Small Non-Coding RNAome of Ageing Chondrocytes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection and Preparation
2.2. RNA Isolation, cDNA Library Preparation, and Small RNA Sequencing (RNA-seq)
2.3. Data Processing
2.4. Pathway Analysis
2.5. qRT-PCR Validation
2.6. tRNA Fragment Analysis
2.7. Novel snoRNA Analysis
2.8. Statistical Analysis
2.9. Human Sample Collection and Preparation
2.10. Removal of tRF Modifications from Human Cartilage RNA
2.11. tRF 3′ and 5′ Adaptor Ligation for Human Cartilage cDNA Synthesis
2.12. tRF and tRNA Human qPCR Arrays
3. Results
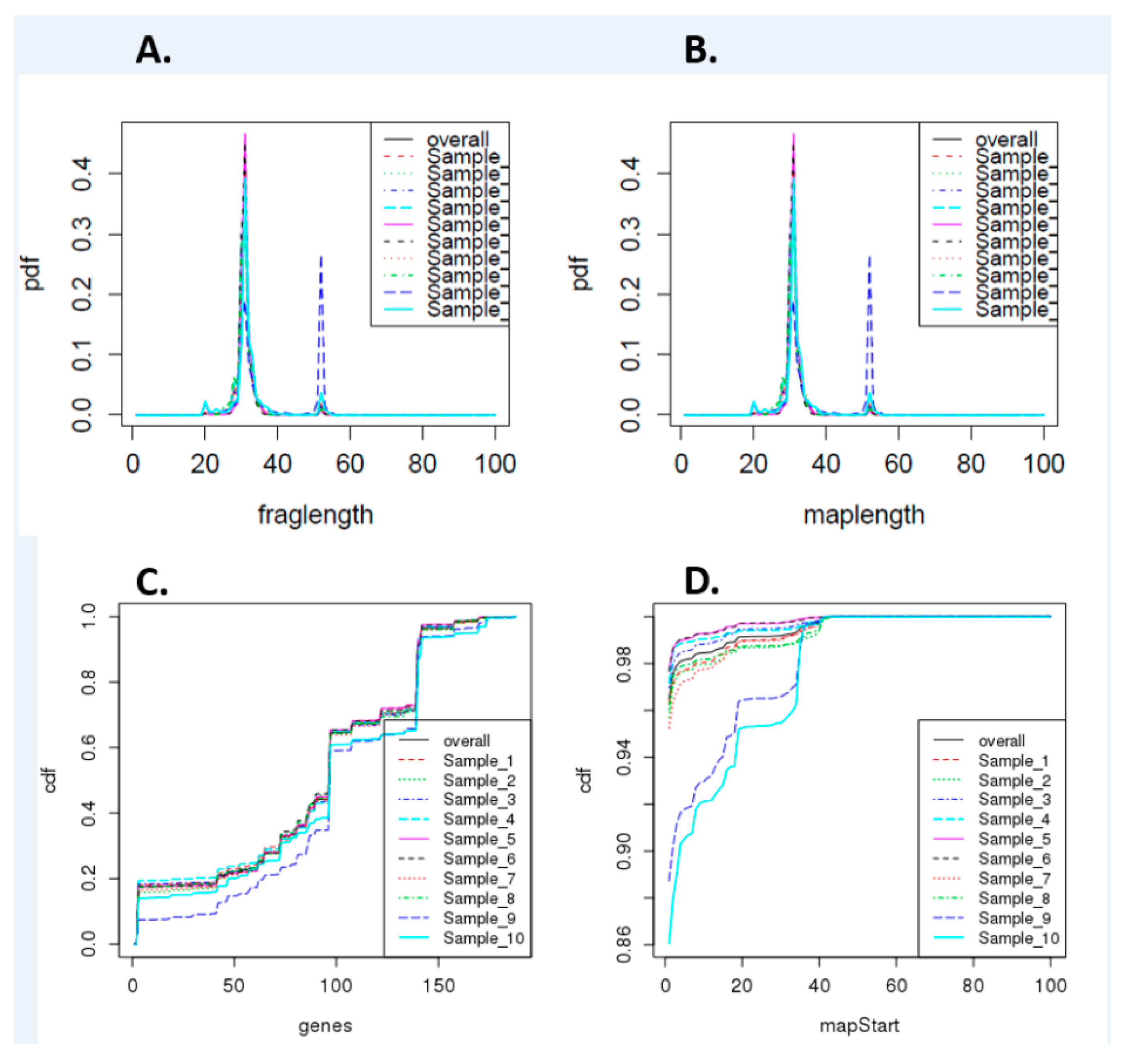
3.1. Preliminary Analysis of Small RNA-seq Data
3.2. Age-Related Differential Small Non-Coding RNA Gene Expression
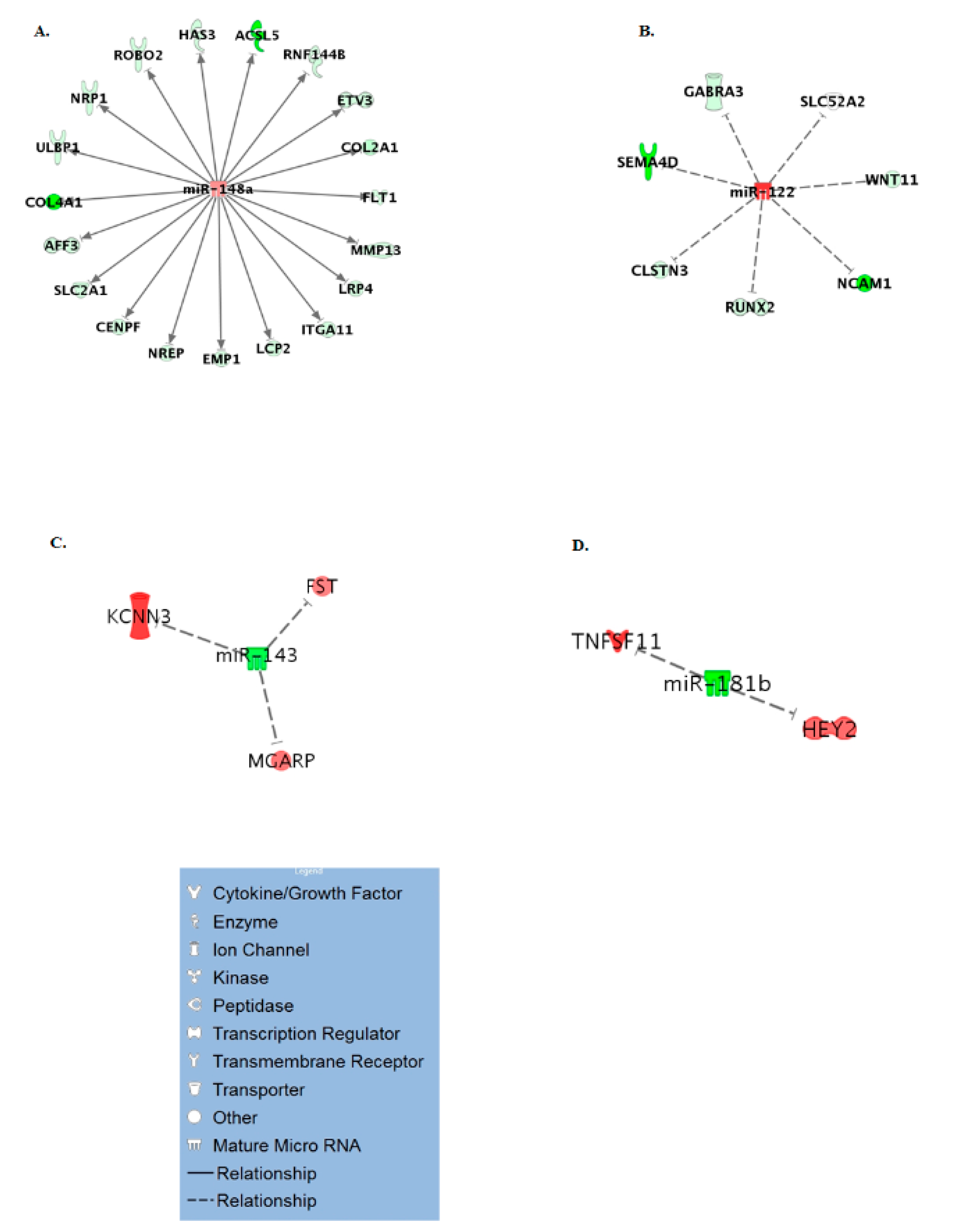
3.3. Age Specific miRNA Interactome
3.4. Confirmation of Differential Gene Expression Using qRT-PCR
3.5. tRNA Fragment Changes in Equine Ageing
3.6. Human tRNA and tRF Profiles Compared to Equine tRNA and tRF Profiles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ECM | extracellular matrix |
| OA | osteoarthritis |
| sncRNAs | small non-coding RNAs |
| miRNAs or miRs | microRNAs |
| snRNAs | small nuclear RNAs |
| snoRNAs | small nucleolar RNAs |
| tRNAs | transfer RNAs |
| mRNA | messenger RNA |
| tiRNAs | tRNA halves |
| ANG | angiogenin |
| tRFs | tRNA-derived small RNA fragments |
| PCA | principle component analysis |
| logFC | log2 fold change |
| FDR | False Discovery Rate |
| IPA | Ingenuity Pathway Analysis |
| RNA-seq | RNA sequencing |
| qRT-PCR | real-time quantitative Polymerase Chain Reaction |
| lncRNAs | long non-coding RNAs |
| PCs | Principle components |
| GP6 | glycoprotein 6 |
| BMP | bone morphogenic protein |
| FST | follistatin |
| TNF | tumour necrosis factor |
| TNFSF11 | TNF ligand superfamily member 11 |
| RANKL | receptor activator of nuclear factor kappa-Β ligand |
| ERK | extracellular signal-regulated kinase |
| SMAD | mothers against decapentaplegic homolog |
| MAPK | mitogen-activated protein kinase |
| TGF-β | transforming growth factor beta |
| IL | interleukin |
| JAK3 | Janus Kinase 3 |
| AGO | Argonaute |
| RISC | RNA-induced silencing complex |
| JAK-STAT | Janus Kinase and Signal Transducer and Activator of Transcription |
References
- Messina, O.D.; Wilman, M.V.; Neira, L.F.V. Nutrition, osteoarthritis and cartilage metabolism. Aging Clin. Exp. Res. 2019, 31, 807–813. [Google Scholar] [CrossRef] [PubMed]
