Transcriptome of the Aedes aegypti Mosquito in Response to Human Complement Proteins
Abstract
:1. Introduction
2. Results
2.1. Sequencing and Read Mapping
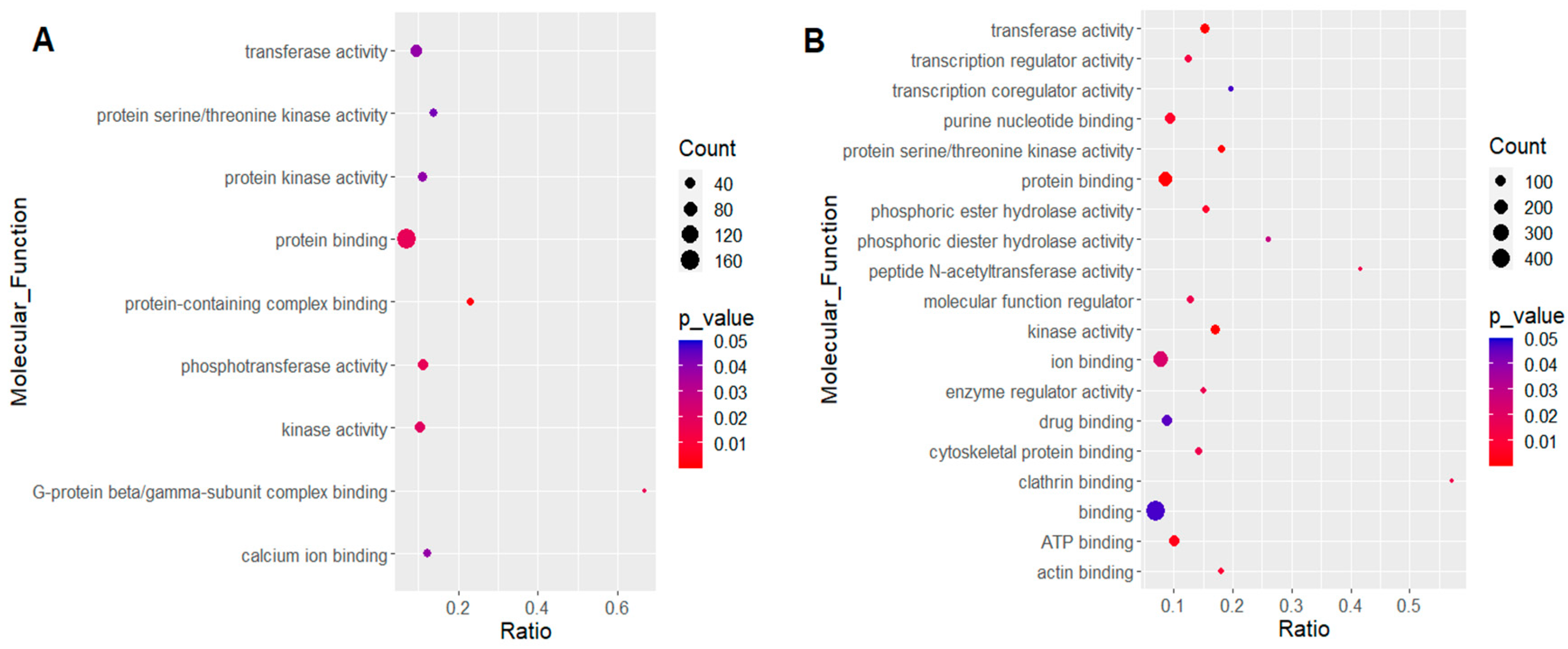
2.2. Transcript Differential Expression in NB and IB Fed Mosquitoes (In Vivo Experiments)
2.3. Transcript Differential Expression in Aag2 Cells upon Treatment with hC3 and hC5a (In Vitro Experiments)
2.4. Changes in the Immune Response Transcripts (IRTs) to Complement in the In Vivo and In Vitro Experiments
2.5. Shared Transcripts between the In Vivo and the In Vitro Assays
3. Discussion
4. Materials and Methods
4.1. Blood Source
4.2. Maintenance of Ae. aegypti Adult Mosquitoes and Cell Line
4.3. Mosquito Blood-Feeding
4.4. Aag2 Cells Exposure to Human Complement Protein hC3 and hC5a
4.5. Mosquito RNA Isolation and Sequencing
4.6. RNA-Seq Differential Expression Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| YFV | Yellow fever virus |
| DENV | Dengue virus |
| CHIKV | Chikungunya virus |
| ZIKV | Zika virus |
| MAC | Membrane attack complex |
| hC3 | Human complement protein C3 |
| hC5a | Human complement protein C5a |
| AaMCR | Ae. aegypti macroglobulin complement-related proteins |
| AMP | Antimicrobial peptides |
| TBV | Transmission blocking vaccines |
| LVP_AGWG | Liverpool Aedes Genome Working Group |
| STAR | Spliced Transcripts Alignment to a Reference |
| RSEM | RNA-Seq by Expectation-Maximization |
| FDR | False discovery rate |
| CPM | Counts per Million |
| TMM | Trimmed Mean of M-values |
| DE | Differentially expressed |
| IB | Blood with heat-inactivated plasma |
| NB | Blood with normal plasma |
| SF | Sugar-fed |
| FC | Fold change |
| GSEA | Gene Set Enrichment Analysis |
| MF | Molecular functions |
| GO | Gene Ontology |
| IRT | Immune response transcripts |
| 5G1 | Late-phase trypsin |
| ACA | Anticomplementary activity |
| C5aR1 | C5a receptor 1 (CD88) |
| C5aR2 | C5a receptor 2 (C5L2) |
| GTP | Guanosine-5’-triphosphate |
| EDTA | Ethylenediaminetetraacetic acid |
| RBC | Red blood cells |
| PBS | Phosphate Buffered Saline |
| BME | Basal Medium Eagle |
| KOAPF | K-State Open Access Publishing Fund |
References
- Powell, J.R. Mosquito-Borne Human Viral Diseases: Why Aedes aegypti? Am. J. Trop. Med. Hyg. 2018, 98, 1563–1565. [Google Scholar] [CrossRef] [PubMed]
- Londono-Renteria, B.; Troupin, A.; Colpitts, T.M. Arbovirosis and potential transmission blocking vaccines. Parasites Vectors 2016, 9, 516. [Google Scholar] [CrossRef] [Green Version]
- Alonso-Palomares, L.A.; Moreno-Garcia, M.; Lanz-Mendoza, H.; Salazar, M.I. Molecular Basis for Arbovirus Transmission by Aedes aegypti Mosquitoes. Intervirology 2018, 61, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Franz, A.W.; Kantor, A.M.; Passarelli, A.L.; Clem, R.J. Tissue Barriers to Arbovirus Infection in Mosquitoes. Viruses 2015, 7, 3741–3767. [Google Scholar] [CrossRef] [PubMed]
