1. Introduction
The normal cellular isoform of prion protein, designated PrP
C, is a glycoprotein tethered to the outer cell membrane via a glycosylphosphatidylinositol (GPI) anchor moiety and expressed most abundantly in the brain, particularly by neurons [
1,
2]. Its conformational conversion into the abnormally folded amyloidogenic isoform, PrP
Sc, which is believed to be a major component of infectious prions, is a key pathogenic event in prion diseases, a group of neurodegenerative disorders, including Creutzfeldt-Jakob disease in humans and scrapie and bovine spongiform encephalopathy (BSE) in animals [
1,
2]. However, the exact conversion mechanism remains largely unknown.
Mice lacking PrP
C (
Prnp0/0) have been shown to be resistant to prion infection, neither producing PrP
Sc or prions in the brain nor developing disease even after intracerebral inoculation with the prions [
3,
4,
5,
6]. Reverse genetic studies using reconstituted
Prnp0/0 mice with various transgenes encoding mutant PrP molecules have revealed that the N-terminal domain plays an important role in the conversion of PrP
C into PrP
Sc after prion infection. Indeed,
Prnp0/0 mice expressing PrP with a deletion of the N-terminal residues 23–31, designated Tg(PrP∆23–31)/
Prnp0/0 mice, were shown to be highly resistant to RML scrapie prions, developing disease only after longer incubation times and showing a slower accumulation of pathogenic PrP or PrP
Sc∆23–31, in the brain after infection with RML scrapie prions [
7]. In contrast, Tg(PrP∆32–80)/
Prnp0/0 mice, which express PrP with a deletion of residues 32–80, developed disease without an elongated incubation time and accumulated PrP
Sc∆23–80 in the brain after infection with RML prions [
8]. Moreover,
Prnp0/0 mice expressing PrP with a deletion of residues 51–90, which correspond to the so-called octapeptide repeat (OR) region, were also susceptible to RML and 22L scrapie prions, developing disease without an elongated incubation time and accumulating PrP
Sc∆OR in the brain [
9,
10]. These results indicate that, in contrast to residues 23–31, residues from position 32 to 90 may be dispensable for the conversion of PrP
C into PrP
Sc, as well as to support prion infection. However, Tg(PrP∆32–93)/
Prnp0/0 mice expressing PrP with a deletion extended to position 93 developed disease with longer incubation times with lower levels of prion infectivity and PrP
Sc∆32–93 in the brain after infection with RML prions [
11]. Furthermore, PrP with a deletion further extended to position 106 or PrP∆32–106, failed to convert into PrP
Sc or support prion pathogenesis in
Prnp0/0 mice after intracerebral inoculation with RML prions [
12]. These results suggest that residues 91–106, which are completely deleted in PrP∆32–106 and partially in PrP∆32–93 but intact in PrP∆32–80 and PrP∆OR, could play a role in the conversion of PrP
C into PrP
Sc and prion pathogenesis after prion infection. However, this remains to be clarified.
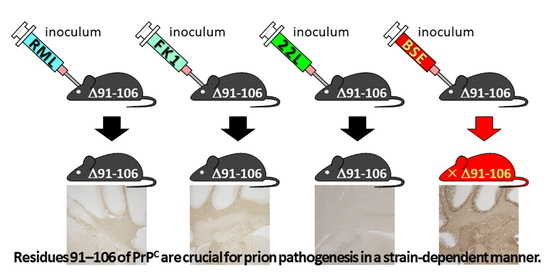
In this study, to evaluate the role of residues 91–106 of PrPC in prion pathogenesis, Tg(PrP∆91–106)/Prnp0/0 mice expressing mouse PrP lacking residues 91–106 or PrP∆91–106, on the Prnp0/0 background were generated and intracerebrally inoculated with mouse-adapted scrapie prions of RML and 22L, mouse-adapted human prions of FK-1 and mouse-adapted BSE prions. Tg(PrP∆91–106)/Prnp0/0 mice were found to be resistant to RML, 22L and FK-1 prions, neither accumulating PrPSc∆91–106 or infectious prions in the brain nor developing disease after inoculation. However, they were found to be marginally susceptible to BSE prions, succumbing to disease after long incubation times, as well as accumulating PrPSc∆91–106 and propagating BSE prions in the brain after inoculation. Furthermore, using an in vitro protein misfolding cyclic amplification (PMCA) assay, recombinant PrP with a smaller deletion of residues 91–104 or PrP∆91–104, was converted into PrPSc∆91–104 after incubation with BSE-PrPSc-prions but not with RML- and 22L-PrPSc-prions. Interestingly, digitonin and heparin stimulated the conversion of PrP∆91–104 into PrPSc∆91–104 even after incubation with RML- and 22L-PrPSc-prions. These results suggest that residues 91–106 or 91–104 of PrPC are crucial for prion pathogenesis in a strain-dependent manner and may play a similar role to digitonin and heparin in the conversion of PrPC into PrPSc.