- Decker, R.S. Articular cartilage and joint development from embryogenesis to adulthood. Semin. Cell Dev. Biol. 2016, 62, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Sacitharan, P.K.; Vincent, T.L. Cellular ageing mechanisms in osteoarthritis. Mamm. Genome 2016, 27, 421–429. [Google Scholar] [CrossRef] [Green Version]
- Philipot, D.; Guérit, D.; Platano, D.; Chuchana, P.; Olivotto, E.; Espinoza, F.; Dorandeu, A.; Pers, Y.M.; Piette, J.; Borzì, R.; et al. p16INK4a and its regulator miR-24 link senescence and chondrocyte terminal differentiation-associated matrix remodeling in osteoarthritis. Arthritis Res. Ther. 2014, 16, R58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verzijl, N.; DeGroot, J.; Ben Zaken, C.; Braun-Benjamin, O.; Maroudas, A.; Bank, R.A.; Mizrahi, J.; Schalkwijk, C.G.; Thorpe, S.R.; Baynes, J.W.; et al. Crosslinking by advanced glycation end products increases the stiffness of the collagen network in human articular cartilage: A possible mechanism through which age is a risk factor for osteoarthritis. Arthritis Rheum. 2002, 46, 114–123. [Google Scholar] [CrossRef]
- Zhang, M.; Theleman, J.L.; Lygrisse, K.A.; Wang, J. Epigenetic Mechanisms Underlying the Aging of Articular Cartilage and Osteoarthritis. Gerontology 2019, 65, 387–396. [Google Scholar] [CrossRef]
- Greene, M.A.; Loeser, R.F. Aging-related inflammation in osteoarthritis. Osteoarthr. Cartil. 2015, 23, 1966–1971. [Google Scholar] [CrossRef] [Green Version]
- Sondag, G.R.; Haqqi, T.M. The Role of MicroRNAs and Their Targets in Osteoarthritis. Curr. Rheumatol. Rep. 2016, 18, 56. [Google Scholar] [CrossRef]
- Steinbusch, M.M.F.; Fang, Y.; Milner, P.I.; Clegg, P.D.; Young, D.A.; Welting, T.J.M.; Peffers, M.J. Serum snoRNAs as biomarkers for joint ageing and post traumatic osteoarthritis. Sci. Rep. 2017, 7, 43558. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Yang, M.; Marks, P.; White, L.; Hurtig, M.; Mi, Q.-S.; Divine, G.; Gibson, G. Serum non-coding RNAs as biomarkers for osteoarthritis progression after ACL injury. Osteoarthr. Cartil. 2012, 20, 1631–1637. [Google Scholar] [CrossRef] [Green Version]
