Ribosome Pausing at Inefficient Codons at the End of the Replicase Coding Region Is Important for Hepatitis C Virus Genome Replication
Abstract
:1. Introduction
2. Results
2.1. Analysis of Ribosome Occupancy of the HCV Genome
2.2. RNA Structure Analyses
2.3. Ribosome Pausing at the HCV Stop Codon Region
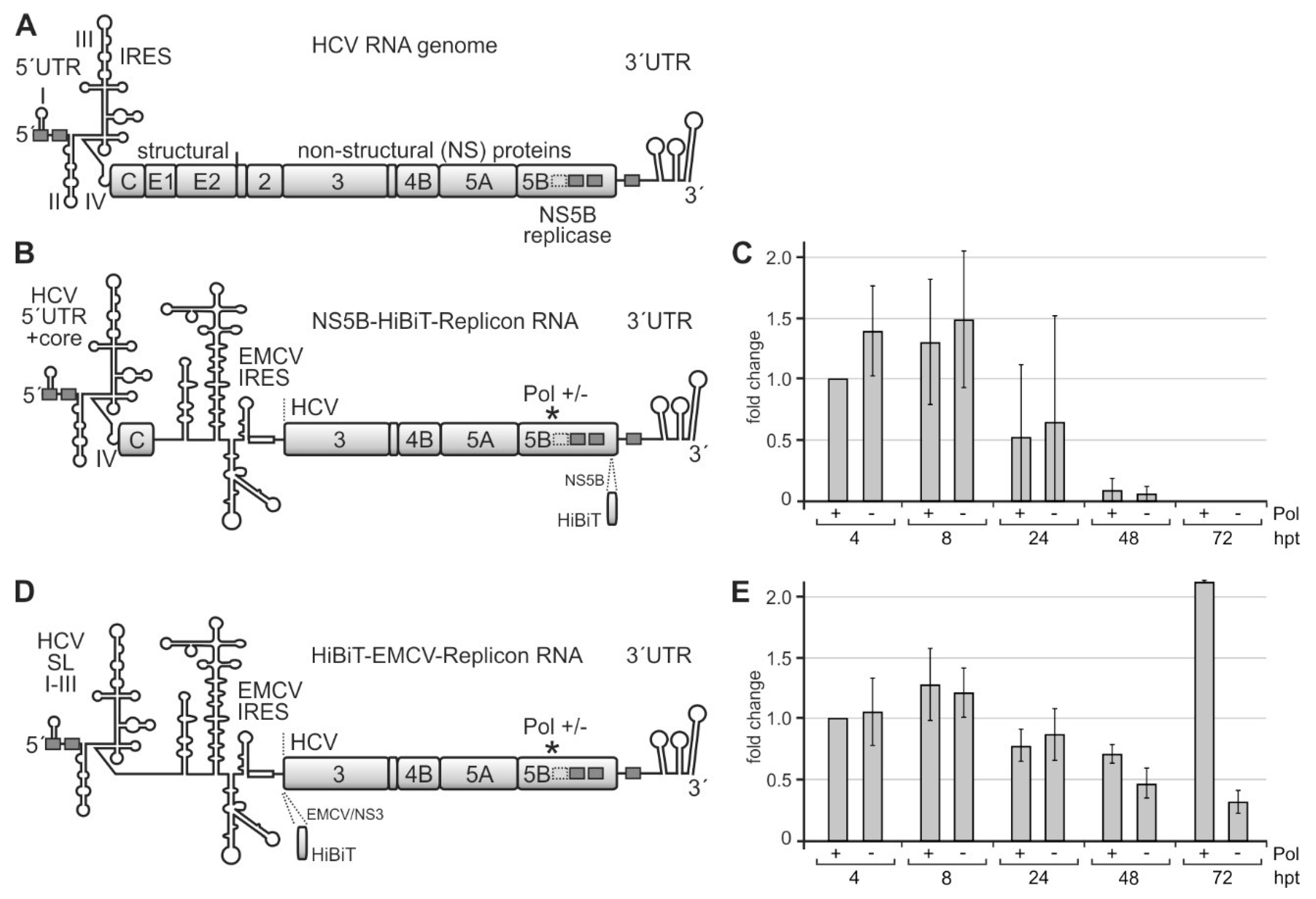
2.4. A Replicon System for the Analysis of Rare Codon Function
2.5. The NS5B Rare Codons Are Important for HCV Genome Replication but Not Translation
3. Discussion
4. Materials and Methods
4.1. Plasmids
4.2. In vitro-Transcription
4.3. RNA Transfection of Replicon RNAs and HiBiT Expression Assay
4.4. Statistical Analysis
4.5. Transfection of HCV Full-Length Genomes and Ribosome Profiling
4.6. RNA Structure Prediction and Visualization
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Colpitts, C.C.; Tsai, P.-L.; Zeisel, M.B. Hepatitis C Virus Entry: An Intriguingly Complex and Highly Regulated Process. Int. J. Mol. Sci. 2020, 21, 2091. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berggren, K.A.; Suzuki, S.; Ploss, A. Animal Models Used in Hepatitis C Virus Research. Int. J. Mol. Sci. 2020, 21, 3869. [Google Scholar] [CrossRef] [PubMed]
- Waziry, R.; Hajarizadeh, B.; Grebely, J.; Amin, J.; Law, M.; Danta, M.; George, J.; Dore, G.J. Hepatocellular carcinoma risk following direct-acting antiviral HCV therapy: A systematic review, meta-analyses, and meta-regression. J. Hepatol. 2017, 67, 1204–1212. [Google Scholar] [CrossRef]
- Virzì, A.; Suarez, A.A.R.; Baumert, T.F.; Lupberger, J. Rewiring Host Signaling: Hepatitis C Virus in Liver Pathogenesis. Cold Spring Harb. Perspect. Med. 2019, 10, a037366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerresheim, G.K.; Roeb, E.; Michel, A.M.; Niepmann, M. Roeb Hepatitis C Virus Downregulates Core Subunits of Oxidative Phosphorylation, Reminiscent of the Warburg Effect in Cancer Cells. Cells 2019, 8, 1410. [Google Scholar] [CrossRef] [Green Version]
- Goto, K.; Suarez, A.A.R.; Wrensch, F.; Baumert, T.F.; Lupberger, J. Hepatitis C Virus and Hepatocellular Carcinoma: When the Host Loses Its Grip. Int. J. Mol. Sci. 2020, 21, 3057. [Google Scholar] [CrossRef]
- Quinkert, D.; Bartenschlager, R.; Lohmann, V. Quantitative Analysis of the Hepatitis C Virus Replication Complex. J. Virol. 2005, 79, 13594–13605. [Google Scholar] [CrossRef] [Green Version]
- Lindenbach, B.D.; Rice, C.M. The ins and outs of hepatitis C virus entry and assembly. Nat. Rev. Genet. 2013, 11, 688–700. [Google Scholar] [CrossRef] [Green Version]
- Luna, J.M.; Scheel, T.K.H.; Danino, T.; Shaw, K.S.; Mele, A.; Fak, J.J.; Nishiuchi, E.; Takacs, C.N.; Catanese, M.T.; De Jong, Y.P.; et al. Hepatitis C virus RNA functionally sequesters miR-122. Cell 2015, 160, 1099–1110. [Google Scholar] [CrossRef] [Green Version]
- Wrensch, F.; Crouchet, E.; Ligat, G.; Zeisel, M.B.; Keck, Z.-Y.; Foung, S.K.H.; Schuster, C.; Baumert, T.F. Hepatitis C Virus (HCV)–Apolipoprotein Interactions and Immune Evasion and Their Impact on HCV Vaccine Design. Front. Immunol. 2018, 9, 9. [Google Scholar] [CrossRef]
