Novel Insight into the Effects of CpxR on Salmonella enteritidis Cells during the Chlorhexidine Treatment and Non-Stressful Growing Conditions
Abstract
:1. Introduction
2. Results
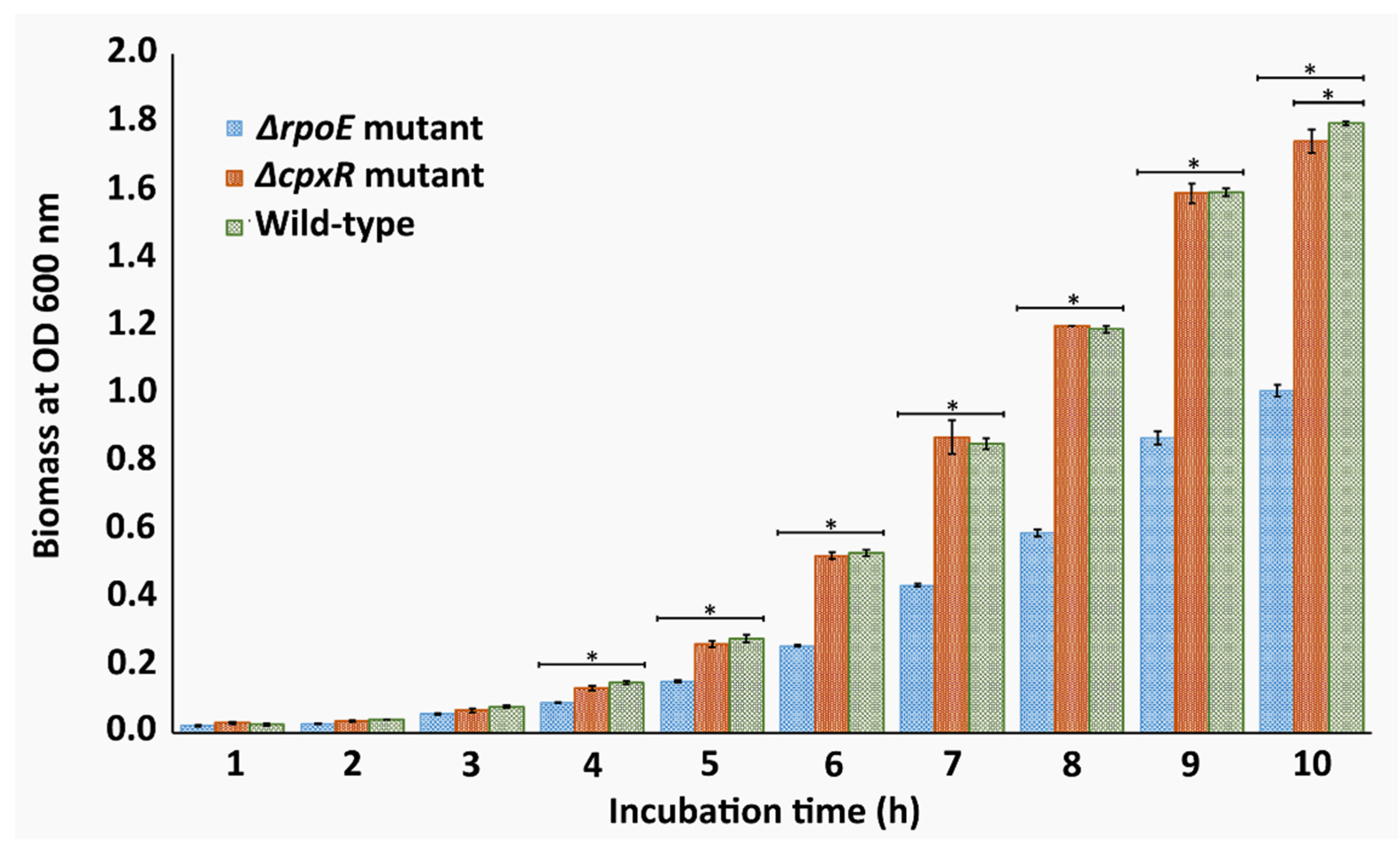
2.1. Growth Kinetics
2.2. Determination of the Minimum Inhibitory Concentration (MIC) of CHX
2.3. Effects of the Drug Treatment on the Viability of the ΔrpoE and ΔcpxR Mutants
2.4. The cpxR Mutation Does Not Cause the Major Perturbation of the Cellular Envelope during the Initial CHX Treatment
2.5. Effect of the CpxR Mutation on the Proteome of S. enteritidis during the Exponential Growth Phase
2.6. Response of the ΔcpxR Mutant to the Minimum Inhibitory Concentration of CHX
2.7. Validation of Proteomic Data
3. Discussion
4. Material and Methods
4.1. Bacterial Strains, Plasmids and Growth Conditions
4.2. Construction of Isogenic ΔrpoE and ΔcpxR Mutants
4.3. Cell Growth Assay
4.4. Determination of the Minimum Inhibitory Concentration of Chlorhexidine
4.5. Drug Treatment Assay
4.6. Complementation Study
4.7. Morphological Analysis
4.8. Samples Preparation for Protein Analysis
4.9. Protein Extraction, Proteolytic Digestion and iTRAQ Labelling
4.10. Peptide LCMS Analysis
4.11. Database Searching with Protein Pilot
4.12. Quantitative Data Analysis
4.13. Validation of Proteomic Data by qRT-PCR
4.14. Experimental Replication and Bioinformatics
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Van Boeckel, T.; Pires, J.; Silvester, R.; Zhao, C.; Song, J.; Criscuolo, N.G.; Gilbert, M.; Bonhoeffer, S.; Laxminarayan, R. Global trends in antimicrobial resistance in animals in low-and middle-income countries. Science 2019, 365. [Google Scholar] [CrossRef] [Green Version]
- Hernando-Amado, S.; Coque, T.M.; Baquero, F.; Martínez, J.L. Antibiotic resistance: Moving from individual health norms to social norms in one health and global health. Front. Microbiol. 2020, 11, 1914. [Google Scholar] [CrossRef] [PubMed]
- An, R.; Alshalchi, S.; Breimhurst, P.; Munoz-Aguayo, J.; Flores-Figueroa, C.; Vidovic, S. Strong influence of livestock environments on the emergence and dissemination of distinct multidrug-resistant phenotypes among the population of non-typhoidal Salmonella. PLoS ONE 2017, 12, e0179005. [Google Scholar] [CrossRef] [Green Version]
- Vidovic, N.; Vidovic, S. Antimicrobial resistance and food animals: Influence of livestock environment on the emergence and dissemination of antimicrobial resistance. Antibiotics 2020, 9, 52. [Google Scholar] [CrossRef] [Green Version]
- World Health Organization. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. 2017. Available online: https://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf (accessed on 18 April 2020).