- Kamaruzaman, H.; Kinghorn, P.; Oppong, R. Cost-effectiveness of surgical interventions for the management of osteoarthritis: A systematic review of the literature. BMC Musculoskelet. Disord. 2017, 18, 183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miyaki, S.; Sato, T.; Inoue, A.; Otsuki, S.; Ito, Y.; Yokoyama, S.; Kato, Y.; Takemoto, F.; Nakasa, T.; Yamashita, S.; et al. MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes Dev. 2010, 24, 1173–1185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peffers, M.; Caron, M.; Cremers, A.; Surtel, D.; Fang, Y.; Dyer, P.; Balaskas, P.; Welting, T. snoRNA signatures in cartilage ageing and osteoarthritis. Osteoarthr. Cartil. 2018, 26, S164. [Google Scholar] [CrossRef] [Green Version]
- Brameier, M.; Herwig, A.; Reinhardt, R.; Walter, L.; Gruber, J. Human box C/D snoRNAs with miRNA like functions: Expanding the range of regulatory RNAs. Nucleic Acids Res. 2011, 39, 675–686. [Google Scholar] [CrossRef]
- Stepanov, G.A.; Filippova, J.A.; Komissarov, A.B.; Kuligina, E.V.; Richter, V.A.; Semenov, D.V. Regulatory Role of Small Nucleolar RNAs in Human Diseases. BioMed Res. Int. 2015, 2015, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peffers, M.J.; Chabronova, A.; Balaskas, P.; Fang, Y.; Dyer, P.; Cremers, A.; Emans, P.J.; Feczko, P.Z.; Caron, M.M.; Welting, T.J.M. SnoRNA signatures in cartilage ageing and osteoarthritis. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Kharaz, Y.A.; Fang, Y.; Welting, T.; Peffers, M.; Comerford, E. Small RNA signatures of the anterior cruciate ligament from patients with knee 1 joint osteoarthritis. medRxiv 2020. [Google Scholar] [CrossRef]
- Raina, M.; Ibba, M. tRNAs as regulators of biological processes. Front. Genet. 2014, 5, 171. [Google Scholar] [CrossRef] [Green Version]
- Yamasaki, S.; Ivanov, P.; Hu, G.-F.; Anderson, P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J. Cell Biol. 2009, 185, 35–42. [Google Scholar] [CrossRef] [Green Version]
- Lee, Y.S.; Shibata, Y.; Malhotra, A.; Dutta, A. A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev. 2009, 23, 2639–2649. [Google Scholar] [CrossRef] [Green Version]
- Takaku, H.; Minagawa, A.; Takagi, M.; Nashimoto, M. A candidate prostate cancer susceptibility gene encodes tRNA 3′ processing endoribonuclease. Nucleic Acids Res. 2003, 31, 2272–2278. [Google Scholar] [CrossRef] [PubMed]