- Vieyres, G.; Pietschmann, T. HCV Pit Stop at the Lipid Droplet: Refuel Lipids and Put on a Lipoprotein Coat before Exit. Cells 2019, 8, 233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerold, G.; Moeller, R.; Pietschmann, T. Hepatitis C Virus Entry: Protein Interactions and Fusion Determinants Governing Productive Hepatocyte Invasion. Cold Spring Harb. Perspect. Med. 2019, 10, a036830. [Google Scholar] [CrossRef] [PubMed]
- Bley, H.; Schöbel, A.; Herker, E. Whole Lotta Lipids—From HCV RNA Replication to the Mature Viral Particle. Int. J. Mol. Sci. 2020, 21, 2888. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heim, M.H.; Thimme, R. Innate and adaptive immune responses in HCV infections. J. Hepatol. 2014, 61, S14–S25. [Google Scholar] [CrossRef] [Green Version]
- Rehermann, B.; Thimme, R. Insights From Antiviral Therapy Into Immune Responses to Hepatitis B and C Virus Infection. Gastroenterology 2019, 156, 369–383. [Google Scholar] [CrossRef]
- Sung, P.S.; Shin, E.-C. Interferon Response in Hepatitis C Virus-Infected Hepatocytes: Issues to Consider in the Era of Direct-Acting Antivirals. Int. J. Mol. Sci. 2020, 21, 2583. [Google Scholar] [CrossRef] [Green Version]
- Hu, P.; Wilhelm, J.; Gerresheim, G.K.; Shalamova, L.A.; Niepmann, M. Lnc-ITM2C-1 and GPR55 Are Proviral Host Factors for Hepatitis C Virus. Viruses 2019, 11, 549. [Google Scholar] [CrossRef] [Green Version]
- Unfried, J.P.; Fortes, P. LncRNAs in HCV Infection and HCV-Related Liver Disease. Int. J. Mol. Sci. 2020, 21, 2255. [Google Scholar] [CrossRef] [Green Version]
- Perales, C.; Quer, J.; Gregori, J.; Esteban, J.I.; Domingo, E. Resistance of Hepatitis C Virus to Inhibitors: Complexity and Clinical Implications. Viruses 2015, 7, 5746–5766. [Google Scholar] [CrossRef] [Green Version]
- Bartenschlager, R.; Baumert, T.F.; Bukh, J.; Houghton, M.; Lemon, S.M.; Lindenbach, B.D.; Lohmann, V.; Moradpour, D.; Pietschmann, T.; Rice, C.M.; et al. Critical challenges and emerging opportunities in hepatitis C virus research in an era of potent antiviral therapy: Considerations for scientists and funding agencies. Virus Res. 2018, 248, 53–62. [Google Scholar] [CrossRef]
- Alazard-Dany, N.; Denolly, S.; Boson, B.; Cosset, F.-L. Overview of HCV Life Cycle with a Special Focus on Current and Possible Future Antiviral Targets. Viruses 2019, 11, 30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choo, Q.; Kuo, G.; Weiner, A.; Overby, L.; Bradley, D.; Houghton, M. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science 1989, 244, 359–362. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niepmann, M.; Gerresheim, G.K. Hepatitis C Virus Translation Regulation. Int. J. Mol. Sci. 2020, 21, 2328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsukiyama-Kohara, K.; Iizuka, N.; Kohara, M.; Nomoto, A. Internal ribosome entry site within hepatitis C virus RNA. J. Virol. 1992, 66, 1476–1483. [Google Scholar] [CrossRef] [Green Version]
- Moradpour, D.; Penin, F. Hepatitis C Virus Proteins: From Structure to Function. Inducible Lymphoid Organs 2013, 369, 113–142. [Google Scholar] [CrossRef]
