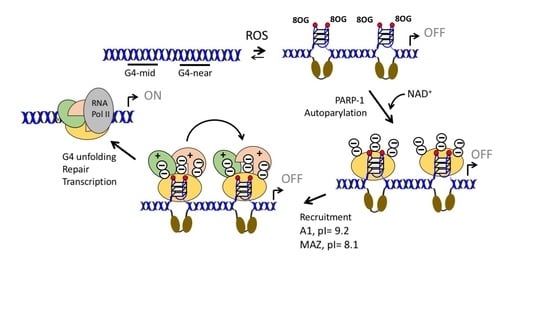
Role of Poly [ADP-ribose] Polymerase 1 in Activating the Kirsten ras (KRAS) Gene in Response to Oxidative Stress
Abstract
1. Introduction
2. Results and Discussion
2.1. G4 Formation in the KRAS Promoter and Guanine Oxidation
2.2. Under Oxidative Stress, PARP-1 Is Recruited to the KRAS Promoter where It Binds to G4 32R and G4 mid1 and G4 mid2
2.3. G4 32R Forms A Multi-Protein Complex with PARP-1, MAZ and hnRNP A1
2.4. PARP-1 Undergoes Autoparylation upon Binding to the KRAS G4
2.5. The Transcription Pre-Initiation Complex Formed at G4 32R Contains Parylated PARP-1
2.6. PARP-1 and G4 Are Essential for KRAS Expression
3. Materials and Methods
3.1. Oligonucleotides
3.2. Cell Culture
3.3. Chromatin Immunoprecipitation and Quantitative PCR
- G4-plus 5′-GTACGCCCGTCTGAAGAAGA-3′ (nucleotides (nt) 4889–4908, 0.2 μM),
- G4-minus 5′-GAGCACACCGATGAGTTCGG-3′ (nt 4958–4977, 0.1 μM),
- Ctr1-plus 5′-ACAAAAAGGTGCTGGGTGAGA-3′ (nt 12–32, 0.2 μM),
- Ctr1-minus 5′-TCCCCTTCCCGGAGACTTAAT-3′ (nt 248–268, 0.2 μM),
- Ctr2-plus 5′-CTCCGACTCTCAGGCTCAAG-3′ (nt 7536–7555, 0.15 μM),
- Ctr2-minus 5′-CAGCACTTTGGGAGGCTTAG-3′ (nt 7692–7711, 0.15 μM).
3.4. Mobility Shift Assays
3.5. Nuclear Extract and Biotin-Streptavidin Pull Down Assay
3.6. Western Blots
3.7. PAR Immunoprecipitation Assay
3.8. PARP-1 AutoPARylation and Inhibition Assays
3.9. Inhibition of KRAS by G4 Decoys, siRNA and Veliparib
3.10. Panc-1 Nuclear Extract
3.11. UV-Melting and CD
3.12. Fluorescence Experiments
3.13. Biotinylated-Anthrathiophenedione Pull-Down Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| KRAS | Kirsten ras gene |
| MAZ | MYC associated zinc finger protein |
| hnRNP A1 | Heterogeneous nuclear ribonucleoprotein A1 |
| PARP-1 | Poly[ADP-ribose] polymerase 1 |
| PDAC | Pancreatic ductal adenocarcinoma |
| 8OG | 8-Oxoguanine |
| TSS | Transcription start site |
| ChIP | Chromatin immunoprecipitation |
| PARylation | Poly(ADP-ribosyl)ation |
References
- Vincent, A.; Herman, J.; Schulick, R.; Hruban, R.H.; Goggins, M. Pancreatic cancer. Lancet 2011, 378, 607–620. [Google Scholar] [CrossRef]
- Collins, M.A.; Bednar, F.; Zhang, Y.; Brisset, J.C.; Galbán, S.; Galbán, C.J.; Rakshit, S.; Flannagan, K.S.; Adsay, N.V.; Pasca Di Magliano, M. Oncogenic Kras is required for both the initiation and maintenance of pancreatic cancer in mice. J. Clin. Investig. 2012, 122, 639–653. [Google Scholar] [CrossRef]
- Ying, H.; Kimmelman, A.C.; Lyssiotis, C.A.; Hua, S.; Chu, G.C.; Fletcher-Sananikone, E.; Locasale, J.W.; Son, J.; Zhang, H.; Coloff, J.L.; et al. Oncogenic kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 2012, 149, 656–670. [Google Scholar] [CrossRef] [PubMed]
- Diehl, F.F.; Lewis, C.A.; Fiske, B.P.; Vander Heiden, M.G. Cellular redox state constrains serine synthesis and nucleotide production to impact cell proliferation. Nat. Metab. 2019, 1, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Son, J.; Lyssiotis, C.A.; Ying, H.; Wang, X.; Hua, S.; Ligorio, M.; Perera, R.M.; Ferrone, C.R.; Mullarky, E.; Shyh-Chang, N.; et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 2013, 496, 101–105. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, I.B.; Joe, A.K. Mechanisms of Disease: Oncogene addiction—A rationale for molecular targeting in cancer therapy. Nat. Clin. Pract. Oncol. 2006, 3, 448–457. [Google Scholar] [CrossRef] [PubMed]
