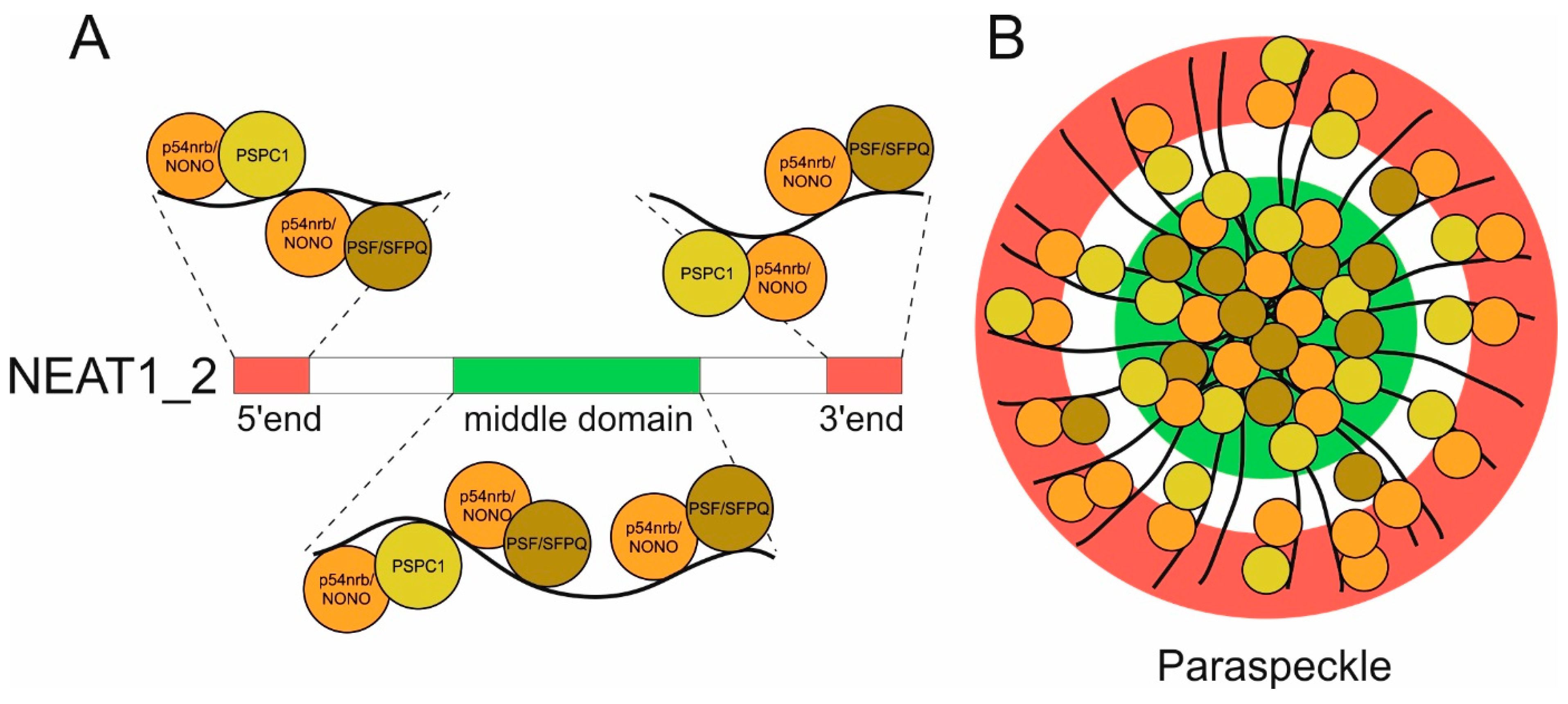
2.2. NEAT1 Domain Architecture
2.3. Cellular Function of NEAT1
2.4. NEAT1 in Physiology
2.5. NEAT1 in Carcinogenesis
3. Immune System and Viral Diseases
3.1. Sepsis and Sepsis-Induced Acute Kidney Injury
3.2. Viral-Induced Diseases
3.2.1. Anti-viral Effects of NEAT1
3.2.2. Pro-viral Effects of NEAT1
4. Neurodegeneration and Neuronal Defects
4.1. Huntington’s Disease
4.2. Multiple Sclerosis
4.3. Amyotrophic Lateral Sclerosis
4.4. Parkinson’s Disease
5. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| A–I | adenosine-to-inosine |
| AD | Alzheimer’s disease |
| ADARs | dsRNA dependent adenosine deaminases |
| AKI | acute kidney injury |
| ALS | amyotrophic lateral sclerosis |
| APP | amyloid-β protein precursor |
| BACE1-AS | β-site APP cleaving enzyme 1 antisense RNA |
| BDNF | brain derived neurotrophic factor |
| ceRNA | competing endogenous RNA |
| CNS | central nervous system |
| DBHS | drosophila behavior human splicing |
| DGCR8 | DiGeorge syndrome critical region 8 |
| FISH | fluorescence in situ hybridization |
| FUS/TLS | fused in sarcoma/translocated in liposarcoma |
| HD | Huntington’s disease |
| HDV | hepatitis D virus |
| hESC | human embryonic stem cells |
| HIV-1 | human immunodeficiency virus 1 |
| Hmx1 | heme oxygenase 1 |
| HTNV | hantavirus |
| IFN | interferon |
| IIM | idiopathic inflammatory myopathy |
| INS elements | instability elements |
| IRAlu | inverted repeated Alu elements |
| KSHV | Kaposi Sarcoma-Associated Herpesvirus |
| LPS | lipopolysaccharide |
| (l)ncRNA | (long) non-coding RNA |
| miRNA | microRNA |
| MPP+ | 1-methyl-4-phenylpyridinium |
| MPTP | 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridin |
| mRNA | messenger RNA |
| MS | multiple sclerosis |
| NEAT1 | nuclear enriched abundant transcript 1 |
| ox-LDL | oxidizied low density lipoprotein |
| p54nrb/NONO | 54 kDa nuclear RNA- and DNA-binding protein/Non-POU domain-containing octamer-binding protein |
| PBMC | peripheral blood mononuclear cells |
| PD | Parkinson’s disease |
| PINK1 | PTEN-induced kinase 1 |
| piRNA | piwi-interacting RNA |
| Pol II | RNA polymerase II |
| PSF/SFPQ | PTB-associated splicing protein/splicing factor proline glutamine rich |
| PSPC1 | paraspeckle component 1 |
| PTB | polypyrimidine tract-binding protein |
| PTEN | phosphatase and tensin homolog |
| RIG-I | retinoic acid inducible gene I |
| SIM | structure illumination microscopy |
| siRNA | short interfering RNA |
| TDP-43 | TAR DNA-binding protein 43 |
| TLR3 | toll-like receptor 3 |
| XPO1 | exportin 1 |
References
- Consortium, E.P. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Vincent, K.; Pichler, M.; Fodde, R.; Berindan-Neagoe, I.; Slack, F.J.; Calin, G.A. Junk DNA and the long non-coding RNA twist in cancer genetics. Oncogene 2015, 34, 5003–5011. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Vagin, V.V.; Sigova, A.; Li, C.; Seitz, H.; Gvozdev, V.; Zamore, P.D. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 2006, 313, 320–324. [Google Scholar] [CrossRef] [PubMed]
- Mello, C.C.; Conte, D., Jr. Revealing the world of RNA interference. Nature 2004, 431, 338–342. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef]
- Stiegelbauer, V.; Vychytilova-Faltejskova, P.; Karbiener, M.; Pehserl, A.M.; Reicher, A.; Resel, M.; Heitzer, E.; Ivan, C.; Bullock, M.; Ling, H.; et al. miR-196b-5p Regulates Colorectal Cancer Cell Migration and Metastases through Interaction with HOXB7 and GALNT5. Clin. Cancer Res. 2017, 23, 5255–5266. [Google Scholar] [CrossRef]
- Pichler, M.; Calin, G.A. MicroRNAs in cancer: From developmental genes in worms to their clinical application in patients. Br. J. Cancer 2015, 113, 569–573. [Google Scholar] [CrossRef]
- Fosselteder, J.; Calin, G.A.; Pichler, M. Long non-coding RNA CCAT2 as a therapeutic target in colorectal cancer. Expert Opin. Targets 2018, 22, 973–976. [Google Scholar] [CrossRef] [PubMed]