- Brackney, D.E.; Foy, B.D.; Olson, K.E. The effects of midgut serine proteases on dengue virus type 2 infectivity of Aedes aegypti. Am. J. Trop. Med. Hyg. 2008, 79, 267–274. [Google Scholar] [CrossRef] [Green Version]
- Raquin, V.; Lambrechts, L. Dengue virus replicates and accumulates in Aedes aegypti salivary glands. Virology 2017, 507, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Pakpour, N.; Riehle, M.A.; Luckhart, S. Effects of ingested vertebrate-derived factors on insect immune responses. Curr. Opin. Insect Sci. 2014, 3, 1–5. [Google Scholar] [CrossRef] [Green Version]
- Londono-Renteria, B.; Grippin, C.; Cardenas, J.C.; Troupin, A.; Colpitts, T.M. Human C5a Protein Participates in the Mosquito Immune Response Against Dengue Virus. J. Med. Entomol. 2016, 53, 505–512. [Google Scholar] [CrossRef] [Green Version]
- Tsuboi, T.; Cao, Y.M.; Torii, M.; Hitsumoto, Y.; Kanbara, H. Murine complement reduces infectivity of Plasmodium yoelii to mosquitoes. Infect. Immun. 1995, 63, 3702–3704. [Google Scholar] [CrossRef] [Green Version]
- Margos, G.; Navarette, S.; Butcher, G.; Davies, A.; Willers, C.; Sinden, R.E.; Lachmann, P.J. Interaction between host complement and mosquito-midgut-stage Plasmodium berghei. Infect. Immun. 2001, 69, 5064–5071. [Google Scholar] [CrossRef] [Green Version]
- Cummings, K.L.; Waggoner, S.N.; Tacke, R.; Hahn, Y.S. Role of complement in immune regulation and its exploitation by virus. Viral Immunol. 2007, 20, 505–524. [Google Scholar] [CrossRef] [PubMed]
- Stoermer, K.A.; Morrison, T.E. Complement and viral pathogenesis. Virology 2011, 411, 362–373. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dunkelberger, J.R.; Song, W.C. Complement and its role in innate and adaptive immune responses. Cell Res. 2010, 20, 34–50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mellors, J.; Tipton, T.; Longet, S.; Carroll, M. Viral Evasion of the Complement System and Its Importance for Vaccines and Therapeutics. Front. Immunol. 2020, 11, 1450. [Google Scholar] [CrossRef] [PubMed]
- Fujita, T.; Matsushita, M.; Endo, Y. The lectin-complement pathway--its role in innate immunity and evolution. Immunol. Rev. 2004, 198, 185–202. [Google Scholar] [CrossRef]
- Sahu, A.; Lambris, J.D. Structure and biology of complement protein C3, a connecting link between innate and acquired immunity. Immunol. Rev. 2001, 180, 35–48. [Google Scholar] [CrossRef]
- Ruseva, M.M.; Heurich, M. Purification and characterization of human and mouse complement C3. Methods Mol. Biol. 2014, 1100, 75–91. [Google Scholar]
- Guo, R.F.; Ward, P.A. Role of C5a in inflammatory responses. Ann. Rev. Immunol. 2005, 23, 821–852. [Google Scholar] [CrossRef]
- Avirutnan, P.; Punyadee, N.; Noisakran, S.; Komoltri, C.; Thiemmeca, S.; Auethavornanan, K.; Jairungsri, A.; Kanlaya, R.; Tangthawornchaikul, N.; Puttikhunt, C.; et al. Vascular leakage in severe dengue virus infections: A potential role for the nonstructural viral protein NS1 and complement. J. Infect. Dis. 2006, 193, 1078–1088. [Google Scholar] [CrossRef] [Green Version]