3. Discussion
In the present study, to determine the role of residues 91–106 of PrP
C in prion pathogenesis, we established a line of Tg(PrP∆91–106)/
Prnp0/0 mice. The mice showed a 0.4-fold expression of mouse PrP with a deletion of residues 91–106 or PrP∆91–106, in the brain compared to PrP
C in WT mice. PrP∆91–106 and PrP
C were expressed similarly at the cell surface, namely preferentially at the raft membrane microdomains, in N2a cells and in mouse brains, indicating that the deletion of residues 91–106 did not alter the subcellular localization of PrP∆91–106 at the raft domains, which are considered a major cellular site for the conversion of PrP
C into PrP
Sc. However, Tg(PrP∆91–106)/
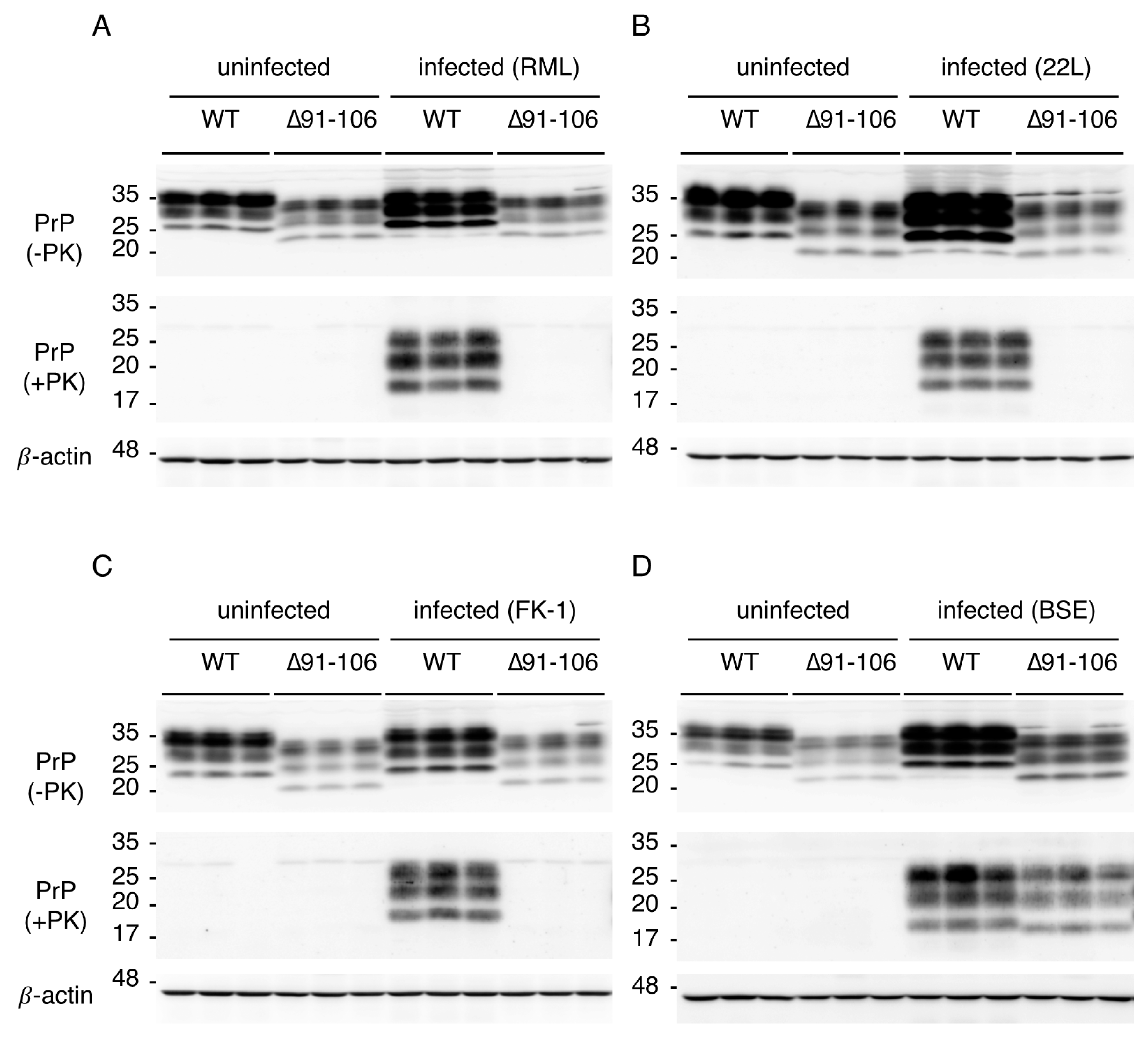
Prnp0/0 mice were found to be resistant to RML, 22L and FK-1 prions, neither accumulating PrP
Sc∆91–106 or prions in the brain nor developing disease after intracerebral inoculation with RML, 22L and FK-1 prions. In contrast, they remained marginally susceptible to BSE prions, accumulating PrP
Sc∆91–106 and prions in the brain and developing disease after long incubation times (538 ± 24 dpi). These results suggest that residues 91–106 of PrP
C are essential for RML, 22L and FK-1 prions but not for BSE prions, to induce the conversion of PrP
C into PrP
Sc and eventually cause prion disease in mice, although the lower expression of PrP∆91–106 in the brain might be partly responsible for the longer incubation times in BSE-infected Tg(PrP∆91–106)/
Prnp0/0 mice. Our results are consistent with those that PrP∆32–80 [
8] and PrP∆OR [
9,
10], both of which contain intact residues 91–106, restored full susceptibility to RML prions in
Prnp0/0 mice, although
Prnp0/0 mice expressing PrP∆32–93 or PrP∆32–106, which lacks residues 91–106 partially or entirely, markedly reduced or completely lost their susceptibility to RML prions, respectively [
11,
12].
Upon conversion into PrP
Sc, PrP
C undergoes marked conformational changes in the 2/3 C-terminal domain to form a PK-resistant structure [
20,
21]. However, the PK-cleavage site varies among PrP
Sc molecules derived from different prion strains. It is thus postulated that PrP
Sc can adopt a conformation in a prion strain-specific way and that the strain-specific pathogenic properties of prions can be enciphered in the strain-specific conformation of PrP
Sc [
20,
21]. PrP
Sc usually has a major PK cleavage site either in the C-terminal end of the OR region or in the post-OR region [