- Pagliarini, R.; Shao, W.; Sellers, W.R. Oncogene addiction: Pathways of therapeutic response, resistance, and road maps toward a cure. EMBO Rep. 2015, 16, 280–296. [Google Scholar] [CrossRef] [PubMed]
- McCormick, F. KRAS as a therapeutic target. Clin. Cancer Res. 2015, 21, 1797–1801. [Google Scholar] [CrossRef] [PubMed]
- Cogoi, S.; Xodo, L.E. G4 DNA in ras genes and its potential in cancer therapy. Biochim. Biophys. Acta 2016, 1859, 663–674. [Google Scholar] [CrossRef]
- Cogoi, S.; Xodo, L.E. G-quadruplex formation within the promoter of the KRAS proto-oncogene and its effect on transcription. Nucleic Acids Res. 2006, 34, 2536–2549. [Google Scholar] [CrossRef]
- Cogoi, S.; Paramasivam, M.; Spolaore, B.; Xodo, L.E. Structural polymorphism within a regulatory element of the human KRAS promoter: Formation of G4-DNA recognized by nuclear proteins. Nucleic Acids Res. 2008, 36, 3765–3780. [Google Scholar] [CrossRef]
- Hänsel-Hertsch, R.; Beraldi, D.; Lensing, S.V.; Marsico, G.; Zyner, K.; Parry, A.; Di Antonio, M.; Pike, J.; Kimura, H.; Narita, M.; et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 2016, 48, 1267–1272. [Google Scholar] [CrossRef]
- Rhodes, D.; Lipps, H.J. Survey and summary G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 2015, 43, 8627–8637. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, J.; Adhikari, S.; Balasubramanian, S. The Structure and Function of DNA G-Quadruplexes. Trends Chem. 2020, 2, 123–136. [Google Scholar] [CrossRef]
- D’Amours, D.; Desnoyers, S.; D’Silva, I.; Poirier, G.G. Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem. J. 1999, 342, 249–268. [Google Scholar] [CrossRef]
- Ji, Y.; Tulin, A.V. The roles of PARP1 in gene control and cell differentiation. Curr. Opin. Genet. Dev. 2010, 20, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Kraus, W.L. Transcriptional control by PARP-1: Chromatin modulation, enhancer-binding, coregulation, and insulation. Curr. Opin. Cell Biol. 2008, 20, 294–302. [Google Scholar] [CrossRef]
- Krishnakumar, R.; Kraus, W.L. The PARP side of the nucleus: Molecular actions, physiological outcomes, and clinical targets. Mol. Cell 2010, 39, 8–24. [Google Scholar] [CrossRef] [PubMed]
- Kraus, W.L.; Hottiger, M.O. PARP-1 and gene regulation: Progress and puzzles. Mol. Asp. Med. 2013, 34, 1109–1123. [Google Scholar] [CrossRef]
- Schreiber, V.; Dantzer, F.; Amé, J.C.; De Murcia, G. Poly(ADP-ribose): Novel functions for an old molecule. Nat. Rev. Mol. Cell Biol. 2006, 7, 517–528. [Google Scholar] [CrossRef]
- Alemasova, E.E.; Lavrik, O.I. Poly(ADP-ribosyl)ation by PARP1: Reaction mechanism and regulatory proteins. Nucleic Acids Res. 2019, 47, 3811–3827. [Google Scholar] [CrossRef]
