SARS-CoV-2: Understanding the Transcriptional Regulation of ACE2 and TMPRSS2 and the Role of Single Nucleotide Polymorphism (SNP) at Codon 72 of p53 in the Innate Immune Response against Virus Infection
Abstract
:1. Introduction
2. Transcriptional Regulation of ACE2 by ISGF3 Assembly Complex in Type I IFNs and the JAK/STAT Pathway

| Accession | Term Name | Term Definition | Gene Count | p Value * |
|---|---|---|---|---|
| GO:0006508 | Proteolysis | The hydrolysis of proteins into smaller polypeptides and/or amino acids by cleavage of their peptide bonds. [GOC:bf, GOC:mah] | 1 | 0.0106 |
| GO:0046718 | Viral entry into host cell | The process that occurs after viral attachment by which a virion, or viral nucleic acid, breaches the plasma membrane or cell envelope and enters the host cell. The process ends when the viral nucleic acid is released into the host cell cytoplasm. [GOC:jl, PMID:12142475] | 1 | 1.3 × 10−4 |
| GO:0046813 | Virion attachment, binding of host cell surface receptor | The process during virion attachment where a virion binds to a host cell receptor, resulting in a conformational change of the virus protein. [ISBN:0879694971] | 1 | 7.02 × 10−5 |
| GO:0042127 | Regulation of cell proliferation | Any process that modulates the frequency, rate, or extent of cell proliferation. [GOC:jl] | 1 | 0.00281 |
| GO:0019229 | Regulation of vasoconstriction | Any process that modulates the frequency, rate, or extent of reductions in the diameter of blood vessels. [GOC:jl] | 1 | 2.81 × 10−4 |
| GO:2000379 | Positive regulation of reactive oxygen species metabolic process | Any process that activates or increases the frequency, rate, or extent of reactive oxygen species metabolic process. [GOC:mah] | 1 | 4.21 × 10−4 |
| GO:0009615 | Response to virus | Any process that results in a change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a stimulus from a virus. [GOC:hb] | 1 | 0.00302 |
| GO:0050727 | Regulation of inflammatory response | Any process that modulates the frequency, rate, or extent of the inflammatory response and the immediate defensive reaction (by vertebrate tissue) to infection or injury caused by chemical or physical agents. [GOC:ai] | 1 | 6.11 × 10−4 |