- Paul, D.; Madan, V.; Bartenschlager, R. Hepatitis C Virus RNA Replication and Assembly: Living on the Fat of the Land. Cell Host Microbe 2014, 16, 569–579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neufeldt, C.J.; Cortese, M.; Acosta, E.G.; Bartenschlager, R. Rewiring cellular networks by members of the Flaviviridae family. Nat. Rev. Genet. 2018, 16, 125–142. [Google Scholar] [CrossRef] [PubMed]
- Bartenschlager, R.; Cosset, F.-L.; Lohmann, V. Hepatitis C virus replication cycle. J. Hepatol. 2010, 53, 583–585. [Google Scholar] [CrossRef] [Green Version]
- Lohmann, V. Hepatitis C Virus RNA Replication. Inducible Lymphoid Organs 2013, 369, 167–198. [Google Scholar] [CrossRef]
- Niepmann, M.; Shalamova, L.A.; Gerresheim, G.K.; Rossbach, O. Signals Involved in Regulation of Hepatitis C Virus RNA Genome Translation and Replication. Front. Microbiol. 2018, 9, 395. [Google Scholar] [CrossRef] [PubMed]
- Romero-López, C.; Berzal-Herranz, A. The Role of the RNA-RNA Interactome in the Hepatitis C Virus Life Cycle. Int. J. Mol. Sci. 2020, 21, 1479. [Google Scholar] [CrossRef] [Green Version]
- Bartel, D.P. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef]
- Sagan, S.M.; Macrae, I.J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell Boil. 2018, 20, 21–37. [Google Scholar] [CrossRef]
- Jopling, C.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of Hepatitis C Virus RNA Abundance by a Liver-Specific MicroRNA. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henke, J.I.; Goergen, D.; Zheng, J.; Song, Y.; Schüttler, C.G.; Fehr, C.; Jünemann, C.; Niepmann, M. microRNA-122 stimulates translation of hepatitis C virus RNA. EMBO J. 2008, 27, 3300–3310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimakami, T.; Yamane, D.; Jangra, R.K.; Kempf, B.J.; Spaniel, C.; Barton, D.J.; Lemon, S.M. Stabilization of hepatitis C virus RNA by an Ago2-miR-122 complex. Proc. Natl. Acad. Sci. USA 2012, 109, 109–941. [Google Scholar] [CrossRef] [Green Version]
- Friebe, P.; Bartenschlager, R. Genetic analysis of sequences in the 3’ nontranslated region of hepatitis C virus that are important for RNA replication. J. Virol. 2002, 76, 5326–5338. [Google Scholar] [CrossRef] [Green Version]
- Yi, M.; Lemon, S.M. 3′ Nontranslated RNA Signals Required for Replication of Hepatitis C Virus RNA. J. Virol. 2003, 77, 3557–3568. [Google Scholar] [CrossRef] [Green Version]
- You, S.; Rice, C.M. 3′ RNA Elements in Hepatitis C Virus Replication: Kissing Partners and Long Poly(U). J. Virol. 2007, 82, 184–195. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Wimmer, E.; Paul, A.V. Cis-acting RNA elements in human and animal plus-strand RNA viruses. Biochim. et Biophys. Acta (BBA) Bioenerg. 2009, 1789, 495–517. [Google Scholar] [CrossRef]
- Chu, D.; Ren, S.; Hu, S.; Wang, W.G.; Subramanian, A.; Contreras, D.; Kanagavel, V.; Chung, E.; Ko, J.; Appadorai, R.S.A.J.; et al. Systematic Analysis of Enhancer and Critical cis-Acting RNA Elements in the Protein-Encoding Region of the Hepatitis C Virus Genome. J. Virol. 2013, 87, 5678–5696. [Google Scholar] [CrossRef] [Green Version]