- Conde, J.N.; Silva, E.M.; Barbosa, A.S.; Mohana-Borges, R. The Complement System in Flavivirus Infections. Front. Microbiol. 2017, 8, 213. [Google Scholar] [CrossRef]
- Nielsen, H.; Svehag, S.R. Detection and differentiation of immune complexes and IgG aggregates by a complement consumption assay. Acta Pathol. Microbiol. Scand. 1976, 84, 261–269. [Google Scholar] [CrossRef] [PubMed]
- Chatchen, S.; Sabchareon, A.; Sirivichayakul, C. Serodiagnosis of asymptomatic dengue infection. Asian Pac. J. Trop. Med. 2017, 10, 11–14. [Google Scholar] [CrossRef] [PubMed]
- Yeo, A.S.; Azhar, N.A.; Yeow, W.; Talbot, C.C., Jr.; Khan, M.A.; Shankar, E.M.; Rathakrishnan, A.; Azizan, A.; Wang, S.M.; Lee, S.K.; et al. Lack of clinical manifestations in asymptomatic dengue infection is attributed to broad down-regulation and selective up-regulation of host defence response genes. PLoS ONE 2014, 9, e92240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cabezas, S.; Bracho, G.; Aloia, A.L.; Adamson, P.J.; Bonder, C.S.; Smith, J.R.; Gordon, D.L.; Carr, J.M. Dengue Virus Induces Increased Activity of the Complement Alternative Pathway in Infected Cells. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [Green Version]
- Shresta, S. Role of complement in dengue virus infection: Protection or pathogenesis? mBio 2012, 3. [Google Scholar] [CrossRef] [Green Version]
- Nascimento, E.J.; Silva, A.M.; Cordeiro, M.T.; Brito, C.A.; Gil, L.H.; Braga-Neto, U.; Marques, E.T. Alternative complement pathway deregulation is correlated with dengue severity. PLoS ONE 2009, 4, e6782. [Google Scholar] [CrossRef]
- Duong, V.; Lambrechts, L.; Paul, R.E.; Ly, S.; Lay, R.S.; Long, K.C.; Huy, R.; Tarantola, A.; Scott, T.W.; Sakuntabhai, A.; et al. Asymptomatic humans transmit dengue virus to mosquitoes. Proc. Natl. Acad. Sci. USA 2015, 112, 14688–14693. [Google Scholar] [CrossRef] [Green Version]
- Xiao, X.; Liu, Y.; Zhang, X.; Wang, J.; Li, Z.; Pang, X.; Wang, P.; Cheng, G. Complement-related proteins control the flavivirus infection of Aedes aegypti by inducing antimicrobial peptides. PLoS Pathog. 2014, 10, e1004027. [Google Scholar] [CrossRef] [Green Version]
- Pereira-Filho, A.A.; Mateus Pereira, R.H.; da Silva, N.C.S.; Ferreira Malta, L.G.; Serravite, A.M.; Carvalho de Almeida, C.G.; Fujiwara, R.T.; Bartholomeu, D.C.; Giunchetti, R.C.; D’Avila Pessoa, G.C.; et al. The gut anti-complement activity of Aedes aegypti: Investigating new ways to control the major human arboviruses vector in the Americas. Insect Biochem. Mol. Biol. 2020, 120, 103338. [Google Scholar] [CrossRef]
- Molina-Cruz, A.; Gupta, L.; Richardson, J.; Bennett, K.; 4th Black, W.; Barillas-Mury, C. Effect of mosquito midgut trypsin activity on dengue-2 virus infection and dissemination in Aedes aegypti. Am. J. Trop. Med. Hyg. 2005, 72, 631–637. [Google Scholar] [CrossRef] [Green Version]