| GO:0003051 | Angiotensin-mediated drinking behavior | The drinking behavior that is mediated by the action of angiotensin in the brain. Angiotensin stimulates the brain centers that control thirst. [GOC:mtg_cardio] | 1 | 4.01 × 10−5 |
| GO:0032800 | Receptor biosynthetic process | The chemical reactions and pathways resulting in the formation of a receptor molecule, a macromolecule in combination with a hormone or neurotransmitter, drug, or intracellular messenger to initiate a change in cell function. [GOC:mah] | 1 | 1.2 × 10−4 |
| GO:0001817 | Regulation of cytokine production | Any process that modulates the frequency, rate, or extent of production of a cytokine. [GOC:add, ISBN:0781735149] | 1 | 1.5 × 10−4 |
| GO:0002005 | Angiotensin catabolic process in blood | The chemical reactions and pathways resulting in the breakdown of angiotensin in the blood. [ISBN:0721643949] | 1 | 3.01 × 10−5 |
| GO:0003081 | Regulation of systemic arterial blood pressure by renin-angiotensin | The process in which renin-angiotensin modulates the force with which blood passes through the circulatory system. [GOC:mtg_cardio] | 1 | 7.02 × 10−5 |
| GO:0042312 | Regulation of vasodilation | Any process that modulates the frequency, rate, or extent of increases in the diameter of blood vessels. [GOC:jl] | 1 | 9.02 × 10−5 |
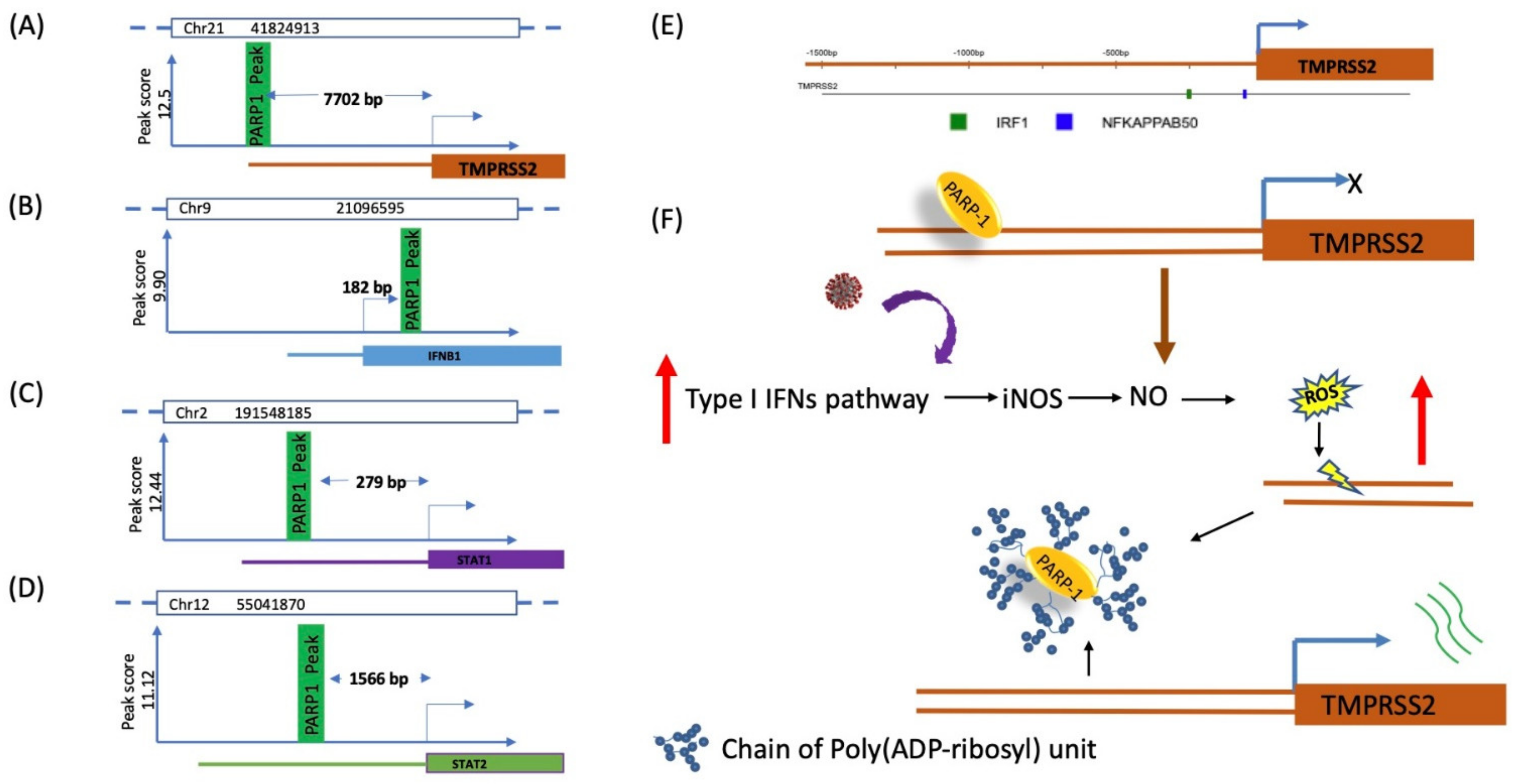
3. Transcriptional Regulation of TMPRSS2 by PARP1 Binding
4. Modulation of the Immune Response to a Virus
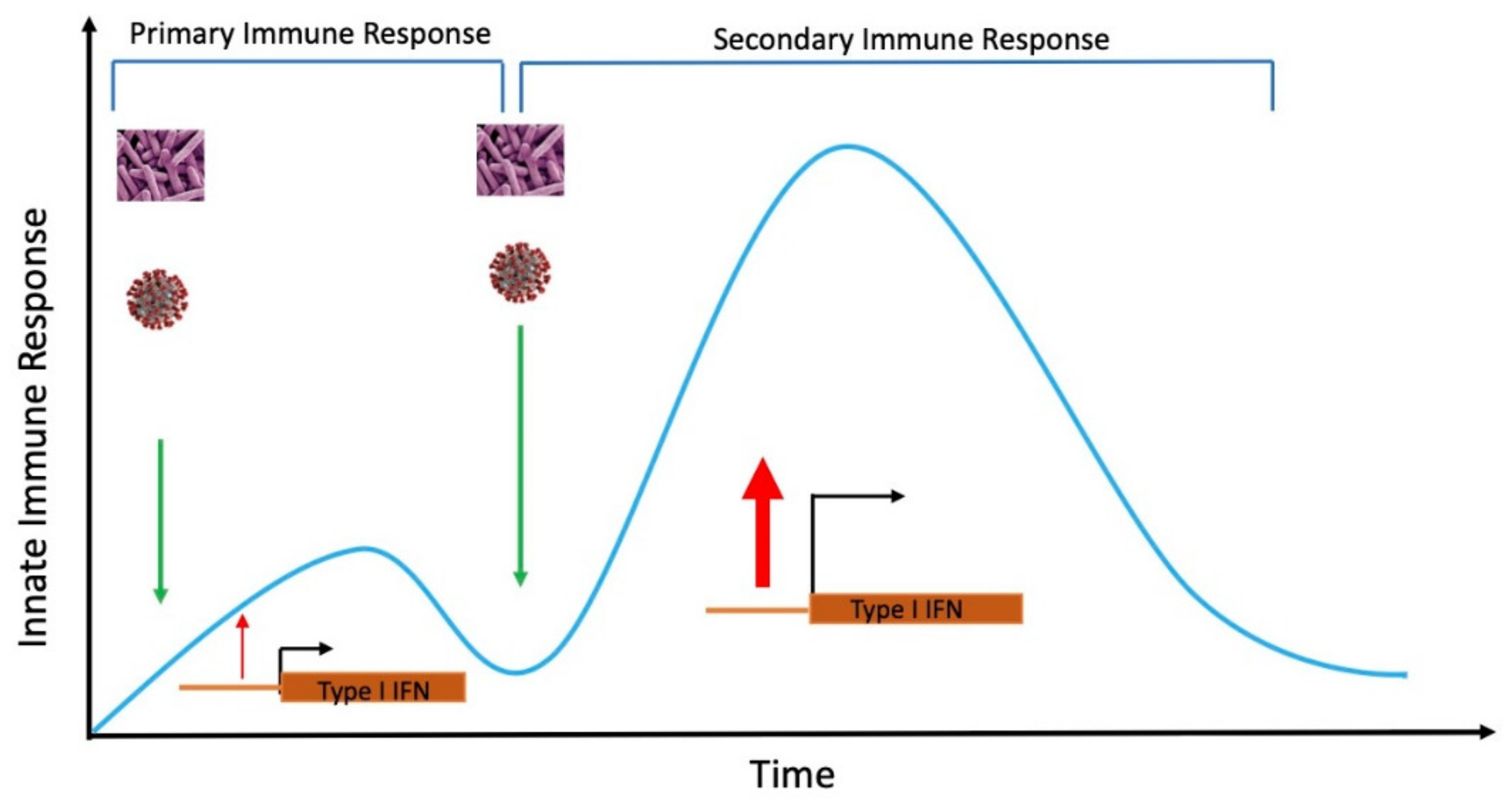
4.1. Immunogenic Response Training to Develop Immunity, Reduce Susceptibility and the Severity of SARS-CoV-2 Infection
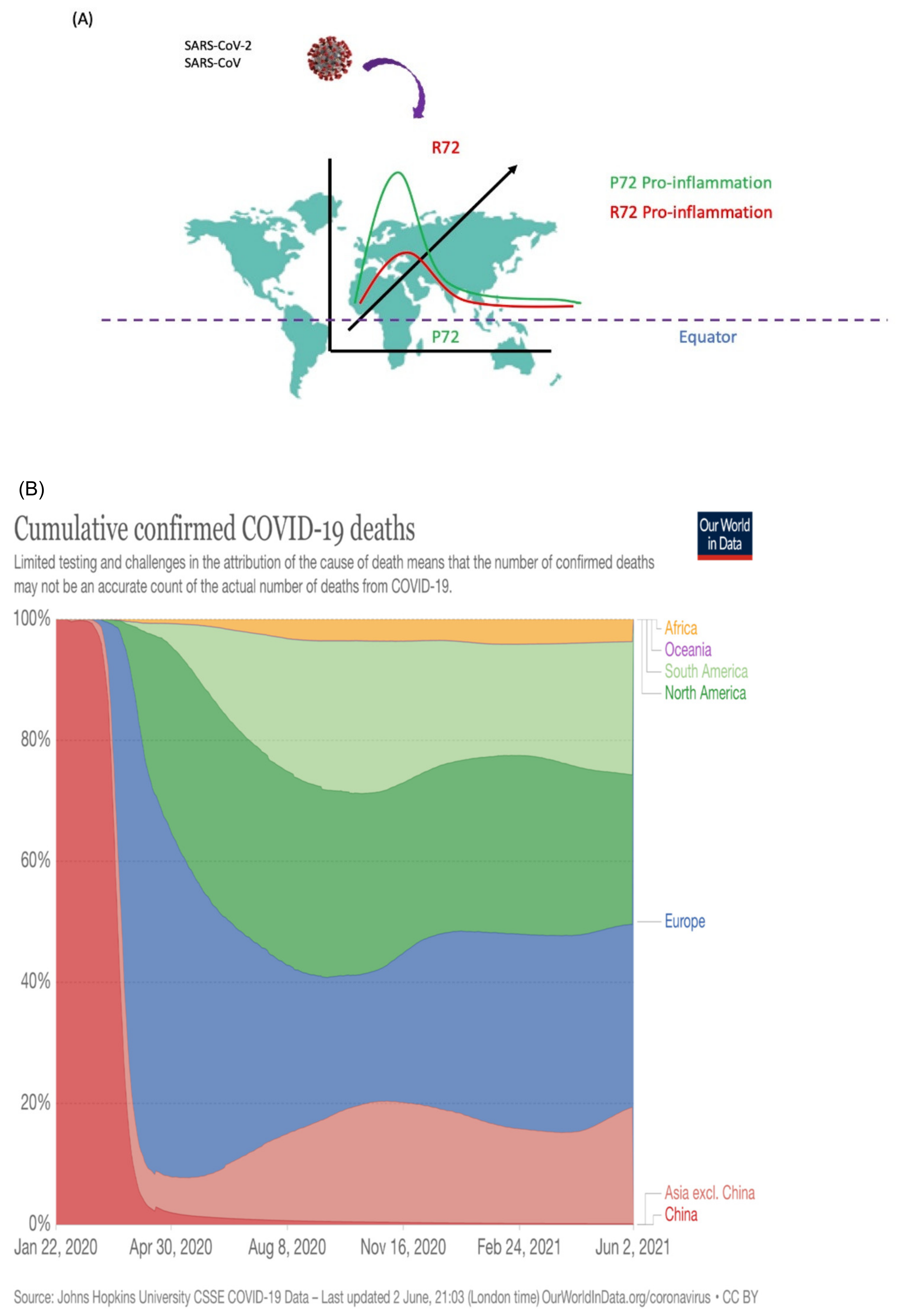
4.2. Gaining Immunity through Genetic Variation during Evolution
5. Modulation of Immune Response Genes to Develop Therapy
6. Prevention and New Targets to Develop Therapy
6.1. Prevention
6.1.1. Vaccination
6.1.2. Codon 72 Polymorphism of p53
6.2. New Targets
6.2.1. PARP1
6.2.2. TLR3, -4, and -9
6.2.3. Type I IFN Receptors
6.2.4. STAT1/2
6.2.5. JAK1/TYK2
6.2.6. IRFs
6.2.7. NFκB
6.2.8. ACE2
6.2.9. TMPRSS2
6.2.10. Viral Genome
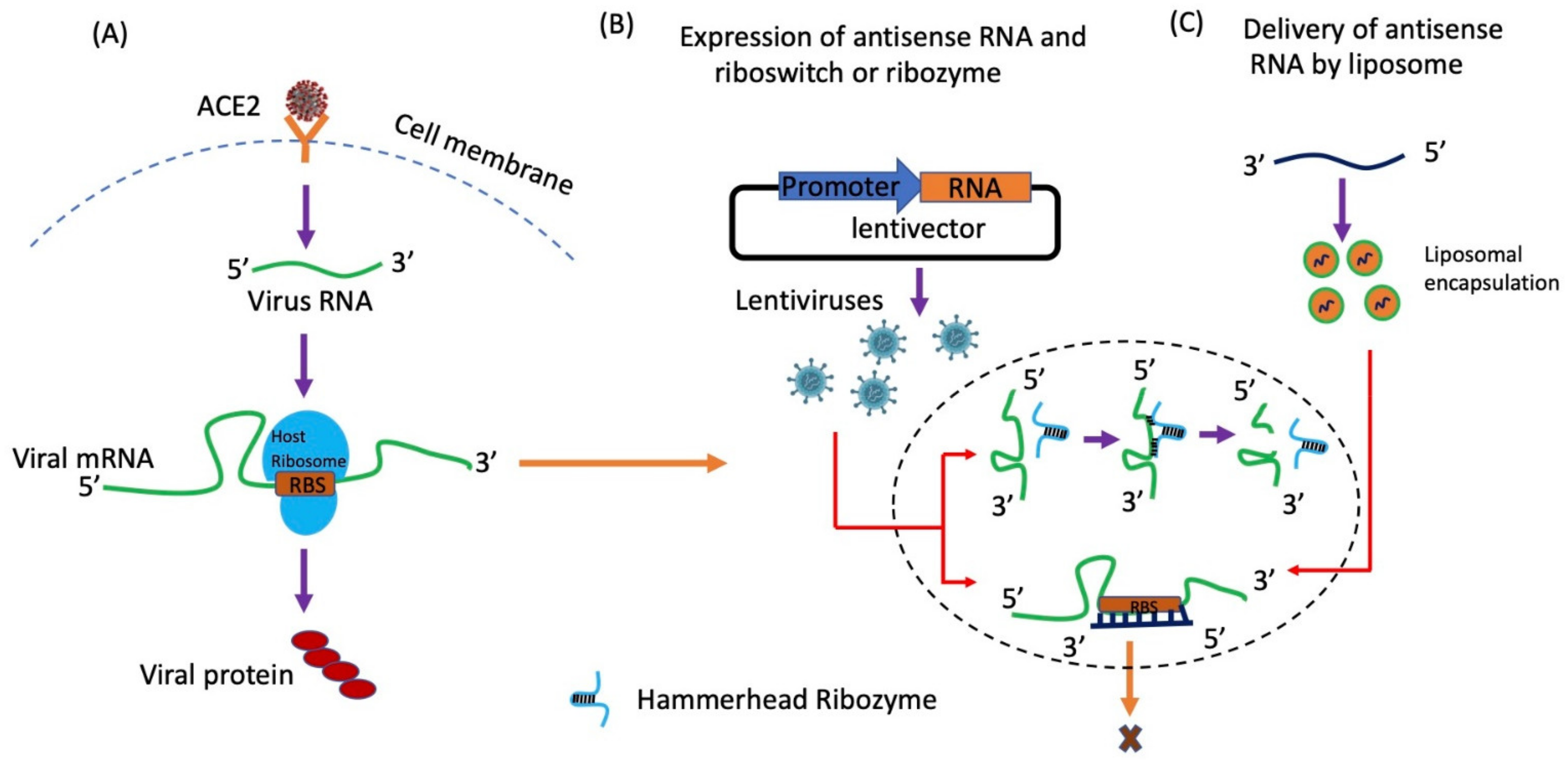
Antisense
- (1)
- (2)
- Complementary oligo hybridizes with viral mRNA to form duplex, after that RNAH enzyme can be used to degrade the duplex. Partially degraded mRNA will be unable to translate in to viral protein; therefore, virus cannot propagate.
