The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance
Abstract
:1. Introduction
2. Domain Structure of CCCH Zinc-Finger Proteins
3. Subcellular Localization of CCCH Zinc-Finger Proteins
4. Transcriptional and Post-Transcriptional Regulation by CCCH Zinc-Finger Proteins
4.1. Transcriptional Regulation
4.2. Post-Transcriptional Regulation
5. Interactions of CCCH Zinc-Finger Proteins with Other Proteins
6. CCCH Zinc-Finger Proteins and Plant Hormones
7. The Roles of CCCH Zinc-Finger Proteins in Abiotic Stress
7.1. Oxidative Stress
7.2. Salt Stress
7.3. Drought Stress
7.4. Flooding Stress
7.5. Cold Stress
7.6. Multiple Stresses
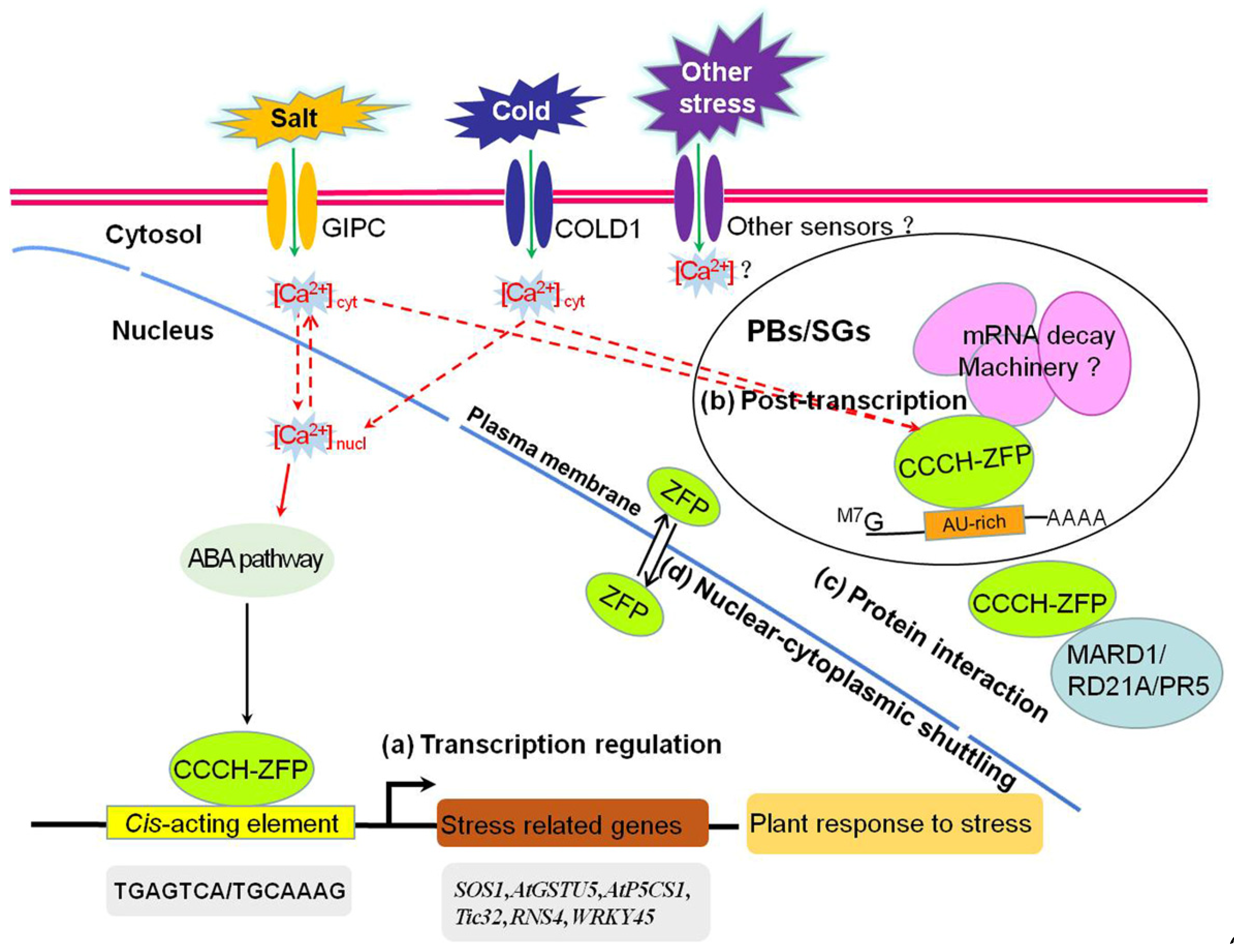
8. Mechanisms That Regulate CCCH Zinc-Finger Proteins under Abiotic Stresses
9. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chen, M.; Yang, Z.; Liu, J.; Zhu, T.; Wei, X.; Fan, H.; Wang, B. Adaptation mechanism of salt excluders under saline conditions and its applications. Int. J. Mol. Sci. 2018, 19, 3668. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, T.; Yang, Z.; Wei, X.; Yuan, F.; Yin, S.; Wang, B. Evaluation of salt-tolerant germplasm and screening of the salt-tolerance traits of sweet sorghum in the germination stage. Funct. Plant Biol. 2018, 45, 1073–1081. [Google Scholar] [CrossRef]
- Li, B.; Gao, K.; Ren, H.; Tang, W. Molecular mechanisms governing plant responses to high temperatures. J. Integr. Plant Biol. 2018, 60, 757–779. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Wang, B.; Gao, S.; Zhang, F.; Terzaghi, W.; Dai, M. Genome-wide association study dissects the genetic bases of salt tolerance in maize seedlings. J. Integr. Plant Biol. 2019, 61, 658–674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rascio, N.; Navari-Izzo, F. Heavy metal hyperaccumulating plants: How and why do they do it? And what makes them so interesting? Plant Sci. 2011, 180, 169–181. [Google Scholar] [CrossRef]
- Yuan, F.; Guo, J.; Shabala, S.; Wang, B. Reproductive Physiology of Halophytes: Current Standing. Front. Plant Sci. 2019, 9, 1954. [Google Scholar] [CrossRef]
- Lu, C.; Feng, Z.; Yuan, F.; Han, G.; Wang, B. The SNARE protein LbSYP61 participates in salt secretion in Limonium bicolor. Environ. Exp. Bot. 2020, 176, 104076. [Google Scholar] [CrossRef]
- Guo, J.; Dong, X.; Li, Y.; Wang, B. NaCl treatment markedly enhanced pollen viability and pollen preservation time of euhalophyte Suaeda salsa via up regulation of pollen development-related genes. J. Plant Res. 2020, 133, 57–71. [Google Scholar] [CrossRef]
- Cheng, S.; Yang, Z.; Wang, M.; Song, J.; Sui, N.; Fan, H. Salinity improves chilling resistance in Suaeda salsa. Acta Physiol. Plant 2014, 36, 1823–1830. [Google Scholar] [CrossRef]
- Jiang, J.; Ma, S.; Ye, N.; Jiang, M.; Cao, J.; Zhang, J. WRKY transcription factors in plant responses to stresses. J. Integr. Plant Biol. 2017, 59, 86–101. [Google Scholar] [CrossRef] [PubMed]
- Zubair, M.; Khan, Q.U.; Mirza, N.; Sarwar, R.; Khan, A.A.; Baloch, M.S.; Fahad, S.; Shah, A.N. Physiological response of spinach to toxic heavy metal stress. Environ. Sci. Pollut. Res. Int. 2019, 26, 31667–31674. [Google Scholar] [CrossRef]
- Song, Y.; Li, J.; Liu, M.; Meng, Z.; Liu, K.; Sui, N. Nitrogen increases drought tolerance in maize seedlings. Funct. Plant Biol. 2019, 46, 350–359. [Google Scholar] [CrossRef]
- Wang, W.; Zheng, H.; Wang, Y.; Han, G.; Sui, N. Overexpression of CCCH zinc finger protein gene delays flowering time and enhances salt tolerance in Arabidopsis by increasing fatty acid unsaturation. Acta Physiol. Plant 2018, 40, 196. [Google Scholar] [CrossRef]
- Li, X.; Zeng, R.; Liao, H. Improving crop nutrient efficiency through root architecture modifications. J. Integr. Plant Biol. 2016, 58, 193–202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, J.; Lu, C.; Zhao, F.; Gao, S.; Wang, B. Improved reproductive growth of euhalophyte Suaeda salsa under salinity is correlated with altered phytohormone biosynthesis and signal transduction. Funct. Plant Biol. 2020, 47, 170–183. [Google Scholar] [CrossRef]
- Liu, Q.; Liu, R.; Ma, Y.; Song, J. Physiological and molecular evidence for Na+ and Cl− exclusion in the roots of two Suaeda salsa populations. Aquat. Bot. 2018, 146, 1–7. [Google Scholar] [CrossRef]
- Qi, Y.; Li, J.; Chen, C.; Li, L.; Zheng, X.; Liu, J.; Zhu, T.; Pang, C.; Wang, B.; Chen, M. Adaptive growth response of exotic Elaeagnus angustifolia L. to indigenous saline soil and its beneficial effects on the soil system in the Yellow River Delta, China. Trees-Struct. Funct. 2018, 32, 1723–1735. [Google Scholar] [CrossRef]
- Wu, Q.; Wang, M.; Shen, J.; Chen, D.; Zheng, Y.; Zhang, W. ZmOST1 mediates abscisic acid regulation of guard cell ion channels and drought stress responses. J. Integr. Plant Biol. 2019, 61, 478–491. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.J.; Wu, W.H.; Wang, Y. ZmHAK5 and ZmHAK1 function in K+ uptake and distribution in maize under low K+ conditions. J. Integr. Plant Biol. 2019, 61, 691–705. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.; Song, Y.; Zheng, H.; Zhang, Y.; Guo, J.; Sui, N. NADP-Malate Dehydrogenase of Sweet Sorghum Improves Salt Tolerance of Arabidopsis thaliana. J. Agric. Food Chem. 2018, 66, 5992–6002. [Google Scholar] [CrossRef] [PubMed]
- Han, G.; Wang, M.; Yuan, F.; Sui, N.; Song, J.; Wang, B. The CCCH zinc finger protein gene AtZFP1 improves salt resistance in Arabidopsis thaliana. Plant Mol. Biol. 2014, 86, 237–253. [Google Scholar] [CrossRef]
- Huang, J.; Yang, M.; Zhang, X. The function of small RNAs in plant biotic stress response. J. Integr. Plant Biol. 2016, 58, 312–327. [Google Scholar] [CrossRef] [Green Version]
- Rozentsvet, O.A.; Nesterov, V.N.; Bogdanova, E.S.; Rozenberg, G.S. Physiological and Biochemical Strategies of Adaptation in Halophytes. Dokl. Biol. Sci. 2020, 492, 83–85. [Google Scholar] [CrossRef]
- Matinzadeh, Z.; Akhani, H.; Abedi, M.; Palacio, S. The elemental composition of halophytes correlates with key morphological adaptations and taxonomic groups. Plant Physiol. Biochem. 2019, 141, 259–278. [Google Scholar] [CrossRef] [Green Version]
