Abstract
We investigated whether likely pathogenic variants co-segregating with gastroschisis through a family-based approach using bioinformatic analyses were implicated in body wall closure. Gene Ontology (GO)/Panther functional enrichment and protein-protein interaction analysis by String identified several biological networks of highly connected genes in UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, AOX1, NOTCH1, HIST1H2BB, RPS3, THBS1, ADCY9, and FGFR4. SVS–PhoRank identified a dominant model in OR10G4 (also as heterozygous de novo), ITIH3, PLEKHG4B, SLC9A3, ITGA2, AOX1, and ALPP, including a recessive model in UGT1A7, UGT1A6, PER2, PTPRD, and UGT1A3. A heterozygous compound model was observed in CDYL, KDM5A, RASGRP1, MYBPC2, PDE4DIP, F5, OBSCN, and UGT1A. These genes were implicated in pathogenetic pathways involving the following GO related categories: xenobiotic, regulation of metabolic process, regulation of cell adhesion, regulation of gene expression, inflammatory response, regulation of vascular development, keratinization, left-right symmetry, epigenetic, ubiquitination, and regulation of protein synthesis. Multiple background modifiers interacting with disease-relevant pathways may regulate gastroschisis susceptibility. Based in our findings and considering the plausibility of the biological pattern of mechanisms and gene network modeling, we suggest that the gastroschisis developmental process may be the consequence of several well-orchestrated biological and molecular mechanisms which could be interacting with gastroschisis predispositions within the first ten weeks of development.
1. Introduction
Gastroschisis constitutes one the leading categories of human birth defects concerning the ventral body wall development [1]. It is characterized by the evisceration of abdominal organs, usually to the right of the umbilical ring, lacking a protective membrane. Emerging evidence suggests that gastroschisis (an epidemiologic and pathogenetic dilemma) could be the consequence of a complex interplay of functionally interacting genetic and non-genetic factors as precipitating elements in its pathogenesis [2,3,4]. These influences have shown to be involved in several crucial biological processes such as blood vessel and epithelium development, cell adhesion, regulation of cytokine biosynthetic process, and regulation of developmental gene expression [2,3,4].
Studying biological processes and pathways through bioinformatic analysis in gastroschisis may have an important clinical relevance. Pregnancies affected with fetal gastroschisis can have multiple complications such as low fetal weight, in utero fetal demise, or preterm delivery [1]. Since this defect develops early in gestation, it is possible that alterations in the regulation of embryonic or fetal vascular, epithelium, immune development, or gene expression may play a role in these related problems [4]. Moreover, it may provide plausible clues to the closure of the ventral body wall.
Given the current paucity of genes with causal implication in human gastroschisis, we leveraged on genes and DNA variations co-segregating with gastroschisis through a family-based approach identified by whole exome sequencing (WES) [4], and investigated whether likely pathogenic variants were implicated in body wall closure through bioinformatic analysis to explore potential pathogenetic pathways with this disease.
2. Results
A total of 429 genes involving heterozygous DNA variations co-segregating with gastroschisis have been identified and selected from the WES gene list for further bioinformatic analyses [4].
2.1. Gene Functional Enrichment Analysis
Gene Ontology (GO) Consortium and Panther Classification System databases [5] identified several highly enriched GO terms (False Discovery Rates, FDR) within genes co-segregating in the family with recurrence for gastroschisis (Table 1).

Table 1.