- Paramasivam, M.; Membrino, A.; Cogoi, S.; Fukuda, H.; Nakagama, H.; Xodo, L.E. Protein hnRNP A1 and its derivative Up1 unfold quadruplex DNA in the human KRAS promoter: Implications for transcription. Nucleic Acids Res. 2009, 37, 2841–2853. [Google Scholar] [CrossRef]
- Cogoi, S.; Zorzet, S.; Rapozzi, V.; Géci, I.; Pedersen, E.B.; Xodo, L.E. MAZ-binding G4-decoy with locked nucleic acid and twisted intercalating nucleic acid modifications suppresses KRAS in pancreatic cancer cells and delays tumor growth in mice. Nucleic Acids Res. 2013, 41, 4049–4064. [Google Scholar] [CrossRef]
- Chu, P.C.; Yang, M.C.; Kulp, S.K.; Salunke, S.B.; Himmel, L.E.; Fang, C.S.; Jadhav, A.M.; Shan, Y.S.; Lee, C.T.; Lai, M.D.; et al. Regulation of oncogenic KRAS signaling via a novel KRAS-integrin-linked kinase-hnRNP A1 regulatory loop in human pancreatic cancer cells. Oncogene 2016, 35, 3897–3908. [Google Scholar] [CrossRef]
- Zhu, X.; Luo, W.; Liang, W.; Tang, F.; Bei, C.; Ren, Y.; Qin, L.; Tan, C.; Zhang, Y.; Tan, S. Overexpression and clinical significance of MYC-associated zinc finger protein in pancreatic carcinoma. Onco Targets Ther. 2016, 9, 7493–7501. [Google Scholar] [CrossRef]
- Cogoi, S.; Shchekotikhin, A.E.; Xodo, L.E. HRAS is silenced by two neighboring G-quadruplexes and activated by MAZ, a zinc-finger transcription factor with DNA unfolding property. Nucleic Acids Res. 2014, 42, 8379–8388. [Google Scholar] [CrossRef]
- Cogoi, S.; Rapozzi, V.; Cauci, S.; Xodo, L.E. Critical role of hnRNP A1 in activating KRAS transcription in pancreatic cancer cells: A molecular mechanism involving G4 DNA. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1389–1398. [Google Scholar] [CrossRef]
- Morgan, R.K.; Batra, H.; Gaerig, V.C.; Hockings, J.; Brooks, T.A. Identification and characterization of a new G-quadruplex forming region within the kRAS promoter as a transcriptional regulator. Biochim. Biophys. Acta 2016, 1859, 235–245. [Google Scholar] [CrossRef]
- Kaiser, C.E.; Van Ert, N.A.; Agrawal, P.; Chawla, R.; Yang, D.; Hurley, L.H. Insight into the Complexity of the i-Motif and G-Quadruplex DNA Structures Formed in the KRAS Promoter and Subsequent Drug-Induced Gene Repression. J. Am. Chem. Soc. 2017, 139, 8522–8536. [Google Scholar] [CrossRef]
- Marquevielle, J.; Robert, C.; Lagrabette, O.; Wahid, M.; Bourdoncle, A.; Xodo, L.E.; Mergny, J.L.; Salgado, G.F. Structure of two G-quadruplexes in equilibrium in the KRAS promoter. Nucleic Acids Res. 2020, gkaa387. [Google Scholar] [CrossRef]
- Cadet, J.; Douki, T.; Ravanat, J.L. Oxidatively generated damage to the guanine moiety of DNA: Mechanistic aspects and formation in cells. Acc. Chem. Res. 2008, 41, 1075–1083. [Google Scholar] [CrossRef] [PubMed]
- Kanvah, S.; Joseph, J.; Schuster, G.B.; Barnett, R.N.; Cleveland, C.L.; Landman, U.Z.I. Oxidation of DNA: Damage to nucleobases. Acc. Chem. Res. 2010, 43, 280–287. [Google Scholar] [CrossRef]
- Loft, S.; Danielsen, P.H.; Mikkelsen, L.; Risom, L.; Forchhammer, L.; Møller, P. Biomarkers of oxidative damage to DNA and repair. Biochem. Soc. Trans. 2008, 36, 1071–1076. [Google Scholar] [CrossRef]