- Mauger, D.M.; Golden, M.; Yamane, D.; Williford, S.; Lemon, S.M.; Martin, D.P.; Weeks, K.M. Functionally conserved architecture of hepatitis C virus RNA genomes. Proc. Natl. Acad. Sci. USA 2015, 112, 3692–3697. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fricke, M.; Dünnes, N.; Zayas, M.; Bartenschlager, R.; Niepmann, M.; Marz, M. Conserved RNA secondary structures and long-range interactions in hepatitis C viruses. RNA 2015, 21, 1219–1232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pirakitikulr, N.; Kohlway, A.; Lindenbach, B.D.; Pyle, A.M. The Coding Region of the HCV Genome Contains a Network of Regulatory RNA Structures. Mol. Cell 2016, 62, 111–120. [Google Scholar] [CrossRef] [PubMed]
- You, S.; Stump, D.D.; Branch, A.D.; Rice, C.M. A cis-Acting Replication Element in the Sequence Encoding the NS5B RNA-Dependent RNA Polymerase Is Required for Hepatitis C Virus RNA Replication. J. Virol. 2004, 78, 1352–1366. [Google Scholar] [CrossRef] [Green Version]
- Cheng, J.C.; Chang, M.F.; Chang, S.C. Specific interaction between the hepatitis C virus NS5B RNA polymerase and the 3’ end of the viral RNA. J. Virol. 1999, 73, 7044–7049. [Google Scholar] [CrossRef] [Green Version]
- Kim, M.; Kim, H.; Cho, S.-P.; Min, M.-K. Template Requirements for De Novo RNA Synthesis by Hepatitis C Virus Nonstructural Protein 5B Polymerase on the Viral X RNA. J. Virol. 2002, 76, 6944–6956. [Google Scholar] [CrossRef] [Green Version]
- Lee, H.; Shin, H.; Wimmer, E.; Paul, A.V. cis-Acting RNA Signals in the NS5B C-Terminal Coding Sequence of the Hepatitis C Virus Genome. J. Virol. 2004, 78, 10865–10877. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Yamada, O.; Sakamoto, T.; Yoshida, H.; Araki, H.; Murata, T.; Shimotohno, K. Inhibition of hepatitis C virus replication by pol III-directed overexpression of RNA decoys corresponding to stem-loop structures in the NS5B coding region. Virology 2005, 342, 276–285. [Google Scholar] [CrossRef] [Green Version]
- Romero-López, C.; Ríos-Marco, P.; Berzal-Herranz, B.; Berzal-Herranz, A. The HCV genome domains 5BSL3.1 and 5BSL3.3 act as managers of translation. Sci. Rep. 2018, 8, 16101. [Google Scholar] [CrossRef]
- Tuplin, A.; Struthers, M.; Cook, J.; Bentley, K.; Evans, D.J. Inhibition of HCV translation by disrupting the structure and interactions of the viral CRE and 3’ X-tail. Nucleic Acids Res. 2015, 43, 2914–2926. [Google Scholar] [CrossRef]
- Ito, T.; Tahara, S.M.; Lai, M.M. The 3’-untranslated region of hepatitis C virus RNA enhances translation from an internal ribosomal entry site. J. Virol. 1998, 72, 8789–8796. [Google Scholar] [CrossRef] [Green Version]
- Song, Y.; Friebe, P.; Tzima, E.; Jünemann, C.; Bartenschlager, R.; Niepmann, M. The hepatitis C virus RNA 3’-untranslated region strongly enhances translation directed by the internal ribosome entry site. J. Virol. 2006, 80, 11579–11588. [Google Scholar] [CrossRef] [Green Version]
- Bai, Y.; Zhou, K.; Doudna, J.A. Hepatitis C virus 3′UTR regulates viral translation through direct interactions with the host translation machinery. Nucleic Acids Res. 2013, 41, 7861–7874. [Google Scholar] [CrossRef] [PubMed]