- Xi, J.J.; Chen, H.Y.; Bai, W.P.; Yang, R.C.; Yang, P.Z.; Chen, R.J.; Hu, T.M.; Wang, S.M. Sodium-Related Adaptations to Drought: New Insights From the Xerophyte Plant Zygophyllum xanthoxylum. Front. Plant Sci. 2018, 9, 1678. [Google Scholar] [CrossRef] [PubMed]
- Boukhris, A.; Laffont-Schwob, I.; Folzer, H.; Rabier, J.; Mezghani, I.; Salducci, M.D.; Tatoni, T.; Chaieb, M. Tolerance strategies of two Mediterranean native xerophytes under fluoride pollution in Tunisia. Environ. Sci. Pollut. Res. Int. 2018, 25, 34753–34764. [Google Scholar] [CrossRef] [PubMed]
- Souri, Z.; Karimi, N.; Sandalio, L.M. Arsenic Hyperaccumulation Strategies: An Overview. Front. Cell Dev. Biol. 2017, 5, 67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, T.; Yan, H.; He, Z. Advances in molecular mechanisms of arsenic hyperaccumulation of Pteris vittata L. Chin. J. Biotechnol. 2020, 36, 397–406. [Google Scholar]
- Sun, X.; Zheng, H.; Sui, N. Regulation mechanism of long non-coding RNA in plant response to stress. Biochem. Biophys. Res. Commun. 2018, 503, 402–407. [Google Scholar] [CrossRef]
- Meng, C.; Sui, N. Overexpression of maize MYB-IF35 increases chilling tolerance in Arabidopsis. Plant Physiol. Biochem. 2019, 135, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Li, J.Q.; Yan, J.Y.; Yuan, J.J.; Li, G.X.; Wu, Y.R.; Xu, J.M.; Huang, R.F.; Harberd, N.P.; Ding, Z.J. Ethylene promotes seed iron storage during Arabidopsis seed maturation via ERF95 transcription factor. J. Integr. Plant Biol. 2020, 62, 1193–1212. [Google Scholar] [CrossRef] [PubMed]
- Tan, Z.; Wen, X.; Wang, Y. Betula platyphylla BpHOX2 transcription factor binds to different cis-acting elements and confers osmotic tolerance. J. Integr. Plant Biol. 2020, 62, 1762–1779. [Google Scholar] [CrossRef]
- Wang, H.; Klatte, M.; Jakoby, M.; Baumlein, H.; Weisshaar, B.; Bauer, P. Iron deficiency-mediated stress regulation of four subgroup Ib BHLH genes in Arabidopsis thaliana. Planta 2007, 226, 897–908. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Gao, W.; Peng, Q.; Zhou, B.; Kong, Q.; Ying, Y.; Shou, H. Two soybean bHLH factors regulate response to iron deficiency. J. Integr. Plant Biol. 2018, 60, 608–622. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Yang, C.; Cao, J.; Chen, H.; Pang, J.; Zhao, Q.; Wang, Z.; Qing Fu, Z.; Liu, J. A bHLH transcription activator regulates defense signaling by nucleo-cytosolic trafficking in rice. J. Integr. Plant Biol. 2020, 62, 1552–1573. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Wang, P.; Hou, L.; Zhao, S.; Zhao, C.; Xia, H.; Li, P.; Zhang, Y.; Bian, X.; Wang, X. Global Analysis of WRKY Genes and Their Response to Dehydration and Salt Stress in Soybean. Front. Plant Sci. 2016, 7, 9. [Google Scholar] [CrossRef] [Green Version]
- Song, Y.; Li, J.; Sui, Y.; Han, G.; Zhang, Y.; Guo, S.; Sui, N. The sweet sorghum SbWRKY50 is negatively involved in salt response by regulating ion homeostasis. Plant Mol. Biol. 2020, 102, 603–614. [Google Scholar] [CrossRef]
- Li, C.X.; Yan, J.Y.; Ren, J.Y.; Sun, L.; Xu, C.; Li, G.X.; Ding, Z.J.; Zheng, S.J. A WRKY transcription factor confers aluminum tolerance via regulation of cell wall modifying genes. J. Integr. Plant Biol. 2020, 62, 1176–1192. [Google Scholar] [CrossRef]
- Zou, M.; Guan, Y.; Ren, H.; Zhang, F.; Chen, F. A bZIP transcription factor, OsABI5, is involved in rice fertility and stress tolerance. Plant Mol. Biol. 2008, 66, 675–683. [Google Scholar] [CrossRef]
- Zong, W.; Yang, J.; Fu, J.; Xiong, L. Synergistic regulation of drought-responsive genes by transcription factor OsbZIP23 and histone modification in rice. J. Integr. Plant Biol. 2020, 62, 723–729. [Google Scholar] [CrossRef]
- Wang, X.; Lu, X.; Malik, W.A.; Chen, X.; Wang, J.; Wang, D.; Wang, S.; Chen, C.; Guo, L.; Ye, W. Differentially expressed bZIP transcription factors confer multi-tolerances in Gossypium hirsutum L. Int. J. Biol. Macromol. 2020, 146, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Zhang, Y.; Tao, J.; Chen, H.; Li, Q.; Zhang, W.; Ma, B.; Lin, Q.; Zhang, J.; Chen, S. The Alfin-like homeodomain finger protein AL5 suppresses multiple negative factors to confer abiotic stress tolerance in Arabidopsis. Plant J. 2015, 81, 871–883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brandt, R.; Cabedo, M.; Xie, Y.; Wenkel, S. Homeodomain leucine-zipper proteins and their role in synchronizing growth and development with the environment. J. Integr. Plant Biol. 2014, 56, 518–526. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.-S.; Jiang, J.; Han, X.-J.; Zhang, Y.-X.; Zhuo, R.-Y. Identification, expression analysis of the Hsf family, and characterization of class A4 in Sedum Alfredii hance under cadmium stress. Int. J. Mol. Sci. 2018, 19, 1216. [Google Scholar] [CrossRef] [Green Version]
- Morton, E.A.; Lamitina, T. Caenorhabditis elegans HSF-1 is an essential nuclear protein that forms stress granule-like structures following heat shock. Aging Cell 2013, 12, 112–120. [Google Scholar] [CrossRef] [Green Version]
- Liu, M.; Zhu, J.; Dong, Z. Immediate transcriptional responses of Arabidopsis leaves to heat shock. J. Integr. Plant Biol. 2020. [Google Scholar] [CrossRef]
- Hong, Y.; Zhang, H.; Huang, L.; Li, D.; Song, F. Overexpression of a Stress-Responsive NAC Transcription Factor Gene ONAC022 Improves Drought and Salt Tolerance in Rice. Front. Plant Sci. 2016, 7, 4. [Google Scholar] [CrossRef]
- Wang, B.; Wei, J.; Song, N.; Wang, N.; Zhao, J.; Kang, Z. A novel wheat NAC transcription factor, TaNAC30, negatively regulates resistance of wheat to stripe rust. J. Integr. Plant Biol. 2018, 60, 432–443. [Google Scholar] [CrossRef]
- Zhao, J.; Liu, J.S.; Meng, F.N.; Zhang, Z.Z.; Long, H.; Lin, W.H.; Luo, X.M.; Wang, Z.Y.; Zhu, S.W. ANAC005 is a membrane-associated transcription factor and regulates vascular development in Arabidopsis. J. Integr. Plant Biol. 2016, 58, 442–451. [Google Scholar] [CrossRef] [Green Version]