- Ferino, A.; Rapozzi, V.; Xodo, L.E. The ROS-KRAS-Nrf2 axis in the control of the redox homeostasis and the intersection with survival-apoptosis pathways: Implications for photodynamic therapy. J. Photochem. Photobiol. B 2020, 202, 111672. [Google Scholar] [CrossRef]
- Cogoi, S.; Ferino, A.; Miglietta, G.; Pedersen, E.B.; Xodo, L.E. The regulatory G4 motif of the Kirsten ras (KRAS) gene is sensitive to guanine oxidation: Implications on transcription. Nucleic Acids Res. 2018, 46, 661–676. [Google Scholar] [CrossRef]
- Saito, I.; Takayama, M.; Sugiyama, H.; Nakatani, K.; Tsuchida, A.; Yamamoto, M. Photoinduced DNA Cleavage via Electron Transfer: Demonstration That Guanine Residues Located 5′ to Guanine Are the Most Electron-Donating Sites. J. Am. Chem. Soc. 1995, 117, 6406–6407. [Google Scholar] [CrossRef]
- Miglietta, G.; Cogoi, S.; Marinello, J.; Capranico, G.; Tikhomirov, A.S.; Shchekotikhin, A.; Xodo, L.E. RNA G-Quadruplexes in Kirsten Ras (KRAS) Oncogene as Targets for Small Molecules Inhibiting Translation. J. Med. Chem. 2017, 60, 9448–9461. [Google Scholar] [CrossRef]
- Ding, Y.; Fleming, A.M.; Burrows, C.J. Sequencing the Mouse Genome for the Oxidatively Modified Base 8-Oxo-7,8-dihydroguanine by OG-Seq. J. Am. Chem. Soc. 2017, 139, 2569–2572. [Google Scholar] [CrossRef]
- Clark, D.W.; Phang, T.; Edwards, M.G.; Geraci, M.W.; Gillespie, M.N. Promoter G-quadruplex sequences are targets for base oxidation and strand cleavage during hypoxia-induced transcription. Free Radic. Biol. Med. 2012, 53, 51–59. [Google Scholar] [CrossRef]
- Vorlícková, M.; Tomasko, M.; Sagi, A.J.; Bednarova, K.; Sagi, J. 8-Oxoguanine in a quadruplex of the human telomere DNA sequence. FEBS J. 2012, 279, 29–39. [Google Scholar] [CrossRef]
- Beckett, J.; Burns, J.; Broxson, C.; Tornaletti, S. Spontaneous DNA lesions modulate DNA structural transitions occurring at nuclease hypersensitive element III1 of the human c-myc proto-oncogene. Biochemistry 2012, 51, 5257–5268. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.J.; Kotova, E.; Andrake, M.; Adolf-Bryfogle, J.; Glaser, R.; Regnard, C.; Tulin, A.V. Kinase-Mediated Changes in Nucleosome Conformation Trigger Chromatin Decondensation via Poly(ADP-Ribosyl)ation. Mol. Cell 2014, 53, 831–842. [Google Scholar] [CrossRef] [PubMed]
- Kraus, W.L.; Lis, J.T. PARP goes transcription. Cell 2003, 113, 677–683. [Google Scholar] [CrossRef]
- Soldatenkov, V.A.; Vetcher, A.A.; Duka, T.; Ladame, S. First evidence of a functional interaction between DNA quadruplexes and poly(ADP-ribose) polymerase-1. ACS Chem. Biol. 2008, 3, 214–219. [Google Scholar] [CrossRef]
- Eustermann, S.; Wu, W.F.; Langelier, M.F.; Yang, J.C.; Easton, L.E.; Riccio, A.A.; Pascal, J.M.; Neuhaus, D. Structural Basis of Detection and Signaling of DNA Single-Strand Breaks by Human PARP-1. Mol. Cell 2015, 60, 742–754. [Google Scholar] [CrossRef] [PubMed]
- Eustermann, S.; Videler, H.; Yang, J.C.; Cole, P.T.; Gruszka, D.; Veprintsev, D.; Neuhaus, D. The DNA-binding domain of human PARP-1 interacts with DNA single-strand breaks as a monomer through its second zinc finger. J. Mol. Biol. 2011, 407, 149–170. [Google Scholar] [CrossRef]