Overrepresentation of enriched Gene Ontology (GO) terms within genes co-segregating with gastroschisis.
Functional biological processes related to xenobiotic glucuronidation, negative regulation of glucuronosyltransferase activity, and flavonoid glucuronidation were the most significantly enriched terms (Subfamily UGT1A involving UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, P FDR 7.99 × 10−11, 1.42 × 10−9, and 2.26 × 10−7, respectively).
Cellular component GO analysis also revealed the integral component of membrane as the most significant functional category (UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, AOX1, NAA30, OMA1, DPYD, PDIA2, ALPP, CDYL, ADPGK, KDM5A, PRDM9, ZFHX3, BCAS3, NOTCH1, TELO2, PLEKHG4B, SPATA17, LRP5, AXIN1, PKD1, SLC9A3, GAL3ST3, RASGRP1, AKAP6, HIST1H2BB, EIF3L, RPS3, RPS6, MRPS6, INPP5B, IRS1, FGFR4, FGFRL1, EFNA5, ATP13A5, TBX21, TLR8, ADCY9, GRM4, GRIN2C, OR13F1, OR10G4, OR4C3, OR2C1, F5, THBS1, MAP2K3, RTN3, EVPL, KLK14, ITGA7, ITGA2B, ITGA2, ITGAD, CTCFL, and RABL6, P FDR 4.95 × 10−3).
The molecular function resulted mainly enriched in glucuronosyltransferase activity (UGT1A, P FDR 1.58 × 10−6), whereas KEGG signaling pathway analysis identified drug metabolism—other enzymes and pentose and glucuronate interconversions, as the most significant functional categories (UGT1A and AOX1, P FDR 1.32 × 10−8 and 2.83 × 10−8, respectively).
2.2. Protein-Protein Interactions Network Analysis
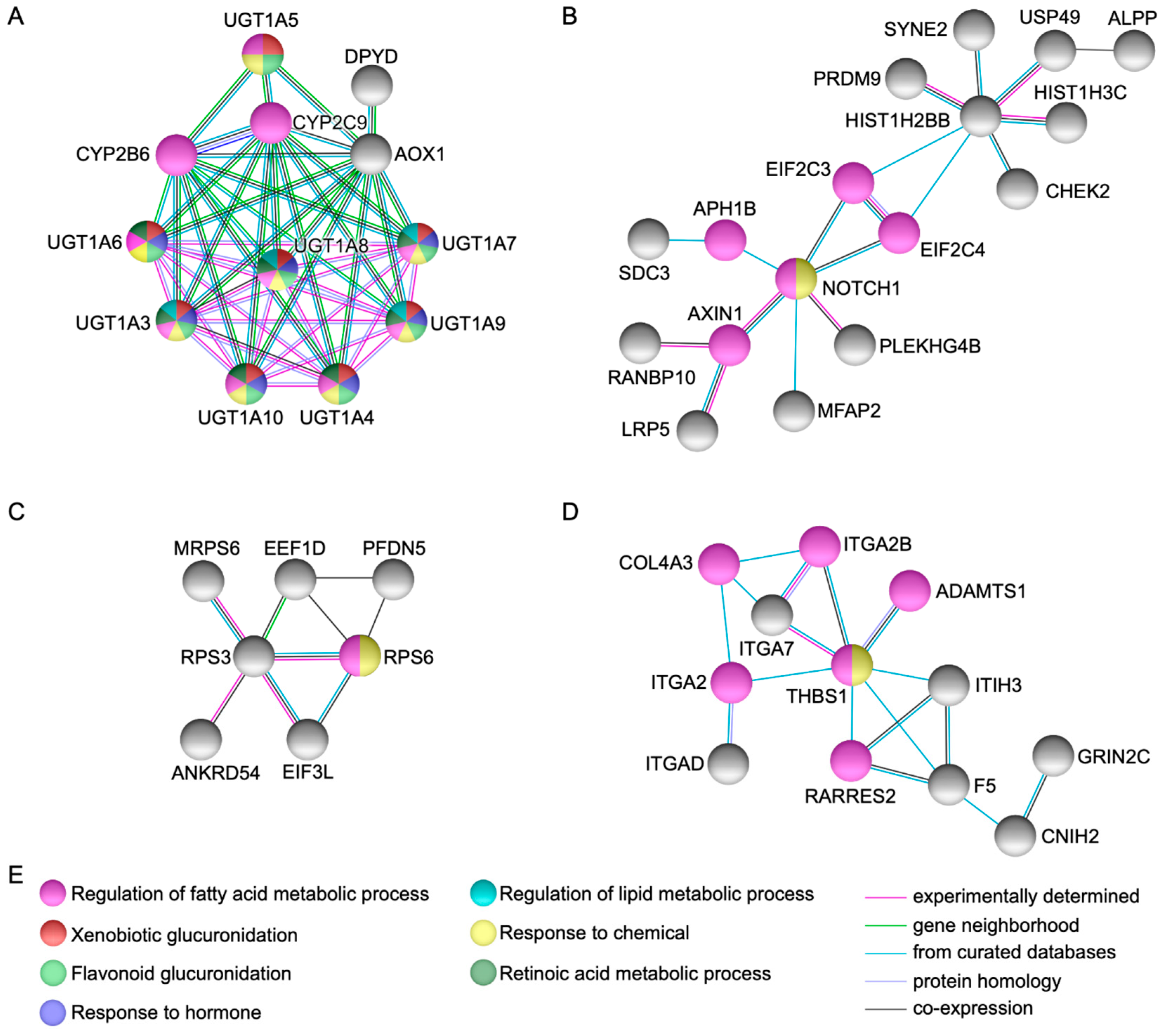
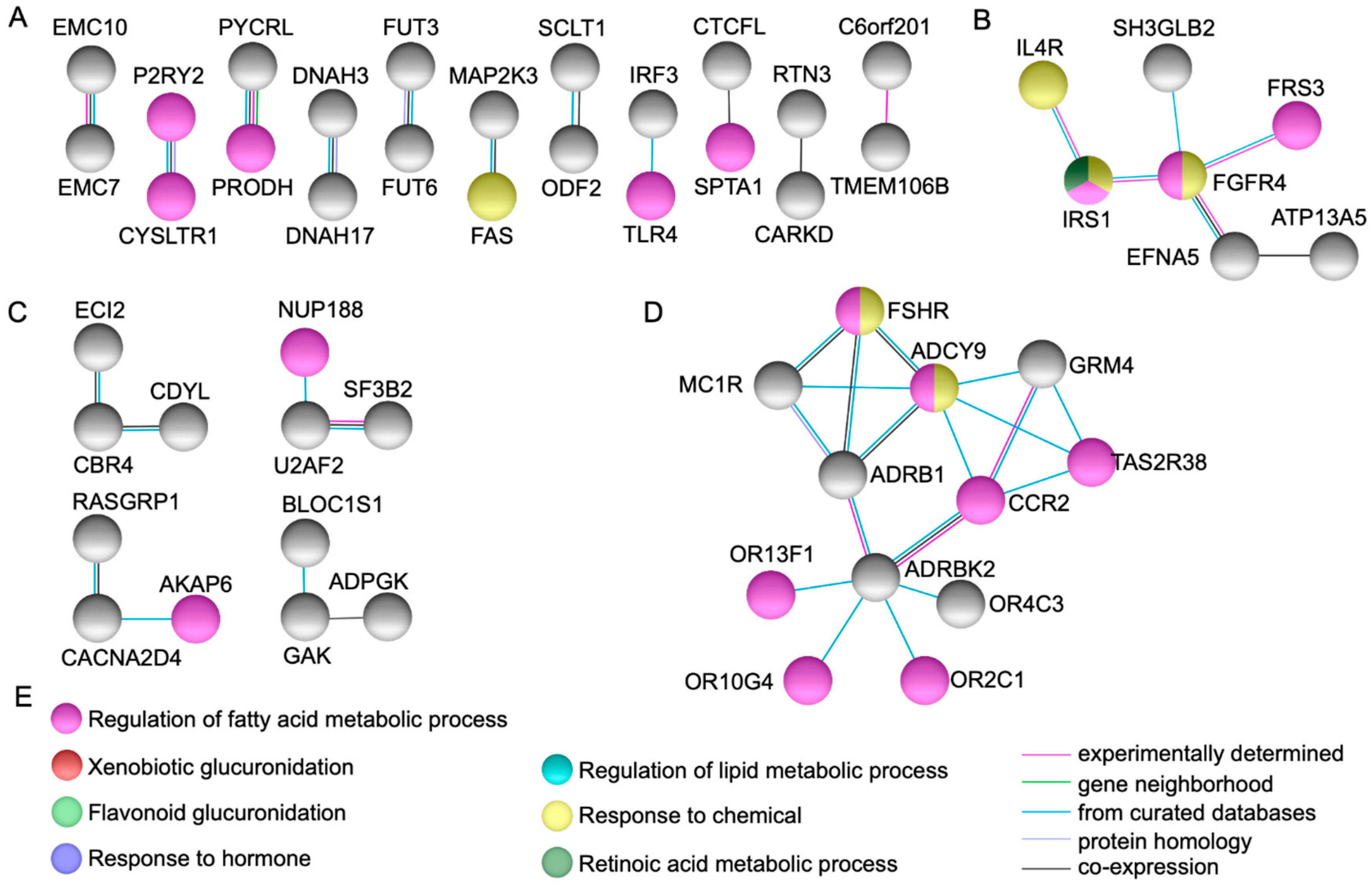
String protein-protein interaction analysis [6] identified fifteen genes represented by UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, AOX1, NOTCH1, HIST1H2BB, RPS3, THBS1, ADCY9, and FGFR4, that showed high connectivity to each other and to other genes in the network via several partners (Figure 1 and Figure 2).
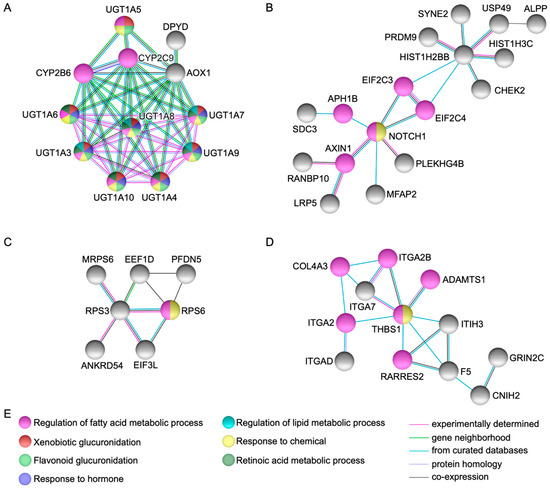
Figure 1.
Network modeling from protein-protein interactions. The figures (A)–(D) show high connectivity and direct protein-protein interactions via several partners in genes UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, AOX1, NOTCH1, HIST1H2BB, RPS3, and THBS1. Figure (E) depicts the biological processes identified in the networks.
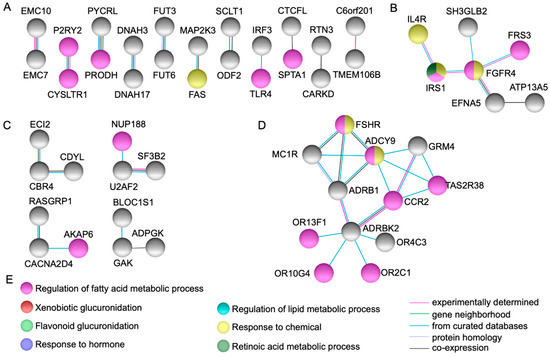
Figure 2.
Network modeling from protein-protein interactions. Figures (A) and (C) show several direct protein-protein dyads and triads. Figures (B) and (D) show high connectivity and direct protein-protein interactions via several partners in genes ADCY9 and FGFR4. Figure (E) depicts the biological processes identified in the networks.
Several protein-protein dyads and triads were observed as depicted in Figure 2. Of note, FGFR4 (p.G388R) (rs351855) was a coding non-synonymous variant identified in the index case and both parents [4].
2.3. Classification by Inheritance Pattern
SVS–PhoRank gene ranking [7] classified the inheritance pattern for all variants (Tables S1–S3). A dominant model score was noted for the following variants: OR10G4 (rs4474449, also as heterozygous de novo variant), ITIH3 (rs2286797), PLEKHG4B (rs114885385), SLC9A3 (rs117027344), ITGA2 (rs1062535), AOX1 (rs3731722), and ALPP (rs13026692). Furthermore, UGT1A7 (rs11692021), UGT1A6 (rs6759892, rs2070959, rs1105879), PER2 (rs934945), PTPRD (rs3824417), and UGT1A3 (rs6431625) were identified within a recessive model score as well as homozygous both.
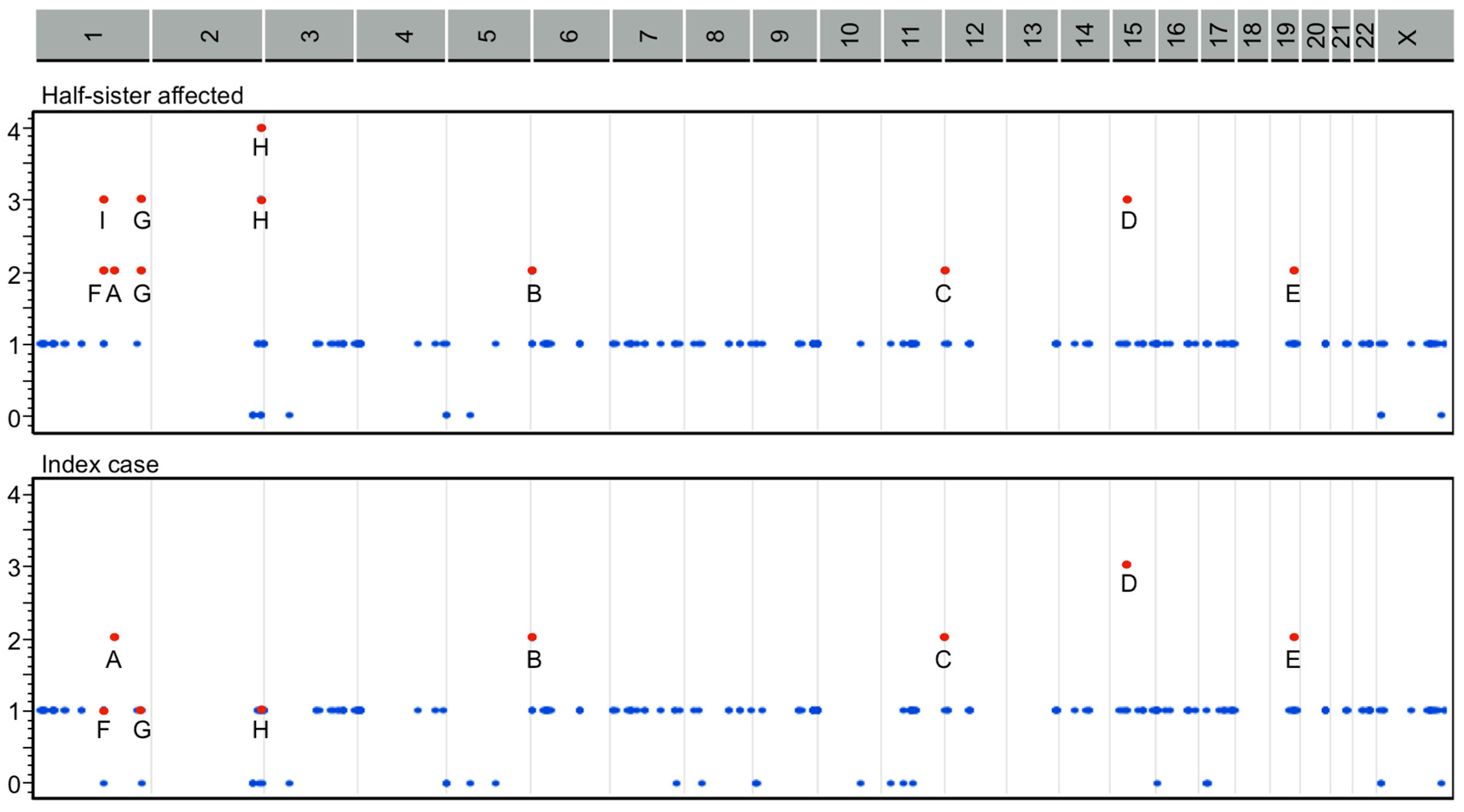
A heterozygous compound model score was observed in the following variants among the mother and both cases: CDYL (rs11965336 and rs11962921), KDM5A (rs148769146 and rs144276601), RASGRP1 (rs56241040, rs55704435, and rs56366451), MYBPC2 (rs75197332 and rs187765595), PDE4DIP (rs61804988, rs144590426, and rs2762779), F5 (rs6027 and rs1800595), OBSCN (rs75280352, rs185523702, and rs72762068), and UGT1A (rs11692021, rs6759892, rs1105879 homozygous genotype in the index case, and rs6431625 homozygous genotype in the half-sister affected) (Figure 3).