- Goodarzi, H.; Liu, X.; Nguyen, H.C.; Zhang, S.; Fish, L.; Tavazoie, S.F. Endogenous tRNA-Derived Fragments Suppress Breast Cancer Progression via YBX1 Displacement. Cell 2015, 161, 790–802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cole, C.; Sobala, A.; Lu, C.; Thatcher, S.R.; Bowman, A.; Brown, J.W.; Green, P.J.; Barton, G.J.; Hutvagner, G. Filtering of deep sequencing data reveals the existence of abundant Dicer-dependent small RNAs derived from tRNAs. RNA 2009, 15, 2147–2160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.; Ender, C.; Meister, G.; Moore, P.S.; Chang, Y.; John, B. Extensive terminal and asymmetric processing of small RNAs from rRNAs, snoRNAs, snRNAs, and tRNAs. Nucleic Acids Res. 2012, 40, 6787–6799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Telonis, A.G.; Loher, P.; Honda, S.; Jing, Y.; Palazzo, J.; Kirino, Y.; Rigoutsos, I. Dissecting tRNA-derived fragment complexities using personalized transcriptomes reveals novel fragment classes and unexpected dependencies. Oncotarget 2015, 6, 24797–24822. [Google Scholar] [CrossRef] [Green Version]
- Jiang, P.; Yan, F. tiRNAs & tRFs Biogenesis and Regulation of Diseases: A Review. Curr. Med. Chem. 2019, 26, 5849–5861. [Google Scholar] [CrossRef]
- Green, J.; Ansari, M.; Ball, H.; Haqqi, T.M. tRNA-derived fragments (tRFs) regulate post-transcriptional gene expression via AGO-dependent mechanism in IL-1β stimulated chondrocytes. Osteoarthr. Cartil. 2020, 28, 1102–1110. [Google Scholar] [CrossRef]
- Ormseth, M.J.; Solus, J.F.; Sheng, Q.; Ye, F.; Song, H.; Wu, Q.; Guo, Y.; Oeser, A.M.; Allen, R.M.; Vickers, K.C.; et al. The Endogenous Plasma Small RNAome of Rheumatoid Arthritis. ACR Open Rheumatol. 2020, 2, 97–105. [Google Scholar] [CrossRef]
- Kawcak, C.; Frisbie, D.; Werpy, N.; Park, R.; McIlwraith, C. Effects of exercise vs experimental osteoarthritis on imaging outcomes. Osteoarthr. Cartil. 2008, 16, 1519–1525. [Google Scholar] [CrossRef] [Green Version]
- Peffers, M.J.; Milner, P.; Tew, S.; Clegg, P.D. Regulation of SOX9 in normal and osteoarthritic equine articular chondrocytes by hyperosmotic loading. Osteoarthr. Cartil. 2010, 18, 1502–1508. [Google Scholar] [CrossRef] [Green Version]
- Peffers, M.; Liu, X.; Clegg, P. Transcriptomic signatures in cartilage ageing. Arthritis Res. Ther. 2013, 15, R98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- User Guide. 2011. Available online: https://emea.support.illumina.com/sequencing/sequencing_software/casava2/questions.html (accessed on 5 August 2020).