- Bottino-Rojas, V.; Talyuli, O.A.; Jupatanakul, N.; Sim, S.; Dimopoulos, G.; Venancio, T.M.; Bahia, A.C.; Sorgine, M.H.; Oliveira, P.L.; Paiva-Silva, G.O. Heme Signaling Impacts Global Gene Expression, Immunity and Dengue Virus Infectivity in Aedes aegypti. PLoS ONE 2015, 10, e0135985. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oliveira, J.H.; Goncalves, R.L.; Lara, F.A.; Dias, F.A.; Gandara, A.C.; Menna-Barreto, R.F.; Edwards, M.C.; Laurindo, F.R.; Silva-Neto, M.A.; Sorgine, M.H.; et al. Blood meal-derived heme decreases ROS levels in the midgut of Aedes aegypti and allows proliferation of intestinal microbiota. PLoS Pathog. 2011, 7, e1001320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mankovich, A.R.; Lee, C.Y.; Heinrich, V. Differential effects of serum heat treatment on chemotaxis and phagocytosis by human neutrophils. PLoS ONE 2013, 8, e54735. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Noguchi, H. The Thermostabile Anticomplementary Constituents of the Blood. J. Exp. Med. 1906, 8, 726–753. [Google Scholar] [CrossRef]
- Soltis, R.D.; Hasz, D.; Morris, M.J.; Wilson, I.D. The effect of heat inactivation of serum on aggregation of immunoglobulins. Immunology 1979, 36, 37–45. [Google Scholar]
- Urban, S.; Dickey, S.W. The rhomboid protease family: A decade of progress on function and mechanism. Genome Biol. 2011, 12, 231. [Google Scholar] [CrossRef] [Green Version]
- McQuibban, G.A.; Lee, J.R.; Zheng, L.; Juusola, M.; Freeman, M. Normal mitochondrial dynamics requires rhomboid-7 and affects Drosophila lifespan and neuronal function. Curr. Biol. 2006, 16, 982–989. [Google Scholar] [CrossRef] [Green Version]
- Nene, V.; Wortman, J.R.; Lawson, D.; Haas, B.; Kodira, C.; Tu, Z.J.; Loftus, B.; Xi, Z.; Megy, K.; Grabherr, M.; et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 2007, 316, 1718–1723. [Google Scholar] [CrossRef] [Green Version]
- Ramirez, J.L.; Souza-Neto, J.; Torres Cosme, R.; Rovira, J.; Ortiz, A.; Pascale, J.M.; Dimopoulos, G. Reciprocal tripartite interactions between the Aedes aegypti midgut microbiota, innate immune system and dengue virus influences vector competence. PLoS Negl. Trop. Dis. 2012, 6, e1561. [Google Scholar] [CrossRef]
- Lowenberger, C.; Charlet, M.; Vizioli, J.; Kamal, S.; Richman, A.; Christensen, B.M.; Bulet, P. Antimicrobial activity spectrum, cDNA cloning, and mRNA expression of a newly isolated member of the cecropin family from the mosquito vector Aedes aegypti. J. Biol. Chem. 1999, 274, 20092–20097. [Google Scholar] [CrossRef] [Green Version]
- Hillyer, J.F.; Schmidt, S.L.; Fuchs, J.F.; Boyle, J.P.; Christensen, B.M. Age-associated mortality in immune challenged mosquitoes (Aedes aegypti) correlates with a decrease in haemocyte numbers. Cell Microbiol. 2005, 7, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Souza-Neto, J.A.; Sim, S.; Dimopoulos, G. An evolutionary conserved function of the JAK-STAT pathway in anti-dengue defense. Proc. Natl. Acad. Sci. USA 2009, 106, 17841–17846. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, Y.; Morton, J.C., Jr.; Ramirez, J.L.; Souza-Neto, J.A.; Dimopoulos, G. The entomopathogenic fungus Beauveria bassiana activate toll and JAK-STAT pathway-controlled effector genes and anti-dengue activity in Aedes aegypti. Insect Biochem. Mol. Biol. 2012, 42, 126–132. [Google Scholar] [CrossRef] [Green Version]