- Wei, Q.; Luo, Q.; Wang, R.; Zhang, F.; He, Y.; Zhang, Y.; Qiu, D.; Li, K.; Chang, J.; Yang, G. A Wheat R2R3-type MYB Transcription Factor TaODORANT1 Positively Regulates Drought and Salt Stress Responses in Transgenic Tobacco Plants. Front. Plant Sci. 2017, 8, 1374. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Liu, G.; Jia, J.; Zhao, G.; Xia, C.; Zhang, L.; Li, F.; Zhang, Q.; Dong, C.; Gao, S. The wheat MYB-related transcription factor TaMYB72 promotes flowering in rice. J. Integr. Plant Biol. 2016, 58, 701–704. [Google Scholar] [CrossRef] [Green Version]
- Guo, H.; Wang, L.; Yang, C.; Zhang, Y.; Zhang, C.; Wang, C. Identification of novel cis-elements bound by BplMYB46 involved in abiotic stress responses and secondary wall deposition. J. Integr. Plant Biol. 2018, 60, 1000–1014. [Google Scholar] [CrossRef] [Green Version]
- Chen, R.; Ma, J.; Luo, D.; Hou, X.; Ma, F.; Zhang, Y.; Meng, Y.; Zhang, H.; Guo, W. CaMADS, a MADS-box transcription factor from pepper, plays an important role in the response to cold, salt, and osmotic stress. Plant Sci. 2019, 280, 164–174. [Google Scholar] [CrossRef]
- Meng, Q.; Li, X.; Zhu, W.; Yang, L.; Liang, W.; Dreni, L.; Zhang, D. Regulatory network and genetic interactions established by OsMADS34 in rice inflorescence and spikelet morphogenesis. J. Integr. Plant Biol. 2017, 59, 693–707. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haas, M.; Schreiber, M.; Mascher, M. Domestication and crop evolution of wheat and barley: Genes, genomics, and future directions. J. Integr. Plant Biol. 2019, 61, 204–225. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.-L.; Zhang, C.-L.; Wang, G.-L.; Wang, Y.-X.; Qi, C.-H.; You, C.-X.; Li, Y.-Y.; Hao, Y.-J. Apple AP2/EREBP transcription factor MdSHINE2 confers drought resistance by regulating wax biosynthesis. Planta 2019, 249, 1627–1643. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Yin, Y.; You, C.; Pan, Q.; Xu, D.; Jin, T.; Zhang, B.; Ma, H. Evolution and protein interactions of AP2 proteins in Brassicaceae: Evidence linking development and environmental responses. J. Integr. Plant Biol. 2016, 58, 549–563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, C.; Zhang, T.Z. Functional diversifications of GhERF1 duplicate genes after the formation of allotetraploid cotton. J. Integr. Plant Biol. 2019, 61, 60–74. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Ni, L.; Liu, Y.; Wang, Y.; Zhang, A.; Tan, M.; Jiang, M. The C2H2-type Zinc Finger Protein ZFP182 is Involved in Abscisic Acid-Induced Antioxidant Defense in Rice F. J. Integr. Plant Biol. 2012, 54, 500–510. [Google Scholar] [CrossRef]
- Yang, L.; Liu, Q.; Liu, Z.; Yang, H.; Wang, J.; Li, X.; Yang, Y. Arabidopsis C3HC4-RING finger E3 ubiquitin ligase AtAIRP4 positively regulates stress-responsive abscisic acid signaling. J. Integr. Plant Biol. 2016, 58, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Chen, H.; Hou, Z.; Li, Y.; Yang, C.; Wang, D.; Song, C.P. Screening of abiotic stress-responsive cotton genes using a cotton full-length cDNA overexpressing Arabidopsis library. J. Integr. Plant Biol. 2020, 62, 998–1016. [Google Scholar] [CrossRef]
- Han, G.; Lu, C.; Guo, J.; Qiao, Z.; Sui, N.; Qiu, N.; Wang, B. C2H2 Zinc Finger Proteins: Master Regulators of Abiotic Stress Responses in Plants. Front. Plant Sci. 2020, 11, 115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, D.; Guo, Y.; Wu, C.; Yang, G.; Li, Y.; Zheng, C. Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice. BMC Genom. 2008, 9, 44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, X.; Zhao, Y.; Cao, J.; Zhang, W.; Jiang, H.; Li, X.; Ma, Q.; Zhu, S.; Cheng, B. CCCH-type zinc finger family in maize: Genome-wide identification, classification and expression profiling under abscisic acid and drought treatments. PLoS ONE 2012, 7, e40120. [Google Scholar] [CrossRef] [Green Version]
- Chai, G.; Hu, R.; Zhang, D.; Qi, G.; Zuo, R.; Cao, Y.; Chen, P.; Kong, Y.; Zhou, G. Comprehensive analysis of CCCH zinc finger family in poplar (Populus trichocarpa). BMC Genom. 2012, 13, 253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Zhang, H.; Zhao, Y.; Jiang, H.; Zhu, S.; Cheng, B.; Xiang, Y. Genome-wide analysis of the CCCH zinc finger gene family in Medicago truncatula. Plant Cell Rep. 2013, 32, 1543–1555. [Google Scholar] [CrossRef] [PubMed]
- Jiang, A.-L.; Xu, Z.-S.; Zhao, G.-Y.; Cui, X.-Y.; Chen, M.; Li, L.-C.; Ma, Y.-Z. Genome-wide analysis of the C3H zinc finger transcription factor family and drought responses of members in Aegilops tauschii. Plant Mol. Biol. Report. 2014, 32, 1241–1256. [Google Scholar] [CrossRef]
- Xu, R. Genome-wide analysis and identification of stress-responsive genes of the CCCH zinc finger family in Solanum lycopersicum. Mol. Genet. Genom. 2014, 289, 965–979. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhong, Y.; Cheng, Z.M. Evolution and Expression Analysis of the CCCH Zinc Finger Gene Family in Vitis vinifera. Plant Genome 2014, 7. [Google Scholar] [CrossRef] [Green Version]
- Pradhan, S.; Kant, C.; Verma, S.; Bhatia, S. Genome-wide analysis of the CCCH zinc finger family identifies tissue specific and stress responsive candidates in chickpea (Cicer arietinum L.). PLoS ONE 2017, 12, e0180469. [Google Scholar] [CrossRef] [PubMed]
- Pi, B.; He, X.; Ruan, Y.; Jang, J.-C.; Huang, Y. Genome-wide analysis and stress-responsive expression of CCCH zinc finger family genes in Brassica rapa. BMC Plant Biol. 2018, 18, 373. [Google Scholar] [CrossRef] [Green Version]
- Kong, Z.; Li, M.; Yang, W.; Xu, W.; Xue, Y. A Novel Nuclear-Localized CCCH-Type Zinc Finger Protein, OsDOS, Is Involved in Delaying Leaf Senescence in Rice. Plant Physiol. 2006, 141, 1376–1388. [Google Scholar] [CrossRef] [Green Version]
- Bogamuwa, S.; Jang, J. The Arabidopsis tandem CCCH zinc finger proteins AtTZF4, 5 and 6 are involved in light-, abscisic acid- and gibberellic acid-mediated regulation of seed germination. Plant Cell Environ. 2013, 36, 1507–1519. [Google Scholar] [CrossRef]
- Chao, Y.; Zhang, T.; Yang, Q.; Kang, J.; Sun, Y.; Gruber, M.Y.; Qin, Z. Expression of the alfalfa CCCH-type zinc finger protein gene MsZFN delays flowering time in transgenic Arabidopsis thaliana. Plant Sci. 2014, 215, 92–99. [Google Scholar] [CrossRef]