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 002832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [Green Version]
- Johnson, N.; Yeoh, J.M.; Coruh, C.; Axtell, M.J. Improved Placement of Multi-mapping Small RNAs. Genes Genomes Genetics 2016, 6, 2103–2111. [Google Scholar] [CrossRef] [Green Version]
- Oliveira, J.V.D.A.; Costa, F.; Backofen, R.; Stadler, P.F.; Walter, M.E.M.T.; Hertel, J. SnoReport 2.0: New features and a refined Support Vector Machine to improve snoRNA identification. BMC Bioinform. 2016, 17, 464–486. [Google Scholar] [CrossRef]
- Pang, Z.; Chong, J.; Li, S.; Xia, J. MetaboAnalystR 3.0: Toward an Optimized Workflow for Global Metabolomics. Metabolites 2020, 10, 186. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Wishart, D.S. Using MetaboAnalyst 3.0 for Comprehensive Metabolomics Data Analysis. Curr. Protoc. Bioinform. 2016, 55, 14.10.1–14.10.91. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Anaya, J.; Mudunuri, S.B.; Dutta, A. Meta-analysis of tRNA derived RNA fragments reveals that they are evolutionarily conserved and associate with AGO proteins to recognize specific RNA targets. BMC Biol. 2014, 12, 78. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Mudunuri, S.B.; Anaya, J.; Dutta, A. tRFdb: A database for transfer RNA fragments. Nucleic Acids Res. 2014, 43, D141–D145. [Google Scholar] [CrossRef] [PubMed]
- Olvedy, M.; Scaravilli, M.; Hoogstrate, Y.; Visakorpi, T.; Jenster, G.; Martens-Uzunova, E. A comprehensive repertoire of tRNA-derived fragments in prostate cancer. Oncotarget 2016, 7, 24766–24777. [Google Scholar] [CrossRef] [PubMed]
- Selitsky, S.R.; Baran-Gale, J.; Honda, M.; Yamane, D.; Masaki, T.; Fannin, E.E.; Guerra, B.; Shirasaki, T.; Shimakami, T.; Kaneko, S.; et al. Small tRNA-derived RNAs are increased and more abundant than microRNAs in chronic hepatitis B and C. Sci. Rep. 2015, 5, 7675. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Lee, I.; Ren, J.; Ajay, S.S.; Lee, Y.S.; Bao, X. Identification and Functional Characterization of tRNA-derived RNA Fragments (tRFs) in Respiratory Syncytial Virus Infection. Mol. Ther. 2013, 21, 368–379. [Google Scholar] [CrossRef] [Green Version]
- Chan, P.P.; Lowe, T.M. GtRNAdb: A database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 2009, 37, D93–D97. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. GtRNAdb 2.0: An expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 2016, 44, D184–D189. [Google Scholar] [CrossRef] [Green Version]
- Jühling, F.; Mörl, M.; Hartmann, R.K.; Sprinzl, M.; Stadler, P.F.; Pütz, J. tRNAdb 2009: Compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 2009, 37, D159–D162. [Google Scholar] [CrossRef] [Green Version]
- Lotz, M.K.; Loeser, R.F. Effects of aging on articular cartilage homeostasis. Bone 2012, 51, 241–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Wei, X.; Zhou, J.; Wei, L. The Age-Related Changes in Cartilage and Osteoarthritis. BioMed Res. Int. 2013, 2013, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.A.; Buckwalter, J.A. Telomere erosion and senescence in human articular cartilage chondrocytes. J. Gerontol. Ser. A Biol. Sci. Med Sci. 2001, 56, B172–B179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jallali, N.; Ridha, H.; Thrasivoulou, C.; Underwood, C.; Butler, P.E.; Cowen, T. Vulnerability to ROS-induced cell death in ageing articular cartilage: The role of antioxidant enzyme activity. Osteoarthr. Cartil. 2005, 13, 614–622. [Google Scholar] [CrossRef] [Green Version]
- Rahmati, M.; Nalesso, G.; Mobasheri, A.; Mozafari, M. Aging and osteoarthritis: Central role of the extracellular matrix. Ageing Res. Rev. 2017, 40, 20–30. [Google Scholar] [CrossRef]
- Hodgson, D.; Rowan, A.D.; Falciani, F.; Proctor, C.J. Systems biology reveals how altered TGFbeta signalling with age reduces protection against pro-inflammatory stimuli. PLoS Comput. Biol. 2019, 15, e1006685. [Google Scholar] [CrossRef]