Bacteriophage-Based Vaccine
Phage DNA Vaccine
CRISPR
7. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| WHO | World Health Organization |
| CDC | Centres for Disease Control and Prevention |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome Coronavirus-2 |
| ACE2 | Angiotensin-converting enzyme 2 |
| TMPRSS2 | Type II transmembrane serine protease |
| TLRs | Toll-Like Receptors |
| DAMPs | Damage-associated molecular patterns |
| ROS | Reactive oxygen species |
| ALI | Acute Lung Injury |
| IFN | Interferon |
| ISGs | Interferon-stimulated genes |
| STAT1 | Signal transducer and activator of transcription 1 |
| IMM | Inflammatory Monocyte-Macrophage |
| AAMEP | Alternatively Activated Macrophages Enhances Pathogenesis |
| PARP1 | Poly(ADP-Ribose) Polymerase 1 |
| SNP | Single Nucleotide Polymorphism |
| ISRE | IFN-stimulated response element |
| ISGF3 | Interferon Stimulated Gene Factor 3 |
| IRF9 | Interferon regulatory factor 9 |
| TSS | Transcription start site |
| ChIP | Chromatin Immunoprecipitation |
| HSP70 | Heat Shock Protein 70 |
| NO | Nitric oxide |
| iNOS | Inducible Nitric oxide synthase |
| BCG | Bacillus Calmette-Guerin |
| PRRs | Pattern Recognition Receptors |
| PAMPs | Pathogen-associated molecular pattern |
| LPS | Lipopolysaccharide |
| APCs | Antigen-presenting cells |
| IRAK | IL-1R-associated protein kinases |
| TRAF6 | TNF receptor-associated factor 6 |
| NFκB | Nuclear factor kappa B |
| IL | Interleukin |
| FDA | Food and Drug Administration |
| NAD | Nicotinamide adenine dinucleotide |
| ICMR | Institute Indian Council of Medical Research |
| DBT | Department of Biotechnology |
| HCQ | Hydroxyquinoline |
| IMMs | Inflammatory monocytes-macrophages |
| SLE | Systemic lupus erythematosus |
| IFNAR1 | Anifrolumab, a monoclonal antibody of IFN receptor |
| RNAi | RNA interference |
| siRNA | small interfering RNA |
| shRNA | small hairpin RNA |
| RDRP | RNA Dependent RNA Polymerase |
| APCs | Antigen-presenting cells |
| MHC | Major histocompatibility complex |
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| NIEHS | National Institute of Environmental Health Sciences |
| LMG | Laboratory of Molecular Genetics |
| HFD | High-fat diet |
| TSC | Tumor Sclerosis Complex |
References
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef] [Green Version]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Fauver, J.R.; Petrone, M.E.; Hodcroft, E.B.; Shioda, K.; Ehrlich, H.Y.; Watts, A.G.; Vogels, C.B.F.; Brito, A.F.; Alpert, T.; Muyombwe, A.; et al. Coast-to-Coast Spread of SARS-CoV-2 during the Early Epidemic in the United States. Cell 2020, 181, 990–996. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Moore, M.J.; Vasllieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greeneugh, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glowacka, I.; Bertram, S.; Muller, M.A.; Allen, P.; Soilleux, E.; Pfefferle, S.; Steffen, I.; Tsegaye, T.S.; He, Y.; Gnirss, K.; et al. Evidence that TMPRSS2 Activates the Severe Acute Respiratory Syndrome Coronavirus Spike Protein for Membrane Fusion and Reduces Viral Control by the Humoral Immune Response. J. Virol. 2011, 85, 4122–4134. [Google Scholar] [CrossRef] [Green Version]
- Matsuyama, S.; Nagata, N.; Shirato, K.; Kawase, M.; Takeda, M.; Taguchi, F. Efficient Activation of the Severe Acute Respiratory Syndrome Coronavirus Spike Protein by the Transmembrane Protease TMPRSS2. J. Virol. 2010, 84, 12658–12664. [Google Scholar] [CrossRef] [Green Version]
- Shulla, A.; Heald-Sargent, T.; Subramanya, G.; Zhao, J.; Perlman, S.; Gallagher, T. A Transmembrane Serine Protease Is Linked to the Severe Acute Respiratory Syndrome Coronavirus Receptor and Activates Virus Entry. J. Virol. 2011, 85, 873–882. [Google Scholar] [CrossRef] [Green Version]
- Stopsack, K.H.; Mucci, L.A.; Antonarakis, E.S.; Nelson, P.S.; Kantoff, P.W. TMPRSS2 and COVID-19: Serendipity or Opportunity for Intervention? Cancer Discov. 2020, 10, 779–782. [Google Scholar] [CrossRef] [Green Version]
- Ziegler, C.G.K.; Allon, S.J.; Nyquist, S.K.; Mbano, I.M.; Miao, V.N.; Tzouanas, C.N.; Cao, Y.; Yousif, A.S.; Bals, J.; Hauser, B.M.; et al. SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets across Tissues. Cell 2020, 181, 1016–1035.e19. [Google Scholar] [CrossRef]
- Medzhitov, R.; Preston-Hurlburt, P.; Janeway, C.A. A human homologue of the Drosophila toll protein signals activation of adaptive immunity. Nature 1997, 388, 394–397. [Google Scholar] [CrossRef]
- Subbarao, K.; Mahanty, S. Respiratory Virus Infections: Understanding COVID-19. Immunity 2020, 52, 905–909. [Google Scholar] [CrossRef] [PubMed]
- Imai, Y.; Kuba, K.; Neely, G.G.; Yaghubian-Malhami, R.; Perkmann, T.; van Loo, G.; Ermolaeva, M.; Veldhuizen, R.; Leung, Y.H.C.; Wang, H.; et al. Identification of Oxidative Stress and Toll-like Receptor 4 Signaling as a Key Pathway of Acute Lung Injury. Cell 2008, 133, 235–249. [Google Scholar] [CrossRef]
- Vardhana, S.A.; Wolchok, J.D. The many faces of the anti-COVID immune response. J. Exp. Med. 2020, 217, e20200678. [Google Scholar] [CrossRef]
- Schneider, W.M.; Chevillotte, M.D.; Rice, C.M. Interferon-stimulated genes: A complex web of host defenses. Annu. Rev. Immunol. 2014, 32, 513–545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dupuis, S.; Jouanguy, E.; Al-Hajjar, S.; Fieschi, C.; Zaid Al-Mohsen, I.; Al-Jumaah, S.; Yang, K.; Chapgier, A.; Eidenschenk, C.; Eid, P.; et al. Impaired response to interferon-α/β and lethal viral disease in human STAT1 deficiency. Nat. Genet. 2003, 33, 388–391. [Google Scholar] [CrossRef]
- Everitt, A.; Clare, S.; Pertel, T.; John, S.; Wash, R.; Smith, S.; Chin, C.; Feeley, E.; Simms, J.; Adams, D.; et al. IFITM3 restricts the morbidity and mortality associated with influenza. Int. J. Infect. Dis. 2012, 16, e79. [Google Scholar] [CrossRef] [Green Version]
- Schoggins, J.W. Interferon-Stimulated Genes: What Do They All Do? Annu. Rev. Virol. 2019, 6, 567–584. [Google Scholar] [CrossRef]
- Utay, N.S.; Douek, D.C. Interferons and HIV Infection: The Good, the Bad, and the Ugly. Pathog. Immun. 2016, 1, 107. [Google Scholar] [CrossRef] [Green Version]
- Broggi, A.; Ghosh, S.; Sposito, B.; Spreafico, R.; Balzarini, F.; Lo Cascio, A.; Clementi, N.; De Santis, M.; Mancini, N.; Granucci, F.; et al. Type III interferons disrupt the lung epithelial barrier upon viral recognition. Science 2020, 3545, eabc3545. [Google Scholar] [CrossRef]
- Jewell, N.A.; Cline, T.; Mertz, S.E.; Smirnov, S.V.; Flaño, E.; Schindler, C.; Grieves, J.L.; Durbin, R.K.; Kotenko, S.V.; Durbin, J.E. Lambda Interferon Is the Predominant Interferon Induced by Influenza A Virus Infection In Vivo. J. Virol. 2010, 84, 11515–11522. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Channappanavar, R.; Fehr, A.R.; Vijay, R.; Mack, M.; Zhao, J.; Meyerholz, D.K.; Perlman, S. Dysregulated Type I Interferon and Inflammatory Monocyte-Macrophage Responses Cause Lethal Pneumonia in SARS-CoV-Infected Mice. Cell Host Microbe 2016, 19, 181–193. [Google Scholar] [CrossRef] [Green Version]