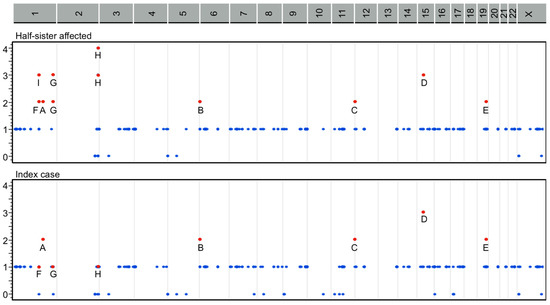
Figure 3.
Heterozygous compound model score regions from gene variants identified in F5 (A), CDYL (B), KDM5A (C), RASGRP1 (D), MYBPC2 (E), PDE4DIP (F), OBSCN (G), UGT1A (H), and 1:144146811–146467744 chromosomal region including PDE4DIP (I) among both cases and the mother (significant heterozygous regions as red points). Vertical scales list the number of heterozygous genotypes inherited from the mother. Horizontal scales depict the chromosomal region involved.
2.4. Pathogenetic Pathways from Selected Genes Co-Segregating with Gastroschisis
According to ToppGene Suite [8], eleven pathogenetic pathways were identified based on manually curated GO categories among selected genes co-segregating in the family with recurrence for gastroschisis (Table 2).

Table 2.
Pathogenetic pathways from GO functional categories among selected genes co-segregating with gastroschisis *.
There was no data available within GO functional categories for SPATA17 and CFAP65. Table S4 includes the complete result list of GO biological processes and pathways.
3. Discussion
In the present study, we employed bioinformatic analyses using genes and genetic variations co-segregating with gastroschisis through a family-based approach identified by WES as a model system, to explore potential pathways implicated in body wall closure. Our findings demonstrate that several functional GO categories were displaying plausibility (Table 1, Figure 1 and Figure 2), whereas novel gene variants within dominant, recessive, and heterozygous compound models (Tables S1–S3, Figure 3) were possibly underlying to gastroschisis as well. We also characterized multiple novel pathogenetic pathways that may have meaningful consequences for the closure of the ventral body wall in gastroschisis (Table 2 and Table S4).
A limitation of the study was that a different filter from bioinformatics tools could prioritize some gene variants and hence, cause their inclusion or exclusion. Thus, findings from bioinformatic algorithms can only be taken as suggestive evidence in the absence of functional validation. However, it must be noted that our results were validated from independent bioinformatics platforms, giving the basis for further validation and reproducibility as well as to obtain more information about the molecular mechanisms involved in gastroschisis.
3.1. Novel Candidate Genes and Pathogenetic Pathways
Several genes could be potential candidates for future studies as those observed with a highly significant functional enrichment, high connectivity/direct protein-protein interactions, and highest score for dominant, recessive, and heterozygous compound models (UGT1A3, UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT1A10, AOX1, NOTCH1, HIST1H2BB, RPS3, THBS1, ADCY9, FGFR4, OR10G4, ITIH3, PLEKHG4B, SLC9A3, ITGA2, ALPP, PER2, PTPRD, CDYL, KDM5A, RASGRP1, MYBPC2, PDE4DIP, F5, and OBSCN) (Table 1 and Tables S1–S3, Figure 1, Figure 2 and Figure 3).
High protein-protein interactions as well as functional enrichment were observed in xenobiotic genes UDP-glucuronosyltransferases (UGT1A at 2q37.1, heterozygous compound model) in UGT1A3 (recessive model and moderate impact), UGT1A4 and UGT1A5 (modifier impact), UGT1A6 (recessive model, moderate and modifier impact), UGT1A7 (recessive model and moderate impact), UGT1A8, UGT1A9, and UGT1A10 (modifier impact). UDPGTs bind to enzyme, carbohydrate, and protein conjugation with subsequent deactivation and elimination of potentially toxic xenobiotics (e.g., drugs, environmental pollutants, dietary chemicals) and endogenous compounds (e.g., bilirubin, steroids, bile acids). Moreover, CYP2C8 along with CYP2C9 and CYP2B6 via cytochrome P450-mediated oxidation may undergo glucuronidation activities (Figure 1A) [9,10]. UDPGTs have been associated with a variety of bilirubin metabolic disorders and their mouse orthologs Ugt1a1, Ugt1a2, Ugt1a5, Ugt1a6b, Ugt1a7c, Ugt1a8, Ugt1a9, Ugt1a10 are mainly expressed in the liver at days 15.5 and 18.5 (Carnegie stages); corresponding to the 6th and 7th week of development in humans (WD, weeks from fertilization) [9,11,12].
AOX1 (Aldehyde Oxidase 1 at 2q33.1, dominant model, moderate and modifier impact), was connected directly and via partners with UGT1A, CYP2C9, CYP2B6, and DPYD; the latter, (moderate and modifier impact) is related to pyrimidine metabolism and nucleotide synthesis. AOX1 is involved in the regulation of reactive oxygen species homeostasis and has been associated to hereditary xanthinuria [9]. The mouse ortholog Aox1 was found expressed in the liver at day 15.5 (6th WD) [12].
Further protein-protein interactions were noted in NOTCH1 (Notch 1 at 9q34.3, modifier impact), PLEKHG4B (Pleckstrin Homology and RhoGEF Domain Containing G4B at 5p15.33, dominant model and modifier impact), and HIST1H2BB (Histone Cluster 1 H2B Family Member B at 6p22.2, modifier impact) (Figure 1B). GO molecular annotations for these genes implies gene expression, Rho guanyl-nucleotide exchange factor activity, core promoter binding, as well as chromatin, DNA, and protein binding [9,11]. Noticeably, Notch1 can be expressed both in trophoblast and embryonic stem cells from preimplantation through all stages of development [9,13], whereas Hist1h2bb was found expressed in the liver and brain between days 15.5 and 18.5 (6th and 7th WD) [12]. Of interest, KDM5A (Lysine Demethylase 5A at 12p13.33) and CDYL (Chromodomain Y Like at 6p25.1) both as heterozygous compound model and modifier impact genes, were closely interacting to this particular network. These genes play a central role in histone code, regulating specific gene transcription and repression of essential biological processes preserving the epigenetic landscape [9]. The mouse orthologs Kdm5a and Cdyl were found expressed in the cell stage embryo, blastomere, and blastocyst (days 1.5 to 4.5, 1st WD) [14,15] as well as in the liver and brain at days 15.5 and 18.5 (6th and 7th WD) [12].
RPS3 (Ribosomal Protein S3 at 11q13.4, modifier impact) was detected interacting with multiple genes implicated in the regulation of protein synthesis (Figure 1C). This gene plays a key role in the protection of DNA, binding with similar affinity to intact and damaged DNA, and when located in the mitochondrion, reduces cellular reactive oxygen species levels and mitochondrial DNA damage. Moreover, it binds and protects TP53/p53 from MDM2-mediated ubiquitination and is involved in the induction of apoptosis through its role in the activation of CASP8 [9]. Rps3 was detected in the liver and brain at day 15.5 (6th WD) [12].