- Weilner, S.; Grillari-Voglauer, R.; Redl, H.; Grillari, J.; Nau, T. The role of microRNAs in cellular senescence and age-related conditions of cartilage and bone. Acta Orthop. 2015, 86, 92–99. [Google Scholar] [CrossRef]
- Goodrich, L.R.; Nixon, A.J. Medical treatment of osteoarthritis in the horse—A review. Veter. J. 2006, 171, 51–69. [Google Scholar] [CrossRef]
- McIlwraith, C.; Frisbie, D.D.; Kawcak, C.E. The horse as a model of naturally occurring osteoarthritis. Bone Jt. Res. 2012, 1, 297–309. [Google Scholar] [CrossRef]
- Liu, Y.; Li, P.; Liu, L.; Zhang, Y. The diagnostic role of miR-122 in drug-induced liver injury. Medicine 2018, 97, e13478. [Google Scholar] [CrossRef]
- Thakral, S.; Ghoshal, K. miR-122 is a unique molecule with great potential in diagnosis, prognosis of liver disease, and therapy both as miRNA mimic and antimir. Curr. Gene Ther. 2015, 15, 142–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mandourah, A.Y.; Ranganath, L.; Barraclough, R.; Vinjamuri, S.; Hof, R.V.; Hamill, S.; Czanner, G.; Dera, A.A.; Wang, D.; Barraclough, D.L. Circulating microRNAs as potential diagnostic biomarkers for osteoporosis. Sci. Rep. 2018, 8, 8421. [Google Scholar] [CrossRef] [PubMed]
- Markopoulos, G.S.; Roupakia, E.; Tokamani, M.; Vartholomatos, G.; Tzavaras, T.; Hatziapostolou, M.; Fackelmayer, F.O.; Sandaltzopoulos, R.; Polytarchou, C.; Kolettas, E. Senescence-associated microRNAs target cell cycle regulatory genes in normal human lung fibroblasts. Exp. Gerontol. 2017, 96, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Burns, D.M.; D’Ambrogio, A.; Nottrott, S.; Richter, J.D. CPEB and two poly(A) polymerases control miR-122 stability and p53 mRNA translation. Nature 2011, 473, 105–108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soriano-Arroquia, A.; McCormick, R.; Molloy, A.P.; McArdle, A.; Goljanek-Whysall, K. Age-related changes in miR-143-3p:Igfbp5 interactions affect muscle regeneration. Aging Cell 2016, 15, 361–369. [Google Scholar] [CrossRef] [Green Version]
- Margolis, L.; Lessard, S.J.; Ezzyat, Y.; Fielding, R.A.; Rivas, D. Circulating MicroRNA Are Predictive of Aging and Acute Adaptive Response to Resistance Exercise in Men. J. Gerontol. Ser. A Biol. Sci. Med. Sci. 2017, 72, 1319–1326. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.; Ren, S.; Zhao, S. Wang. LncRNA MALAT1/MiR-145 Adjusts IL-1beta-Induced Chondrocytes Viability and Cartilage Matrix Degradation by Regulating ADAMTS5 in Human Osteoarthritis. Yonsei Med. J. 2019, 60, 1081–1092. [Google Scholar] [CrossRef]
- Hu, G.; Zhao, X.; Wang, C.; Geng, Y.; Zhao, J.; Xu, J.; Zuo, B.; Zhao, C.; Wang, C.; Zhang, X.-L. MicroRNA-145 attenuates TNF-α-driven cartilage matrix degradation in osteoarthritis via direct suppression of MKK4. Cell Death Dis. 2017, 8, e3140. [Google Scholar] [CrossRef]
- Mercken, E.M.; Majounie, E.; Ding, J.; Guo, R.; Kim, J.; Bernier, M.; Mattison, J.; Cookson, M.R.; Gorospe, M.; Decabo, R.; et al. Age-associated miRNA Alterations in Skeletal Muscle from Rhesus Monkeys reversed by caloric restriction. Aging 2013, 5, 692–703. [Google Scholar] [CrossRef] [Green Version]
- Collins, J.A.; Diekman, B.O.; Loeser, R. Targeting aging for disease modification in osteoarthritis. Curr. Opin. Rheumatol. 2018, 30, 101–107. [Google Scholar] [CrossRef]
- Thielen, N.G.M.; van der Kraan, P.M.; van Caam, A.P.M. TGFbeta/BMP Signaling Pathway in Cartilage Homeostasis. Cells 2019, 8, 969. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tardif, G.; Hum, D.; Pelletier, J.-P.; Boileau, C.; Ranger, P.; Martel-Pelletier, J. Differential gene expression and regulation of the bone morphogenetic protein antagonists follistatin and gremlin in normal and osteoarthritic human chondrocytes and synovial fibroblasts. Arthritis Rheum. 2004, 50, 2521–2530. [Google Scholar] [CrossRef] [PubMed]