- Dawicki-McKenna, J.M.; Langelier, M.F.; De Nizio, J.E.; Riccio, A.A.; Cao, C.D.; Karch, K.R.; McCauley, M.; Steffen, J.D.; Black, B.E.; Pascal, J.M. PARP-1 Activation Requires Local Unfolding of an Autoinhibitory Domain. Mol. Cell 2015, 60, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Gupte, R.; Liu, Z.; Kraus, W.L. Parps and adp-ribosylation: Recent advances linking molecular functions to biological outcomes. Genes Dev. 2017, 31, 101–126. [Google Scholar] [CrossRef] [PubMed]
- Langelier, M.F.; Eisemann, T.; Riccio, A.A.; Pascal, J.M. PARP family enzymes: Regulation and catalysis of the poly(ADP-ribose) posttranslational modification. Curr. Opin. Struct. Biol. 2018, 53, 187–198. [Google Scholar] [CrossRef]
- Jain, P.G.; Patel, B.D. Medicinal chemistry approaches of poly ADP-Ribose polymerase 1 (PARP1) inhibitors as anticancer agents—A recent update. Eur. J. Med. Chem. 2019, 165, 198–215. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Ding, Y.; Burrows, C.J. Oxidative DNA damage is epigenetic by regulating gene transcription via base excision repair. Proc. Natl. Acad. Sci. USA 2017, 114, 2604–2609. [Google Scholar] [CrossRef] [PubMed]
- Ba, X.; Boldogh, I. 8-oxoguanine DNA glycosylase 1: Beyond repair of oxidatively modified base lesion. Redox Biol. 2018, 14, 669–678. [Google Scholar] [CrossRef] [PubMed]
- Giorgio, M.; Dellino, G.I.; Gambino, V.; Roda, N.; Pelicci, P.G. On the epigenetic role of guanosine oxidation. Redox Biol. 2020, 29, 101398. [Google Scholar] [CrossRef] [PubMed]
- Gibson, B.A.; Zhang, Y.; Jiang, H.; Hussey, K.M.; Shrimp, J.M.; Lin, H.; Schwede, F.; Yu, Y.; Kraus, W.L. Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation. Science 2016, 353, 45–50. [Google Scholar] [CrossRef]
- Gunderson, C.C.; Moore, K.N. Olaparib: An oral PARP-1 and PARP-2 inhibitor with promising activity in ovarian cancer. Future Oncol. 2015, 11, 747–757. [Google Scholar] [CrossRef]
- Bodepudi, V.; Shibutani, S.; Johnson, F. Synthesis of 2′-Deoxy-7,8-dihydro-8-oxoguanosine and 2′-Deoxy-7,8-dihydro-8-oxoadenosine and Their Incorporation into Oligomeric DNA. Chem. Res. Toxicol. 1992, 5, 608–617. [Google Scholar] [CrossRef]
- Markwell, M.A.K.; Haas, S.M.; Bieber, L.L.; Tolbert, N.E. A modification of the Lowry procedure to simplify protein determination in membrane and lipoprotein samples. Anal. Biochem. 1978, 87, 206–210. [Google Scholar] [CrossRef]
- Fleming, A.M.; Burrows, C.J. Interplay of guanineoxidation and G-quadruplex folding in gene promoters. J. Am. Chem. Soc. 2020, 142, 1115–1136. [Google Scholar] [CrossRef]
- Sajadimajd, S.; Khazaei, M. Oxidative Stress and Cancer: The Role of Nrf2. Curr. Cancer Drug Targets 2018, 18, 538–557. [Google Scholar] [CrossRef]
- Lee, S.B.; Sellers, B.N.; DeNicola, G.M. The Regulation of NRF2 by Nutrient-Responsive Signaling and Its Role in Anabolic Cancer Metabolism. Antioxid. Redox Signal. 2018, 29, 1774–1791. [Google Scholar] [CrossRef]