- Friebe, P.; Boudet, J.; Simorre, J.P.; Bartenschlager, R. Kissing-loop interaction in the 3’ end of the hepatitis C virus genome essential for RNA replication. J. Virol. 2005, 79, 380–392. [Google Scholar] [CrossRef] [Green Version]
- Pietschmann, T.; Kaul, A.; Koutsoudakis, G.; Shavinskaya, A.; Kallis, S.; Steinmann, E.; Abid, K.; Negro, F.; Dreux, M.; Cosset, F.-L.; et al. Construction and characterization of infectious intragenotypic and intergenotypic hepatitis C virus chimeras. Proc. Natl. Acad. Sci. USA 2006, 103, 7408–7413. [Google Scholar] [CrossRef] [Green Version]
- Gerresheim, G.K.; Bathke, J.; Michel, A.M.; Andreev, D.E.; Shalamova, L.A.; Rossbach, O.; Hu, P.; Glebe, D.; Fricke, M.; Marz, M.; et al. Cellular Gene Expression during Hepatitis C Virus Replication as Revealed by Ribosome Profiling. Int. J. Mol. Sci. 2019, 20, 1321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Binder, M.; Quinkert, D.; Bochkarova, O.; Klein, R.; Kezmic, N.; Bartenschlager, R.; Lohmann, V. Identification of Determinants Involved in Initiation of Hepatitis C Virus RNA Synthesis by Using Intergenotypic Replicase Chimeras. J. Virol. 2007, 81, 5270–5283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerresheim, G.K.; Dünnes, N.; Nieder-Röhrmann, A.; Shalamova, L.A.; Fricke, M.; Hofacker, I.; Höner Zu Siederdissen, C.; Marz, M.; Niepmann, M. microRNA-122 target sites in the hepatitis C virus RNA NS5B coding region and 3’ untranslated region: Function in replication and influence of RNA secondary structure. Cell. Mol. Life Sci. 2017, 74, 747–760. [Google Scholar] [CrossRef] [PubMed]
- Fricke, M.; Gerst, R.; Ibrahim, B.; Niepmann, M.; Marz, M. Global importance of RNA secondary structures in protein-coding sequences. Bioinformatics 2019, 35, 579–583. [Google Scholar] [CrossRef]
- Roseman, M.A. Hydrophilicity of polar amino acid side-chains is markedly reduced by flanking peptide bonds. J. Mol. Boil. 1988, 200, 513–522. [Google Scholar] [CrossRef]
- Tuller, T.; Carmi, A.; Vestsigian, K.; Navon, S.; Dorfan, Y.; Zaborske, J.; Pan, T.; Dahan, O.; Furman, I.; Pilpel, Y. An Evolutionarily Conserved Mechanism for Controlling the Efficiency of Protein Translation. Cell 2010, 141, 344–354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Firth, A.E.; Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012, 93, 1385–1409. [Google Scholar] [CrossRef] [PubMed]
- Meydan, S.; Klepacki, R.; Karthikeyan, S.; Margus, T.; Thomas, P.M.; Jones, J.E.; Khan, Y.A.; Briggs, J.; Dinman, J.D.; Vázquez-Laslop, N.; et al. Programmed Ribosomal Frameshifting Generates a Copper Transporter and a Copper Chaperone from the Same Gene. Mol. Cell 2017, 65, 207–219. [Google Scholar] [CrossRef] [Green Version]
- Atkins, J.F.; Loughran, G.; Baranov, P.V. A [Cu]rious Ribosomal Profiling Pattern Leads to the Discovery of Ribosomal Frameshifting in the Synthesis of a Copper Chaperone. Mol. Cell 2017, 65, 203–204. [Google Scholar] [CrossRef] [Green Version]
- Choi, J.; O’Loughlin, S.; Atkins, J.F.; Puglisi, J.D. The energy landscape of -1 ribosomal frameshifting. Sci. Adv. 2020, 6, eaax6969. [Google Scholar] [CrossRef] [Green Version]
- Available online: http://www.gcua.de (accessed on 19 June 2018).