- Rigoutsos, I.; Lee, S.K.; Nam, S.Y.; Anfossi, S.; Pasculli, B.; Pichler, M.; Jing, Y.; Rodriguez-Aguayo, C.; Telonis, A.G.; Rossi, S.; et al. N-BLR, a primate-specific non-coding transcript leads to colorectal cancer invasion and migration. Genome Biol. 2017, 18, 98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kung, J.T.; Colognori, D.; Lee, J.T. Long noncoding RNAs: Past, present, and future. Genetics 2013, 193, 651–669. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Richtig, G.; Haemmerle, M.; Pichler, M. From biomarkers to therapeutic targets-the promises and perils of long non-coding RNAs in cancer. Cancer Metastasis Rev. 2018, 37, 83–105. [Google Scholar] [CrossRef] [PubMed]
- Smolle, M.A.; Bauernhofer, T.; Pummer, K.; Calin, G.A.; Pichler, M. Current Insights into Long Non-Coding RNAs (LncRNAs) in Prostate Cancer. Int. J. Mol. Sci. 2017, 18, 473. [Google Scholar] [CrossRef] [PubMed]
- Del Vecchio, F.; Lee, G.H.; Hawezi, J.; Bhome, R.; Pugh, S.; Sayan, E.; Thomas, G.; Packham, G.; Primrose, J.; Pichler, M.; et al. Long non-coding RNAs within the tumour microenvironment and their role in tumour-stroma cross-talk. Cancer Lett. 2018, 421, 94–102. [Google Scholar] [CrossRef] [PubMed]
- Smolle, M.A.; Pichler, M. The Role of Long Non-Coding RNAs in Osteosarcoma. Non-Coding RNA 2018, 4, 7. [Google Scholar] [CrossRef] [PubMed]
- Klec, C.; Prinz, F.; Pichler, M. Involvement of the long non-coding RNA NEAT1 in carcinogenesis. Mol. Oncol. 2018. [Google Scholar] [CrossRef]
- Hutchinson, J.N.; Ensminger, A.W.; Clemson, C.M.; Lynch, C.R.; Lawrence, J.B.; Chess, A. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genom. 2007, 8, 39. [Google Scholar] [CrossRef] [PubMed]
- Guru, S.C.; Agarwal, S.K.; Manickam, P.; Olufemi, S.E.; Crabtree, J.S.; Weisemann, J.M.; Kester, M.B.; Kim, Y.S.; Wang, Y.; Emmert-Buck, M.R.; et al. A transcript map for the 2.8-Mb region containing the multiple endocrine neoplasia type 1 locus. Genome Res. 1997, 7, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, Y.T.; Ideue, T.; Sano, M.; Mituyama, T.; Hirose, T. MENepsilon/beta noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc. Natl. Acad. Sci. USA 2009, 106, 2525–2530. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, S.; Naganuma, T.; Shioi, G.; Hirose, T. Paraspeckles are subpopulation-specific nuclear bodies that are not essential in mice. J. Cell Biol. 2011, 193, 31–39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clemson, C.M.; Hutchinson, J.N.; Sara, S.A.; Ensminger, A.W.; Fox, A.H.; Chess, A.; Lawrence, J.B. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell 2009, 33, 717–726. [Google Scholar] [CrossRef] [PubMed]
- Prasanth, K.V.; Prasanth, S.G.; Xuan, Z.; Hearn, S.; Freier, S.M.; Bennett, C.F.; Zhang, M.Q.; Spector, D.L. Regulating gene expression through RNA nuclear retention. Cell 2005, 123, 249–263. [Google Scholar] [CrossRef] [PubMed]
- Fox, A.H.; Lam, Y.W.; Leung, A.K.; Lyon, C.E.; Andersen, J.; Mann, M.; Lamond, A.I. Paraspeckles: A novel nuclear domain. Curr. Biol. 2002, 12, 13–25. [Google Scholar] [CrossRef]
- Bond, C.S.; Fox, A.H. Paraspeckles: Nuclear bodies built on long noncoding RNA. J. Cell Biol. 2009, 186, 637–644. [Google Scholar] [CrossRef]
- Zhang, Z.; Carmichael, G.G. The fate of dsRNA in the nucleus: A p54(nrb)-containing complex mediates the nuclear retention of promiscuously A-to-I edited RNAs. Cell 2001, 106, 465–475. [Google Scholar] [CrossRef]
- Chen, L.L.; Carmichael, G.G. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: Functional role of a nuclear noncoding RNA. Mol. Cell 2009, 35, 467–478. [Google Scholar] [CrossRef]
- Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 2002, 71, 817–846. [Google Scholar] [CrossRef]
- Torres, M.; Becquet, D.; Blanchard, M.P.; Guillen, S.; Boyer, B.; Moreno, M.; Franc, J.L.; Francois-Bellan, A.M. Paraspeckles as rhythmic nuclear mRNA anchorages responsible for circadian gene expression. Nucleus 2017, 8, 249–254. [Google Scholar] [CrossRef]
- Fox, A.H.; Bond, C.S.; Lamond, A.I. P54nrb forms a heterodimer with PSP1 that localizes to paraspeckles in an RNA-dependent manner. Mol. Biol. Cell 2005, 16, 5304–5315. [Google Scholar] [CrossRef] [PubMed]