- Page, C.; Goicochea, L.; Matthews, K.; Zhang, Y.; Klover, P.; Holtzman, M.J.; Hennighausen, L.; Frieman, M. Induction of Alternatively Activated Macrophages Enhances Pathogenesis during Severe Acute Respiratory Syndrome Coronavirus Infection. J. Virol. 2012, 86, 13334–13349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lucas, C.; Wong, P.; Klein, J.; Castro, T.B.R.; Silva, J.; Sundaram, M.; Ellingson, M.K.; Mao, T.; Oh, J.E.; Israelow, B.; et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 2020, 584, 463–469. [Google Scholar] [CrossRef]
- Perlman, S. COVID-19 poses a riddle for the immune system. Nature 2020, 584, 345–346. [Google Scholar] [CrossRef]
- Levy, D.E.; Kessler, D.S.; Pine, R.; Reich, N.; Darnell, J.E. Interferon-induced nuclear factors that bind a shared promoter element correlate with positive and negative transcriptional control. Genes Dev. 1988, 2, 383–393. [Google Scholar] [CrossRef] [Green Version]
- Qureshi, S.A.; Salditt-Georgieff, M.; Darnell, J.E. Tyrosine-phosphorylated Stat1 and Stat2 plus a 48-kDa protein all contact DNA in forming interferon-stimulated-gene factor 3. Proc. Natl. Acad. Sci. USA 1995, 92, 3829–3833. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Au-Yeung, N.; Mandhana, R.; Horvath, C.M. Transcriptional regulation by STAT1 and STAT2 in the interferon JAK-STAT pathway. Jak-Stat 2013, 2, e23931. [Google Scholar] [CrossRef] [Green Version]
- Kuan, T.C.; Yang, T.H.; Wen, C.H.; Chen, M.Y.; Lee, I.L.; Lin, C.S. Identifying the regulatory element for human angiotensin-converting enzyme 2 (ACE2) expression in human cardiofibroblasts. Peptides 2011, 32, 1832–1839. [Google Scholar] [CrossRef]
- Michalska, A.; Blaszczyk, K.; Wesoly, J.; Bluyssen, H.A.R. A positive feedback amplifier circuit that regulates interferon (IFN)-stimulated gene expression and controls type I and type II IFN responses. Front. Immunol. 2018, 9, 1135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wan, Y.; Shang, J.; Graham, R.; Baric, R.S.; Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: An Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020, 94, e00127-20. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; Ren, L.; Zhang, L.; Zhong, J.; Xiao, Y.; Jia, Z.; Guo, L.; Yang, J.; Wang, C.; Jiang, S.; et al. Heightened Innate Immune Responses in the Respiratory Tract of COVID-19 Patients. Cell Host Microbe 2020, 27, 883–890. [Google Scholar] [CrossRef] [PubMed]
- Israelow, B.; Song, E.; Mao, T.; Lu, P.; Meir, A.; Liu, F.; Alfajaro, M.M.; Wei, J.; Dong, H.; Homer, R.J.; et al. Mouse model of SARS-CoV-2 reveals inflammatory role of type I interferon signaling. J. Exp. Med. 2020, 217, e20201241. [Google Scholar] [CrossRef]
- Barker, H.; Parkkila, S. Bioinformatic characterization of angiotensin-converting enzyme 2, the entry receptor for SARS-CoV-2. PLoS ONE 2020, 15, e0240647. [Google Scholar] [CrossRef] [PubMed]
- Matys, V.; Fricke, E.; Geffers, R.; Gößling, E.; Haubrock, M.; Hehl, R.; Hornischer, K.; Karas, D.; Kel, A.E.; Kel-Margoulis, O.V.; et al. TRANSFAC®: Transcriptional regulation, from patterns to profiles. Nucleic Acids Res. 2003, 31, 374–378. [Google Scholar] [CrossRef] [PubMed]
- Rivals, I.; Personnaz, L.; Taing, L.; Potier, M.C. Enrichment or depletion of a GO category within a class of genes: Which test? Bioinformatics 2007, 23, 401–407. [Google Scholar] [CrossRef]
- Tavazoie, S.; Hughes, J.D.; Campbell, M.J.; Cho, R.J.; Church, G.M. Systematic determination of genetic network architecture. Nat. Genet. 1999, 22, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Lodhi, N.; Kossenkov, A.V.; Tulin, A.V. Bookmarking promoters in mitotic chromatin: Poly(ADP-ribose)polymerase-1 as an epigenetic mark. Nucleic Acids Res. 2014, 42, 7028–7038. [Google Scholar] [CrossRef] [Green Version]
- Lodhi, N.; Ji, Y.; Tulin, A.V. Mitotic bookmarking: Maintaining post-mitotic reprogramming of transcription reactivation. Curr. Mol. Biol. Rep. 2016, 2, 10–16. [Google Scholar] [CrossRef] [Green Version]
- Caravaca, J.M.; Donahue, G.; Becker, J.S.; He, X.; Vinson, C.; Zaret, K.S. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev. 2013, 27, 251–260. [Google Scholar] [CrossRef] [Green Version]
- Kadauke, S.; Udugama, M.I.; Pawlicki, J.M.; Achtman, J.C.; Jain, D.P.; Cheng, Y.; Hardison, R.C.; Blobel, G.A. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 2012, 150, 725–737. [Google Scholar] [CrossRef] [Green Version]
- Schiewer, M.J.; Knudsen, K.E. Transcriptional roles of PARP1 in cancer. Mol. Cancer Res. 2014, 12, 1069–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fehr, A.R.; Singh, S.A.; Kerr, C.M.; Mukai, S.; Higashi, H.; Aikawa, M. The impact of PARPs and ADP-ribosylation on inflammation and host-pathogen interactions. Genes Dev. 2020, 34, 341–359. [Google Scholar] [CrossRef]
- Gupte, R.; Nandu, T.; Kraus, W.L. Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macro phages. Nat. Commun. 2021, 12, 3931. [Google Scholar] [CrossRef] [PubMed]
- Kotova, E.; Lodhi, N.; Jarnik, M.; Pinnola, A.D.; Ji, Y.; Tulin, A.V. Drosophila histone H2A variant (H2Av) controls poly(ADP-ribose) polymerase 1 (PARP1) activation. Proc. Natl. Acad. Sci. USA 2011, 108, 6205–6210. [Google Scholar] [CrossRef] [Green Version]
- Lomvardas, S.; Thanos, D. Modifying gene expression programs by altering core promoter chromatin architecture. Cell 2002, 110, 261–271. [Google Scholar] [CrossRef] [Green Version]
- Thomas, C.J.; Kotova, E.; Andrake, M.; Adolf-Bryfogle, J.; Glaser, R.; Regnard, C.; Tulin, A.V. Kinase-mediated changes in nucleosome conformation trigger chromatin decondensation via poly(ADP-ribosyl)ation. Mol. Cell 2014, 53, 831–842. [Google Scholar] [CrossRef] [Green Version]
- Krishnakumar, R.; Kraus, W.L. PARP-1 Regulates Chromatin Structure and Transcription through a KDM5B-Dependent Pathway. Mol. Cell 2010, 39, 736–749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tulin, A.; Stewart, D.; Spradling, A.C. The Drosophila heterochromatic gene encoding poly(ADP-ribose) polymerase (PARP) is required to modulate chromatin structure during development. Genes Dev. 2002, 16, 2108–2119. [Google Scholar] [CrossRef] [Green Version]
- Boamah, E.K.; Kotova, E.; Garabedian, M.; Jarnik, M.; Tulin, A.V. Poly(ADP-ribose) polymerase 1 (PARP-1) regulates ribosomal biogenesis in Drosophila nucleoli. PLoS Genet. 2012, 8, e1002442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meder, V.S.; Boeglin, M.; de Murcia, G.; Schreiber, V. PARP-1 and PARP-2 interact with nucleophosmin/B23 and accumulate in transcriptionally active nucleoli. J. Cell Sci. 2005, 118, 211–222. [Google Scholar] [CrossRef] [Green Version]
- Ji, Y.; Tulin, A.V. The roles of PARP1 in gene control and cell differentiation. Curr. Opin. Genet. Dev. 2010, 20, 512–518. [Google Scholar] [CrossRef] [Green Version]
- Tulin, A.; Spradling, A. Chromatin loosening by poly(ADP)-ribose polymerase (PARP) at Drosophila puff loci. Science 2003, 299, 560–562. [Google Scholar] [CrossRef]