- Available online: https://www.kazusa.or.jp/codon/ (accessed on 23 August 2018).
- Sabi, R.; Tuller, T. Computational analysis of nascent peptides that induce ribosome stalling and their proteomic distribution in Saccharomyces cerevisiae. RNA 2017, 23, 983–994. [Google Scholar] [CrossRef] [Green Version]
- Wruck, F.; Katranidis, A.; Nierhaus, K.H.; Büldt, G.; Hegner, M. Translation and folding of single proteins in real time. Proc. Natl. Acad. Sci. USA 2017, 114, E4399–E4407. [Google Scholar] [CrossRef] [Green Version]
- Holcomb, D.D.; Alexaki, A.; Katneni, U.; Kimchi-Sarfaty, C. The Kazusa codon usage database, CoCoPUTs, and the value of up-to-date codon usage statistics. Infect. Genet. Evol. 2019, 73, 266–268. [Google Scholar] [CrossRef]
- Quax, T.E.F.; Claassens, N.J.; Söll, D.; Van Der Oost, J. Codon Bias as a Means to Fine-Tune Gene Expression. Mol. Cell 2015, 59, 149–161. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; McManus, J.; Kingsford, C. Accurate Recovery of Ribosome Positions Reveals Slow Translation of Wobble-Pairing Codons in Yeast. J. Comput. Boil. 2016, 24, 486–500. [Google Scholar] [CrossRef] [PubMed]
- Schuller, A.P.; Green, R. Roadblocks and resolutions in eukaryotic translation. Nat. Rev. Mol. Cell Boil. 2018, 19, 526–541. [Google Scholar] [CrossRef] [PubMed]
- Feng, Q.; Shao, S. In vitro reconstitution of translational arrest pathways. Methods 2018, 137, 20–36. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Ignatova, Z. Folding at the birth of the nascent chain: Coordinating translation with co-translational folding. Curr. Opin. Struct. Boil. 2011, 21, 25–31. [Google Scholar] [CrossRef]
- Yu, C.-H.; Dang, Y.; Zhou, Z.; Wu, C.; Zhao, F.; Sachs, M.S.; Liu, Y. Codon Usage Influences the Local Rate of Translation Elongation to Regulate Co-translational Protein Folding. Mol. Cell 2015, 59, 744–754. [Google Scholar] [CrossRef] [Green Version]
- Oh-Hashi, K.; Furuta, E.; Fujimura, K.; Hirata, Y. Application of a novel HiBiT peptide tag for monitoring ATF4 protein expression in Neuro2a cells. Biochem. Biophys. Rep. 2017, 12, 40–45. [Google Scholar] [CrossRef]
- Schwinn, M.K.; Machleidt, T.; Zimmerman, K.; Eggers, C.T.; Dixon, A.S.; Hurst, R.; Hall, M.P.; Encell, L.P.; Binkowski, B.F.; Wood, K.V. CRISPR-Mediated Tagging of Endogenous Proteins with a Luminescent Peptide. ACS Chem. Boil. 2017, 13, 467–474. [Google Scholar] [CrossRef]
- Mosley, R.T.; E Edwards, T.; Murakami, E.; Lam, A.M.; Grice, R.L.; Du, J.; Sofia, M.J.; Furman, P.A.; Otto, M.J. Structure of Hepatitis C Virus Polymerase in Complex with Primer-Template RNA. J. Virol. 2012, 86, 6503–6511. [Google Scholar] [CrossRef] [Green Version]
- Lohmann, V.; Körner, F.; Herian, U.; Bartenschlager, R. Biochemical properties of hepatitis C virus NS5B RNA-dependent RNA polymerase and identification of amino acid sequence motifs essential for enzymatic activity. J. Virol. 1997, 71, 8416–8428. [Google Scholar] [CrossRef] [Green Version]
- Binder, M.; Kochs, G.; Bartenschlager, R.; Lohmann, V. Hepatitis C virus escape from the interferon regulatory factor 3 pathway by a passive and active evasion strategy. Hepatology 2007, 46, 1365–1374. [Google Scholar] [CrossRef]