- Murthy, U.M.; Rangarajan, P.N. Identification of protein interaction regions of VINC/NEAT1/Men epsilon RNA. FEBS Lett. 2010, 584, 1531–1535. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mito, M.; Kawaguchi, T.; Hirose, T.; Nakagawa, S. Simultaneous multicolor detection of RNA and proteins using super-resolution microscopy. Methods 2016, 98, 158–165. [Google Scholar] [CrossRef] [PubMed]
- West, J.A.; Mito, M.; Kurosaka, S.; Takumi, T.; Tanegashima, C.; Chujo, T.; Yanaka, K.; Kingston, R.E.; Hirose, T.; Bond, C.; et al. Structural, super-resolution microscopy analysis of paraspeckle nuclear body organization. J. Cell Biol. 2016, 214, 817–830. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamazaki, T.; Souquere, S.; Chujo, T.; Kobelke, S.; Chong, Y.S.; Fox, A.H.; Bond, C.S.; Nakagawa, S.; Pierron, G.; Hirose, T. Functional Domains of NEAT1 Architectural lncRNA Induce Paraspeckle Assembly through Phase Separation. Mol. Cell 2018. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Harvey, A.R.; Hodgetts, S.I.; Fox, A.H. Functional dissection of NEAT1 using genome editing reveals substantial localization of the NEAT1_1 isoform outside paraspeckles. RNA 2017, 23, 872–881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, Y.; Schmidt, B.F.; Bruchez, M.P.; McManus, C.J. Structural analyses of NEAT1 lncRNAs suggest long-range RNA interactions that may contribute to paraspeckle architecture. Nucleic Acids Res. 2018, 46, 3742–3752. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zampetaki, A.; Albrecht, A.; Steinhofel, K. Long Non-coding RNA Structure and Function: Is There a Link? Front. Physiol. 2018, 9, 1201. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Shao, C.; Wu, Q.J.; Chen, G.; Zhou, J.; Yang, B.; Li, H.; Gou, L.T.; Zhang, Y.; Wang, Y.; et al. NEAT1 scaffolds RNA-binding proteins and the Microprocessor to globally enhance pri-miRNA processing. Nat. Struct. Mol. Biol. 2017, 24, 816–824. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.L.; DeCerbo, J.N.; Carmichael, G.G. Alu element-mediated gene silencing. EMBO J. 2008, 27, 1694–1705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirose, T.; Virnicchi, G.; Tanigawa, A.; Naganuma, T.; Li, R.; Kimura, H.; Yokoi, T.; Nakagawa, S.; Benard, M.; Fox, A.H.; et al. NEAT1 long noncoding RNA regulates transcription via protein sequestration within subnuclear bodies. Mol. Biol. Cell 2014, 25, 169–183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakagawa, S.; Shimada, M.; Yanaka, K.; Mito, M.; Arai, T.; Takahashi, E.; Fujita, Y.; Fujimori, T.; Standaert, L.; Marine, J.C.; et al. The lncRNA Neat1 is required for corpus luteum formation and the establishment of pregnancy in a subpopulation of mice. Development 2014, 141, 4618–4627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Standaert, L.; Adriaens, C.; Radaelli, E.; Van Keymeulen, A.; Blanpain, C.; Hirose, T.; Nakagawa, S.; Marine, J.C. The long noncoding RNA Neat1 is required for mammary gland development and lactation. RNA 2014, 20, 1844–1849. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barry, G.; Briggs, J.A.; Hwang, D.W.; Nayler, S.P.; Fortuna, P.R.; Jonkhout, N.; Dachet, F.; Maag, J.L.; Mestdagh, P.; Singh, E.M.; et al. The long non-coding RNA NEAT1 is responsive to neuronal activity and is associated with hyperexcitability states. Sci. Rep. 2017, 7, 40127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Lun, L.; Li, H.; Wang, Q.; Lin, J.; Tian, R.; Pan, H.; Zhang, H.; Chen, X. The Value of lncRNA NEAT1 as a Prognostic Factor for Survival of Cancer Outcome: A Meta-Analysis. Sci. Rep. 2017, 7, 13080. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Wang, H.; Yang, P.; He, Z.Y. Prognostic role of long noncoding RNA NEAT1 in various carcinomas: A meta-analysis. Oncotargets Ther. 2017, 10, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.J.; Lin, Q.; Ma, J.X.; Shi, W.W.; Yang, B.; Li, F. Long non-coding RNA NEAT1 acts as oncogene in NSCLC by regulating the Wnt signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 504–510. [Google Scholar]