- Kim, W.C.; Kim, J.Y.; Ko, J.H.; Kang, H.; Kim, J.; Han, K.H. AtC3H14, a plant-specific tandem CCCH zinc-finger protein, binds to its target mRNAs in a sequence-specific manner and affects cell elongation in Arabidopsis thaliana. Plant J. 2014, 80, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Chai, G.; Qi, G.; Cao, Y.; Wang, Z.; Yu, L.; Tang, X.; Yu, Y.; Wang, D.; Kong, Y.; Zhou, G. Poplar PdC3H17 and PdC3H18 are direct targets of PdMYB3 and PdMYB21, and positively regulate secondary wall formation in Arabidopsis and poplar. New Phytol. 2014, 203, 520–534. [Google Scholar] [CrossRef]
- Chen, Y.; Sun, A.; Wang, M.; Zhu, Z.; Ouwerkerk, P.B.F. Functions of the CCCH type zinc finger protein OsGZF1 in regulation of the seed storage protein GluB-1 from rice. Plant Mol. Biol. 2014, 84, 621–634. [Google Scholar] [CrossRef] [PubMed]
- Chai, G.; Kong, Y.; Zhu, M.; Yu, L.; Qi, G.; Tang, X.; Wang, Z.; Cao, Y.; Yu, C.; Zhou, G. Arabidopsis C3H14 and C3H15 have overlapping roles in the regulation of secondary wall thickening and anther development. J. Exp. Bot. 2015, 66, 2595–2609. [Google Scholar] [CrossRef] [Green Version]
- Guo, D.; Yi, H.-Y.; Li, H.-L.; Liu, C.; Yang, Z.-P.; Peng, S.-Q. Molecular characterization of HbCZF1, a Hevea brasiliensis CCCH-type zinc finger protein that regulates hmg1. Plant Cell Rep. 2015, 34, 1569–1578. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Z.; Xiong, T.; Xiong, X.; Wu, X.; Guan, C.; Xiao, G. The CCCH-type transcription factor BnZFP1 is a positive regulator to control oleic acid levels through the expression of diacylglycerol O-acyltransferase 1 gene in Brassica napus. Plant Physiol. Biochem. 2018, 132, 633–640. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Gao, X.; Zhi, Y.; Li, X.; Zhang, Q.; Niu, J.; Wang, J.; Zhai, H.; Zhao, N.; Li, J. A non-tandem CCCH-type zinc-finger protein, IbC3H18, functions as a nuclear transcriptional activator and enhances abiotic stress tolerance in sweet potato. New Phytol. 2019, 223, 1918–1936. [Google Scholar] [CrossRef]
- Han, G.; Wei, X.; Dong, X.; Wang, C.; Sui, N.; Guo, J.; Yuan, F.; Gong, Z.; Li, X.; Zhang, Y. Arabidopsis ZINC FINGER PROTEIN1 Acts Downstream of GL2 to Repress Root Hair Initiation and Elongation by Directly Suppressing bHLH Genes. Plant Cell 2020, 32, 206–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, A.; Wu, M.; Zhao, Y.; Zhang, A.; Liu, B.; Schiefelbein, J.; Gan, Y. Involvement of C2H2 zinc finger proteins in the regulation of epidermal cell fate determination in Arabidopsis. J. Integr. Plant Biol. 2014, 56, 1112–1117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Y.H.; Yu, Y.P.; Wang, D.; Wu, C.A.; Yang, G.D.; Huang, J.G.; Zheng, C.C. GhZFP1, a novel CCCH-type zinc finger protein from cotton, enhances salt stress tolerance and fungal disease resistance in transgenic tobacco by interacting with GZIRD21A and GZIPR5. New Phytol. 2009, 183, 62–75. [Google Scholar] [CrossRef]
- Matthews, J.M.; Sunde, M. Zinc fingers--folds for many occasions. IUBMB Life 2002, 54, 351–355. [Google Scholar] [CrossRef]
- Brown, R.S. Zinc finger proteins: Getting a grip on RNA. Curr. Opin. Struct. Biol. 2005, 15, 94–98. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Song, W.; Tromp, G.; Kolattukudy, P.E.; Fu, M. Genome-wide survey and expression profiling of CCCH-zinc finger family reveals a functional module in macrophage activation. PLoS ONE 2008, 3, e2880. [Google Scholar] [CrossRef]
- Blackshear, P.J. Tristetraprolin and other CCCH tandem zinc-finger proteins in the regulation of mRNA turnover. Biochem. Soc. Trans. 2001, 30, 945–952. [Google Scholar] [CrossRef]
- Berg, J.M.; Shi, Y. The galvanization of biology: A growing appreciation for the roles of zinc. Science 1996, 271, 1081–1085. [Google Scholar] [CrossRef]
- Zhuang, Y.; Wang, C.; Zhang, Y.; Chen, S.; Wang, D.; Liu, Q.; Zhou, G.; Chai, G. Overexpression of PdC3H17 Confers Tolerance to Drought Stress Depending on Its CCCH Domain in Populus. Front. Plant Sci. 2020, 10, 1748. [Google Scholar] [CrossRef] [Green Version]
- Bogamuwa, S.; Jang, J. Tandem CCCH Zinc Finger Proteins in Plant Growth, Development and Stress Response. Plant Cell Physiol. 2014, 55, 1367–1375. [Google Scholar] [CrossRef]
- Blackshear, P.J.; Phillips, R.S.; Lai, W.S. Tandem CCCH zinc finger proteins in mRNA binding. In Zinc Finger Proteins; Springer: Berlin/Heidelberg, Germany, 2005; pp. 80–90. [Google Scholar]
- Pomeranz, M.; Hah, C.; Lin, P.; Kang, S.G.; Finer, J.J.; Blackshear, P.J.; Jang, J. The Arabidopsis Tandem Zinc Finger Protein AtTZF1 Traffics between the Nucleus and Cytoplasmic Foci and Binds Both DNA and RNA. Plant Physiol. 2010, 152, 151–165. [Google Scholar] [CrossRef] [Green Version]
- D’Orso, F.; De Leonardis, A.M.; Salvi, S.; Gadaleta, A.; Ruberti, I.; Cattivelli, L.; Morelli, G.; Mastrangelo, A.M. Conservation of AtTZF1, AtTZF2, and AtTZF3 homolog gene regulation by salt stress in evolutionarily distant plant species. Front. Plant Sci. 2015, 6, 394. [Google Scholar] [PubMed] [Green Version]
- Seok, H.; Nguyen, L.V.; Park, H.; Tarte, V.N.; Ha, J.; Lee, S.; Moon, Y. Arabidopsis non-TZF gene AtC3H17 functions as a positive regulator in salt stress response. Biochem. Biophys. Res. Commun. 2018, 498, 954–959. [Google Scholar] [CrossRef] [PubMed]
- Murata, T.; Yoshino, Y.; Morita, N.; Kaneda, N. Identification of nuclear import and export signals within the structure of the zinc finger protein TIS11. Biochem. Biophys. Res. Commun. 2002, 293, 1242–1247. [Google Scholar] [CrossRef]
- Bogamuwa, S.; Jang, J. Plant Tandem CCCH Zinc Finger Proteins Interact with ABA, Drought, and Stress Response Regulators in Processing-Bodies and Stress Granules. PLoS ONE 2016, 11, e0151574. [Google Scholar] [CrossRef] [Green Version]
- Xu, L.; Liu, T.; Xiong, X.; Liu, W.; Yu, Y.; Cao, J. AtC3H18L is a stop-codon read-through gene and encodes a novel non-tandem CCCH zinc-finger protein that can form cytoplasmic foci similar to mRNP granules. Biochem. Biophys. Res. Commun. 2020, 528, 140–145. [Google Scholar] [CrossRef]