22]. BSE-PrP
Sc has a major PK cleavage site between residues 95 and 96 in the post-OR region [
22], resulting in a shorter PK-resistant C-terminal fragment in western blotting. By contrast, RML-, 22L and FK-1-PrP
Sc have a longer PK-resistant C-terminal fragment, indicating that the post-OR region is conserved in the PK-resistant core of RML-, 22L- and FK-1-PrP
Sc. Thus, RML, 22L and FK-1 prions may favor the post-OR residues more than BSE prions for the conversion of PrP
C into PrP
Sc. As a result, PrP∆91-106 is able to convert into PrP
Sc∆91–106 in Tg(PrP∆91–106)/
Prnp0/0 mice inoculated with BSE prions but not with RML, 22L and FK-1 prions. Alternatively, according to the conformational selection model, inoculated PrP
Sc selects host PrP
C as a substrate for conversion on the basis of its conformational compatibility with the host PrP
C, thereby converting the host PrP
C into PrP
Sc in a strain-dependent manner [
20,
21]. It is therefore also possible that, due to the deletion of residues 91–106, PrP∆91–106 may adopt a conformation that is not compatible with RML-, 22L- and FK-1-PrP
Sc but which is marginally compatible with BSE-PrP
Sc. As a result, PrP∆91–106 may be slightly more convertible into PrP
Sc∆91–106 in Tg(PrP∆91–106)/
Prnp0/0 mice after inoculation with BSE-PrP
Sc but not with RML-, 22L- and FK-1-PrP
Sc. Moreover, we observed that secondary inoculation with PrP
Sc∆91–106-BSE prions into Tg(PrP∆91–106)/
Prnp0/0 mice caused disease much earlier than Tg(PrP∆91–106)/
Prnp0/0 mice inoculated with WT PrP
Sc-BSE prions (343 ± 28 vs. 538 ± 24 dpi). In contrast, WT mice inoculated with PrP
Sc∆91–106-BSE prions developed disease much later than WT mice inoculated with WT PrP
Sc-BSE prions (326 ± 29 vs 174 ± 4 dpi). These results suggest that PrP∆91–106 and WT PrP
Sc prions create so-called prion transmission barriers, which are often observed between the PrP
C in recipient animals and PrP
Sc in an inoculum when their primary sequences vary [
23]. It is thus also possible that the prion transmission barrier created between PrP∆91–106 and WT BSE-PrP
Sc prions may be substantial but not so high, compared to those between PrP∆91–106 and WT RML-, 22L- or FK-1-PrP
Sc-prions. This results in PrP∆91–106 being able to support BSE infection but not infection with RML, 22L and FK-1 prions. To further characterize BSE-PrP
Sc∆91–106 prions, it would be interesting to perform the third inoculation of BSE-PrP
Sc∆91–106 prions into Tg(PrP∆91–106)/
Prnp0/0 mice.