- Jen, J.; Tang, Y.A.; Lu, Y.H.; Lin, C.C.; Lai, W.W.; Wang, Y.C. Oct4 transcriptionally regulates the expression of long non-coding RNAs NEAT1 and MALAT1 to promote lung cancer progression. Mol. Cancer 2017, 16, 104. [Google Scholar] [CrossRef]
- Zhang, M.; Wu, W.B.; Wang, Z.W.; Wang, X.H. lncRNA NEAT1 is closely related with progression of breast cancer via promoting proliferation and EMT. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 1020–1026. [Google Scholar]
- Wu, F.; Mo, Q.; Wan, X.; Dan, J.; Hu, H. NEAT1/has-mir-98-5p/MAPK6 axis is involved in non-small-cell lung cancer (NSCLC) development. J. Cell. Biochem. 2017. [Google Scholar] [CrossRef]
- Ke, H.; Zhao, L.; Feng, X.; Xu, H.; Zou, L.; Yang, Q.; Su, X.; Peng, L.; Jiao, B. NEAT1 is Required for Survival of Breast Cancer Cells Through FUS and miR-548. Gene Regul. Syst. Biol. 2016, 10, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Lo, P.K.; Zhang, Y.; Wolfson, B.; Gernapudi, R.; Yao, Y.; Duru, N.; Zhou, Q. Dysregulation of the BRCA1/long non-coding RNA NEAT1 signaling axis contributes to breast tumorigenesis. Oncotarget 2016, 7, 65067–65089. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, W.; Zhang, Z.; Liu, X.; Cheng, X.; Zhang, Y.; Han, X.; Zhang, Y.; Liu, S.; Yang, J.; Xu, B.; et al. The FOXN3-NEAT1-SIN3A repressor complex promotes progression of hormonally responsive breast cancer. J. Clin. Investig. 2017, 127, 3421–3440. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Sun, J.; Pan, Z.; Song, Y.; Zhong, L.; Zhang, Y.; Liu, Y.; Zheng, X.; Huang, P. Long non-coding RNA NEAT1 promotes hepatocellular carcinoma cell proliferation through the regulation of miR-129-5p-VCP-IkappaB. Am. J. Physiol. Gastrointest. Liver Physiol. 2017, 313, G150–G156. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zou, Q.; Song, M.; Chen, J. NEAT1 promotes cell proliferation and invasion in hepatocellular carcinoma by negative regulating miR-613 expression. Biomed. Pharmacother. Biomed. Pharmacother. 2017, 94, 612–618. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.N.; Zhou, J.; Lu, X.J. The long noncoding RNA NEAT1 contributes to hepatocellular carcinoma development by sponging miR-485 and enhancing the expression of the STAT3. J. Cell. Physiol. 2018, 233, 6733–6741. [Google Scholar] [CrossRef]
- Tu, J.; Zhao, Z.; Xu, M.; Lu, X.; Chang, L.; Ji, J. NEAT1 upregulates TGF-beta1 to induce hepatocellular carcinoma progression by sponging hsa-mir-139-5p. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef]
- Chen, Z.J.; Zhang, Z.; Xie, B.B.; Zhang, H.Y. Clinical significance of up-regulated lncRNA NEAT1 in prognosis of ovarian cancer. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3373–3377. [Google Scholar]
- Ding, N.; Wu, H.; Tao, T.; Peng, E. NEAT1 regulates cell proliferation and apoptosis of ovarian cancer by miR-34a-5p/BCL2. Oncotargets Ther. 2017, 10, 4905–4915. [Google Scholar] [CrossRef]
- An, J.; Lv, W.; Zhang, Y. LncRNA NEAT1 contributes to paclitaxel resistance of ovarian cancer cells by regulating ZEB1 expression via miR-194. Oncotargets Ther. 2017, 10, 5377–5390. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Y.; Fu, X.; Lu, Z. Long non-coding RNA NEAT1 promoted ovarian cancer cells’ metastasis via regulating of miR-382-3p/ROCK1 axial. Cancer Sci. 2018. [Google Scholar] [CrossRef]
- Xiong, W.; Huang, C.; Deng, H.; Jian, C.; Zen, C.; Ye, K.; Zhong, Z.; Zhao, X.; Zhu, L. Oncogenic non-coding RNA NEAT1 promotes the prostate cancer cell growth through the SRC3/IGF1R/AKT pathway. Int. J. Biochem. Cell Biol. 2018, 94, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, X.; Song, W.; Xu, H.; Huang, R.; Wang, Y.; Zhao, W.; Xiao, Z.; Yang, X. Oncogenic properties of NEAT1 in prostate cancer cells depend on the CDC5L-AGRN transcriptional regulation circuit. Cancer Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Zhang, D.H.; Wu, N.; Xiao, J.H.; Wang, X.; Ma, W. ceRNA in cancer: Possible functions and clinical implications. J. Med Genet. 2015, 52, 710–718. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Geng, H.; Tan, X.D. Functional diversity of long non-coding RNAs in immune regulation. Genes Dis. 2016, 3, 72–81. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smolle, M.A.; Calin, H.N.; Pichler, M.; Calin, G.A. Noncoding RNAs and immune checkpoints-clinical implications as cancer therapeutics. FEBS J. 2017, 284, 1952–1966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, G.; Tang, Q.; Sharma, S.; Yu, F.; Escobar, T.M.; Muljo, S.A.; Zhu, J.; Zhao, K. Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat. Immunol. 2013, 14, 1190–1198. [Google Scholar] [CrossRef] [Green Version]