- Yeats, T.H.; Bacic, A.; Johnson, K.L. Plant glycosylphosphatidylinositol anchored proteins at the plasma membrane-cell wall nexus. J. Integr. Plant Biol. 2018, 60, 649–669. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wei, X.; Jiao, G.; Chen, W.; Wu, Y.; Sheng, Z.; Hu, S.; Xie, L.; Wang, J.; Tang, S.; et al. GBSS-BINDING PROTEIN, encoding a CBM48 domain-containing protein, affects rice quality and yield. J. Integr. Plant Biol. 2020, 62, 948–966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, J.; Jiang, H.; Xu, Y.; Li, H.; Wu, X.; Xie, Q.; Li, C. The CCCH-Type Zinc Finger Proteins AtSZF1 and AtSZF2 Regulate Salt Stress Responses in Arabidopsis. Plant Cell Physiol. 2007, 48, 1148–1158. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Jia, J.; Yan, X.; Shi, H.; Han, Y. Arabidopsis KHZ1 and KHZ2, two novel non-tandem CCCH zinc-finger and K-homolog domain proteins, have redundant roles in the regulation of flowering and senescence. Plant Mol. Biol. 2017, 95, 549–565. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Fang, R.; Chen, F.; Han, J.; Liu, Y.; Chen, L.; Zhu, Q. A novel CCCH-type zinc-finger protein SAW1 activates OsGA20ox3 to regulate gibberellin homeostasis and anther development in rice. J. Integr. Plant Biol. 2020, 62, 1594–1606. [Google Scholar] [CrossRef]
- Seong, S.Y.; Shim, J.S.; Bang, S.W.; Kim, J.K. Overexpression of OsC3H10, a CCCH-Zinc Finger, Improves Drought Tolerance in Rice by Regulating Stress-Related Genes. Plants 2020, 9, 1298. [Google Scholar] [CrossRef]
- Huang, P.; Chung, M.; Ju, H.; Na, H.; Lee, D.J.; Cheong, H.; Kim, C.S. Physiological characterization of the Arabidopsis thaliana Oxidation-related Zinc Finger 1, a plasma membrane protein involved in oxidative stress. J. Plant Res. 2011, 124, 699–705. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Ju, H.; Min, J.; Zhang, X.; Chung, J.; Cheong, H.; Kim, C.S. Molecular and Physiological Characterization of the Arabidopsis thaliana Oxidation-Related Zinc Finger 2, a Plasma Membrane Protein Involved in ABA and Salt Stress Response Through the ABI2-Mediated Signaling Pathway. Plant Cell Physiol. 2012, 53, 193–203. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frederick, E.D.; Ramos, S.B.V.; Blackshear, P.J. A Unique C-terminal Repeat Domain Maintains the Cytosolic Localization of the Placenta-specific Tristetraprolin Family Member ZFP36L3. J. Biol. Chem. 2008, 283, 14792–14800. [Google Scholar] [CrossRef] [Green Version]
- Matsushita, K.; Takeuchi, O.; Standley, D.M.; Kumagai, Y.; Kawagoe, T.; Miyake, T.; Satoh, T.; Kato, H.; Tsujimura, T.; Nakamura, H. Zc3h12a is an RNase essential for controlling immune responses by regulating mRNA decay. Nature 2009, 458, 1185–1190. [Google Scholar] [CrossRef]
- Lee, S.-j.; Jung, H.J.; Kang, H.; Kim, S.Y. Arabidopsis Zinc Finger Proteins AtC3H49/AtTZF3 and AtC3H20/AtTZF2 are Involved in ABA and JA Responses. Plant Cell Physiol. 2012, 53, 673–686. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xu, Y.; Zhang, C.; Ma, Q.; Joo, S.; Kim, S.; Xu, Z.; Chong, K. OsLIC, a Novel CCCH-Type Zinc Finger Protein with Transcription Activation, Mediates Rice Architecture via Brassinosteroids Signaling. PLoS ONE 2008, 3, e3521. [Google Scholar] [CrossRef] [Green Version]
- Blanvillain, R.; Wei, S.; Wei, P.; Kim, J.H.; Ow, D.W. Stress tolerance to stress escape in plants: Role of the OXS2 zinc-finger transcription factor family. EMBO J. 2011, 30, 3812–3822. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phillips, R.S.; Ramos, S.B.; Blackshear, P.J. Members of the tristetraprolin family of tandem CCCH zinc finger proteins exhibit CRM1-dependent nucleocytoplasmic shuttling. J. Biol. Chem. 2002, 277, 11606–11613. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Chen, G.; Ji, X.; Gao, G. ZAP is a CRM1-dependent nucleocytoplasmic shuttling protein. Biochem. Biophys. Res. Commun. 2004, 321, 517–523. [Google Scholar] [CrossRef] [PubMed]
- Jang, J. Arginine-rich motif-tandem CCCH zinc finger proteins in plant stress responses and post-transcriptional regulation of gene expression. Plant Sci. 2016, 252, 118–124. [Google Scholar] [CrossRef] [Green Version]
- Gao, H.; Zheng, X.; Fei, G.; Chen, J.; Jin, M.; Ren, Y.; Wu, W.; Zhou, K.; Sheng, P.; Zhou, F. Ehd4 Encodes a Novel and Oryza-Genus-Specific Regulator of Photoperiodic Flowering in Rice. PLoS Genet. 2013, 9, e1003281. [Google Scholar] [CrossRef]
- Li, Z.; Thomas, T.L. PEI1, an Embryo-Specific Zinc Finger Protein Gene Required for Heart-Stage Embryo Formation in Arabidopsis. Plant Cell 1998, 10, 383–398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seok, H.-Y.; Woo, D.-H.; Park, H.-Y.; Lee, S.-Y.; Tran, H.T.; Lee, E.-H.; Vu Nguyen, L.; Moon, Y.-H. AtC3H17, a Non-Tandem CCCH Zinc Finger Protein, Functions as a Nuclear Transcriptional Activator and Has Pleiotropic Effects on Vegetative Development, Flowering and Seed Development in Arabidopsis. Plant Cell Physiol. 2016, 57, 603–615. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, D.; Xu, Z.; Cao, S.; Chen, K.; Li, S.; Liu, X.; Gao, C.; Zhang, B.; Zhou, Y. An Uncanonical CCCH-Tandem Zinc-Finger Protein Represses Secondary Wall Synthesis and Controls Mechanical Strength in Rice. Mol. Plant 2018, 11, 163–174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barreau, C.; Paillard, L.; Osborne, H.B. AU-rich elements and associated factors: Are there unifying principles? Nucleic Acids Res. 2005, 33, 7138–7150. [Google Scholar] [CrossRef]
- Jan, A.; Maruyama, K.; Todaka, D.; Kidokoro, S.; Abo, M.; Yoshimura, E.; Shinozaki, K.; Nakashima, K.; Yamaguchi-Shinozaki, K. OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress-related genes. Plant Physiol. 2013, 161, 1202–1216. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.Y.; Shyu, A.B. AU-rich elements: Characterization and importance in mRNA degradation. Trends Biochem. Sci. 1995, 20, 465–470. [Google Scholar] [CrossRef]
- Bakheet, T.; Frevel, M.; Williams, B.R.; Greer, W.; Khabar, K.S. ARED: Human AU-rich element-containing mRNA database reveals an unexpectedly diverse functional repertoire of encoded proteins. Nucleic Acids Res. 2001, 29, 246–254. [Google Scholar] [CrossRef]
- Bakheet, T.; Williams, B.R.; Khabar, K.S. ARED 2.0: An update of AU-rich element mRNA database. Nucleic Acids Res. 2003, 31, 421–423. [Google Scholar] [CrossRef]