Additional direct and via partners protein-protein interactions were observed in THBS1 (Thrombospondin 1 at 15q14, moderate and modifier impact), ITGA2 (Integrin Subunit Alpha 2 at 5q11.2, dominant model and modifier impact), ITIH3 (Inter-Alpha-Trypsin Inhibitor Heavy Chain 3 at 3p21.1, dominant model and modifier impact), and F5 (Coagulation Factor V at 1q24.2, heterozygous compound model, moderate and modifier impact) (Figure 1D). These genes have been associated with a variety of thrombotic abnormalities, as they play a central role in hemostasis and cell adhesion, mediating and stabilizing the adhesion of platelets (including other cell types such as collagen) to the extracellular matrix [9]. Expression analysis of mouse Thbs1 was identified in the yolk sac and embryo mesenchyme at days 9.5 and 14.5 (4th and 5th WD), respectively [16,17], whereas Itga2 was found expressed in the embryo, epidermis, blood vessel endothelium, and epidermis stratum basale at days 7.5, 16.5, and 18.5 (3rd, 6th, and 7th WD), respectively [18,19]. Itih3 was found expressed in the liver at day 15.5 (6th WD) [12] and F5 was detected in the embryo and notochord at days 7.5 to 14.5 (3rd to 5th WD) and 8.0 to 8.5 (3rd WD), respectively [20,21].
ADCY9 (Adenylate Cyclase 9 at 16p13.3, modifier impact) was noted to interact with multiple genes implicated in G protein-coupled receptor activities, including catecholamine-induced activation, chemotaxis and migration, follitropin, melanogenesis, and signal transduction of odorant and taste molecules (Figure 2D). This gene contributes to signaling cascades activated by the corticotropin-releasing factor, corticosteroids, and beta-adrenergic receptors, implicated in the production of platelet-derived and vascular endothelial growth factors [9]. Murine Adcy9 was detected in the cardiovascular system, axial skeleton, skin, and hair at day 14.5 (5th WD) [22].
FGFR4 (Fibroblast Growth Factor Receptor 4 at 5q35.2) was observed interacting with several genes involved in the regulation of tyrosine kinases activities and cell surface receptors for fibroblast growth factors (Figure 2B). These genes are crucial for diverse developmental, metabolic, immune, and cancer processes, as they regulate cell proliferation, differentiation, and migration, including the regulation of lipid metabolism, bile acid biosynthesis, glucose uptake, vitamin D metabolism, and phosphate homeostasis [9]. Murine Fgfr4 was detected in the cell stage embryo and blastocyst at days 2 and 3.5, respectively (1st WD) [15,23], as well as in the neural ectoderm, embryo mesenchyme, extra embryonic ectoderm at day 7.5 (3rd WD) and the somite, yolk sac endoderm, and endoderm of the developing gut from days 8.5 to 14.5 (3rd to 5th WD) [24,25]. Moreover, it became restricted at low levels of expression to the liver and the central nervous system from day 11.5 (4th WD) onward [23].
Bioinformatic classification by inheritance pattern also identified PER2 (Period Circadian Regulator 2 at 2q37.3, recessive model and moderate impact). This gene regulates circadian rhythms in gene expression, which are translated into rhythms in metabolism and behavior, including a wide array of physiological functions such as sleep, body temperature, blood pressure, endocrine, immune, cardiovascular, and renal functions [9]. Per2 was identified in the future spinal cord and central nervous system at days 11.5 and 13.5 (4th WD) [26,27], the blood vessel and cardiovascular system at day 14.5 (5th WD) [22], and the liver at day 15.5 (6th WD) [12].
SLC9A3 (Solute Carrier family 9 member A3 at 5p15.33, dominant model and modifier impact), is an epithelial brush border Na/H exchanger that eliminates acid-toxicity generated by active metabolism or counters adverse environmental conditions [9]. The mouse ortholog Slc9a3 was found expressed from the cell stage embryo to trophectoderm (days 1.0 to 3.5, 1st WD) [28,29] as well as in the metanephros at day 18.5 (7th WD) [30].
RASGRP1 (RAS Guanyl Releasing Protein 1 at 15q14, heterozygous compound model and modifier impact) and PTPRD (Protein Tyrosine Phosphatase Receptor type D at 9p24.1-p23, recessive model and modifier impact) are crucial signaling molecules critical for a variety of cellular processes including cell growth, differentiation, mitotic cycle, and oncogenic transformation, by regulating T-cell/B-cell development, NK cell cytotoxicity, and ITAM-dependent cytokine production by activation of Ras-mediated ERK/MAPK and JNK pathways [9]. Mouse orthologs Rasgrp1 and Ptprd were detected in the olfactory lobe/epithelium at day 14.5 (5th WD) as well as in the liver and central nervous system at days 15.5 and 16.5 (6th WD) [12,17,31].
MYBPC2 (Myosin Binding Protein C, Fast Type at 19q13.33, heterozygous compound model and modifier impact) and OBSCN (Obscurin at 1q42.13, heterozygous compound model, moderate and modifier impact) are key signaling proteins that play a role in the regulation of myofibrillogenesis and modulation of muscle contraction, as they bind and modify the assembly of actin-activated myosin ATPase into sarcomeric A bands in striated muscle [9]. Murine Mybpc2 was found expressed in the hindlimb at day 18 (7th WD) [32], whereas Obscn was detected in the diaphragm and cardiomyocytes at days 14.5 and 17 (5th and 6th WD), respectively [17,33].
Overall, these interacting genes (together with previous high impact variants and gene variants segregating among both cases and the mother) [4], demonstrated significant representation of xenobiotic, regulation of metabolic processes, regulation of cell adhesion, regulation of gene expression, inflammatory response, regulation of vascular development, keratinization, left-right symmetry, epigenetic, ubiquitination, and regulation of protein synthesis related GO categories (Table 2 and Table S4). Thus, based on our results, multiple novel pathogenetic pathways could be implicated in gastroschisis pathogenesis. Indeed, our findings point to a complex interplay of functionally interacting genetic and non-genetic factors; thereby, we propose that each of these interacting genes are likely substantial influences to its development, underscoring major gaps concerning the current knowledge of genetics and developmental biology of the defect.
3.2. Theories on the Failure of Body Wall Closure in Human and its Implication for Gastroschisis
As enlisted in Table S2, a considerable number of GO categories evoking related pathogenetic theories in gastroschisis were identified [34,35,36,37,38,39,40,41,42,43]. These hypotheses along with our proposed “multigenic/multifactorial model” [4], highlight novel susceptibility gene variants within pathogenetic pathways that could be possibly implicated in the pathogenesis of gastroschisis (Table 2).
Perhaps the chief explanation for an experimental scarcity regarding the biological or molecular mechanism of human ventral body wall development, particularly in gastroschisis, is linked to the assumption that such defect does not have a genetic basis. Recently, we published research suggesting that this phenotype may result from complex gene-gene or gene-environment interactions [1,2,3,4,44,45]. Accordingly, it is possible that several of the above pathogenetic pathways do indeed lead to a disruptive closure of the ventral body wall, as several well-orchestrated biological and molecular mechanisms could be interacting with gastroschisis genetic predisposition within the first ten weeks of human development.
Previous reports from ortholog expression tissues assays involving the above-described genes indicated that reduced but significant gene predispositions (as those implicated in the regulation of gene expression) are expressed in cell-embryo and implantation stages [13,14,15,22,23,28,29], whereas several genetic influences implicated in the regulation of metabolic processes, regulation of vascular development, regulation of protein synthesis, and ubiquitination may lead to a disruptive closure of the ventral body wall in a fast-growing or massive fashion presumably between the 3rd and 5th WD [13,16,17,18,20,21,22,23,24,25,26,27]. Notably, multiple factors and gene interactions (background modifiers) can aggravate or mask the expected phenotype through gain- or loss-of-function mutations in genes or gene pathways involved in related disease-relevant cells [46,47,48], perhaps giving an isolated anomaly. In these stages, folding of the embryo due to a rapid increase in the proliferation of the neuroectoderm and the underlying mesoderm occurs, giving rise to the four folds of the abdominal ventral body wall as well as the thoracic, sternum, diaphragm, and cloacal membrane. These morphogenetic events are determined due to growth arrest caused by cell death at the umbilical ring [41,49,50,51].
On the other hand, predominant and significant gene predispositions (as those implicated in xenobiotic, regulation of metabolic processes, regulation of cell adhesion, inflammatory response, keratinization, and epigenetic) may lead to a disruptive closure of the ventral body wall as an enlarged umbilical ring presumably between 6th and 10th WD [12,13,19,23,30,31,32,33]. Between these stages, the umbilical ring emerges from the four body folds and is located in the center of the abdomen (ventral surface) performing proper transition between the body wall and the amnion, whereas the latter remains in continuity with the periderm throughout the embryonic period. During the 10th and 11th WD, the epithelial tissues from either side of the embryo meet and fuse at the midline. Concurrently, an intermediate layer rises from the stratum basale/germinativum including collagenous and elastic fibers (periderm keratinizes and desquamates); whereas cell death and cell deposition prevents the growth of the umbilical ring maintaining approximately the same diameter throughout development [41,49,50,51,52].