- Ranzani, V.; Rossetti, G.; Panzeri, I.; Arrigoni, A.; Bonnal, R.J.; Curti, S.; Gruarin, P.; Provasi, E.; Sugliano, E.; Marconi, M.; et al. The long intergenic noncoding RNA landscape of human lymphocytes highlights the regulation of T cell differentiation by linc-MAF-4. Nat. Immunol. 2015, 16, 318–325. [Google Scholar] [CrossRef] [Green Version]
- Venkatraman, A.; He, X.C.; Thorvaldsen, J.L.; Sugimura, R.; Perry, J.M.; Tao, F.; Zhao, M.; Christenson, M.K.; Sanchez, R.; Yu, J.Y.; et al. Maternal imprinting at the H19-Igf2 locus maintains adult haematopoietic stem cell quiescence. Nature 2013, 500, 345–349. [Google Scholar] [CrossRef]
- Wang, P.; Xue, Y.; Han, Y.; Lin, L.; Wu, C.; Xu, S.; Jiang, Z.; Xu, J.; Liu, Q.; Cao, X. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 2014, 344, 310–313. [Google Scholar] [CrossRef] [PubMed]
- Kotzin, J.J.; Spencer, S.P.; McCright, S.J.; Kumar, D.B.U.; Collet, M.A.; Mowel, W.K.; Elliott, E.N.; Uyar, A.; Makiya, M.A.; Dunagin, M.C.; et al. The long non-coding RNA Morrbid regulates Bim and short-lived myeloid cell lifespan. Nature 2016, 537, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Spurlock, C.F., 3rd; Tossberg, J.T.; Guo, Y.; Collier, S.P.; Crooke, P.S., 3rd; Aune, T.M. Expression and functions of long noncoding RNAs during human T helper cell differentiation. Nat. Commun. 2015, 6, 6932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carpenter, S.; Aiello, D.; Atianand, M.K.; Ricci, E.P.; Gandhi, P.; Hall, L.L.; Byron, M.; Monks, B.; Henry-Bezy, M.; Lawrence, J.B.; et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 2013, 341, 789–792. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.G.; Satpathy, A.T.; Chang, H.Y. Gene regulation in the immune system by long noncoding RNAs. Nat. Immunol. 2017, 18, 962–972. [Google Scholar] [CrossRef] [PubMed]
- Zeng, C.; Xu, Y.; Xu, L.; Yu, X.; Cheng, J.; Yang, L.; Chen, S.; Li, Y. Inhibition of long non-coding RNA NEAT1 impairs myeloid differentiation in acute promyelocytic leukemia cells. BMC Cancer 2014, 14, 693. [Google Scholar] [CrossRef]
- Giza, D.E.; Fuentes-Mattei, E.; Bullock, M.D.; Tudor, S.; Goblirsch, M.J.; Fabbri, M.; Lupu, F.; Yeung, S.J.; Vasilescu, C.; Calin, G.A. Cellular and viral microRNAs in sepsis: Mechanisms of action and clinical applications. Cell Death Differ. 2016, 23, 1906–1918. [Google Scholar] [CrossRef]
- Fuentes-Mattei, E.; Giza, D.E.; Shimizu, M.; Ivan, C.; Manning, J.T.; Tudor, S.; Ciccone, M.; Kargin, O.A.; Zhang, X.; Mur, P.; et al. Plasma Viral miRNAs Indicate a High Prevalence of Occult Viral Infections. EBioMedicine 2017, 20, 182–192. [Google Scholar] [CrossRef]
- Huang, S.; Qian, K.; Zhu, Y.; Huang, Z.; Luo, Q.; Qing, C. Diagnostic Value of the lncRNA NEAT1 in Peripheral Blood Mononuclear Cells of Patients with Sepsis. Dis. Markers 2017, 2017, 7962836. [Google Scholar] [CrossRef]
- Chen, Y.; Qiu, J.; Chen, B.; Lin, Y.; Chen, Y.; Xie, G.; Qiu, J.; Tong, H.; Jiang, D. Long non-coding RNA NEAT1 plays an important role in sepsis-induced acute kidney injury by targeting miR-204 and modulating the NF-kappaB pathway. Int. Immunopharmacol. 2018, 59, 252–260. [Google Scholar] [CrossRef]
- Li, X.Y.; Zhang, K.; Jiang, Z.Y.; Cai, L.H. MiR-204/miR-211 downregulation contributes to candidemia-induced kidney injuries via derepression of Hmx1 expression. Life Sci. 2014, 102, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Abraham, E. Nuclear factor-kappaB and its role in sepsis-associated organ failure. J. Infect. Dis. 2003, 187 (Suppl. 2), S364–S369. [Google Scholar] [CrossRef]