- Hiratsu, K.; Amemura, M.; Nashimoto, H.; Shinagawa, H.; Makino, K. The rpoE gene of Escherichia coli, which encodes sigma E, is essential for bacterial growth at high temperature. J. Bacteriol. 1995, 177, 2918–2922. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grabowicz, M.; Silhavy, T.J. Envelope stress responses: An interconnected safety net. Trends Biochem. Sci. 2017, 42, 232–242. [Google Scholar] [CrossRef] [Green Version]
- Rhodius, V.A.; Suh, W.C.; Nonaka, G.; West, J.; Gross, C.A. Conserved and variable functions of the σE stress response in related genomes. PLoS Biol. 2006, 4, e2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mecsas, J.; Rouviere, P.E.; Erickson, J.W.; Donohue, T.J.; Gross, C.A. The activity of δE, an Escherichia coli heat-inducible δ-factor, is modulated by expression of outer membrane proteins. Genes Dev. 1993, 7, 2618. [Google Scholar] [CrossRef] [Green Version]
- Bernal-Cabas, M.; Ayala, J.A.; Raivio, T.L. The Cpx envelope stress response modifies peptidoglycan cross-linking via the L,D-transpeptidase LdtD and the novel protein YgaU. J. Bacteriol. 2015, 197, 603–614. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guest, R.L.; Wang, J.; Wong, J.L.; Raivio, T.L. A bacterial stress response regulates respiratory protein complexes to control envelope stress adaptation. J. Bacteriol. 2017, 199, e00153-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimohata, N.; Chiba, S.; Saikawa, N.; Ito, K.; Akiyama, Y. The Cpx stress response system of Escherichia coli senses plasma membrane proteins and controls HtpX, a membrane protease with a cytosolic active site. Genes Cells 2002, 7, 653–662. [Google Scholar] [CrossRef] [PubMed]
- Hung, D.L.; Raivio, T.L.; Jones, C.H.; Silhavy, T.J.; Hultgren, S.J. Cpx signalling pathway monitors biogenesis and affects assembly and expression of P pili. EMBO J. 2001, 20, 1508–1518. [Google Scholar] [CrossRef]
- Batchelor, E.; Walthers, D.; Kenney, L.J.; Goulian, M. The Escherichia coli CpxA-CpxR envelope stress response system regulates expression of the porins ompF and ompC. J. Bacteriol. 2005, 187, 5723–5731. [Google Scholar] [CrossRef] [Green Version]
- De Wulf, P.; McGuire, A.M.; Liu, X.; Lin, E.C. Genome-wide profiling of promoter recognition by the two-component response regulator CpxR-P in Escherichia coli. J. Biol. Chem. 2002, 277, 26652–26661. [Google Scholar] [CrossRef] [Green Version]
- Rattanama, P.; Thompson, J.R.; Kongkerd, M.; Srinitiwarawong, K.; Vuddhakul, V.; Mekalanos, J.J. Sigma E regulators control haemolytic activity and virulence in a shrimp pathogenic Vibrio harveyi. PLoS ONE 2012, 7, e32523. [Google Scholar] [CrossRef]