- Lambert, S.; Langley, D.R.; Garnett, J.A.; Angell, R.; Hedgethorne, K.; Meanwell, N.A.; Matthews, S.J. The crystal structure of NS5A domain 1 from genotype 1a reveals new clues to the mechanism of action for dimeric HCV inhibitors. Protein Sci. 2014, 23, 723–734. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shanmugam, S.; Nichols, A.K.; Saravanabalaji, D.; Welsch, C.; Yi, M. HCV NS5A dimer interface residues regulate HCV replication by controlling its self-interaction, hyperphosphorylation, subcellular localization and interaction with cyclophilin A. PLOS Pathog. 2018, 14, e1007177. [Google Scholar] [CrossRef] [Green Version]
- Stern-Ginossar, N.; Weisburd, B.; Michalski, A.; Le-Trilling, V.T.K.; Hein, M.Y.; Huang, S.-X.; Ma, M.; Shen, B.; Qian, S.-B.; Hengel, H.; et al. Decoding Human Cytomegalovirus. Science 2012, 338, 1088–1093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andreev, D.E.; O’Connor, P.B.F.; Loughran, G.; Dmitriev, S.E.; Baranov, P.V.; Shatsky, I.N. Insights into the mechanisms of eukaryotic translation gained with ribosome profiling. Nucleic Acids Res. 2016, 45, 513–526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andreev, D.E.; O’Connor, P.B.; Fahey, C.; Kenny, E.M.; Terenin, I.M.; Dmitriev, S.E.; Cormican, P.; Morris, D.W.; Shatsky, I.N.; Baranov, P.V. Translation of 5’ leaders is pervasive in genes resistant to eIF2 repression. eLife 2015, 4, e3971. [Google Scholar] [CrossRef]
- Brass, V.; Gouttenoire, J.; Wahl, A.; Pal, Z.; Blum, H.E.; Penin, F.; Moradpour, D. Hepatitis C Virus RNA Replication Requires a Conserved Structural Motif within the Transmembrane Domain of the NS5B RNA-Dependent RNA Polymerase. J. Virol. 2010, 84, 11580–11584. [Google Scholar] [CrossRef] [Green Version]
- Harrus, D.; Ahmed-El-Sayed, N.; Simister, P.C.; Miller, S.; Triconnet, M.; Hagedorn, C.H.; Mahias, K.; Rey, F.A.; Astier-Gin, T.; Bressanelli, S. Further Insights into the Roles of GTP and the C Terminus of the Hepatitis C Virus Polymerase in the Initiation of RNA Synthesis. J. Boil. Chem. 2010, 285, 32906–32918. [Google Scholar] [CrossRef] [Green Version]
- Appleby, T.C.; Perry, J.K.; Murakami, E.; Barauskas, O.; Feng, J.; Cho, A.; Fox, D.; Wetmore, D.R.; McGrath, M.E.; Ray, A.S.; et al. Structural basis for RNA replication by the hepatitis C virus polymerase. Science 2015, 347, 771–775. [Google Scholar] [CrossRef]
- Reich, S.; Golbik, R.P.; Geissler, R.; Lilie, H.; Behrens, S.-E. Mechanisms of Activity and Inhibition of the Hepatitis C Virus RNA-dependent RNA Polymerase. J. Boil. Chem. 2010, 285, 13685–13693. [Google Scholar] [CrossRef] [Green Version]
- Karam, P.; Powdrill, M.H.; Liu, H.-W.; Vasquez, C.; Mah, W.; Bernatchez, J.A.; Götte, M.; Cosa, G. Dynamics of Hepatitis C Virus (HCV) RNA-dependent RNA Polymerase NS5B in Complex with RNA. J. Boil. Chem. 2014, 289, 14399–14411. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Jiang, W.W.; Pratt, J.; Rockway, T.; Harris, K.; Vasavanonda, S.; Tripathi, R.; Pithawalla, R.; Kati, W.M. Mechanistic Study of HCV Polymerase Inhibitors at Individual Steps of the Polymerization Reaction. Biochemistry 2006, 45, 11312–11323. [Google Scholar] [CrossRef] [PubMed]