We showed that BSE-PrP
Sc but not RML- and 22L-PrP
Sc, converted PrP∆91–104 into PrP
Sc∆91–104 in PMCA, while digitonin and heparin stimulated the conversion of PrP∆91–104 into PrP
Sc∆91–104 in PMCA even with RML- and 22L-PrP
Sc. These results suggest that the deletion of residues 91–104 in PrP∆91–106 may be responsible for the prion strain-dependent resistance of Tg(PrP∆91–106)/
Prnp0/0 mice and that digitonin and heparin may play a role similar to that of residues 91–104 in the conversion of PrP
C to PrP
Sc. Digitonin, a non-ionic detergent, has been shown to enhance the conversion of PrP
C into PrP
Sc in PMCA, as well as prevent PrP
C from forming autonomous aggregates, thereby increasing the levels of free, soluble PrP
C and facilitating the conversion of PrP
C into PrP
Sc in PMCA [
17]. Thus, it is possible that residues 91–106 may also be involved in the structural stability of PrP
C, preventing PrP
C from becoming structurally unstable enough to form autonomous aggregates. Heparin has been also reported to enhance the conversion of PrP
C into PrP
Sc in PMCA [
18]. Heparin binds to residues 23–52, 53–93 and 110–128 of PrP
C and to the PK-resistant core of PrP
Sc via sulfated groups [
18,
24]. A previous study reported that heparin promoted the formation of amyloid fibrils from a human PrP peptide comprised of residues 106–126, indicating that heparin may function as a scaffold for the formation of amyloid fibers from peptides [
25]. Since all of the heparin-binding sites of PrP∆91–104 are intact, it is possible that PrP∆91–104 is assembled on the heparin scaffold and converted into PrP
Sc∆91–104 in PMCA and that residues 91–106 may be involved in the assembly of the PrP
C molecules themselves or in PrP
C and PrP
Sc upon the conversion of PrP
C into PrP
Sc.
Residues 91–106 encompass the majority of the positively-charged central region, which is rich in lysine residues. This region was previously demonstrated to be involved in the lipid-induced conversion of recombinant PrP into PK-resistant PrP [
26]. It is thus possible that the positive charge in residues 91–106 may be important for PrP
C molecules either to be structurally stable and soluble or to assemble with themselves and/or with PrP
Sc. Alternatively, residues 91–106 contain proline residues, which are mutated to leucine residues in human hereditary prion diseases (P101L and P104L in mouse PrP homologous to P102L and P105L in human PrP). These mutations are considered to trigger structural changes in the surrounding region, eventually causing the structural instability of the mutant PrPs and thereby triggering the conversion of mutant PrPs into pathogenic PrPs [
27]. Therefore, it is possible that residues 91–106 play an important role in the structural integrity of PrP
C. Furthermore, the substitution of copper-binding histidine (H) residue to glycine (G) residue at position 95 in mouse PrP was reported to accelerate prion disease in mice after infection with RML prions [
28], suggesting that copper-binding at H95 might also structurally stabilize PrP
C and thereby play an inhibitory role in the conversion of PrP
C into PrP
Sc.