- Krishnakumar, R.; Kraus, W.L. PARP-1 Regulates Chromatin Structure and Transcription through a KDM5B-Dependent Pathway. Mol. Cell 2010, 39, 736–749. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Yadav, V.K.; Baral, A.; Kumar, P.; Saha, D.; Chowdhury, S. Zinc-finger transcription factors are associated with guanine quadruplex motifs in human, chimpanzee, mouse and rat promoters genome-wide. Nucleic Acids Res. 2011, 39, 8005–8016. [Google Scholar] [CrossRef] [PubMed]










| Oligonucleotide | 5′ → 3′ | 8OG |
|---|---|---|
| G4 32R | AGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG | |
| 92 | AGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG (1) (3) | 1 |
| 96 | AGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG (1) (3) | 2 |
| G4 mid | CGGGGAGAAGGAGGGGGCCGGGCCGGGCCGGCGGGGGAGGAGCGGGGGCCGGGC | |
| G4 mid1 | CGGGGAGAAGGAGGGGGCCGGGCCGGGC | |
| G4 mid2 | CGGGCCGGCGGGGGAGGAGCGGGGGCCGGGC | |
| b-32R | b-TTTTAGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG (2) | |
| b-92 | b-TTTTAGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG (1) (2) (3) | 1 |
| b-96 | b-TTTTAGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG (1) (2) (3) | 2 |
| b-mid | b-TTTTCGGGGAGAAGGAGGGGGCCGGGCCGGGCCGGCG--GGGGAGGAGCGGGGGCCGGGC (2) | |
| b-mid1 | b-TTTTCGGGGAGAAGGAGGGGGCCGGGCCGGGC (2) | |
| b-mid2 | b-TTTTCGGGCCGGCGGGGGAGGAGCGGGGGCCGGG (2) | |
| b-midOX | b-TTTTCGGGGAGAAGGAGGGGGCCGGGCCGGGCCGGCG- -GGGGAGGAGCGGGGGCCGGGC (1) (2) | 2 |
| b-mid1OX | b-TTTTCGGGGAGAAGGAGGGGGCCGGGCCGGGC (1) (2) | 1 |
| b-mid2OX | b-TTTTCGGGCCGGCGGGGGAGGAGCGGGGGCCGGGC (1) (2) | 2 |
| Cy5.5-32R | Cy5.5-AGGGCGGTGTGGGAAGAGGGAAGAGGGGGAGG | |
| Cy5.5-mid1 | Cy5.5-CGGGGAGAAGGAGGGGGCCGGGCCGGGC | |
| Cy5.5-mid2 | Cy5.5-CGGGCCGGCGGGGGAGGAGCGGGGGCCGGGC | |
| Cy5.5-mid | Cy5.5-CGGGGAGAAGGAGGGGGCCGGGCCGGGCCGGCGGGGGAGGAGCGGGGGCCGGGC | |
| 32Y | CCTCCCCCTCTTCCCTCTTCCCACACCGCCCT | |
| G4 mid Y | GCCCGGCCCCCGCTCCTCCCCCGCCGGCCCGGCCCGGCCCCCTCCTTCTCCCCG | |
| Oligo dT | TTTTTTTTTTTTTTTT | |
| ODN1 | Cy5.5-CATCAGAAGGCTAGCAATCA | |
| ODN2 | Cy5.5-AATAGTAATTGCTTAGCCTG | |
| ODN3 | Cy5.5-CCTAATGCTGCTAAACTCCC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cinque, G.; Ferino, A.; Pedersen, E.B.; Xodo, L.E. Role of Poly [ADP-ribose] Polymerase 1 in Activating the Kirsten ras (KRAS) Gene in Response to Oxidative Stress. Int. J. Mol. Sci. 2020, 21, 6237. https://doi.org/10.3390/ijms21176237
Cinque G, Ferino A, Pedersen EB, Xodo LE. Role of Poly [ADP-ribose] Polymerase 1 in Activating the Kirsten ras (KRAS) Gene in Response to Oxidative Stress. International Journal of Molecular Sciences. 2020; 21(17):6237. https://doi.org/10.3390/ijms21176237
Chicago/Turabian StyleCinque, Giorgio, Annalisa Ferino, Erik B. Pedersen, and Luigi E. Xodo. 2020. "Role of Poly [ADP-ribose] Polymerase 1 in Activating the Kirsten ras (KRAS) Gene in Response to Oxidative Stress" International Journal of Molecular Sciences 21, no. 17: 6237. https://doi.org/10.3390/ijms21176237
APA StyleCinque, G., Ferino, A., Pedersen, E. B., & Xodo, L. E. (2020). Role of Poly [ADP-ribose] Polymerase 1 in Activating the Kirsten ras (KRAS) Gene in Response to Oxidative Stress. International Journal of Molecular Sciences, 21(17), 6237. https://doi.org/10.3390/ijms21176237