- Peng, S.S.; Chen, C.Y.; Shyu, A.B. Functional characterization of a non-AUUUA AU-rich element from the c-jun proto-oncogene mRNA: Evidence for a novel class of AU-rich elements. Mol. Cell. Biol. 1996, 16, 1490–1499. [Google Scholar] [CrossRef] [Green Version]
- Qu, J.; Kang, S.G.; Wang, W.; Musier-Forsyth, K.; Jang, J.C. The Arabidopsis thaliana tandem zinc finger 1 (AtTZF1) protein in RNA binding and decay. Plant J. 2014, 78, 452–467. [Google Scholar] [CrossRef] [Green Version]
- Maris, C.; Dominguez, C.; Allain, F.H.T. The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression. FEBS J. 2005, 272, 2118–2131. [Google Scholar] [CrossRef]
- Maldonadobonilla, L.D.; Eschenlippold, L.; Gagozachert, S.; Tabassum, N.; Bauer, N.; Scheel, D.; Lee, J. The Arabidopsis Tandem Zinc Finger 9 Protein Binds RNA and Mediates Pathogen-Associated Molecular Pattern-Triggered Immune Responses. Plant Cell Physiol. 2014, 55, 412–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hudson, B.P.; Martinez-Yamout, M.A.; Dyson, H.J.; Wright, P.E. Recognition of the mRNA AU-rich element by the zinc finger domain of TIS11d. Nat. Struct. Mol. Biol. 2004, 11, 257. [Google Scholar] [CrossRef]
- Cheng, Y.; Kato, N.; Wang, W.; Li, J.; Chen, X. Two RNA Binding Proteins, HEN4 and HUA1, Act in the Processing of AGAMOUS Pre-mRNA in Arabidopsis thaliana. Dev. Cell 2003, 4, 53–66. [Google Scholar] [CrossRef] [Green Version]
- Rodriguezcazorla, E.; Ortunomiquel, S.; Candela, H.; Baileysteinitz, L.J.; Yanofsky, M.F.; Martinezlaborda, A.; Ripoll, J.J.; Vera, A. Ovule identity mediated by pre-mRNA processing in Arabidopsis. PLoS Genet. 2018, 14, e1007182. [Google Scholar]
- Li, J.; Jia, D.; Chen, X. HUA1, a Regulator of Stamen and Carpel Identities in Arabidopsis, Codes for a Nuclear RNA Binding Protein. Plant Cell 2001, 13, 2269–2281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmitz, R.J.; Hong, L.; Michaels, S.; Amasino, R.M. FRIGIDA-ESSENTIAL 1 interacts genetically with FRIGIDA and FRIGIDA-LIKE 1 to promote the winter-annual habit of Arabidopsis thaliana. Development 2005, 132, 5471–5478. [Google Scholar] [CrossRef] [Green Version]
- Delaney, K.J.; Xu, R.; Zhang, J.; Li, Q.Q.; Yun, K.-Y.; Falcone, D.L.; Hunt, A.G. Calmodulin interacts with and regulates the RNA-binding activity of an Arabidopsis polyadenylation factor subunit. Plant Physiol. 2006, 140, 1507–1521. [Google Scholar] [CrossRef] [Green Version]
- Yan, Z.; Shi, H.; Liu, Y.; Jing, M.; Han, Y. KHZ1 and KHZ2, novel members of the autonomous pathway, repress the splicing efficiency of FLC pre-mRNA in Arabidopsis. J. Exp. Bot. 2020, 71, 1375–1386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Song, Z.; Bai, J.; Liu, X.; Nauwynck, H.; Jiang, P. ZAP, a CCCH-Type Zinc Finger Protein, Inhibits Porcine Reproductive and Respiratory Syndrome Virus Replication and Interacts with Viral Nsp9. J. Virol. 2019, 93, e00001-19. [Google Scholar] [CrossRef] [Green Version]
- Brooks, S.A.; Blackshear, P.J. Tristetraprolin (TTP): Interactions with mRNA and proteins, and current thoughts on mechanisms of action. Biochim. Biophys. Acta 2013, 1829, 666–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raja, V.; Majeed, U.; Kang, H.; Andrabi, K.I.; John, R. Abiotic stress: Interplay between ROS, hormones and MAPKs. Environ. Exp. Bot. 2017, 137, 142–157. [Google Scholar] [CrossRef]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.H.; Yamaguchi, S.; Lim, S.; Oh, E.; Park, J.; Hanada, A.; Kamiya, Y.; Choi, G. SOMNUS, a CCCH-Type Zinc Finger Protein in Arabidopsis, Negatively Regulates Light-Dependent Seed Germination Downstream of PIL5. Plant Cell 2008, 20, 1260–1277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, P.; Pomeranz, M.; Jikumaru, Y.; Kang, S.G.; Hah, C.; Fujioka, S.; Kamiya, Y.; Jang, J. The Arabidopsis tandem zinc finger protein AtTZF1 affects ABA- and GA-mediated growth, stress and gene expression responses. Plant J. 2011, 65, 253–268. [Google Scholar] [CrossRef]
- Wang, W.; Liu, B.; Xu, M.; Jamil, M.; Wang, G. ABA-induced CCCH tandem zinc finger protein OsC3H47 decreases ABA sensitivity and promotes drought tolerance in Oryza sativa. Biochem. Biophys. Res. Commun. 2015, 464, 33–37. [Google Scholar] [CrossRef]
- Zhou, T.; Yang, X.; Wang, L.; Xu, J.; Zhang, X. GhTZF1 regulates drought stress responses and delays leaf senescence by inhibiting reactive oxygen species accumulation in transgenic Arabidopsis. Plant Mol. Biol. 2014, 85, 163–177. [Google Scholar] [CrossRef] [PubMed]
- Hung, K.T.; Hsu, Y.T.; Kao, C.H. Hydrogen peroxide is involved in methyl jasmonate-induced senescence of rice leaves. Physiol. Plant 2006, 127, 293–303. [Google Scholar] [CrossRef]
- Nadarajah, K.K. ROS Homeostasis in Abiotic Stress Tolerance in Plants. Int J. Mol. Sci. 2020, 21, 5208. [Google Scholar] [CrossRef]
- Choudhury, F.K.; Rivero, R.M.; Blumwald, E.; Mittler, R. Reactive oxygen species, abiotic stress and stress combination. Plant J. 2017, 90, 856–867. [Google Scholar] [CrossRef]
- Gong, Z.; Xiong, L.; Shi, H.; Yang, S.; Herrera-Estrella, L.R.; Xu, G.; Chao, D.Y.; Li, J.; Wang, P.Y.; Qin, F.; et al. Plant abiotic stress response and nutrient use efficiency. Sci. China. Life Sci. 2020, 63, 635–674. [Google Scholar] [CrossRef]
- Guo, J.; Li, Y.; Han, G.; Song, J.; Wang, B. NaCl markedly improved the reproductive capacity of the euhalophyte Suaeda salsa. Funct. Plant Biol. 2017, 45, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Wang, B. Using euhalophytes to understand salt tolerance and to develop saline agriculture: Suaeda salsa as a promising model. Ann. Bot. 2015, 115, 541–553. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sui, N.; Tian, S.; Wang, W.; Wang, M.; Fan, H. Overexpression of Glycerol-3-Phosphate Acyltransferase from Suaeda salsa Improves Salt Tolerance in Arabidopsis. Front. Plant Sci. 2017, 8, 1337. [Google Scholar] [CrossRef]
- Deng, Y.; Bao, J.; Yuan, F.; Liang, X.; Feng, Z.; Wang, B. Exogenous hydrogen sulfide alleviates salt stress in wheat seedlings by decreasing Na+ content. Plant Growth Regul. 2016, 79, 391–399. [Google Scholar] [CrossRef]