In summary, we showed that Tg(PrP∆91–106)/Prnp0/0 mice were resistant to RML, 22L and FK-1 prions and remained marginally susceptible to BSE prions, suggesting that residues 91–106 of PrPC are crucially involved in prion pathogenesis in a strain-dependent manner. We also showed that these residues might have a similar role to that of digitonin and heparin in the conversion of PrPC into PrPSc. The precise role of residues 91–106 in the conversion of PrPC into PrPSc would be helpful not just to improve our understanding of the mechanism underlying the conversion of PrPC into PrPSc but also for the development of therapeutic agents in prion diseases.
4. Materials and Methods
4.1. Ethics Statements
The Ethics Committees of Animal Care and Experimentation of the University of Occupational and Environmental Health (approval number AE08-013, 18 March 2019), Kyoto University (approval number S-13-10-2, 1 April 2017) and the Institutional Animal Care and Use Committee at the National Institute of Animal Health (approval number 13-005, 13 April 2013) approved this study. Animals were cared for in accordance with The Guiding Principle for Animal Care and Experimentation of the University of Occupational and Environmental Health, Kyoto University, the National Institute of Animal Health and Japanese Law for Animal Welfare and Care. Every effort was made to reduce distress and the number of animals used.
4.2. Generation of Tg(PrP∆91–106)/Prnp0/0 Mice
A DNA fragment encoding residues 1–90 fused with residues 107–111 of mouse PrP was amplified by polymerase chain reaction (PCR) with a sense primer (5′-ccc
gtcgacctcgagatggcgaaccttggc-3′; underlined sequence,
Sal I and
Xho I sites; bold sequence, start codon) and an antisense primer (5′-
cacatgcttgaggttttggccccatccacc-3′; underlined sequence, residues 107–111; italic sequence, residues 86–90) using mouse WT PrP
C-encoding pcDNA3.1-moPrP [
29] as a template. The resulting DNA fragment was then utilized as a 5′ primer to amplify another DNA fragment encoding PrP∆91–106 together with an antisense primer (5′-ccc
gtcgacctcgagtcatcccacgatcag-3′; underlined sequence,
Sal I and
Xho I sites; bold sequence, stop codon) using pcDNA3.1-moPrP as a template. The amplified DNA was then cloned into pCR2.1-TOPO TA vector (Thermo Fisher Scientific, Waltham, MA, USA), resulting in pCR2.1-TOPO TA-PrP∆91–106. After DNA sequence confirmation, it was inserted into a unique
Sal I site of the Syrian hamster PrP cosmid vector, CosSHa.tet (InPro Biotechnology, Inc., South San Francisco, CA, USA), to construct PrP∆91–106 transgene. The transgene was injected into the zygotes of C57BL/6 mice, as described elsewhere [
30,
31], after the removal of the cosmid-derived sequences, resulting in a line of Tg(PrP∆91–106) mice. The Tg(PrP∆91–106) mice were successively mated with
Prnp0/0 mice, which had been intercrossed with C57BL/6 mice more than 10 times [
32,
33], to produce Tg(PrP∆91–106)/
Prnp0/0 mice.
4.3. Prion Inoculation
Brains from terminally ill C57BL/6 mice infected with RML, 22L, FK-1 and BSE prions were homogenized (10%, w/v) in phosphate-buffered saline (PBS) by passing through 18 to 26 gauge needles and then diluted to 1% with PBS. Four to five week-old Tg(PrP∆91–106)/Prnp0/0 and C57BL/6 mice (CLEA Japan, Tokyo, Japan) were intracerebrally inoculated with a 20 μL-aliquot of the homogenates.
4.4. Establishment of N2a∆PrP/PrP∆91–106 Cells
The
BamH I/
Xba I DNA fragment including mouse PrP∆91–106 from pCR2.1-TOPO TA-PrP∆91–106 was inserted into pcDNA3.1(+) (Thermo Fisher Scientific) to construct an expression vector for PrP∆91–106, designated pcDNA3.1(+)-PrP∆91–106. The vector was transfected into PrP
C-knockout N2a∆PrP cells [
14] using Lipofectamine 2000 (Thermo Fisher Scientific). The transfected cells were then subjected to neomycin selection followed by a limiting dilution to produce N2a∆PrP/PrP∆91–106 cells.