- Tardif, G.; Pelletier, J.-P.; Boileau, C.; Martel-Pelletier, J. The BMP antagonists follistatin and gremlin in normal and early osteoarthritic cartilage: An immunohistochemical study. Osteoarthr. Cartil. 2009, 17, 263–270. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, N.; Yin, Y.; Wang, H.; Zhou, Z.; Sheng, X.; Fu, H.; Guo, R.; Wang, H.; Yang, J.; Gong, P.; et al. Telomere dysfunction impairs epidermal stem cell specification and differentiation by disrupting BMP/pSmad/P63 signaling. PLoS Genet. 2019, 15, e1008368. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, J.; Venton, L.; Sakata, T.; Halloran, B.P. Expression of RANKL and OPG Correlates With Age-Related Bone Loss in Male C57BL/6 Mice. J. Bone Miner. Res. 2003, 18, 270–277. [Google Scholar] [CrossRef]
- Martínez-Calatrava, M.J.; Prieto-Potín, I.; Roman-Blas, J.A.; Tardío, L.; Largo, R.; Herrero-Beaumont, G. RANKL synthesized by articular chondrocytes contributes to juxta-articular bone loss in chronic arthritis. Arthritis Res. Ther. 2012, 14, R149. [Google Scholar] [CrossRef] [Green Version]
- Upton, A.R.; Holding, C.A.; Dharmapatni, A.A.S.S.K.; Haynes, D. The expression of RANKL and OPG in the various grades of osteoarthritic cartilage. Rheumatol. Int. 2012, 32, 535–540. [Google Scholar] [CrossRef]
- Dieci, G.; Preti, M.; Montanini, B. Eukaryotic snoRNAs: A paradigm for gene expression flexibility. Genomics 2009, 94, 83–88. [Google Scholar] [CrossRef] [Green Version]
- Dupuis-Sandoval, F.; Poirier, M.; Scott, M.S. The emerging landscape of small nucleolar RNAs in cell biology. Wiley Interdiscip. Rev. RNA 2015, 6, 381–397. [Google Scholar] [CrossRef]
- Kishore, S.; Gruber, A.R.; Jedlinski, D.J.; Syed, A.P.; Jorjani, H.; Zavolan, M. Insights into snoRNA biogenesis and processing from PAR-CLIP of snoRNA core proteins and small RNA sequencing. Genome Biol. 2013, 14, R45. [Google Scholar] [CrossRef] [Green Version]
- Khanna, A.; Stamm, S. Regulation of alternative splicing by short non-coding nuclear RNAs. RNA Biol. 2010, 7, 480–485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Michel, C.I.; Holley, C.L.; Scruggs, B.S.; Sidhu, R.; Brookheart, R.T.; Listenberger, L.L.; A Behlke, M.; Ory, D.S.; Schaffer, J.E. Small Nucleolar RNAs U32a, U33, and U35a Are Critical Mediators of Metabolic Stress. Cell Metab. 2011, 14, 33–44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, C.; Shi, J.; Guo, Y.; Huang, W.; Huang, S.; Ming, S.; Wu, X.; Zhang, R.; Ding, J.; Zhao, W.; et al. A snoRNA modulates mRNA 3′ end processing and regulates the expression of a subset of mRNAs. Nucleic Acids Res. 2017, 45, 8647–8660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montanaro, L.; Treré, D.; Derenzini, M. Nucleolus, Ribosomes, and Cancer. Am. J. Pathol. 2008, 173, 301–310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jorjani, H.; Kehr, S.; Jedlinski, D.J.; Gumienny, R.; Hertel, J.; Stadler, P.F.; Zavolan, M.; Gruber, A.R. An updated human snoRNAome. Nucleic Acids Res. 2016, 44, 5068–5082. [Google Scholar] [CrossRef]
- Xu, G.; Yang, F.; Ding, C.-L.; Zhao, L.-J.; Ren, H.; Zhao, P.; Wang, W.; Qi, Z. Small nucleolar RNA 113–1 suppresses tumorigenesis in hepatocellular carcinoma. Mol. Cancer 2014, 13, 216. [Google Scholar] [CrossRef] [Green Version]
- Goldring, M.B. The role of cytokines as inflammatory mediators in osteoarthritis: Lessons from animal models. Connect. Tissue Res. 1999, 40, 1–11. [Google Scholar] [CrossRef]
- Kapoor, M.; Martel-Pelletier, J.; Lajeunesse, D.; Pelletier, J.-P.; Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 2011, 7, 33–42. [Google Scholar] [CrossRef]
- Emara, M.M.; Ivanov, P.; Hickman, T.; Dawra, N.; Tisdale, S.; Kedersha, N.; Hu, G.-F.; Anderson, P. Angiogenin-induced tRNA-derived Stress-induced RNAs Promote Stress-induced Stress Granule Assembly. J. Biol. Chem. 2010, 285, 10959–10968. [Google Scholar] [CrossRef] [Green Version]
- Ali, S.A.; Alman, B. RNA extraction from human articular cartilage by chondrocyte isolation. Anal. Biochem. 2012, 429, 39–41. [Google Scholar] [CrossRef]





| Gene/Transcript Name | Gene Biotype | Log2 Fold Change |
|---|---|---|
| eca-miR-143 | miRNA | −1.3 |
| eca-miR-145 | miRNA | −1.8 |
| eca-miR-181b | miRNA | −1.8 |
| eca-miR-122 | miRNA | 2.3 |
| eca-miR-148a | miRNA | 1.3 |
| snora71 | snoRNA | −3.2 |
| snord113 | snoRNA | −2.4 |