- Han, G.; Yuan, F.; Guo, J.; Zhang, Y.; Sui, N.; Wang, B. AtSIZ1 improves salt tolerance by maintaining ionic homeostasis and osmotic balance in Arabidopsis. Plant Sci. 2019, 285, 55–67. [Google Scholar] [CrossRef]
- Liu, S.; Wang, W.; Li, M.; Wan, S.; Sui, N. Antioxidants and unsaturated fatty acids are involved in salt tolerance in peanut. Acta Physiol. Plant 2017, 39, 207. [Google Scholar] [CrossRef]
- Yang, Z.; Li, J.; Liu, L.; Xie, Q.; Sui, N. Photosynthetic Regulation Under Salt Stress and Salt-Tolerance Mechanism of Sweet Sorghum. Front. Plant Sci. 2020, 10, 1722. [Google Scholar] [CrossRef]
- Feng, Z.; Deng, Y.; Fan, H.; Sun, Q.; Sui, N.; Wang, B. Effects of NaCl stress on the growth and photosynthetic characteristics of Ulmus pumila L. seedlings in sand culture. Photosynthetica 2014, 52, 313–320. [Google Scholar] [CrossRef]
- Sui, N.; Yang, Z.; Liu, M.; Wang, B. Identification and transcriptomic profiling of genes involved in increasing sugar content during salt stress in sweet sorghum leaves. BMC Genom. 2015, 16, 534. [Google Scholar] [CrossRef] [Green Version]
- Yuan, F.; Leng, B.; Wang, B. Progress in Studying Salt Secretion from the Salt Glands in Recretohalophytes: How Do Plants Secrete Salt? Front. Plant Sci. 2016, 7, 977. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Lin, L.; Sui, N. Regulation mechanism of microRNA in plant response to abiotic stress and breeding. Mol. Biol. Rep. 2019, 46, 1447–1457. [Google Scholar] [CrossRef]
- Wu, H.; Guo, J.; Wang, C.; Li, K.; Zhang, X.; Yang, Z.; Li, M.; Wang, B. An Effective Screening Method and a Reliable Screening Trait for Salt Tolerance of Brassica napus at the Germination Stage. Front. Plant Sci 2019, 10, 530. [Google Scholar] [CrossRef]
- Yang, Z.; Zheng, H.; Wei, X.; Song, J.; Wang, B.; Sui, N. Transcriptome analysis of sweet Sorghum inbred lines differing in salt tolerance provides novel insights into salt exclusion by roots. Plant Soil 2018, 430, 423–439. [Google Scholar] [CrossRef]
- Zhao, Y.; Yang, Y.; Song, Y.; Li, Q.; Song, J. Analysis of storage compounds and inorganic ions in dimorphic seeds of euhalophyte Suaeda salsa. Plan. Physiol. Biochem. 2018, 130, 511–516. [Google Scholar] [CrossRef]
- Yuan, F.; Leng, B.; Zhang, H.; Wang, X.; Han, G.; Wang, B. A WD40-Repeat Protein From the Recretohalophyte Limonium bicolor Enhances Trichome Formation and Salt Tolerance in Arabidopsis. Front. Plant Sci. 2019, 10, 1456. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Guo, Y. Unraveling salt stress signaling in plants. J. Integr. Plant Biol. 2018, 60, 796–804. [Google Scholar] [CrossRef] [Green Version]
- Jamil, M.; Iqbal, W.; Bangash, A.; Rehman, S.U.; Imran, Q.M.; Rha, E.S. Constitutive expression of OSC3H33, OSC3H50 and OSC3H37 genes in rice under salt stress. Pak. J. Bot 2010, 42, 4003–4009. [Google Scholar]
- Jiang, M.; Jiang, J.; Miao, L.; He, C. Over-expression of a C3H-type zinc finger gene contributes to salt stress tolerance in transgenic broccoli plants. Plant Cell Tissue Organ. Cult. 2017, 130, 239–254. [Google Scholar] [CrossRef]
- Ma, Y.; Yang, Y.; Liu, R.; Li, Q.; Song, J. Adaptation of euhalophyte Suaeda salsa to nitrogen starvation under salinity. Plant Physiol. Biochem. 2020, 146, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.; Zhao, L.; Ren, Y.; Yang, S.; Zhu, J.K.; Zhao, C. The transcription factor ICE1 functions in cold stress response by binding to the promoters of CBF and COR genes. J. Integr. Plant Biol. 2020, 62, 258–263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qi, J.; Song, C.P.; Wang, B.; Zhou, J.; Kangasjärvi, J.; Zhu, J.K.; Gong, Z. Reactive oxygen species signaling and stomatal movement in plant responses to drought stress and pathogen attack. J. Integr. Plant Biol. 2018, 60, 805–826. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Wang, Y.; Sui, N. Transcriptional regulation of bHLH during plant response to stress. Biochem. Biophys. Res. Commun. 2018, 503, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Yang, Z.; Han, G.; Zhao, X.; Yin, S.; Yuan, F.; Wang, B. The developmental dynamics of the sweet sorghum root transcriptome elucidate the differentiation of apoplastic barriers. Plant Signal. Behav. 2020, 15, 1724465. [Google Scholar] [CrossRef]
- Abid, M.; Ali, S.; Qi, L.K.; Zahoor, R.; Tian, Z.; Jiang, D.; Snider, J.L.; Dai, T. Physiological and biochemical changes during drought and recovery periods at tillering and jointing stages in wheat (Triticum aestivum L.). Sci. Rep. 2018, 8, 4615. [Google Scholar] [CrossRef]
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.W.; Saud, S. Crop production under drought and heat stress: Plant responses and management options. Front. Plant Sci. 2017, 8, 1147. [Google Scholar] [CrossRef] [Green Version]
- Zheng, H.; Yang, Z.; Wang, W.; Guo, S.; Li, Z.; Liu, K.; Sui, N. Transcriptome analysis of maize inbred lines differing in drought tolerance provides novel insights into the molecular mechanisms of drought responses in roots. Plant Physiol. Biochem. 2020, 149, 11–26. [Google Scholar] [CrossRef]
- Chang, Y.N.; Zhu, C.; Jiang, J.; Zhang, H.; Zhu, J.K.; Duan, C.G. Epigenetic regulation in plant abiotic stress responses. J. Integr. Plant Biol. 2020, 62, 563–580. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Han, G.; Meng, Z.; Lin, L.; Sui, N. Roles of malic enzymes in plant development and stress responses. Plant Signal. Behav. 2019, 14, e1644596. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Liu, H.-L.; Wang, K.; Gao, Y.-M.; Wu, M.; Xiang, Y. Identification of CCCH Zinc Finger Proteins Family in Moso Bamboo (Phyllostachys edulis), and PeC3H74 Confers Drought Tolerance to Transgenic Plants. Front. Plant Sci. 2020, 11, 1697. [Google Scholar] [CrossRef] [PubMed]
- Selvaraj, M.G.; Jan, A.; Ishizaki, T.; Valencia, M.; Dedicova, B.; Maruyama, K.; Ogata, T.; Todaka, D.; Yamaguchi-Shinozaki, K.; Nakashima, K.; et al. Expression of the CCCH-tandem zinc finger protein gene OsTZF5 under a stress-inducible promoter mitigates the effect of drought stress on rice grain yield under field conditions. Plant Biotechnol. J. 2020, 18, 1711–1721. [Google Scholar] [CrossRef] [Green Version]