Our results contribute to the growing evidence linking gastroschisis to the embryonic dysgenesis theory [3,41,49,50,53]. Aberration on the ectodermal placodes leads to disturbances in the cell deposition process and hence, the mesoderm of the ventral body wall could be underdeveloped resulting in an enlarged umbilical ring (mesodermal deficiency) [41,49,50]. An ectodermal placode expressly at the umbilical ring involves pivotal signaling either to proper transition between the amnion and the umbilical ring or to cell-cell communication from the ectoderm to the mesoderm, depositing mesoectodermal cells contributing to the lateral and ventral body wall of the embryo. Moreover, due to a combined mechanism of apoptosis and cell proliferation of the underlying ectodermal cells, this cell deposition process can be added into the mesodermal compartment that lies below, including tissues associated with ventral body wall closure [41,49,50,51]. Although significant differences take place featuring the human and mouse embryology [51], these molecular mechanisms have been successfully elucidated in rat embryos cultured in vivo and AP-2α knockout mouse [54,55].
Taken together, our findings favors the recently proposed “mediolateral growth in the dorsolateral body wall” as well as the “differential growth” models [56,57]. According to these models, cell migration is unnecessary for the development of new structures, as a robust increase in body size of the experimented embryos contrasted with minimal changes in size of the umbilical ring and diastasis of the rectus muscles observed from the 6th to 10th WD. These observations harbor the hypothesis that though bones and ventral body wall muscles may have undergone normal differentiation, ventral midline defects could originate from an insufficient dorsoventral growth [56]. In this sense, the development of the ventral body wall is consistent with a variety of orchestrated interactions involving a balanced adhesome phenomena for which conserved cell-matrix adhesions in vertebrate muscle morphogenesis are paramount [58,59]. Although the type of defect would depend on the degree and location of the insufficiency, such a period is consistent with the absence of associated anomalies [3,38,53,60]. Furthermore, it must be noted that previous observations in gastroschisis support failure in normal attachment between the umbilical cord and umbilical ring [37,38,53], as well as a secondary rupture of the umbilical cord following normal attachment [60].
Alternatively, the classical “lateral somitic frontier” model hypothesizes that somitic muscle cells are patterned independently migrating or translocating into the lateral plate area, as migration of ribs and muscles through the body wall occurs in a ventral direction [61]. This model recalls other theories linked to the disruption of the basic organization of embryonic development, adverse/teratogenic embryonic environment provided by young mothers, and high concurrence risk from intrauterine vascular disruption between the 3rd to 5th WD [34,35,36,39,40,42,43,62,63]. Embryonic disruption as well as poor timing of vascular transition involves essential processes such as angiogenesis, apoptosis, and cell migration. From our perspective, this biological scenario might be demonstrative for those cases displaying a constellation of co-occurring anomalies [3,64,65], or perhaps fetal demise, particularly when associated with maternal toxic exposures, given that more than one disruptive event may occur, even after the embryonic period [66]. Of note, cell deposition may also be massively reduced between the abdominal wall and the amnion, which could lead to an extensive defect of the abdominal wall along with detrimental consequences to the abdominal organs and cord [49].
3.3. Gastroschisis Clinical Susceptibility
In our view, all possible genetic and non-genetic factors including their direct regulators contributing to the gastroschisis complex landscape should be considered [4]. Thus, considering our results and possible variable disease presentation [3], several complex mechanisms such as epistatic mutational effect, synergistic heterozygosity or multigenic inheritance, variants in modifier genes, or effects due to overall rare variant load, may cause enhancement or repression of the phenotype [48]. Additionally, defects in pathways causing cumulative effects could lead to susceptibility or phenotypic variability (severe/unusual or milder/isolated presentation) [46,47].
Accordingly, the reason for a prevailing association of gastroschisis with young maternal age (perhaps along with low pre-pregnancy body mass index or environmental noxious influences), could be orchestrated by genes involved in several well-synchronized biological and molecular mechanisms of xenobiotic and regulation of metabolic processes. These genes may contribute towards gastroschisis risk due to an endo- or xenobiotic, flavonoid, fatty acid, drug, hormone, carbohydrate, chemical, or lipid metabolism-related toxicity. Furthermore, they also regulate the response to growth factors, detection and response to stimulus, folate biosynthesis, as well as a negative co-regulation of skeletal muscle, fibroblasts, and subcutaneous adipose tissue [9,11]; notwithstanding that epigenetic-related influences could also be interacting with a particular plethora of non-genetic risk factors such as parental alcohol intake and cigarette smoking. This appears to be the case for an increasing proportion of affected fetuses from young mothers, which may be well considered as a vulnerable population associated with a limited optimal pre/peri-conceptional care, as well as estrogen-related disruptors (decreased maternal age, nulliparity, decreased body mass, alcohol intake) [1,43].
Given the above, it follows that genes involved in the inflammatory response may play a key role for maternal immunological factors and possibly be responsible for susceptibility to genitourinary infections and the association of a change in sex partner with gastroschisis [44,45,67]. Altogether with the genes involved in the regulation of gene expression, epigenetic, regulation of protein synthesis, and ubiquitination, fetal immune development regulating genes may also play a central role in preterm birth and low fetal weight. In this context, patients with gastroschisis have shown significant high systemic levels of inflammatory cytokines and chemokines, including an earlier activation of CD4+ and CD8+ effector and memory T cells [68].
The mechanism/cause of prenatal death in gastroschisis is controversial and we do not offer any explanation for this related complication. Morphological studies provide support for a possible effect involving the umbilical cord compression due to extra-abdominal bowel herniation leading to hypoxia and eventually to fetal death [53,69]. Cytokine-mediated inflammatory response has been equally incriminated [70]. Further studies investigating vascular or cell adhesion phenomena as well as chronic inflammation/immune response could be valuable for understanding this detrimental consequence.
CFAP65, CROCC, NOTCH1, AXIN1, LRP5, HHIP, PKD1, NR1H3, and IFT140 were identified to possibly influence the left-right symmetry of gastroschisis cases [4]. These genes may reflect an asymmetric arrest of vascular supply (3rd to 5th WD) [13,14,17,22,27,71,72,73,74,75,76], asymmetric cell deposition at the umbilical ring (6th to 10th WD) [53,74,75,76], or amniotic rupture along the umbilical cord in its pars flaccida (8th to 11th WD) [19,60,75], similar to the suggested weak point left after involution of the right umbilical vein between the 5th and 10th WD [37,39]. Interestingly, an ectopic expression of Pitx2 in the right side of experimental embryos affected the left-right asymmetry of the heart, gut, and vitelline vein, resulting in phenotypes similar to those associated with Shh and nodal misexpression at days 10 to 16 (4th to 6th WD) [77]. These genes may also serve as critical transcription targets that mediates left-right asymmetry as well as left-to-rightward shunts between the left umbilical vein, the right hepatocardiac channel, and the right vitelline vein [78]. Furthermore, differential left-right expression of genes during embryonic and organogenesis stages may impact cell proliferation, cell-cell communication, and other morphogenetic processes of specific laterality development; particularly when an unfavorable/teratogenic embryonic environment or adverse genetic influences are involved [79].
Finally, a protective or risk factor owing to race/ethnicity has also been associated with gastroschisis, which may differ based on maternal nativity [80,81,82]. The effect of Hispanic ethnicity in the present study may add to this particular association as heritable factors and genetic susceptibility among the Hispanic population may play a role in the pathogenesis of gastroschisis [2,3,44,45]. Yet, when considering the lacking genomic research on gastroschisis, including this association, the proportion of genetic variation that has been evaluated requires the validation of these findings among additional populations along with its interracial and cultural variation.
In conclusion, we identified multiple novel pathogenetic pathways implicated in the closure of the ventral body wall from genes co-segregating with gastroschisis. Based on our findings and considering the plausibility of the biological pathogenetic pattern of mechanisms and gene network modeling, we suggest that the gastroschisis developmental process may be the consequence of several well-orchestrated biological and molecular mechanisms, which could be interacting with gastroschisis predispositions within the first ten weeks of human development. These genes highlight a role for xenobiotic, regulation of metabolic processes, regulation of cell adhesion, regulation of gene expression, inflammatory response, regulation of vascular development, keratinization, left-right symmetry, epigenetic, ubiquitination, and regulation of protein synthesis in gastroschisis pathogenesis.
4. Materials and Methods
4.1. Study Participants and Identification of Gene Variants from WES
The present study was approved by the Institutional Ethics Committee from the School of Medicine and University Hospital “Dr. José Eleuterio González”, Universidad Autónoma de Nuevo León, México (Approval: 21 September 2017, GN17-00002). Written informed consent was obtained from the parents. A Mexican family (two affected half-sisters with gastroschisis, mother, and father of the proband) was assessed for WES. Screening, identification, and data analyses of genes and gene variants have been previously described [4].
4.2. Gene Functional Enrichment Analysis
A functional enrichment analysis was performed using the GO Consortium and Panther Classification System databases, which contain comprehensive information on the evolution and function of protein-coding genes from 104 completely sequenced genomes [5]. A P value of less than 0.05 was the selection criteria for significantly enriched biological processes and pathways which include a “hierarchical view” as a structure of the most significant classifications and ontologies of the human genes and E-value statistics (FDR).
4.3. Gene and Protein Network Analysis
A protein-protein interaction network including physical, functional, and biological processes associated with genes co-segregating with gastroschisis was created using String database with a confidence score of 0.4 [6]. A gene-gene pairwise network was constructed using the protein-protein interactions. Gene pairs detected in two or more of the protein-protein interactions including co-expression, protein homology, curated databases, gene neighborhood, or experimentally determined data sets were selected and included in the network analysis using String database 10.5 [6].