| snora46 | snoRNA | 1.0 |
| snora77 | snoRNA | 2.0 |
| snora47 | snoRNA | 2.5 |
| snord29 | snoRNA | 2.5 |
| ECABCGRLG0000003960 | novel snoRNA | −2.6 |
| ECABCGRLG0000002980 | novel snoRNA | −2.5 |
| ECABCGRLG0000006050 | novel snoRNA | −1.6 |
| ECABCGRLG0000000640 | novel snoRNA | −1.1 |
| ECABCGRLG0000002800 | novel snoRNA | −1.0 |
| ECABCGRLG0000007680 | novel snoRNA | 1.8 |
| ECABCGRLG0000008070 | novel snoRNA | 2.0 |
| ECABCGRLG0000007010 | novel snoRNA | 2.5 |
| ECABCGRLG0000008090 | novel snoRNA | 2.9 |
| ECABCGRLG0000004470 | novel snoRNA | 3.0 |
| ECABCGRLG0000005680 | novel snoRNA | 3.1 |
| LOC111775808 | snRNA | −1.3 |
| LOC111773055 | snRNA | 2.2 |
| LOC111772636 | snRNA | 2.5 |
| LOC111770368 | lncRNA | −3.4 |
| LOC111768432 | lncRNA | −3.2 |
| LOC102148414 | lncRNA | −3.2 |
| LOC102149168 | lncRNA | −2.9 |
| LOC111772155 | lncRNA | −2.5 |
| LOC106783307 | lncRNA | −2.5 |
| LOC111775319 | lncRNA | −2.5 |
| LOC102148711 | lncRNA | −2.5 |
| LOC111775759 | lncRNA | −2.5 |
| LOC111774351 | lncRNA | −2.5 |
| LOC102150704 | lncRNA | −2.5 |
| LOC111775994 | lncRNA | −2.5 |
| LOC102150027 | lncRNA | −2.5 |
| LOC106781358 | lncRNA | −2.5 |
| LOC102147393 | lncRNA | −2.5 |
| LOC111776203 | lncRNA | −2.5 |
| LOC111771286 | lncRNA | −1.3 |
| LOC102149893 | lncRNA | 1.7 |
| LOC111775969 | lncRNA | 1.7 |
| LOC106781629 | lncRNA | 1.9 |
| LOC111773181 | lncRNA | 2.0 |
| LOC102149863 | lncRNA | 2.5 |
| LOC102149361 | lncRNA | 2.5 |
| LOC111770630 | lncRNA | 2.5 |
| LOC102147707 | lncRNA | 2.5 |
| LOC106783385 | lncRNA | 2.5 |
| LOC102149569 | lncRNA | 2.5 |
| LOC106782740 | lncRNA | 2.5 |
| LOC111770896 | lncRNA | 2.5 |
| LOC102150024 | lncRNA | 2.9 |
| LOC102150338 | lncRNA | 2.9 |
| TRNAR-ACG | tRNA | −3.4 |
| TRNAR-CCU | tRNA | −3.2 |
| TRNAS-AGA | tRNA | −3.2 |
| TRNAS-GCU | tRNA | −3.1 |
| TRNAA-UGC | tRNA | −3.0 |
| TRNAP-AGG | tRNA | −2.9 |
| TRNAQ-UUG | tRNA | −2.9 |
| TRNAS-AGA | tRNA | −2.9 |
| TRNAA-UGC | tRNA | −2.9 |
| TRNAF-GAA | tRNA | −2.5 |
| TRNAP-UGG | tRNA | −2.5 |
| TRNAE-UUC | tRNA | −2.5 |
| TRNAF-GAA | tRNA | −2.2 |
| TRNAV-AAC | tRNA | −2.2 |
| TRNAM-CAU | tRNA | −2.1 |
| TRNAT-AGU | tRNA | −1.9 |
| TRNAY-GUA | tRNA | −1.9 |
| TRNAQ-CUG | tRNA | −1.4 |
| TRNAN-GUU | tRNA | −1.4 |
| TRNAY-GUA | tRNA | −1.4 |
| TRNAY-GUA | tRNA | −1.2 |
| TRNAV-AAC | tRNA | 1.9 |
| TRNAP-CGG | tRNA | 2.2 |
| TRNAA-UGC | tRNA | 2.4 |
| TRNAC-GCA | tRNA | 2.5 |
| TRNAI-AAU | tRNA | 2.9 |
| TRNAD-GUC | tRNA | 3.4 |
| a. Top Molecular and Cellular Functions | p-Value Range | Number of Molecules |
|---|---|---|
| Cellular Movement | 1.41 × 10−3–4.23 × 10−10 | 21 |
| Cell Morphology | 1.37 × 10−3–3.52 × 10−8 | 16 |
| Cellular Development | 1.42 × 10−3–4.94 × 10−7 | 24 |
| Cell Death and Survival | 1.37 × 10−3–1.08 × 10−5 | 18 |
| Cell-To-Cell Signalling and Interaction | 1.37 × 10−3–1.88 × 10−5 | 20 |
| b. Top Diseases and Disorders | ||
| Skeletal and Muscular Disorders | 1.50 × 10−3–1.22 × 10-9 | 16 |
| Connective Tissue Disorders | 1.50 × 10−3–3.52 × 10-8 | 16 |
| Organismal Injury and Abnormalities | 1.50 × 10−3–3.52 × 10-8 | 31 |
| Cancer | 1.49 × 10−3–5.30 × 10-7 | 31 |
| Developmental Disorder | 1.37 × 10−3–5.30 × 10-7 | 17 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balaskas, P.; Green, J.A.; Haqqi, T.M.; Dyer, P.; Kharaz, Y.A.; Fang, Y.; Liu, X.; Welting, T.J.M.; Peffers, M.J. Small Non-Coding RNAome of Ageing Chondrocytes. Int. J. Mol. Sci. 2020, 21, 5675. https://doi.org/10.3390/ijms21165675
Balaskas P, Green JA, Haqqi TM, Dyer P, Kharaz YA, Fang Y, Liu X, Welting TJM, Peffers MJ. Small Non-Coding RNAome of Ageing Chondrocytes. International Journal of Molecular Sciences. 2020; 21(16):5675. https://doi.org/10.3390/ijms21165675
Chicago/Turabian StyleBalaskas, Panagiotis, Jonathan A. Green, Tariq M. Haqqi, Philip Dyer, Yalda A. Kharaz, Yongxiang Fang, Xuan Liu, Tim J.M. Welting, and Mandy J. Peffers. 2020. "Small Non-Coding RNAome of Ageing Chondrocytes" International Journal of Molecular Sciences 21, no. 16: 5675. https://doi.org/10.3390/ijms21165675