- Baileyserres, J.; Voesenek, L.A.C.J. Flooding Stress: Acclimations and Genetic Diversity. Annu. Rev. Plant Biol. 2008, 59, 313–339. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Shi, G.; Gao, B.; Fan, H.; Wang, B. Waterlogging and salinity effects on two Suaeda salsa populations. Physiol. Plant 2011, 141, 343–351. [Google Scholar] [CrossRef]
- Wang, L.; Gao, J.; Zhang, Z.; Liu, W.; Cheng, P.; Mu, W.; Su, T.; Chen, S.; Chen, F.; Jiang, J. Overexpression of CmSOS1 confers waterlogging tolerance in Chrysanthemum. J. Integr. Plant Biol. 2020, 62, 1059–1064. [Google Scholar] [CrossRef]
- Yang, J.C.; Li, M.; Xie, X.Z.; Han, G.L.; Sui, N.; Wang, B.S. Deficiency of phytochrome B alleviates chilling-induced photoinhibition in rice. Am. J. Bot. 2013, 100, 1860–1870. [Google Scholar] [CrossRef]
- Zhou, Y.; Tan, W.J.; Xie, L.J.; Qi, H.; Yang, Y.C.; Huang, L.P.; Lai, Y.X.; Tan, Y.F.; Zhou, D.M.; Yu, L.J.; et al. Polyunsaturated linolenoyl-CoA modulates ERF-VII-mediated hypoxia signaling in Arabidopsis. J. Integr. Plant Biol. 2020, 62, 330–348. [Google Scholar] [CrossRef] [Green Version]
- Pandey, D.M.; Kim, S. Identification and expression analysis of hypoxia stress inducible CCCH-type zinc finger protein genes in rice. J. Plant Biol. 2012, 55, 489–497. [Google Scholar] [CrossRef]
- Sui, N. Photoinhibition of Suaeda salsa to chilling stress is related to energy dissipation and water-water cycle. Photosynthetica 2015, 53, 207–212. [Google Scholar] [CrossRef]
- Ding, Y.; Lv, J.; Shi, Y.; Gao, J.; Hua, J.; Song, C.; Gong, Z.; Yang, S. EGR2 phosphatase regulates OST1 kinase activity and freezing tolerance in Arabidopsis. EMBO J. 2019, 38, e99819. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Zhou, Y.; Fan, F.; Peng, J.; Zhang, J. Coordination of light, circadian clock with temperature: The potential mechanisms regulating chilling tolerance in rice. J. Integr. Plant Biol. 2020, 62, 737–760. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Liu, D.; Chong, K. Cold signaling in plants: Insights into mechanisms and regulation. J. Integr. Plant Biol. 2018, 60, 745–756. [Google Scholar] [CrossRef] [Green Version]
- Sui, N.; Wang, Y.; Liu, S.; Yang, Z.; Wang, F.; Wan, S. Transcriptomic and physiological evidence for the relationship between unsaturated fatty acid and salt stress in peanut. Front. Plant Sci. 2018, 9, 7. [Google Scholar] [CrossRef] [Green Version]
- Zhang, D.; Guo, X.; Xu, Y.; Li, H.; Ma, L.; Yao, X.; Weng, Y.; Guo, Y.; Liu, C.M.; Chong, K. OsCIPK7 point-mutation leads to conformation and kinase-activity change for sensing cold response. J. Integr. Plant Biol. 2019, 61, 1194–1200. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Wang, L.; Wang, J.; Jin, J.; Zhang, N.; Lei, L.; Gao, T.; Jing, T.; Zhang, S.; Wu, Y. Induction of priming by cold stress via inducible volatile cues in neighboring tea plants. J. Integr. Plant Biol. 2020, 62, 1461–1468. [Google Scholar] [CrossRef]
- Liu, Y.; Xu, C.; Zhu, Y.; Zhang, L.; Chen, T.; Zhou, F.; Chen, H.; Lin, Y. The calcium-dependent kinase OsCPK24 functions in cold stress responses in rice. J. Integr. Plant Biol. 2018, 60, 173–188. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, H.; Lin, P.; Li, X.; Liao, X.; Wan, L.; Yang, X.; Luo, Y.; Zhang, L.; Zhang, F.; Liu, S. DgC3H1, a CCCH zinc finger protein gene, confers cold tolerance in transgenic chrysanthemum. Sci. Hortic. 2021, 281, 109901. [Google Scholar] [CrossRef]
- Xie, Z.; Lin, W.; Yu, G.; Cheng, Q.; Xu, B.; Huang, B. Improved cold tolerance in switchgrass by a novel CCCH-type zinc finger transcription factor gene, PvC3H72, associated with ICE1–CBF–COR regulon and ABA-responsive genes. Biotechnol. Biofuels 2019, 12, 224. [Google Scholar] [CrossRef]
- Meng, X.; Zhou, J.; Sui, N. Mechanisms of salt tolerance in halophytes: Current understanding and recent advances. Open Life Sci. 2018, 13, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tergel, T.; Chen, J.; Yang, J.; Kang, Y.; Qi, Z. Arabidopsis transcriptional response to extracellular Ca2+ depletion involves a transient rise in cytosolic Ca2+. J. Integr. Plant Biol. 2015, 57, 138–150. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Zhou, X.; Tao, M.; Yuan, F.; Liu, L.; Wu, F.; Wu, X.; Xiang, Y.; Niu, Y.; Liu, F. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx. Nature 2019, 572, 341–346. [Google Scholar] [CrossRef]
- Yuan, F.; Yang, H.; Xue, Y.; Kong, D.; Ye, R.; Li, C.; Zhang, J.; Theprungsirikul, L.; Shrift, T.; Krichilsky, B. Corrigendum: OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis. Nature 2015, 519, 378. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Dai, X.; Xu, Y.; Luo, W.; Zheng, X.; Zeng, D.; Pan, Y.; Lin, X.; Liu, H.; Zhang, D. COLD1 Confers Chilling Tolerance in Rice. Cell 2015, 160, 1209–1221. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Shi, Y.; Yang, S. Insights into the regulation of C-repeat binding factors in plant cold signaling. J. Integr. Plant Biol. 2018, 60, 780–795. [Google Scholar] [CrossRef]
- Zhang, C.; Lei, Y.; Lu, C.; Wang, L.; Wu, J. MYC2, MYC3, and MYC4 function additively in wounding-induced jasmonic acid biosynthesis and catabolism. J. Integr. Plant Biol. 2020, 62, 1159–1175. [Google Scholar] [CrossRef]
- Marchese, D.; de Groot, N.S.; Lorenzo Gotor, N.; Livi, C.M.; Tartaglia, G.G. Advances in the characterization of RNA-binding proteins. Wiley Interdiscip. Rev. RNA 2016, 7, 793–810. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, G.; Qiao, Z.; Li, Y.; Wang, C.; Wang, B. The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 8327. https://doi.org/10.3390/ijms22158327
Han G, Qiao Z, Li Y, Wang C, Wang B. The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance. International Journal of Molecular Sciences. 2021; 22(15):8327. https://doi.org/10.3390/ijms22158327
Chicago/Turabian StyleHan, Guoliang, Ziqi Qiao, Yuxia Li, Chengfeng Wang, and Baoshan Wang. 2021. "The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance" International Journal of Molecular Sciences 22, no. 15: 8327. https://doi.org/10.3390/ijms22158327