- Zhang, Q.; Chen, C.Y.; Yedavalli, V.S.; Jeang, K.T. NEAT1 long noncoding RNA and paraspeckle bodies modulate HIV-1 posttranscriptional expression. mBio 2013, 4, e00596-12. [Google Scholar] [CrossRef] [PubMed]
- Budhiraja, S.; Liu, H.; Couturier, J.; Malovannaya, A.; Qin, J.; Lewis, D.E.; Rice, A.P. Mining the human complexome database identifies RBM14 as an XPO1-associated protein involved in HIV-1 Rev function. J. Virol. 2015, 89, 3557–3567. [Google Scholar] [CrossRef] [PubMed]
- Pandey, A.D.; Goswami, S.; Shukla, S.; Das, S.; Ghosal, S.; Pal, M.; Bandyopadhyay, B.; Ramachandran, V.; Basu, N.; Sood, V.; et al. Correlation of altered expression of a long non-coding RNA, NEAT1, in peripheral blood mononuclear cells with dengue disease progression. J. Infect. 2017, 75, 541–554. [Google Scholar] [CrossRef] [PubMed]
- Adriaens, C.; Standaert, L.; Barra, J.; Latil, M.; Verfaillie, A.; Kalev, P.; Boeckx, B.; Wijnhoven, P.W.; Radaelli, E.; Vermi, W.; et al. p53 induces formation of NEAT1 lncRNA-containing paraspeckles that modulate replication stress response and chemosensitivity. Nat. Med. 2016, 22, 861–868. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, M.; Kikuchi, M.; Natsukawa, T.; Shinobu, N.; Imaizumi, T.; Miyagishi, M.; Taira, K.; Akira, S.; Fujita, T. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 2004, 5, 730–737. [Google Scholar] [CrossRef] [PubMed]
- Loo, Y.M.; Gale, M., Jr. Immune signaling by RIG-I-like receptors. Immunity 2011, 34, 680–692. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Han, P.; Ye, W.; Chen, H.; Zheng, X.; Cheng, L.; Zhang, L.; Yu, L.; Wu, X.; Xu, Z.; et al. The Long Noncoding RNA NEAT1 Exerts Antihantaviral Effects by Acting as Positive Feedback for RIG-I Signaling. J. Virol. 2017, 91, JVI-02250. [Google Scholar] [CrossRef] [PubMed]
- Beeharry, Y.; Goodrum, G.; Imperiale, C.J.; Pelchat, M. The Hepatitis Delta Virus accumulation requires paraspeckle components and affects NEAT1 level and PSP1 localization. Sci. Rep. 2018, 8, 6031. [Google Scholar] [CrossRef] [PubMed]
- Imamura, K.; Imamachi, N.; Akizuki, G.; Kumakura, M.; Kawaguchi, A.; Nagata, K.; Kato, A.; Kawaguchi, Y.; Sato, H.; Yoneda, M.; et al. Long noncoding RNA NEAT1-dependent SFPQ relocation from promoter region to paraspeckle mediates IL8 expression upon immune stimuli. Mol. Cell 2014, 53, 393–406. [Google Scholar] [CrossRef] [PubMed]
- Viollet, C.; Davis, D.A.; Tekeste, S.S.; Reczko, M.; Ziegelbauer, J.M.; Pezzella, F.; Ragoussis, J.; Yarchoan, R. RNA Sequencing Reveals that Kaposi Sarcoma-Associated Herpesvirus Infection Mimics Hypoxia Gene Expression Signature. PLoS Pathog. 2017, 13, e1006143. [Google Scholar] [CrossRef]
- Wang, Z.; Fan, P.; Zhao, Y.; Zhang, S.; Lu, J.; Xie, W.; Jiang, Y.; Lei, F.; Xu, N.; Zhang, Y. NEAT1 modulates herpes simplex virus-1 replication by regulating viral gene transcription. Cell. Mol. Life Sci. 2017, 74, 1117–1131. [Google Scholar] [CrossRef] [PubMed]
- Choudhry, H.; Albukhari, A.; Morotti, M.; Haider, S.; Moralli, D.; Smythies, J.; Schodel, J.; Green, C.M.; Camps, C.; Buffa, F.; et al. Tumor hypoxia induces nuclear paraspeckle formation through HIF-2alpha dependent transcriptional activation of NEAT1 leading to cancer cell survival. Oncogene 2015, 34, 4546. [Google Scholar] [CrossRef]
- Riva, P.; Ratti, A.; Venturin, M. The Long Non-Coding RNAs in Neurodegenerative Diseases: Novel Mechanisms of Pathogenesis. Curr. Alzheimer Res. 2016, 13, 1219–1231. [Google Scholar] [CrossRef]
- Li, D.; Yang, M.Q. Identification and characterization of conserved lncRNAs in human and rat brain. BMC Bioinform. 2017, 18, 489. [Google Scholar] [CrossRef]
- D’Haene, E.; Jacobs, E.Z.; Volders, P.J.; De Meyer, T.; Menten, B.; Vergult, S. Identification of long non-coding RNAs involved in neuronal development and intellectual disability. Sci. Rep. 2016, 6, 28396. [Google Scholar] [CrossRef] [PubMed]