- Simister, P.; Schmitt, M.; Geitmann, M.; Wicht, O.; Danielson, U.H.; Klein, R.; Bressanelli, S.; Lohmann, V. Structural and Functional Analysis of Hepatitis C Virus Strain JFH1 Polymerase. J. Virol. 2009, 83, 11926–11939. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, W.; Ferrari, E.; Lesburg, C.A.; Maag, D.; Ghosh, S.K.B.; Cameron, C.E.; Lau, J.Y.; Hong, Z. Template/Primer Requirements and Single Nucleotide Incorporation by Hepatitis C Virus Nonstructural Protein 5B Polymerase. J. Virol. 2000, 74, 9134–9143. [Google Scholar] [CrossRef] [Green Version]
- Zhong, W.; Uss, A.S.; Ferrari, E.; Lau, J.Y.N. De Novo Initiation of RNA Synthesis by Hepatitis C Virus Nonstructural Protein 5B Polymerase. J. Virol. 2000, 74, 2017–2022. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, T.; Friedrich, S.; Golbik, R.P.; Behrens, S.-E. NF90–NF45 is a selective RNA chaperone that rearranges viral and cellular riboswitches: Biochemical analysis of a virus host factor activity. Nucleic Acids Res. 2017, 45, 12441–12454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liutkute, M.; Samatova, E.; Rodnina, M.V. Cotranslational Folding of Proteins on the Ribosome. Biomolecules 2020, 10, 97. [Google Scholar] [CrossRef] [Green Version]
- Kaminski, A.; Belsham, G.J.; Jackson, R.J. Translation of encephalomyocarditis virus RNA: Parameters influencing the selection of the internal initiation site. EMBO J. 1994, 13, 1673–1681. [Google Scholar] [CrossRef]
- Blight, K.J.; McKeating, J.A.; Rice, C.M. Highly Permissive Cell Lines for Subgenomic and Genomic Hepatitis C Virus RNA Replication. J. Virol. 2002, 76, 13001–13014. [Google Scholar] [CrossRef] [Green Version]
- Ingolia, N.T.; Brar, G.A.; Rouskin, S.; McGeachy, A.M.; Weissman, J.S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 2012, 7, 1534–1550. [Google Scholar] [CrossRef]
- Ingolia, N.T.; Ghaemmaghami, S.; Newman, J.R.S.; Weissman, J.S. Genome-Wide Analysis in Vivo of Translation with Nucleotide Resolution Using Ribosome Profiling. Science 2009, 324, 218–223. [Google Scholar] [CrossRef] [Green Version]
- Lorenz, R.; Bernhart, S.H.; Zu Siederdissen, C.H.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Boil. 2011, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Darty, K.; Denise, A.; Ponty, Y. VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 2009, 25, 1974–1975. [Google Scholar] [CrossRef] [PubMed] [Green Version]





© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gerresheim, G.K.; Hess, C.S.; Shalamova, L.A.; Fricke, M.; Marz, M.; Andreev, D.E.; Shatsky, I.N.; Niepmann, M. Ribosome Pausing at Inefficient Codons at the End of the Replicase Coding Region Is Important for Hepatitis C Virus Genome Replication. Int. J. Mol. Sci. 2020, 21, 6955. https://doi.org/10.3390/ijms21186955
Gerresheim GK, Hess CS, Shalamova LA, Fricke M, Marz M, Andreev DE, Shatsky IN, Niepmann M. Ribosome Pausing at Inefficient Codons at the End of the Replicase Coding Region Is Important for Hepatitis C Virus Genome Replication. International Journal of Molecular Sciences. 2020; 21(18):6955. https://doi.org/10.3390/ijms21186955
Chicago/Turabian StyleGerresheim, Gesche K., Carolin S. Hess, Lyudmila A. Shalamova, Markus Fricke, Manja Marz, Dmitri E. Andreev, Ivan N. Shatsky, and Michael Niepmann. 2020. "Ribosome Pausing at Inefficient Codons at the End of the Replicase Coding Region Is Important for Hepatitis C Virus Genome Replication" International Journal of Molecular Sciences 21, no. 18: 6955. https://doi.org/10.3390/ijms21186955