- Ricklin, D.; Reis, E.S.; Mastellos, D.C.; Gros, P.; Lambris, J.D. Complement component C3—The “Swiss Army Knife” of innate immunity and host defense. Immunol. Rev. 2016, 274, 33–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merle, N.S.; Paule, R.; Leon, J.; Daugan, M.; Robe-Rybkine, T.; Poillerat, V.; Torset, C.; Fremeaux-Bacchi, V.; Dimitrov, J.D.; Roumenina, L.T. P-selectin drives complement attack on endothelium during intravascular hemolysis in TLR-4/heme-dependent manner. Proc. Natl. Acad. Sci. USA 2019, 116, 6280–6285. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schoengraf, P.; Lambris, J.D.; Recknagel, S.; Kreja, L.; Liedert, A.; Brenner, R.E.; Huber-Lang, M.; Ignatius, A. Does complement play a role in bone development and regeneration? Immunobiology 2013, 218, 1–9. [Google Scholar] [CrossRef]
- Klos, A.; Tenner, A.J.; Johswich, K.O.; Ager, R.R.; Reis, E.S.; Kohl, J. The role of the anaphylatoxins in health and disease. Mol. Immunol. 2009, 46, 2753–2766. [Google Scholar] [CrossRef] [Green Version]
- Huber-Lang, M.; Kovtun, A.; Ignatius, A. The role of complement in trauma and fracture healing. Semin. Immunol. 2013, 25, 73–78. [Google Scholar] [CrossRef] [Green Version]
- Wiegner, R.; Chakraborty, S.; Huber-Lang, M. Complement-coagulation crosstalk on cellular and artificial surfaces. Immunobiology 2016, 221, 1073–1079. [Google Scholar] [CrossRef]
- Cui, Y.; Grant, D.G.; Lin, J.; Yu, X.; Franz, A.W.E. Zika Virus Dissemination from the Midgut of Aedes aegypti is Facilitated by Bloodmeal-Mediated Structural Modification of the Midgut Basal Lamina. Viruses 2019, 11, 1056. [Google Scholar] [CrossRef] [Green Version]
- Dong, S.; Balaraman, V.; Kantor, A.M.; Lin, J.; Grant, D.G.; Held, N.L.; Franz, A.W.E. Chikungunya virus dissemination from the midgut of Aedes aegypti is associated with temporal basal lamina degradation during bloodmeal digestion. PLoS Negl. Trop. Dis. 2017, 11, e0005976. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barletta, A.B.; Silva, M.C.; Sorgine, M.H. Validation of Aedes aegypti Aag-2 cells as a model for insect immune studies. Parasites Vectors 2012, 5, 148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whistler, J.L.; Gerber, B.O.; Meng, E.C.; Baranski, T.J.; von Zastrow, M.; Bourne, H.R. Constitutive activation and endocytosis of the complement factor 5a receptor: Evidence for multiple activated conformations of a G protein-coupled receptor. Traffic 2002, 3, 866–877. [Google Scholar] [CrossRef] [PubMed]