- Petesch, S.J.; Lis, J.T. Activator-induced spread of poly(ADP-Ribose) polymerase promotes nucleosome Loss at Hsp70. Mol. Cell 2012, 45, 64–74. [Google Scholar] [CrossRef] [Green Version]
- Kim, M.Y.; Mauro, S.; Gévry, N.; Lis, J.T.; Kraus, W.L. NAD+-dependent modulation of chromatin structure and transcription by nucleosome binding properties of PARP-1. Cell 2004, 119, 803–814. [Google Scholar] [CrossRef] [Green Version]
- Espey, M.G.; Miranda, K.M.; Pluta, R.M.; Wink, D.A. Nitrosative capacity of macrophages is dependent on nitric-oxide synthase induction signals. J. Biol. Chem. 2000, 275, 11341–11347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wink, D.A.; Hines, H.B.; Cheng, R.Y.S.; Switzer, C.H.; Flores-Santana, W.; Vitek, M.P.; Ridnour, L.A.; Colton, C.A. Nitric oxide and redox mechanisms in the immune response. J. Leukoc. Biol. 2011, 89, 873–891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, W.; Zheng, S.; Dweik, R.A.; Erzurum, S.C. Role of epithelial nitric oxide in airway viral infection. Free Radic. Biol. Med. 2006, 41, 19–28. [Google Scholar] [CrossRef]
- Forrester, S.J.; Kikuchi, D.S.; Hernandes, M.S.; Xu, Q.; Griendling, K.K. Reactive oxygen species in metabolic and inflammatory signaling. Circ. Res. 2018, 122, 877–902. [Google Scholar] [CrossRef]
- Xia, Y.; Roman, L.J.; Masters, B.S.S.; Zweier, J.L. Inducible nitric-oxide synthase generates superoxide from the reductase domain. J. Biol. Chem. 1998, 273, 22635–22639. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pou, S.; Keaton, L.; Surichamorn, W.; Rosen, G.M. Mechanism of superoxide generation by neuronal nitric-oxide synthase. J. Biol. Chem. 1999, 274, 9573–9580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Villa, V.; Thellung, S.; Bajetto, A.; Gatta, E.; Robello, M.; Novelli, F.; Tasso, B.; Tonelli, M.; Florio, T. Novel celecoxib analogues inhibit glial production of prostaglandin E2, nitric oxide, and oxygen radicals reverting the neuroinflammatory responses induced by misfolded prion protein fragment 90–231 or lipopolysaccharide. Pharmacol. Res. 2016, 113, 500–514. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Song, M.; Zhang, B.; Zhang, Y. Reactive Oxygen Species Regulate T Cell Immune Response in the Tumor Microenvironment. Oxid. Med. Cell. Longev. 2016, 2016, 1580967. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flescher, E.; Ledbetter, J.A.; Schieven, G.L.; Vela-Roch, N.; Fossum, D.; Dang, H.; Ogawa, N.; Talal, N. Longitudinal exposure of human T lymphocytes to weak oxidative stress suppresses transmembrane and nuclear signal transduction. J. Immunol. 1994, 153, 4880–4889. [Google Scholar]
- Lahdenpohja, N.; Hurme, M. CD28-mediated activation in CD45RA+ and CD45RO+ T cells: Enhanced levels of reactive oxygen intermediates and c-Rel nuclear translocation in CD45RA+ cells. J. Leukoc. Biol. 1998, 63, 775–780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dayal, D.; Gupta, S. Connecting BCG Vaccination and COVID-19: Additional Data. medRxiv 2020. [Google Scholar] [CrossRef]
- Wardhana; Datau, E.A.; Sultana, A.; Mandang, V.V.; Jim, E. The efficacy of Bacillus Calmette-Guerin vaccinations for the prevention of acute upper respiratory tract infection in the elderly. Acta Med. Indones. 2011, 43, 185–190. [Google Scholar]
- Ohrui, T.; Nakayama, K.; Fukushima, T.; Chiba, H.; Sasaki, H. Prevention of elderly pneumonia by pneumococcal, influenza and BCG vaccinations. Jpn. J. Geriatr. 2005, 42, 34–36. [Google Scholar] [CrossRef] [Green Version]
- Freyne, B.; Marchant, A.; Curtis, N. BCG-associated heterologous immunity, a historical perspective: Intervention studies in animal models of infectious diseases. Trans. R. Soc. Trop. Med. Hyg. 2014, 109, 52–61. [Google Scholar] [CrossRef]
- Spencer, J.C.; Ganguly, R.; Waldman, R.H. Nonspecific protection of mice against influenza virus infection by local or systemic immunization with Bacille Calmette-Guérin. J. Infect. Dis. 1977, 136, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Moorlag, S.J.C.F.M.; Arts, R.J.W.; van Crevel, R.; Netea, M.G. Non-specific effects of BCG vaccine on viral infections. Clin. Microbiol. Infect. 2019, 25, 1473–1478. [Google Scholar] [CrossRef]
- Chen, Y.; Shen, Z. Gene polymorphisms in the folate metabolism and their association with MTX-related adverse events in the treatment of ALL. Tumor Biol. 2015, 36, 4913–4921. [Google Scholar] [CrossRef] [PubMed]
- De Iuliis, F.; Salerno, G.; Taglieri, L.; Lanza, R.; Scarpa, S. On and off metronomic oral vinorelbine in elderly women with advanced breast cancer. Tumori 2015, 101, 30–35. [Google Scholar] [CrossRef]
- Deng, N.; Zhou, H.; Fan, H.; Yuan, Y. Single nucleotide polymorphisms and cancer susceptibility. Oncotarget 2017, 8, 110635–110649. [Google Scholar] [CrossRef] [Green Version]
- Yue, J.X.; Li, J.; Aigrain, L.; Hallin, J.; Persson, K.; Oliver, K.; Bergström, A.; Coupland, P.; Warringer, J.; Lagomarsino, M.C.; et al. Contrasting evolutionary genome dynamics between domesticated and wild yeasts. Nat. Genet. 2017, 49, 913–924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Webster, G.A.; Perkins, N.D. Transcriptional Cross Talk between NF-κB and p53. Mol. Cell. Biol. 1999, 19, 3485–3495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niu, G.; Wright, K.L.; Ma, Y.; Wright, G.M.; Huang, M.; Irby, R.; Briggs, J.; Karras, J.; Cress, W.D.; Pardoll, D.; et al. Role of Stat3 in Regulating p53 Expression and Function. Mol. Cell. Biol. 2005, 25, 7432–7440. [Google Scholar] [CrossRef] [Green Version]
- Beckman, G.; Birgander, R.; Sjalander, A.; Saha, N.; Holmberg, P.A.; Kivela, A.; Beckman, L. Is p53 polymorphism maintained by natural selection? Hum. Hered. 1994, 44, 266–270. [Google Scholar] [CrossRef] [PubMed]
- Murphy, M.E. Polymorphic variants in the p53 pathway. Cell Death Differ. 2006, 13, 916–920. [Google Scholar] [CrossRef]
- Whibley, C.; Pharoah, P.D.P.; Hollstein, M. p53 polymorphisms: Cancer implications. Nat. Rev. Cancer 2009, 9, 95–107. [Google Scholar] [CrossRef]
- Matlashewski, G.J.; Tuck, S.; Pim, D.; Lamb, P.; Schneider, J.; Crawford, L. V Primary structure polymorphism at amino acid residue 72 of human p53. Mol. Cell. Biol. 1987, 7, 961–963. [Google Scholar] [CrossRef] [Green Version]
- Leu, J.I.-J.; Murphy, M.E.; George, D.L. The p53 Codon 72 Polymorphism Modifies the Cellular Response to Inflammatory Challenge in the Liver. J. Liver 2013, 2, 117. [Google Scholar]
- Jeong, B.; Hu, W.; Belyi, V.; Rabadan, R.; Levine, A.J. Differential levels of transcription of p53-regulated genes by the arginine/proline polymorphism: p53 with arginine at codon 72 favors apoptosis. FASEB J. 2010, 24, 1347–1353. [Google Scholar] [CrossRef] [PubMed]
- Ørsted, D.D.; Bojesen, S.E.; Tybjærg-Hansen, A.; Nordestgaard, B.G. Tumor suppressor p53 Arg72Pro polymorphism and longevity, cancer survival, and risk of cancer in the general population. J. Exp. Med. 2007, 204, 1295–1301. [Google Scholar] [CrossRef]
- Van Heemst, D.; Mooijaart, S.P.; Beekman, M.; Schreuder, J.; De Craen, A.J.M.; Brandt, B.W.; Eline Slagboom, P.; Westendorp, R.G.J. Variation in the human TP53 gene affects old age survival and cancer mortality. Exp. Gerontol. 2005, 40, 11–15. [Google Scholar] [CrossRef]