4.4. Classification of Gene Variants by Inheritance Pattern
SVS–PhoRank gene ranking (Golden Helix®, Inc., Bozeman, MT, USA) was used to classify the inheritance pattern for all variants, which ranks genes based on their relevance to user-specified phenotypes as defined by GO and Human Phenotype Ontology biomedical ontologies [7]. A sum of the standard score above 1.333 was considered for dominant and recessive models, whereas a heterozygous compound model score was considered from the detection of at least two heterozygous genotypes within the same gene inherited from the mother at a minimum of two different loci (recessive model). The classification also included the following models: maternal de novo, paternal de novo, heterozygous de novo, homozygous de novo, heterozygous either, heterozygous maternal, heterozygous paternal, and homozygous both.
4.5. Pathogenetic Pathways from GO Functional Categories
ToppGene Suite (Cincinnati, OH, USA) [8] was used to classify the pathogenetic pathways (manually curated) based on the identification of related GO functional categories (biological processes and pathways) from selected genes co-segregating with gastroschisis (e.g., high impact variants and gene variants segregating among both cases and the mother, as well as further genes resulting from our analyses) [4]. GO terms were selected based on their proximity and plausibility to the phenotype, including previous pathways associated to gastroschisis [2,3].
Supplementary Materials
Supplementary materials can be found at https://www.mdpi.com/1422-0067/20/9/2295/s1.
Author Contributions
Conceptualization, V.M.S.-T., and L.E.M.d.V.; Recruitment and examination of patients, V.M.S.-T., and R.A.S.-T.; Formal analysis and interpretation of data, V.M.S.-T., R.A.S.-T., J.J.L.-T., and H.L.G.-B.; Writing original draft, V.M.S.-T.; Review and editing, H.L.G.-B., R.A.S.-T., R.M.C.-F., D.Z.V.-M., and L.E.M.d.V.; All authors read and approved the final manuscript.
Funding
This research received no external funding.
Acknowledgments
The authors are grateful to the participating family for their valuable cooperation.
Conflicts of Interest
The authors declared no potential conflicts of interest and received no financial support concerning the research, authorship, or publication of this article.
Abbreviations
| ADCY9 | Adenylate cyclase 9 gene |
| ALPP | Alkaline phosphatase placental gene |
| AOX1 | Aldehyde oxidase gene |
| BCAS3 | Microtubule associated cell migration factor gene |
| BP | Biological process |
| CC | Cellular component |
| CDYL | Chromodomain Y like gene |
| CEACAM5 | Carcinoembryonic antigen related cell adhesion molecule gene |
| CFAP65 | Cilia and flagella associated protein 65 gene |
| COL6A3 | Collagen type VI alpha 3 chain gene |
| CROCC | Ciliary rootlet coiled-coil, rootletin gene |
| DPYD | Dihydropyrimidine dehydrogenase gene |
| EVPL | Envoplakin gene |
| F5 | Coagulation factor V gene |
| FDR | False discovery rate |
| FGFR4 | Fibroblast growth factor receptor 4 gene |
| FGFRL1 | Fibroblast growth factor receptor like 1 gene |
| GO | Gene ontology |
| HHIP | Hedgehog interacting protein gene |
| HIST1H2BB | Histone cluster 1 H2B family member B gene |
| IFT140 | Intraflagellar transport 140 gene |
| ITGA2 | Integrin subunit alpha 2 gene |
| ITIH3 | Inter-alpha-trypsin inhibitor heavy chain 3 gene |
| KDM5A | Lysine demethylase 5A gene |
| KP | Kegg pathway |
| KLK14 | Kallikrein related peptidase 14 gene |
| MAP2K3 | Mitogen-activated protein kinase kinase 3 gene |
| MF | Molecular function |
| MYBPC2 | Myosin binding protein C, fast type gene |
| NOTCH1 | Notch 1 gene |
| OBSCN | Obscurin gene |
| OR2C1 | Olfactory receptor family 2 subfamily C member 1 gene |
| OR4C3 | Olfactory receptor family 4 subfamily C member 3 gene |
| OR10G4 | Olfactory receptor family 10 subfamily G member 4 gene |
| OR13F1 | Olfactory receptor family 13 subfamily F member 1 gene |
| PDE4DIP | Phosphodiesterase 4D interacting protein gene |
| PER2 | Period circadian regulator 2 gene |
| PKD1 | Polycystin 1 gene |
| PLEKHG4B | Pleckstrin homology and RhoGEF domain containing G4B gene |
| PLOD1 | Procollagen-lysine, 2-oxoglutarate 5-dioxygenase 1 gene |
| PTPRD | Protein tyrosine phosphatase receptor type D gene |
| RASGRP1 | RAS Guanyl releasing protein 1 gene |
| RAPGEF1 | Rap guanine nucleotide exchange factor 1 gene |
| RPS3 | Ribosomal protein S3 gene |
| SGCD | Sarcoglycan delta gene |
| SLC9A3 | Solute carrier family 9 member A3 gene |
| SPATA17 | Spermatogenesis associated 17 gene |
| SVS | SNP and variation suite |
| THBS1 | Thrombospondin 1 gene |
| TLR8 | Toll like receptor 8 gene |
| UBE2NL | Ubiquitin conjugating enzyme E2 N like gene |
| UGT1A3 | UDP Glucuronosyltransferase family 1 member A3 gene |
| UGT1A4 | UDP Glucuronosyltransferase family 1 member A4 gene |
| UGT1A5 | UDP Glucuronosyltransferase family 1 member A5 gene |
| UGT1A6 | UDP Glucuronosyltransferase family 1 member A6 gene |
| UGT1A7 | UDP Glucuronosyltransferase family 1 member A7 gene |
| UGT1A8 | UDP Glucuronosyltransferase family 1 member A8 gene |
| UGT1A9 | UDP Glucuronosyltransferase family 1 member A9 gene |
| UGT1A10 | UDP Glucuronosyltransferase family 1 member A10 gene |
| WD | Week of development |
| WES | Whole exome sequence |
| ZFHX3 | Zinc finger homeobox 3 gene |
| ZNF717 | Zinc finger protein 717 gene |
References
- Salinas-Torres, V.M.; Salinas-Torres, R.A.; Cerda-Flores, R.M.; Martínez-de-Villarreal, L.E. Prevalence, mortality, and spatial distribution of gastroschisis in Mexico. J. Pediatr. Adolesc. Gynecol. 2018, 31, 232–237. [Google Scholar] [CrossRef] [PubMed]
- Salinas-Torres, V.M.; Salinas-Torres, R.A.; Cerda-Flores, R.M.; Martínez-de-Villarreal, L.E. Genetic variants conferring susceptibility to gastroschisis: A phenomenon restricted to the interaction with the environment? Pediatr. Surg. Int. 2018, 34, 505–514. [Google Scholar] [CrossRef] [PubMed]
- Salinas-Torres, V.M.; Salinas-Torres, R.A.; Cerda-Flores, R.M.; Gallardo-Blanco, H.L.; Martínez-de-Villarreal, L.E. A clinical-pathogenetic approach on associated anomalies and chromosomal defects supports novel candidate critical regions and genes for gastroschisis. Pediatr. Surg. Int. 2018, 34, 931–943. [Google Scholar] [CrossRef] [PubMed]
- Salinas-Torres, V.M.; Gallardo-Blanco, H.L.; Salinas-Torres, R.A.; Martínez de Villarreal, L.E. Dataset for Genes and Gene Variants from Familial Gastroschisis. Available online: https://doi.org/10.5281/zenodo.2673050 (accessed on 7 May 2019).
- Mi, H.; Huang, X.; Muruganujan, A.; Tang, H.; Mills, C.; Kang, D.; Thomas, P.D. PANTHER version 11: Expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2017, 45, D183–D189. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- SNP & Variation Suite ™ (Version 8.x); Golden Helix, Inc.: Bozeman, MT, USA; Available online: http://www.goldenhelix.com (accessed on 31 August 2018).