- Burg, M.; Martin, U.; Bock, D.; Rheinheimer, C.; Kohl, J.; Bautsch, W.; Klos, A. Differential regulation of the C3a and C5a receptors (CD88) by IFN-gamma and PMA in U937 cells and related myeloblastic cell lines. J. Immunol. 1996, 157, 5574–5581. [Google Scholar]
- Tam, J.C.; Bidgood, S.R.; McEwan, W.A.; James, L.C. Intracellular sensing of complement C3 activates cell autonomous immunity. Science 2014, 345, 1256070. [Google Scholar] [CrossRef] [Green Version]
- Liszewski, M.K.; Kolev, M.; Le Friec, G.; Leung, M.; Bertram, P.G.; Fara, A.F.; Subias, M.; Pickering, M.C.; Drouet, C.; Meri, S.; et al. Intracellular complement activation sustains T cell homeostasis and mediates effector differentiation. Immunity 2013, 39, 1143–1157. [Google Scholar] [CrossRef] [Green Version]
- Baudino, L.; Sardini, A.; Ruseva, M.M.; Fossati-Jimack, L.; Cook, H.T.; Scott, D.; Simpson, E.; Botto, M. C3 opsonization regulates endocytic handling of apoptotic cells resulting in enhanced T-cell responses to cargo-derived antigens. Proc. Natl. Acad. Sci. USA 2014, 111, 1503–1508. [Google Scholar] [CrossRef] [Green Version]
- Walker, T.; Jeffries, C.L.; Mansfield, K.L.; Johnson, N. Mosquito cell lines: History, isolation, availability and application to assess the threat of arboviral transmission in the United Kingdom. Parasites Vectors 2014, 7, 382. [Google Scholar] [CrossRef] [Green Version]
- Giraldo-Calderon, G.I.; Emrich, S.J.; MacCallum, R.M.; Maslen, G.; Dialynas, E.; Topalis, P.; Ho, N.; Gesing, S.; VectorBase, C.; Madey, G.; et al. VectorBase: An updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res. 2015, 43, D707–D713. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tarazona, S.; Furio-Tari, P.; Turra, D.; Pietro, A.D.; Nueda, M.J.; Ferrer, A.; Conesa, A. Data quality aware analysis of differential expression in RNA-seq with NOISeq R/Bioc package. Nucleic Acids Res. 2015, 43, e140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Durinck, S.; Spellman, P.T.; Birney, E.; Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 2009, 4, 1184–1191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Durinck, S.; Moreau, Y.; Kasprzyk, A.; Davis, S.; De Moor, B.; Brazma, A.; Huber, W. BioMart and Bioconductor: A powerful link between biological databases and microarray data analysis. Bioinformatics 2005, 21, 3439–3440. [Google Scholar] [CrossRef] [Green Version]
- Bonizzoni, M.; Dunn, W.A.; Campbell, C.L.; Olson, K.E.; Dimon, M.T.; Marinotti, O.; James, A.A. RNA-seq analyses of blood-induced changes in gene expression in the mosquito vector species, Aedes aegypti. BMC Genom. 2011, 12, 82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maag, J.L.V. gganatogram: An R package for modular visualisation of anatograms and tissues based on ggplot2. F1000Research 2018, 7, 1576. [Google Scholar] [CrossRef]
- Ito, K.; Murphy, D. Application of ggplot2 to Pharmacometric Graphics. CPT Pharmacomet. Syst. Pharmacol. 2013, 2, e79. [Google Scholar] [CrossRef]