- Azzam, G.A.; Frank, A.K.; Hollstein, M.; Murphy, M.E. Tissue-specific apoptotic effects of the p53 codon 72 polymorphism in a mouse model. Cell Cycle 2011, 10, 1352–1355. [Google Scholar] [CrossRef] [Green Version]
- Frank, A.K.; Leu, J.I.-J.; Zhou, Y.; Devarajan, K.; Nedelko, T.; Klein-Szanto, A.; Hollstein, M.; Murphy, M.E. The Codon 72 Polymorphism of p53 Regulates Interaction with NF- B and Transactivation of Genes Involved in Immunity and Inflammation. Mol. Cell. Biol. 2011, 31, 1201–1213. [Google Scholar] [CrossRef] [Green Version]
- Kung, C.P.; Liu, Q.; Murphy, M.E. The codon 72 polymorphism of p53 influences cell fate following nutrient deprivation. Cancer Biol. Ther. 2017, 18, 484–491. [Google Scholar] [CrossRef] [Green Version]
- Kung, C.P.; Leu, J.I.J.; Basu, S.; Khaku, S.; Anokye-Danso, F.; Liu, Q.; George, D.L.; Ahima, R.S.; Murphy, M.E. The P72R Polymorphism of p53 Predisposes to Obesity and Metabolic Dysfunction. Cell Rep. 2016, 14, 2413–2425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Wu, L.; Yue, X.; Zhang, C.; Wang, J.; Li, J.; Sun, X.; Zhu, Y.; Feng, Z.; Hu, W. A polymorphism in the tumor suppressor p53 affects aging and longevity in mouse models. eLife 2018, 7, e34701. [Google Scholar] [CrossRef] [PubMed]
- Bojesen, S.E.; Nordestgaard, B.G. The common germline Arg72Pro polymorphism of p53 and increased longevity in humans. Cell Cycle 2008, 7, 158–163. [Google Scholar] [CrossRef] [Green Version]
- Cho, J.H.; Patel, B.; Bonala, S.; Mansouri, H.; Manne, S.; Vadrevu, S.K.; Ghouse, S.; Kung, C.P.; Murphy, M.E.; Astrinidis, A.; et al. The Codon 72 TP53Polymorphism Contributes to TSC Tumorigenesis through the Notch-Nodal Axis. Mol. Cancer Res. 2019, 17, 1639–1651. [Google Scholar] [CrossRef]
- Schuck-Paim, C.; Viboud, C.; Simonsen, L.; Miller, M.A.; Moura, F.E.A.; Fernandes, R.M.; Carvalho, M.L.; Alonso, W.J. Were equatorial regions less affected by the 2009 influenza pandemic? The Brazilian experience. PLoS ONE 2012, 7, e41918. [Google Scholar] [CrossRef] [Green Version]
- Gage, Z.O.; Vasou, A.; Gray, D.W.; Randall, R.E.; Adamson, C.S. Identification of novel inhibitors of the type i interferon induction pathway using cell-based high-throughput screening. J. Biomol. Screen. 2016, 21, 978–988. [Google Scholar] [CrossRef] [Green Version]
- Kawai, T.; Akira, S. The role of pattern-recognition receptors in innate immunity: Update on toll-like receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Celhar, T.; Magalhães, R.; Fairhurst, A.M. TLR7 and TLR9 in SLE: When sensing self goes wrong. Immunol. Res. 2012, 53, 58–77. [Google Scholar] [CrossRef] [PubMed]
- Roh, J.S.; Sohn, D.H. Origin and List of Damps. Immune Netw. 2018, 18, e27. [Google Scholar] [CrossRef]
- Blasius, A.L.; Beutler, B. Intracellular Toll-like Receptors. Immunity 2010, 32, 305–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jefferies, C.A.; Fitzgerald, K.A. Interferon gene regulation: Not all roads lead to Tolls. Trends Mol. Med. 2005, 11, 403–411. [Google Scholar] [CrossRef]
- Bowie, A.G.; Haga, I.R. The role of Toll-like receptors in the host response to viruses. Mol. Immunol. 2005, 42, 859–867. [Google Scholar] [CrossRef]
- Philbin, V.J.; Iqbal, M.; Boyd, Y.; Goodchild, M.J.; Beal, R.K.; Bumstead, N.; Young, J.; Smith, A.L. Identification and characterization of a functional, alternatively spliced Toll-like receptor 7 (TLR7) and genomic disruption of TLR8 in chickens. Immunology 2005, 114, 507–521. [Google Scholar] [CrossRef]
- Gorden, K.B.; Gorski, K.S.; Gibson, S.J.; Kedl, R.M.; Kieper, W.C.; Qiu, X.; Tomai, M.A.; Alkan, S.S.; Vasilakos, J.P. Synthetic TLR Agonists Reveal Functional Differences between Human TLR7 and TLR8. J. Immunol. 2005, 174, 1259–1268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mancuso, G.; Gambuzza, M.; Midiri, A.; Biondo, C.; Papasergi, S.; Akira, S.; Teti, G.; Beninati, C. Bacterial recognition by TLR7 in the lysosomes of conventional dendritic cells. Nat. Immunol. 2009, 10, 587–594. [Google Scholar] [CrossRef] [PubMed]
- Guiducci, C.; Gong, M.; Cepika, A.M.; Xu, Z.; Tripodo, C.; Bennett, L.; Crain, C.; Quartier, P.; Cush, J.J.; Pascual, V.; et al. RNA recognition by human TLR8 can lead to autoimmune inflammation. J. Exp. Med. 2013, 210, 2903–2919. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Sato, S.; Ishii, K.J.; Coban, C.; Hemmi, H.; Yamamoto, M.; Terai, K.; Matsuda, M.; Inoue, J.I.; Uematsu, S.; et al. Interferon-α induction through Toll-like receptors involves a direct interaction of IRF7 with MyD88 and TRAF6. Nat. Immunol. 2004, 5, 1061–1068. [Google Scholar] [CrossRef]
- Coban, C.; Igari, Y.; Yagi, M.; Reimer, T.; Koyama, S.; Aoshi, T.; Ohata, K.; Tsukui, T.; Takeshita, F.; Sakurai, K.; et al. Immunogenicity of Whole-Parasite Vaccines against Plasmodium falciparum Involves Malarial Hemozoin and Host TLR9. Cell Host Microbe 2010, 7, 50–61. [Google Scholar] [CrossRef] [Green Version]
- Alexopoulou, L.; Holt, A.; Medzhitov, R.; Flavell, R. Recognition of double-stranded RNA and activation of NF-kappaB by Toll. Nature 2001, 413, 732–738. [Google Scholar] [CrossRef] [PubMed]
- Fitzgerald, K.A.; Kagan, J.C. Toll-like Receptors and the Control of Immunity. Cell 2020, 180, 1044–1066. [Google Scholar] [CrossRef]
- El-Zayat, S.R.; Sibaii, H.; Mannaa, F.A. Toll-like receptors activation, signaling, and targeting: An overview. Bull. Natl. Res. Cent. 2019, 43, 187. [Google Scholar] [CrossRef] [Green Version]
- Kumar, A.; Negi, G.; Sharma, S.S. JSH-23 targets nuclear factor-kappa B and reverses various deficits in experimental diabetic neuropathy: Effect on neuroinflammation and antioxidant defence. Diabetes Obes. Metab. 2011, 13, 750–758. [Google Scholar] [CrossRef]
- Gay, N.J.; Symmons, M.F.; Gangloff, M.; Bryant, C.E. Assembly and localization of Toll-like receptor signalling complexes. Nat. Rev. Immunol. 2014, 14, 546–558. [Google Scholar] [CrossRef]
- Zhao, G.N.; Jiang, D.S.; Li, H. Interferon regulatory factors: At the crossroads of immunity, metabolism, and disease. Biochim. Biophys. Acta Mol. Basis Dis. 2015, 1852, 365–378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turner, M.D.; Nedjai, B.; Hurst, T.; Pennington, D.J. Cytokines and chemokines: At the crossroads of cell signalling and inflammatory disease. BBA Mol. Cell Res. 2014, 1843, 2563–2582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kotenko, S.V.; Gallagher, G.; Baurin, V.V.; Lewis-Antes, A.; Shen, M.; Shah, N.K.; Langer, J.A.; Sheikh, F.; Dickensheets, H.; Donnelly, R.P. IFN-λs mediate antiviral protection through a distinct class II cytokine receptor complex. Nat. Immunol. 2003, 4, 69–77. [Google Scholar] [CrossRef]