- Ramos, A.D.; Diaz, A.; Nellore, A.; Delgado, R.N.; Park, K.Y.; Gonzales-Roybal, G.; Oldham, M.C.; Song, J.S.; Lim, D.A. Integration of genome-wide approaches identifies lncRNAs of adult neural stem cells and their progeny in vivo. Cell Stem Cell 2013, 12, 616–628. [Google Scholar] [CrossRef]
- Quan, Z.; Zheng, D.; Qing, H. Regulatory Roles of Long Non-Coding RNAs in the Central Nervous System and Associated Neurodegenerative Diseases. Front. Cell Neurosci. 2017, 11, 175. [Google Scholar] [CrossRef]
- Scheele, C.; Petrovic, N.; Faghihi, M.A.; Lassmann, T.; Fredriksson, K.; Rooyackers, O.; Wahlestedt, C.; Good, L.; Timmons, J.A. The human PINK1 locus is regulated in vivo by a non-coding natural antisense RNA during modulation of mitochondrial function. BMC Genom. 2007, 8, 74. [Google Scholar] [CrossRef] [PubMed]
- Chiba, M.; Kiyosawa, H.; Hiraiwa, N.; Ohkohchi, N.; Yasue, H. Existence of Pink1 antisense RNAs in mouse and their localization. Cytogenet. Genome Res. 2009, 126, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Morais, V.A.; Verstreken, P.; Roethig, A.; Smet, J.; Snellinx, A.; Vanbrabant, M.; Haddad, D.; Frezza, C.; Mandemakers, W.; Vogt-Weisenhorn, D.; et al. Parkinson’s disease mutations in PINK1 result in decreased Complex I activity and deficient synaptic function. Embo Mol. Med. 2009, 1, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Noble, E.E.; Billington, C.J.; Kotz, C.M.; Wang, C. The lighter side of BDNF. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2011, 300, R1053–R1069. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Y.; Hayden, M.R.; Xu, B. BDNF overexpression in the forebrain rescues Huntington’s disease phenotypes in YAC128 mice. J. Neurosci. 2010, 30, 14708–14718. [Google Scholar] [CrossRef]
- Wan, P.; Su, W.; Zhuo, Y. The Role of Long Noncoding RNAs in Neurodegenerative Diseases. Mol. Neurobiol. 2017, 54, 2012–2021. [Google Scholar] [CrossRef] [PubMed]
- Michelhaugh, S.K.; Lipovich, L.; Blythe, J.; Jia, H.; Kapatos, G.; Bannon, M.J. Mining Affymetrix microarray data for long non-coding RNAs: Altered expression in the nucleus accumbens of heroin abusers. J. Neurochem. 2011, 116, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Jiang, L.; Huang, Z.; Zhang, H.; Cheng, C.; Liu, H.; He, J.; Wu, J.; Darwazeh, R.; Wu, Y.; et al. The long non-coding RNA Neat1 is an important mediator of the therapeutic effect of bexarotene on traumatic brain injury in mice. Brain Behav. Immun. 2017, 65, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Bates, G.P. History of genetic disease: The molecular genetics of Huntington disease—A history. Nat. Rev. Genet. 2005, 6, 766–773. [Google Scholar] [CrossRef]
- Sunwoo, J.S.; Lee, S.T.; Im, W.; Lee, M.; Byun, J.I.; Jung, K.H.; Park, K.I.; Jung, K.Y.; Lee, S.K.; Chu, K.; et al. Altered Expression of the Long Noncoding RNA NEAT1 in Huntington’s Disease. Mol. Neurobiol. 2017, 54, 1577–1586. [Google Scholar] [CrossRef]
- Johnson, R. Long non-coding RNAs in Huntington’s disease neurodegeneration. Neurobiol. Dis. 2012, 46, 245–254. [Google Scholar] [CrossRef]
- Goldenberg, M.M. Multiple sclerosis review. P T 2012, 37, 175–184. [Google Scholar] [PubMed]
- Santoro, M.; Nociti, V.; Lucchini, M.; De Fino, C.; Losavio, F.A.; Mirabella, M. Expression Profile of Long Non-Coding RNAs in Serum of Patients with Multiple Sclerosis. J. Mol. Neurosci. 2016, 59, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Lund, B.T.; Ashikian, N.; Ta, H.Q.; Chakryan, Y.; Manoukian, K.; Groshen, S.; Gilmore, W.; Cheema, G.S.; Stohl, W.; Burnett, M.E.; et al. Increased CXCL8 (IL-8) expression in Multiple Sclerosis. J. Neuroimmunol. 2004, 155, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Bsibsi, M.; Bajramovic, J.J.; Vogt, M.H.; van Duijvenvoorden, E.; Baghat, A.; Persoon-Deen, C.; Tielen, F.; Verbeek, R.; Huitinga, I.; Ryffel, B.; et al. The microtubule regulator stathmin is an endogenous protein agonist for TLR3. J. Immunol. 2010, 184, 6929–6937. [Google Scholar] [CrossRef] [PubMed]
- Zarei, S.; Carr, K.; Reiley, L.; Diaz, K.; Guerra, O.; Altamirano, P.F.; Pagani, W.; Lodin, D.; Orozco, G.; Chinea, A. A comprehensive review of amyotrophic lateral sclerosis. Surg. Neurol. Int. 2015, 6, 171. [Google Scholar] [CrossRef] [PubMed]