4.5. Western Blotting
Tissues were homogenized (10%, w/v) using a Multi-beads shocker (Yasui Kikai Co., Osaka, Japan) in lysis buffer (150 mM NaCl, 50 mM Tris-HCl, pH 7.5, 0.5% Triton X-100, 0.5% sodium deoxycholate, 1 mM EDTA) containing protease inhibitor mixture (Nakalai Tesque Co., Kyoto, Japan) and protein concentrations were determined using the BCA protein assay kit (Pierce, Rockford, IL, USA). The homogenates were then subjected to an SDS-polyacrylamide gel electrophoresis after treatment with or without PK (Wako Pure Chemical Industries, Ltd., Osaka, Japan) at 20 µg/mL for 30 min at 37 °C and then the total proteins were electrically transferred to an Immobilon-P PVDF membrane (Millipore Corp., MA, USA). The membrane was blocked using TBST (0.1% Tween-20, 100 mM NaCl, 10 mM Tris-HCl, pH7.6) containing 5% non-fat dry milk for 1 h at room temperature (RT), incubated with rabbit polyclonal anti-PrP antibody (18635; Immuno Biological Laboratories, Gunma, Japan), SAF61 mouse monoclonal antibody (Bertin Pharma, Montigny le Bretonneux, France), rabbit polyclonal anti-Flottilin-2 antibody (Cell Signaling, MA, USA) or anti-β-actin monoclonal antibody (Sigma-Aldrich, St. Louis, MO, USA) for 2 h at RT or overnight at 4 °C in TBST containing 1% non-fat dry milk and washed in TBST several times. Chemiluminescent signals were detected using horseradish peroxidase (HRP)-conjugated anti-mouse IgG antibodies (Amersham Biosciences, Piscataway, NJ, USA), anti-goat IgG antibodies (CHEMICON International., Temecula, CA, USA) and Immobilon Western Chemiluminescent HRP substrate (Millipore), with a chemiluminescence image analyzer (LAS-4000 mini; Fujifilm Co., Tokyo, Japan).
4.6. HE Staining
Samples embedded in paraffin were sliced at 5 μm and the sliced samples were then deparaffinized, rehydrated, stained with Mayer’s hematoxylin solution (Wako Pure Chemical Industries, Osaka, Japan) and 1% Eosin Y solution (Wako Pure Chemical Industries) and mounted with Softmount (Wako Pure Chemical Industries).
4.7. Immunohistochemistry
Paraffin-embedded samples were sectioned, deparaffinized and rehydrated. The samples were autoclaved in 1 mM HCl at 105 °C for 5 min and subsequently washed with PBS. For the detection of PrPSc, the samples were digested with 50 μg/mL PK in PBS at 37 °C for 30 min, treated with 3 M guanidine thiocyanate for 10 min and washed with PBS. After blocking with 5% FBS in PBS for 1 h, the samples were incubated with SAF83 (Bertin Pharma, Montigny le Bretonneux, France) anti-PrP Ab or rabbit polyclonal anti-GFAP antibody (Shima laboratories Co., Ltd., Tokyo, Japan) for 2 h, washed with PBS and treated with ImmPRESS REAGENT Anti-Mouse IgG (Vector Laboratories, Burlingame, CA, USA) for 30 min. After washing with PBS, the samples were incubated with ImmPACT DAB (Vector Laboratories) for 180 s for staining of PrPSc or 60 s for staining of GFAP.
Cells grown on coverslips were fixed with 3% PFA for 10 min at room temperature and permeabilized with 0.1% Triton X-100 in PBS for 4 min at room temperature. After blocking with 5% FBS in PBS for 1 h, the cells were stained with SAF83 anti-PrP Ab (Bertin pharma) for 2 h, washed with PBS and stained with Alexa Fluoro 488-conjugated anti-mouse IgG Abs (Thermo Fisher Scientific) and 1 μg/mL DAPI (Sigma-Aldrich, St Louis, MO, USA) for 2 h at room temperature.
4.8. Fractionation of Membrane Microdomains
Cells grown to ~80% confluency in a 60-mm tissue culture dish were suspended in 250 μL of MBS buffer (25 mM MES-NaOH (pH 6.5), 0.15 M NaCl) containing 1% (w/v) Triton X-100 and homogenized by being passed through a 21G-needle 15 times. After centrifugation at 500× g for 5 min at 4 °C, 220 μL of the supernatant was transferred to a new tube and mixed with 220 μL of MBS buffer containing 80% (w/v) sucrose to make 40% (w/v) sucrose. Then, 200 μL of sample was placed at the bottom of a discontinuous sucrose gradient consisting of 600 μL of 30% (w/v) sucrose and 200 μL of 5% (w/v) sucrose and centrifuged at 140,000× g for 24 h at 4 °C in an S55S rotor (Hitachi Koki, Tokyo, Japan). Ten fractions (100 μL/fraction) were collected from the top.