| Transcript_ID | Gene Description (Gene Ontology Molecular Function and Description) | Log2FC | ||
|---|---|---|---|---|
| hC3/Ctl | hC5a/Ctl | NB/IB | ||
| AAEL001698-RB | Charged multivesicular body protein 4b (GO:0007034, vacuolar transport) | 2.80 | 2.06 | 5.46 |
| AAEL007290-RE | None (None) | 2.93 | 2.90 | 3.06 |
| AAEL001447-RG | None | 3.40 | 4.26 | 3.91 |
| AAEL002557-RF | Cationic amino acid transporter (GO:0022857, transmembrane transporter activity) | 3.41 | 2.91 | 5.27 |
| AAEL005460-RD | alpha-galactosidase/alpha-n-acetylgalactosaminidase (GO:0003824;GO:0016787;GO:0016798;GO:0004553, catalytic activity; hydrolase activity; hydrolase activity, acting on glycosyl bonds) | 3.78 | 3.75 | 3.54 |
| AAEL025517-RB | None (GO:0005524;GO:0004672, ATP binding; protein kinase activity) | 3.79 | 3.47 | 5.92 |
| AAEL012899-RB | Meiotic coiled-coil protein, putative GO:0003690, double-stranded DNA binding) | 3.96 | 3.58 | 6.30 |
| AAEL026355-RA | None (GO:0005515, protein binding) | 4.25 | 3.23 | 4.85 |
| AAEL008574-RB | Acyl-CoA oxidase (GO:0071949;GO:0003997;GO:0050660;GO:0016627, FAD binding; acyl-CoA oxidase activity; flavin adenine dinucleotide binding) | 4.45 | 4.36 | 5.09 |
| AAEL009249-RC | Coronin (GO:0005515, protein binding) | 4.58 | 5.13 | 4.97 |
| AAEL009212-RP | Lola (GO:0005515, protein binding) | 4.86 | 4.88 | 6.08 |
| AAEL007845-RF | Rab5 (GO:0005525;GO:0003924, GTP binding; GTPase activity) | 4.88 | 5.16 | 6.89 |
| AAEL001557-RC | Mucolipin (GO:0005261, cation channel activity) | 4.89 | 2.70 | 5.76 |
| AAEL005022-RB | Mediator of RNA polymerase II transcription subunit 15 (Med15) (GO:0003712, transcription coregulator activity) | 5.01 | 5.26 | 8.12 |
| AAEL003762-RD | Protoporphyrinogen IX oxidase (GO:0016491;GO:0004729, oxidoreductase activity; oxygen-dependent protoporphyrinogen oxidase activity) | 5.15 | 6.23 | 3.13 |
| AAEL015598-RA | None (GO:0016787, hydrolase activity) | 5.21 | 2.97 | 2.24 |
| AAEL019780-RB | None (GO:0008410, CoA-transferase activity) | 5.27 | 5.19 | 2.62 |
| AAEL005023-RD | Palmitoyltransferase (None) | 5.33 | 6.30 | 5.14 |
| AAEL021339-RF | None (None) | 5.50 | 3.79 | 3.93 |
| AAEL004931-RA | Beta-hexosaminidase b (GO:0102148;GO:0004563;GO:0016787;GO:0016798;GO:0004553, N-acetyl-beta-D-galactosaminidase activity; beta-N-acetylhexosaminidase activity) | 5.67 | 3.41 | 6.73 |
| AAEL024186-RC | None (GO:0003779;GO:0030276;GO:0005543, actin binding; clathrin binding; phospholipid binding) | 5.87 | 3.00 | 6.26 |
| AAEL006633-RB | Inhibitor of Apoptosis (IAP) (None) | −7.22 | −7.08 | −4.47 |
| AAEL006510-RC | None (GO:0003723;GO:0008190, RNA binding; eukaryotic initiation factor 4E binding) | −6.61 | −6.04 | −6.91 |
| AAEL012000-RA | None (None) | −6.42 | −6.29 | −4.70 |
| AAEL009059-RB | Arp2/3 complex 16 kd subunit (P16-arc) (None) | −5.86 | −5.74 | −5.88 |
| AAEL001041-RN | Guanine nucleotide-binding protein beta 3 (g protein beta3) (GO:0005515, protein binding) | −5.72 | −5.00 | −4.14 |