- Muller, M.; Briscoe, J.; Laxton, C.; Guschin, D.; Ziemiecki, A.; Silvennoinen, O.; Harpur, A.G.; Barbieri, G.; Witthuhn, B.A.; Schindler, C.; et al. The protein tyrosine kinase JAK1 complements defects in interferon-alpha/beta and -gamma signal transduction. Nature 1993, 366, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Dumoutier, L.; Lejeune, D.; Hor, S.; Fickenscher, H.; Renauld, J.C. Cloning of a new type II cytokine receptor activating signal transducer and activator of transcription (STAT)1, STAT2 and STAT3. Biochem. J. 2003, 370, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Ivashkiv, L.B.; Donlin, L.T. Donlin1 Regulation of type I interferon responses Lionel. Nat. Rev. Immunol. 2014, 14, 36–49. [Google Scholar] [CrossRef] [Green Version]
- Stark, G.R.; Darnell, J.E. The JAK-STAT Pathway at Twenty. Immunity 2012, 36, 503–514. [Google Scholar] [CrossRef] [Green Version]
- Pestka, S.; Krause, C.D.; Walter, M.R. Interferons, interferon-like cytokines, and their receptors. Immunol. Rev. 2004, 202, 8–32. [Google Scholar] [CrossRef]
- Hamiel, U.; Kozer, E.; Youngster, I. SARS-CoV-2 Rates in BCG-Vaccinated and Unvaccinated Young Adults. JAMA J. Am. Med. Assoc. 2020, 323, 2340–2341. [Google Scholar] [CrossRef]
- Yu, Y.; Jin, H.; Chen, Z.; Yu, Q.L.; Ma, Y.J.; Sun, X.L.; Wang, B. Children’s vaccines do not induce cross reactivity against SARS-CoV. J. Clin. Pathol. 2007, 60, 208–211. [Google Scholar] [CrossRef] [Green Version]
- Netea, M.G.; Giamarellos-Bourboulis, E.J.; Domínguez-Andrés, J.; Curtis, N.; van Crevel, R.; van de Veerdonk, F.L.; Bonten, M. Trained Immunity: A Tool for Reducing Susceptibility to and the Severity of SARS-CoV-2 Infection. Cell 2020, 181, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Karpova, Y.; Wu, C.; Divan, A.; McDonnell, M.E.; Hewlett, E.; Makhov, P.; Gordon, J.; Ye, M.; Reitz, A.B.; Childers, W.E.; et al. Non-NAD-like PARP-1 inhibitors in prostate cancer treatment. Biochem. Pharmacol. 2019, 167, 149–162. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.; Ji, Y.; Lodhi, N.; Kotova, E.; Pinnola, A.D.; Golovine, K.; Makhov, P.; Pechenkina, K.; Kolenko, V.; Tulin, A.V. Non-NAD-Like poly(ADP-Ribose) Polymerase-1 Inhibitors effectively Eliminate Cancer in vivo. EBioMedicine 2016, 13, 90–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brotherton, H.; Usuf, E.; Nadjm, B.; Forrest, K.; Bojang, K.; Samateh, A.L.; Bittaye, M.; Roberts, C.A.P.; Alessandro, U.; Roca, A. Correspondence Dexamethasone for COVID-19: Data needed from randomised clinical trials in Africa. Lancet Glob. Health 2020, 8, e1125–e1126. [Google Scholar] [CrossRef]
- Riggs, J.M.; Hanna, R.N.; Rajan, B.; Zerrouki, K.; Karnell, J.L.; Sagar, D.; Vainshtein, I.; Farmer, E.; Rosenthal, K.; Morehouse, C.; et al. Characterisation of anifrolumab, a fully human anti-interferon receptor antagonist antibody for the treatment of systemic lupus erythematosus. Lupus Sci. Med. 2018, 5, e000261. [Google Scholar] [CrossRef]
- Ho, J.; Pelzel, C.; Begitt, A.; Mee, M.; Elsheikha, H.M.; Scott, D.J.; Vinkemeier, U. STAT2 Is a Pervasive Cytokine Regulator due to Its Inhibition of STAT1 in Multiple Signaling Pathways. PLoS Biol. 2016, 14, e2000117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Menet, C.J. A Dual Inhibition, a Better Solution: Development of a JAK1/TYK2 inhibitor. J. Med. Chem. 2018, 61, 8594–8596. [Google Scholar] [CrossRef] [Green Version]
- Fensome, A.; Ambler, C.M.; Arnold, E.; Banker, M.E.; Brown, M.F.; Chrencik, J.; Clark, J.D.; Dowty, M.E.; Efremov, I.V.; Flick, A.; et al. Dual Inhibition of TYK2 and JAK1 for the Treatment of Autoimmune Diseases: Discovery of ((S)-2,2-Difluorocyclopropyl)((1 R,5 S)-3-(2-((1-methyl-1 H-pyrazol-4-yl)amino)pyrimidin-4-yl)-3,8-diazabicyclo[3.2.1]octan-8-yl)methanone (PF-06700841). J. Med. Chem. 2018, 61, 8597–8612. [Google Scholar] [CrossRef]
- Ikushima, H.; Negishi, H.; Taniguchi, T. The IRF family transcription factors at the interface of innate and adaptive immune responses. Cold Spring Harb. Symp. Quant. Biol. 2013, 78, 105–116. [Google Scholar] [CrossRef] [Green Version]
- Honda, K.; Takaoka, A.; Taniguchi, T. Type I Inteferon Gene Induction by the Interferon Regulatory Factor Family of Transcription Factors. Immunity 2006, 25, 349–360. [Google Scholar] [CrossRef] [Green Version]
- Honda, K.; Taniguchi, T. IRFs: Master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat. Rev. Immunol. 2006, 6, 644–658. [Google Scholar] [CrossRef] [PubMed]
- Negishi, H.; Ohba, Y.; Yanai, H.; Takaoka, A.; Honma, K.; Yui, K.; Matsuyama, T.; Taniguchi, T.; Honda, K. Negative regulation of Toll-like-receptor signaling by IRF-4. Proc. Natl. Acad. Sci. USA 2005, 102, 15989–15994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szelag, M.; Piaszyk-Borychowska, A.; Plens-Galaska, M.; Wesoly, J.; Bluyssen, H.A.R. Targeted inhibition of STATs and IRFs as a potential treatment strategy in cardiovascular disease. Oncotarget 2016, 7, 48788–48812. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Antonczyk, A.; Krist, B.; Sajek, M.; Michalska, A.; Piaszyk-Borychowska, A.; Plens-Galaska, M.; Wesoly, J.; Bluyssen, H.A.R. Direct inhibition of IRF-dependent transcriptional regulatory mechanisms associated with disease. Front. Immunol. 2019, 10, 1176. [Google Scholar] [CrossRef] [Green Version]
- Liu, T.; Zhang, L.; Joo, D.; Sun, S.C. NF-κB signaling in inflammation. Signal Transduct. Target. Ther. 2017, 2, 17023. [Google Scholar] [CrossRef] [Green Version]
- Taniguchi, K.; Karin, M. NF-B, inflammation, immunity and cancer: Coming of age. Nat. Rev. Immunol. 2018, 18, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Lowe, J.M.; Menendez, D.; Bushel, P.R.; Shatz, M.; Kirk, E.L.; Troester, A.; Garantziotis, S.; Fessler, M.B.; Resnick, M.A. p53 and NF-kappaB Co-Regulate Pro-Inflammatory Gene Responses in Human Macrophages. Cancer Res. 2014, 74, 2182–2192. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Dawson, V.L.; Dawson, T.M.; Snyder, S.H. Nitric oxide activation of poly(ADP-ribose) synthetase in neurotoxicity. Science 1994, 263, 687–689. [Google Scholar] [CrossRef]
- Zingarelli, B.; O’Connor, M.; Wong, H.; Salzman, A.L.; Szabó, C. Peroxynitrite-mediated DNA strand breakage activates poly-adenosine diphosphate ribosyl synthetase and causes cellular energy depletion in macrophages stimulated with bacterial lipopolysaccharide. J. Immunol. 1996, 156, 350–358. [Google Scholar]
- Le Page, C.; Sanceau, J.; Drapier, J.C.; Wietzerbin, J. Inhibitors of ADP-Ribosylation impair inducible nitric oxide synthase gene transcription through inhibition of NF kappa B activation. Biochem. Biophys. Res. Commun. 1998, 243, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Gilmore, T.D.; Herscovitch, M. Inhibitors of NF-κB signaling: 785 and counting. Oncogene 2006, 25, 6887–6899. [Google Scholar] [CrossRef] [Green Version]
- Herrington, F.D.; Carmody, R.J.; Goodyear, C.S. Modulation of NF-κB Signaling as a Therapeutic Target in Autoimmunity. J. Biomol. Screen. 2016, 21, 223–242. [Google Scholar] [CrossRef] [Green Version]
- Bupp, K.; Roth, M.J. Alteration and Analyses of Viral Entry with Library-Derived Peptides. Adv. Virus Res. 2005, 65, 147–172. [Google Scholar] [CrossRef]