- Chen, J.; Bardes, E.E.; Aronow, B.J.; Jegga, A.G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 2009, 37, W305–W311. [Google Scholar] [CrossRef] [PubMed]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Stein, T.I.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards suite: From gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 2016, 54, 1.30.1–1.30.33. [Google Scholar] [PubMed]
- Rowbotham, S.E.; Illingworth, N.A.; Daly, A.K.; Veal, G.J.; Boddy, A.V. Role of UDP-glucuronosyltransferase isoforms in 13-cis retinoic acid metabolism in humans. Drug Metab. Dispos. 2010, 38, 1211–1217. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, B.M.; Rudolph, K.L.; Karagianni, P.; Fonseca, N.A.; White, R.J.; Talianidis, I.; Odom, D.T.; Marioni, J.C.; Kutter, C. High-resolution mapping of transcriptional dynamics across tissue development reveals a stable mRNA-tRNA interface. Genome Res. 2014, 24, 1797–1807. [Google Scholar] [CrossRef] [PubMed]
- Cormier, S.; Vandormael-Pournin, S.; Babinet, C.; Cohen-Tannoudji, M. Developmental expression of the Notch signaling pathway genes during mouse preimplantation development. Gene Expr. Patterns 2004, 4, 713–717. [Google Scholar] [CrossRef] [PubMed]
- Guo, G.; Huss, M.; Tong, G.Q.; Wang, C.; Li Sun, L.; Clarke, N.D.; Robson, P. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 2010, 18, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Barbacioru, C.; Nordman, E.; Bao, S.; Lee, C.; Wang, X.; Tuch, B.B.; Heard, E.; Lao, K.; Surani, M.A. Deterministic and stochastic allele specific gene expression in single mouse blastomeres. PLoS ONE 2011, 6, e21208. [Google Scholar] [CrossRef]
- Redmond, L.C.; Dumur, C.I.; Archer, K.J.; Haar, J.L.; Lloyd, J.A. Identification of erythroid-enriched gene expression in the mouse embryonic yolk sac using microdissected cells. Dev. Dyn. 2008, 237, 436–446. [Google Scholar] [CrossRef] [PubMed]
- Diez-Roux, G.; Banfi, S.; Sultan, M.; Geffers, L.; Anand, S.; Rozado, D.; Magen, A.; Canidio, E.; Pagani, M.; Peluso, I.; et al. A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol. 2011, 9, e1000582. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, A.E.; Calarco, P.G.; Damsky, C.H. Developmental regulation of integrin expression at the time of implantation in the mouse embryo. Development 1993, 119, 1175–1186. [Google Scholar] [PubMed]
- Wu, J.E.; Santoro, S.A. Complex patterns of expression suggest extensive roles for the alpha 2 beta 1 integrin in murine development. Dev. Dyn. 1994, 199, 292–314. [Google Scholar] [CrossRef] [PubMed]
- Ong, K.; Horsfall, W.; Conway, E.M.; Schuh, A.C. Early embryonic expression of murine coagulation system components. Thromb. Haemost. 2000, 84, 1023–1030. [Google Scholar] [CrossRef]
- Tamplin, O.J.; Cox, B.J.; Rossant, J. Integrated microarray and ChIP analysis identifies multiple Foxa2 dependent target genes in the notochord. Dev. Biol. 2011, 360, 415–425. [Google Scholar] [CrossRef]
- Visel, A.; Thaller, C.; Eichele, G. GenePaint.org: An atlas of gene expression patterns in the mouse embryo. Nucleic Acids Res. 2004, 32, D552–D556. [Google Scholar] [CrossRef]
- Thomas, T.; Voss, A.K.; Petrou, P.; Gruss, P. The murine gene, Traube, is essential for the growth of preimplantation embryos. Dev. Biol. 2000, 227, 324–342. [Google Scholar] [CrossRef]
- Wright, T.J.; Hatch, E.P.; Karabagli, H.; Karabagli, P.; Schoenwolf, G.C.; Mansour, S.L. Expression of mouse fibroblast growth factor and fibroblast growth factor receptor genes during early inner ear development. Dev. Dyn. 2003, 228, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Stark, K.L.; McMahon, J.A.; McMahon, A.P. FGFR-4, a new member of the fibroblast growth factor receptor family, expressed in the definitive endoderm and skeletal muscle lineages of the mouse. Development 1991, 113, 641–651. [Google Scholar] [PubMed]
- Magdaleno, S.; Jensen, P.; Brumwell, C.L.; Seal, A.; Lehman, K.; Asbury, A.; Cheung, T.; Cornelius, T.; Batten, D.M.; Eden, C.; et al. BGEM: An in situ hybridization database of gene expression in the embryonic and adult mouse nervous system. PLoS Biol. 2006, 4, e86. [Google Scholar] [CrossRef] [PubMed]
- Gray, P.A.; Fu, H.; Luo, P.; Zhao, Q.; Yu, J.; Ferrari, A.; Tenzen, T.; Yuk, D.I.; Tsung, E.F.; Cai, Z.; et al. Mouse brain organization revealed through direct genome-scale TF expression analysis. Science 2004, 306, 2255–2257. [Google Scholar] [CrossRef] [PubMed]
- Siyanov, V.; Baltz, J.M. NHE1 is the sodium-hydrogen exchanger active in acute intracellular pH regulation in preimplantation mouse embryos. Biol. Reprod. 2013, 88, 157. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Barr, K.J.; Garrill, A.; Jones, D.H.; Orlowski, J.; Kidder, G.M. Contributions of Na+/H+ exchanger isoforms to preimplantation development of the mouse. Mol. Reprod. Dev. 1998, 50, 146–153. [Google Scholar] [CrossRef]
- Nakamura, K.; Stokes, J.B.; McCray, P.B., Jr. Endogenous and exogenous glucocorticoid regulation of ENaC mRNA expression in developing kidney and lung. Am. J. Physiol. Cell Physiol. 2002, 283, C762–C772. [Google Scholar] [CrossRef][Green Version]
- Schaapveld, R.Q.; Schepens, J.T.; Bächner, D.; Attema, J.; Wieringa, B.; Jap, P.H.; Hendriks, W.J. Developmental expression of the cell adhesion molecule-like protein tyrosine phosphatases LAR, RPTPdelta and RPTPsigma in the mouse. Mech. Dev. 1998, 77, 59–62. [Google Scholar] [CrossRef]
- Kurasawa, M.; Sato, N.; Matsuda, A.; Koshida, S.; Totsuka, T.; Obinata, T. Differential expression of C-protein isoforms in developing and degenerating mouse striated muscles. Muscle Nerve. 1999, 22, 196–207. [Google Scholar] [CrossRef]
- Giudice, J.; Xia, Z.; Wang, E.T.; Scavuzzo, M.A.; Ward, A.J.; Kalsotra, A.; Wang, W.; Wehrens, X.H.; Burge, C.B.; Li, W.; et al. Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development. Nat. Commun. 2014, 5, 3603. [Google Scholar] [CrossRef] [PubMed]
- Moore, T.C.; Stokes, G.E. Gastroschisis; report of two cases treated by a modification of the gross operation for omphalocele. Surgery 1953, 33, 112–120. [Google Scholar] [PubMed]
- Duhamel, B. Embryology of exomphalos and allied malformations. Arch. Dis. Child. 1963, 38, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Feldkamp, M.L.; Carey, J.C.; Sadler, T.W. Development of gastroschisis: Review of hypotheses, a novel hypothesis, and implications for research. Am. J. Med. Genet. A 2007, 143A, 639–652. [Google Scholar] [CrossRef] [PubMed]
- Shaw, A. The myth of gastroschisis. J. Pediatr. Surg. 1975, 10, 235–244. [Google Scholar] [CrossRef]
- Thomas, D.F.; Atwell, J.D. The embryology and surgical management of gastroschisis. Br. J. Surg. 1976, 63, 893–897. [Google Scholar] [CrossRef] [PubMed]
- deVries, P.A. The pathogenesis of gastroschisis and omphalocele. J. Pediatr. Surg. 1980, 15, 245–251. [Google Scholar] [CrossRef]
- Hoyme, H.E.; Higginbottom, M.C.; Jones, K.L. The vascular pathogenesis of gastroschisis: Intrauterine interruption of the omphalomesenteric artery. J. Pediatr. 1981, 98, 228–231. [Google Scholar] [CrossRef]
- Vermeij-Keers, C.; Hartwig, N.G.; van der Werff, J.F. Embryonic development of the ventral body wall and its congenital malformations. Semin. Pediatr. Surg. 1996, 5, 82–89. [Google Scholar] [PubMed]
- Stevenson, R.E.; Rogers, R.C.; Chandler, J.C.; Gauderer, M.W.; Hunter, A.G. Escape of the yolk sac: A hypothesis to explain the embryogenesis of gastroschisis. Clin. Genet. 2009, 75, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Lubinsky, M. A vascular and thrombotic model of gastroschisis. Am. J. Med. Genet. A 2014, 164A, 915–917. [Google Scholar] [CrossRef]
- Salinas-Torres, V.M.; Salinas-Torres, R.A.; Cerda-Flores, R.M.; Martínez-de-Villarreal, L.E. Evaluation of familial factors in a Mexican population-based setting with gastroschisis: Further evidence for an underlying genetic susceptibility. J. Pediatr. Surg. 2018, 53, 521–524. [Google Scholar] [CrossRef] [PubMed]