| AAEL000234-RN | Class B Scavenger Receptor (CD36 domain) (None) | −5.50 | −3.75 | −3.05 |
| AAEL012410-RB | Eukaryotic translation initiation factor 2C (GO:0003676;GO:0005515, nucleic acid binding; protein binding) | −5.50 | −4.20 | −3.32 |
| AAEL008542-RC | Kinesin heavy chain subunit (GO:0005524;GO:0008017;GO:0003777, ATP binding; microtubule binding; microtubule motor activity) | −5.04 | −4.91 | −4.09 |
| AAEL023848-RA | None (None) | −4.87 | −2.81 | −6.23 |
| AAEL007874-RC | None (GO:0047372, acylglycerol lipase activity) | −4.76 | −4.63 | −4.37 |
| AAEL004496-RD | Glutamate transporter (GO:0015293, symporter activity) | −4.66 | −4.52 | −5.88 |
| AAEL010373-RK | Dullard protein (GO:0016791, phosphatase activity) | −4.29 | −4.16 | −4.68 |
| AAEL006364-RC | None (GO:0045127, N-acetylglucosamine kinase activity) | −4.24 | −4.11 | −7.08 |
| AAEL004420-RC | None (None) | −4.19 | −4.06 | −5.29 |
| AAEL011858-RD | Transcription factor IIIA (GO:0003676;GO:0008270, nucleic acid binding; zinc ion binding) | −4.19 | −4.07 | −3.54 |
| AAEL006982-RG | Lipase (GO:0052689, carboxylic ester hydrolase activity) | −4.18 | −4.07 | −2.50 |
| AAEL025570-RE | None (GO:0005524;GO:0003774;GO:0005515, ATP binding; motor activity; protein binding) | −4.12 | −6.57 | −2.59 |
| AAEL004385-RB | UGA suppressor tRNA-associated antigenic protein (GO:0003824;GO:0016740;GO:0016785, catalytic activity; transferase activity; transferase activity) | −4.11 | −3.98 | −5.09 |
| AAEL018032-RE | None (None) | −3.97 | −3.43 | −6.07 |
| AAEL000549-RE | Fibulin 1 (GO:0005509;GO:0005515, calcium ion binding; protein binding) | −3.75 | −3.62 | −4.69 |
| AAEL000354-RB | Dimeric dihydrodiol dehydrogenase (GO:0016491, oxidoreductase activity) | −3.69 | −3.56 | −3.09 |
| AAEL006556-RA | None (None) | −2.47 | −2.43 | −5.21 |
| AAEL003471-RA | None (GO:0003676;GO:0008270, nucleic acid binding; zinc ion binding) | −2.44 | −2.40 | −3.73 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giraldo-Calderón, G.I.; Calle-Tobón, A.; Rozo-López, P.; Colpitts, T.M.; Park, Y.; Rua-Uribe, G.L.; Londono-Renteria, B. Transcriptome of the Aedes aegypti Mosquito in Response to Human Complement Proteins. Int. J. Mol. Sci. 2020, 21, 6584. https://doi.org/10.3390/ijms21186584
Giraldo-Calderón GI, Calle-Tobón A, Rozo-López P, Colpitts TM, Park Y, Rua-Uribe GL, Londono-Renteria B. Transcriptome of the Aedes aegypti Mosquito in Response to Human Complement Proteins. International Journal of Molecular Sciences. 2020; 21(18):6584. https://doi.org/10.3390/ijms21186584
Chicago/Turabian StyleGiraldo-Calderón, Gloria I., Arley Calle-Tobón, Paula Rozo-López, Tonya M. Colpitts, Yoonseong Park, Guillermo L. Rua-Uribe, and Berlin Londono-Renteria. 2020. "Transcriptome of the Aedes aegypti Mosquito in Response to Human Complement Proteins" International Journal of Molecular Sciences 21, no. 18: 6584. https://doi.org/10.3390/ijms21186584