- Nishimoto, Y.; Nakagawa, S.; Hirose, T.; Okano, H.J.; Takao, M.; Shibata, S.; Suyama, S.; Kuwako, K.; Imai, T.; Murayama, S.; et al. The long non-coding RNA nuclear-enriched abundant transcript 1_2 induces paraspeckle formation in the motor neuron during the early phase of amyotrophic lateral sclerosis. Mol. Brain 2013, 6, 31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kabashi, E.; Valdmanis, P.N.; Dion, P.; Spiegelman, D.; McConkey, B.J.; Vande Velde, C.; Bouchard, J.P.; Lacomblez, L.; Pochigaeva, K.; Salachas, F.; et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 2008, 40, 572–574. [Google Scholar] [CrossRef]
- Kwiatkowski, T.J., Jr.; Bosco, D.A.; Leclerc, A.L.; Tamrazian, E.; Vanderburg, C.R.; Russ, C.; Davis, A.; Gilchrist, J.; Kasarskis, E.J.; Munsat, T.; et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 2009, 323, 1205–1208. [Google Scholar] [CrossRef]
- Chesselet, M.F. Dopamine and Parkinson’s disease: Is the killer in the house? Mol. Psychiatry 2003, 8, 369–370. [Google Scholar] [CrossRef]
- Pickrell, A.M.; Youle, R.J. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 2015, 85, 257–273. [Google Scholar] [CrossRef]
- Yan, W.; Chen, Z.Y.; Chen, J.Q.; Chen, H.M. LncRNA NEAT1 promotes autophagy in MPTP-induced Parkinson’s disease through stabilizing PINK1 protein. Biochem. Biophys. Res. Commun. 2018, 496, 1019–1024. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lu, Z. Long non-coding RNA NEAT1 mediates the toxic of Parkinson’s disease induced by MPTP/MPP+ via regulation of gene expression. Clin. Exp. Pharmacol. Physiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xu, Y.; Zhu, Y.C.; Wang, Y.K.; Li, J.; Li, X.Y.; Ji, T.; Bai, S.J. LncRNA NEAT1 promotes extracellular matrix accumulation and epithelial-to-mesenchymal transition by targeting miR-27b-3p and ZEB1 in diabetic nephropathy. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xia, J.W.; Ke, Z.P.; Zhang, B.H. Blockade of NEAT1 represses inflammation response and lipid uptake via modulating miR-342-3p in human macrophages THP-1 cells. J. Cell. Physiol. 2019, 234, 5319–5326. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.D.; Hui, L.L.; Zhang, X.C.; Chang, Q. NEAT1 contributes to ox-LDL-induced inflammation and oxidative stress in macrophages through inhibiting miR-128. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef] [PubMed]

| NEAT1 Expression | Viral Disease | Pro-viral or Anti-viral | Pathway | Literature |
|---|---|---|---|---|
| Upregulation | HIV-1 | anti-viral | Rev-dependent nuclear export | [83,84] |
| Mild dengue | [85] | |||
| Herpes simplex | pro-viral | P54nrb, PSPC1 | [91,93] | |
| Hantavirus | anti-viral | RIG-I-signaling | [88,89] | |
| Hepatitis D | anti-viral | IL-8 induction | [90] | |
| Influenza | anti-viral | IL-8 induction by SFPQ inhibition | [91] | |
| Down-regulation | Severe dengue | anti-viral | p53 induced apoptosis | [85] |
| NEAT1 Expression | Neurodegenerative Disease | Co-players | Literature |
|---|---|---|---|
| Upregulation | Huntington’s Disease | REST, p53 | [109,110] |
| Multiple Sclerosis | IL-8, stathmin & TLR3 | [112,113,114] | |
| Amyotrophic Lateral Sclerosis | TDP-43, FUS/TLS | [116] | |
| Parkinson’s Disease | α-synuclein | [121,122] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prinz, F.; Kapeller, A.; Pichler, M.; Klec, C. The Implications of the Long Non-Coding RNA NEAT1 in Non-Cancerous Diseases. Int. J. Mol. Sci. 2019, 20, 627. https://doi.org/10.3390/ijms20030627
Prinz F, Kapeller A, Pichler M, Klec C. The Implications of the Long Non-Coding RNA NEAT1 in Non-Cancerous Diseases. International Journal of Molecular Sciences. 2019; 20(3):627. https://doi.org/10.3390/ijms20030627
Chicago/Turabian StylePrinz, Felix, Anita Kapeller, Martin Pichler, and Christiane Klec. 2019. "The Implications of the Long Non-Coding RNA NEAT1 in Non-Cancerous Diseases" International Journal of Molecular Sciences 20, no. 3: 627. https://doi.org/10.3390/ijms20030627