- Salinas-Torres, V.M.; Salinas-Torres, R.A.; Cerda-Flores, R.M.; Martínez-de-Villarreal, L.E. Familial occurrence of gastroschisis: A population-based overview on recurrence risk, sex-dependent influence, and geographical distribution. Pediatr. Surg. Int. 2018, 34, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; van Leeuwen, J.; Andrews, B.J.; Boone, C. Genetic network complexity shapes background-dependent phenotypic expression. Trends Genet. 2018, 34, 578–586. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.A.; Li, Y.I.; Pritchard, J.K. An expanded view of complex traits: From polygenic to omnigenic. Cell 2017, 169, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Chakravorty, S.; Hegde, M. Inferring the effect of genomic variation in the new era of genomics. Hum. Mutat. 2018, 39, 756–773. [Google Scholar] [CrossRef] [PubMed]
- Hartwig, N.G.; Vermeij-Keers, C.; De Vries, H.E.; Kagie, M.; Kragt, H. Limb body wall malformation complex: An embryologic etiology? Hum. Pathol. 1989, 20, 1071–1077. [Google Scholar] [CrossRef]
- Hartwig, N.G.; Steffelaar, J.W.; Van de Kaa, C.; Schueler, J.A.; Vermeij-Keers, C. Abdominal wall defect associated with persistent cloaca. The embryologic clues in autopsy. Am. J. Clin. Pathol. 1991, 96, 640–647. [Google Scholar] [CrossRef]
- Brewer, S.; Williams, T. Finally, a sense of closure? Animal models of human ventral body wall defects. Bioessays 2004, 26, 1307–1321. [Google Scholar] [CrossRef] [PubMed]
- O’Rahilly, R.; Müller, F. Human Embryology and Teratology, 3rd ed.; Wiley-Liss: New York, NY, USA, 2001. [Google Scholar]
- Rittler, M.; Vauthay, L.; Mazzitelli, N. Gastroschisis is a defect of the umbilical ring: Evidence from morphological evaluation of stillborn fetuses. Birth Defects Res. A Clin. Mol. Teratol. 2013, 97, 198–209. [Google Scholar] [CrossRef] [PubMed]
- Smits-van Prooije, A.E.; Vermeij-Keers, C.; Poelmann, R.E.; Mentink, M.M.; Dubbeldam, J.A. The formation of mesoderm and mesectoderm in 5- to 41-somite rat embryos cultured in vitro, using WGA-Au as a marker. Anat. Embryol. (Berl.) 1988, 177, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Brewer, S.; Williams, T. Loss of AP-2alpha impacts multiple aspects of ventral body wall development and closure. Dev. Biol. 2004, 267, 399–417. [Google Scholar] [CrossRef] [PubMed]
- Mekonen, H.K.; Hikspoors, J.P.; Mommen, G.; Köhler, S.E.; Lamers, W.H. Development of the ventral body wall in the human embryo. J. Anat. 2015, 227, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Gasser, R.F. Evidence that some events of mammalian embryogenesis can result from differential growth, making migration unnecessary. Anat. Rec. B New. Anat. 2006, 289, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Hussain, F.; Decuzzi, P. Role of differential adhesion in cell cluster evolution: From vasculogenesis to cancer metastasis. Comput. Methods Biomech. Biomed. Eng. 2015, 18, 282–292. [Google Scholar] [CrossRef] [PubMed]
- Goody, M.F.; Sher, R.B.; Henry, C.A. Hanging on for the ride: Adhesion to the extracellular matrix mediates cellular responses in skeletal muscle morphogenesis and disease. Dev. Biol. 2015, 401, 75–91. [Google Scholar] [CrossRef]
- Bargy, F.; Beaudoin, S. Comprehensive developmental mechanisms in gastroschisis. Fetal Diagn. Ther. 2014, 36, 223–330. [Google Scholar] [CrossRef] [PubMed]
- Shearman, R.M.; Burke, A.C. The lateral somitic frontier in ontogeny and phylogeny. J. Exp. Zool. B. Mol. Dev. Evol. 2009, 312, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Streeter, G.L. Focal deficiencies in fetal tissues and their relation to intra-uterine amputation. Contrib. Embryol. Carnegie Inst. 1930, 22, 1–44. [Google Scholar]
- Bamforth, J.S. Amniotic band sequence: Streeter’s hypothesis reexamined. Am. J. Med. Genet. 1992, 44, 280–287. [Google Scholar] [CrossRef] [PubMed]
- Salinas-Torres, V.M.; De La O-Espinoza, E.A.; Salinas-Torres, R.A. Severe intrauterine amputations in one dichorionic twin with pentalogy of Cantrell: Further evidence and consideration for mechanical teratogenesis. Pediatr. Dev. Pathol. 2017, 20, 440–443. [Google Scholar] [CrossRef]
- Salinas-Torres, V.M. Fetus with Casamassima-Morton-Nance syndrome and limb-body wall defect: Presentation of a novel association and review of the phenotype. Pediatr. Dev. Pathol. 2016, 19, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Folkerth, R.D.; Habbe, D.M.; Boyd, T.K.; McMillan, K.; Gromer, J.; Sens, M.A.; Elliott, A.J. Prenatal Alcohol, SIDS and Stillbirth (PASS) Research Network. Gastroschisis, destructive brain lesions, and placental infarction in the second trimester suggest a vascular pathogenesis. Pediatr. Dev. Pathol. 2013, 16, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Chambers, C.D.; Chen, B.H.; Kalla, K.; Jernigan, L.; Jones, K.L. Novel risk factor in gastroschisis: Change of paternity. Am. J. Med. Genet. A 2007, 143A, 653–659. [Google Scholar] [CrossRef]
- Frascoli, M.; Jeanty, C.; Fleck, S.; Moradi, P.W.; Keating, S.; Mattis, A.N.; Tang, Q.; MacKenzie, T.C. Heightened immune activation in fetuses with gastroschisis may be blocked by targeting IL-5. J. Immunol. 2016, 196, 4957–4966. [Google Scholar] [CrossRef]
- Kalache, K.D.; Bierlich, A.; Hammer, H.; Bollmann, R. Is unexplained third trimester intrauterine death of fetuses with gastroschisis caused by umbilical cord compression due to acute extra-abdominal bowel dilatation? Prenat. Diagn. 2002, 22, 715–717. [Google Scholar] [CrossRef] [PubMed]
- Luton, D.; de Lagausie, P.; Guibourdenche, J.; Oury, J.; Sibony, O.; Vuillard, E.; Boissinot, C.; Aigrain, Y.; Beaufils, F.; Navarro, J.; et al. Effect of amnioinfusion on the outcome of prenatally diagnosed gastroschisis. Fetal Diagn. Ther. 1999, 14, 152–155. [Google Scholar] [CrossRef] [PubMed]
- McClintock, T.S.; Glasser, C.E.; Bose, S.C.; Bergman, D.A. Tissue expression patterns identify mouse cilia genes. Physiol. Genomics 2008, 32, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Stauber, M.; Weidemann, M.; Dittrich-Breiholz, O.; Lobschat, K.; Alten, L.; Mai, M.; Beckers, A.; Kracht, M.; Gossler, A. Identification of FOXJ1 effectors during ciliogenesis in the foetal respiratory epithelium and embryonic left-right organiser of the mouse. Dev. Biol. 2017, 423, 170–188. [Google Scholar] [CrossRef] [PubMed]
- Olsen, C.L.; Hsu, P.P.; Glienke, J.; Rubanyi, G.M.; Brooks, A.R. Hedgehog-interacting protein is highly expressed in endothelial cells but down-regulated during angiogenesis and in several human tumors. BMC Cancer 2004, 4, 43. [Google Scholar] [CrossRef] [PubMed]
- Guillaume, R.; D’Agati, V.; Daoust, M.; Trudel, M. Murine Pkd1 is a developmentally regulated gene from morula to adulthood: Role in tissue condensation and patterning. Dev. Dyn. 1999, 214, 337–348. [Google Scholar] [CrossRef]
- Weinmaster, G.; Roberts, V.J.; Lemke, G. A homolog of Drosophila Notch expressed during mammalian development. Development 1991, 113, 199–205. [Google Scholar] [PubMed]
- Schmidts, M.; Frank, V.; Eisenberger, T.; Al Turki, S.; Bizet, A.A.; Antony, D.; Rix, S.; Decker, C.; Bachmann, N.; Bald, M.; et al. Combined NGS approaches identify mutations in the intraflagellar transport gene IFT140 in skeletal ciliopathies with early progressive kidney Disease. Hum. Mutat. 2013, 34, 714–724. [Google Scholar] [CrossRef]
- Ryan, A.K.; Blumberg, B.; Rodriguez-Esteban, C.; Yonei-Tamura, S.; Tamura, K.; Tsukui, T.; de la Peña, J.; Sabbagh, W.; Greenwald, J.; Choe, S.; et al. Pitx2 determines left-right asymmetry of internal organs in vertebrates. Nature 1998, 394, 545–551. [Google Scholar] [CrossRef] [PubMed]
- Hikspoors, J.P.J.M.; Peeters, M.M.J.P.; Mekonen, H.K.; Kruepunga, N.; Mommen, G.M.C.; Cornillie, P.; Köhler, S.E.; Lamers, W.H. The fate of the vitelline and umbilical veins during the development of the human liver. J. Anat. 2017, 231, 718–735. [Google Scholar] [CrossRef] [PubMed]
- Ramsdell, A.F.; Yost, H.J. Molecular mechanisms of vertebrate left-right development. Trends Genet. 1998, 14, 459–465. [Google Scholar] [CrossRef]
- Benjamin, B.G.; Ethen, M.K.; Van Hook, C.L.; Myers, C.A.; Canfield, M.A. Gastroschisis prevalence in Texas 1999–2003. Birth Defects Res. A Clin. Mol. Teratol. 2010, 88, 178–185. [Google Scholar] [PubMed]
- Khodr, Z.G.; Lupo, P.J.; Canfield, M.A.; Chan, W.; Cai, Y.; Mitchell, L.E. Hispanic ethnicity and acculturation, maternal age and the risk of gastroschisis in the National Birth Defects Prevention Study. Birth Defects Res. A Clin. Mol. Teratol. 2013, 97, 538–545. [Google Scholar] [CrossRef] [PubMed]
- Friedman, A.M.; Ananth, C.V.; Siddiq, Z.; D’Alton, M.E.; Wright, J.D. Gastroschisis: Epidemiology and mode of delivery, 2005–2013. Am. J. Obstet. Gynecol. 2016, 215, e1–e9. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).