4.9. Preparation of Insect Cell Suspension Expressing Recombinant WT PrP and PrP∆91–104
A baculovirus expression vector for mouse WT PrP, termed pFastBac-moPrP, was used as constructed elsewhere [
34] and a baculovirus expression vector for PrP∆91–104, pFastBac-moPrP∆91–104, was constructed in the present study. In brief, a DNA fragment encoding residues 1–90 fused with residues 105–109 of mouse PrP was amplified by PCR with a Met forward sense primer (5′-cg
ggatccgccacc
atggcgaaccttggctac-3′; underlined sequence,
BamH I site; bold sequence, start codon) and a moPrP∆91–104 antisense primer (5′-
gaggttggttttttggccccat-3′; underlined sequence, residues 105–108; italic sequence, residues 88–90) using pFastBac-moPrP as a template. The resulting DNA fragment was then utilized as a 5′ primer to amplify another DNA fragment encoding PrP∆91–104 together with a Stop reverse antisense primer (5′-ccc
aagctttcatcccacgatcaggaag-3′; underlined sequence,
Hind III site; bold sequence, stop codon) using pFastBac-moPrP as a template. The amplified DNA were cloned into pCR2.1-TOPO TA vector (Thermo Fisher Scientific). After confirming the DNA sequence of the fragments, these were inserted into the
BamH I/
Hind III-digested baculovirus expression vector pFastBac1 (Thermo Fisher Scientific).
Recombinant WT PrP and PrP∆91–104 were prepared according to the manufacturer’s protocol (Thermo Fisher Scientific). pFastBac-moPrP(3F4) and pFastBac-moPrP∆91–104 were first introduced into DH10 Bac E. coli to obtain recombinant bacmid encoding each molecule. The bacmid was then transfected into Spodoptera frugiperda 21 insect cells with a bacmid DNA-cellfectin mixture (Thermo Fisher Scientific) to obtain recombinant baculovirus encoding WT PrP or PrP∆91–104. HighFive cells (Thermo Fisher Scientific) were infected with the recombinant baculovirus encoding each molecule for 72 h at 27 °C to produce recombinant WT PrP or PrP∆91–104. The cells were suspended in distilled water at a concentration of 5 × 104 cells/µL for PMCA. The yield of recombinant WT PrP or PrP∆91–104 in the cells was approximately 16 µg/106 cells.
4.10. PMCA
The brains from RML-, 22L- or BSE-infected, terminal ill ICR mice were homogenized at a 20% (
w/
v) concentration in PBS containing a complete protease inhibitor cocktail (Roche) and then diluted to 10% by adding an equal volume of 2× PMCA buffer (2× PBS, 8 mM EDTA, 2% Triton X-100). Uninfected ICR mouse brains were prepared in a similar manner as a control. ICR mice were purchased from Japan SLC Inc. (Hamamatsu, Japan). Briefly, 5 µL of cell suspension (5 × 10
4 cells/µL) producing WT PrP or PrPΔ91–104 was added to 95 µL of each of the 10% brain homogenates and subjected to PMCA procedures. PMCA was performed using automatic cross-ultrasonic protein activating apparatus (Elestein 070-GOT; Elekon Science Corp., Chiba, Japan), as previously reported [
34]. Amplification was performed using 32 cycles of sonication (pulse oscillation for 3 s repeated five times at 0.1-s intervals), followed by incubation at 37 °C for 30 min with gentle agitation. The amplified products of the first round of amplification were diluted 1:10 with the WT PrP or PrP∆91–104 cell suspension and subjected to a second round of amplification. This process was repeated up to 6 rounds. The products from each round were treated with 40 µg/mL of PK for 1 h at 37 °C and subjected to western blotting with HRP-conjugated T2 anti-PrP antibody [
19]. Signals were detected using SuperSignal West Dura Extended Duration Substrate (Pierce) with a ChemiImager (Alpha InnoTec, San Leandro, CA, USA).