- Perlman, S.; Netland, J. Coronaviruses post-SARS: Update on replication and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 439–450. [Google Scholar] [CrossRef] [Green Version]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Simmons, G.; Gosalia, D.N.; Rennekamp, A.J.; Reeves, J.D.; Diamond, S.L.; Bates, P. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc. Natl. Acad. Sci. USA 2005, 102, 11876–11881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dimitrov, D.S. The Secret Life of ACE2 as a Receptor for the SARS Virus. Cell 2003, 115, 652–653. [Google Scholar] [CrossRef] [Green Version]
- Bittmann, S.; Weissenstein, A.; Villalon, G.; Moschuring-Alieva, E.; Luchter, E. Simultaneous Treatment of COVID-19 With Serine Protease Inhibitor Camostat and/or Cathepsin L Inhibitor? J. Clin. Med. Res. 2020, 12, 320–322. [Google Scholar] [CrossRef]
- Shen, L.W.; Mao, H.J.; Wu, Y.L.; Tanaka, Y.; Zhang, W. TMPRSS2: A potential target for treatment of influenza virus and coronavirus infections. Biochimie 2017, 142, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Lucas, J.M.; Heinlein, C.; Kim, T.; Hernandez, S.A.; Malik, M.S.; True, L.D.; Morrissey, C.; Corey, E.; Montgomery, B.; Mostaghel, E.; et al. The androgen-regulated protease TMPRSS2 activates a proteolytic cascade involving components of the tumor microenvironment and promotes prostate cancer metastasis. Cancer Discov. 2014, 4, 1310–1325. [Google Scholar] [CrossRef] [Green Version]
- Bestle, D.; Heindl, M.R.; Limburg, H.; Van Lam van, T.; Pilgram, O.; Moulton, H.; Stein, D.A.; Hardes, K.; Eickmann, M.; Dolnik, O.; et al. TMPRSS2 and furin are both essential for proteolytic activation and spread of SARS-CoV-2 in human airway epithelial cells and provide promising drug targets. bioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Yamamoto, M.; Matsuyama, S.; Li, X.; Takeda, M.; Kawaguchi, Y.; Inoue, J. Respiratory Syndrome Coronavirus S Protein-Mediated Membrane. Antimicrob. Agents Chemother. 2016, 60, 6532–6539. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crooke, S.T.; Witztum, J.L.; Bennett, C.F.; Baker, B.F. RNA-Targeted Therapeutics. Cell Metab. 2018, 27, 714–739. [Google Scholar] [CrossRef] [Green Version]
- Barrey, E.; Burzio, V.; Dhorne-pollet, S.; Delmas, B. Think Different with RNA Therapy: Can Antisense Oligonucleotides Be Used to Inhibit Replication and Transcription of SARS-Cov-2 ? Prepints 2020, 2020040412. [Google Scholar] [CrossRef] [Green Version]
- Neuman, B.W.; Stein, D.A.; Kroeker, A.D.; Bestwick, R.K.; Iversen, P.L.; Moulton, H.M.; Buchmeier, M.J. Inhibition and escape of SARS-CoV treated with antisense morpholino oligomers. Adv. Exp. Med. Biol. 2006, 581, 567–571. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.; Singh, R.P.; Rabadia, N.; Patel, G.; Panchal, H. Antisense Technology. Int. J. Pharm. Sci. Rev. 2011, 9, 38–45. [Google Scholar]
- Mohammed, K.-S.; Scanlon, K.J. Antisense and Ribozyme Therapy. In The Cancer Handbook; John Wiley & Sons: Hoboken, NJ, USA, 2005. [Google Scholar] [CrossRef]
- Wang, S.; White, K.A. Riboswitching on RNA virus replication. Proc. Natl. Acad. Sci. USA 2007, 104, 10406–10411. [Google Scholar] [CrossRef] [Green Version]
- Rodrigo, G.; Prakash, S.; Cordero, T.; Kushwaha, M.; Jaramillo, A. Functionalization of an Antisense Small RNA. J. Mol. Biol. 2016, 428, 889–892. [Google Scholar] [CrossRef] [Green Version]
- Bao, Q.; Li, X.; Han, G.; Zhu, Y.; Mao, C.; Yang, M. Phage-based vaccines. Adv. Drug Deliv. Rev. 2019, 145, 40–56. [Google Scholar] [CrossRef]
- Wojewodzic, M.W. Bacteriophages Could Be a Potential Game Changer in the Trajectory of Coronavirus Disease (COVID-19) Bacteriophages Used for Accelerated Therapeutic Antibody Production. Ther. Appl. Res. 2020, 1, 3237. [Google Scholar]
- Abbott, T.R.; Dhamdhere, G.; Liu, Y.; Lin, X.; Goudy, L.; Zeng, L.; Chemparathy, A.; Chmura, S.; Heaton, N.S.; Debs, R.; et al. Development of CRISPR as an Antiviral Strategy to Combat SARS-CoV-2 and Influenza. Cell 2020, 181, 865–876.e12. [Google Scholar] [CrossRef]
- Nakad, R.; Schumacher, B. DNA damage response and immune defense: Links and mechanisms. Front. Genet. 2016, 21, 7504. [Google Scholar] [CrossRef] [Green Version]
- Morales, A.J.; Carrero, J.A.; Hung, P.J.; Tubbs, A.T.; Andrews, J.M.; Edelson, B.T.; Calderon, B.; Innes, C.L.; Paules, R.S.; Payton, J.E.; et al. A type I IFN-dependent DNA damage response regulates the genetic program and inflammasome activation in macrophages. eLife 2017, 6, e24655. [Google Scholar] [CrossRef] [PubMed]
- Dexamethasone in Hospitalized Patients with Covid-19. N. Engl. J. Med. 2021, 384, 693–704. [CrossRef] [PubMed]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Kleine-Tebbe, J.; Klimek, L.; Hamelmann, E.; Pfaar, O.; Taube, C.; Wagenmann, M.; Werfel, T.; Worm, M. Severe allergic reactions to the COVID-19 vaccine—Statement and practical consequences. Allergol. Sel. 2021, 5, 26–28. [Google Scholar] [CrossRef] [PubMed]
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: An interim analysis of four randomised controlled trials in Brazil, South Africa, and the UK. Lancet 2021, 397, 99–111. [Google Scholar] [CrossRef]
- Lauring, A.S.; Hodcroft, E.B. Genetic Variants of SARS-CoV-2—What Do They Mean? JAMA J. Am. Med. Assoc. 2021, 325, 529–531. [Google Scholar] [CrossRef]
- Walensky, R.P.; Walke, H.T.; Fauci, A.S. SARS-CoV-2 Variants of Concern in the United States—Challenges and Opportunities. JAMA 2021, 325, 1037–1038. [Google Scholar] [CrossRef]
- Zimmer, K. A Guide to Emerging SARS-CoV-2 Variants|The Scientist Magazine®. The Scientist, 2021; 1–8. Available online: https://www.the-scientist.com/news-opinion/a-guide-to-emerging-sars-cov-2-variants-68387(accessed on 9 August 2021).
- Konings, F.; Perkins, M.D.; Kuhn, J.H.; Pallen, M.J.; Alm, E.J.; Archer, B.N.; Barakat, A.; Bedford, T.; Bhiman, J.N.; Caly, L.; et al. SARS-CoV-2 Variants of Interest and Concern naming scheme conducive for global discourse. Nat. Microbiol. 2021, 6, 821–823. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lodhi, N.; Singh, R.; Rajput, S.P.; Saquib, Q. SARS-CoV-2: Understanding the Transcriptional Regulation of ACE2 and TMPRSS2 and the Role of Single Nucleotide Polymorphism (SNP) at Codon 72 of p53 in the Innate Immune Response against Virus Infection. Int. J. Mol. Sci. 2021, 22, 8660. https://doi.org/10.3390/ijms22168660
Lodhi N, Singh R, Rajput SP, Saquib Q. SARS-CoV-2: Understanding the Transcriptional Regulation of ACE2 and TMPRSS2 and the Role of Single Nucleotide Polymorphism (SNP) at Codon 72 of p53 in the Innate Immune Response against Virus Infection. International Journal of Molecular Sciences. 2021; 22(16):8660. https://doi.org/10.3390/ijms22168660
Chicago/Turabian StyleLodhi, Niraj, Rubi Singh, Satya Prakash Rajput, and Quaiser Saquib. 2021. "SARS-CoV-2: Understanding the Transcriptional Regulation of ACE2 and TMPRSS2 and the Role of Single Nucleotide Polymorphism (SNP) at Codon 72 of p53 in the Innate Immune Response against Virus Infection" International Journal of Molecular Sciences 22, no. 16: 8660. https://doi.org/10.3390/ijms22168660