- Matter, L.B.; Ares, M.A.; Abundes-Gallegos, J.; Cedillo, M.L.; Yáñez, J.A.; Martinez-Laguna, Y.; De laCruz, M.A.; Girón, J.A. The CpxRA stress response system regulates virulence features of avian pathogenic Escherichia coli. Environ. Microbiol. 2018, 20, 3363–3377. [Google Scholar] [CrossRef]
- Lima, S.; Guo, M.S.; Chaba, R.; Gross, C.A.; Sauer, R.T. Dual molecular signals mediate the bacterial response to outer-membrane stress. Science 2013, 340, 837. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jones, C.H.; Danese, P.N.; Pinkner, J.S.; Silhavy, T.J.; Hultgren, S.J. The chaperone-assisted membrane release and folding pathway is sensed by two signal transduction systems. EMBO J. 1997, 16, 6394–6406. [Google Scholar] [CrossRef]
- Gerken, H.; Charlson, E.S.; Cicirelli, E.M.; Kenney, L.J.; Misra, R. MzrA: A novel modulator of the EnvZ/OmpRtwo-component regulon. Mol. Microbiol. 2009, 72, 1408–1422. [Google Scholar] [CrossRef]
- Grabowicz, M.; Koren, D.; Silhavy, T.J. The CpxQ sRNA negatively regulates Skp to prevent mistargeting of β-barrel outer membrane proteins into the cytoplasmic membrane. mBio 2016, 7, e00312–e00316. [Google Scholar] [CrossRef] [Green Version]
- Crouch, M.L.; Becker, L.A.; Bang, I.S.; Tanabe, H.; Ouellette, A.J.; Fang, F.C. The alternative sigma factor σE is required for resistance of Salmonella enterica serovar Typhimurium to anti-microbial peptides. Mol. Microbiol. 2005, 56, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, S.; Medihala, P.; Dynes, J.J.; Daida, P.; Vujanovic, V.; Hitchcock, A.P.; Shitty, D.; Zhang, H.; Brown, D.R.; Lawrence, J.R.; et al. Importance of the RpoE regulon in maintaining the lipid bilayer during antimicrobial treatment with the polycationic agent, chlorhexidine. Proteomics 2018, 18, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Sun, Y.; Yan, L.; Pan, Y.; Gao, Y.; Ma, C.; Hu, G. Regulation of the two-component regulator CpxR on aminoglycosides and ß-lactams resistance in Salmonella enterica serovar Typhimurium. Front. Microbiol. 2016, 7, 604. [Google Scholar] [CrossRef] [Green Version]
- Tian, Z.X.; Yi, X.X.; Cho, A.; O’Gara, F.; Wang, Y.P. CpxR activates MexAB-OprM pump expression and enhances antibiotic resistance in both laboratory and clinical nalB-type isolates of Pseudomonas aeruginosa. PLoS Pthog. 2016, 12, e1005932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raivio, T.L.; Leblanc, S.K.D.; Price, N.L. The Escherichia coli Cpx envelope stress response regulates genes of diverse function that impact antibiotic resistance and membrane integrity. J. Bacteriol. 2013, 195, 2755–2767. [Google Scholar] [CrossRef] [Green Version]
- Otto, K.; Silhavy, T.J. Surface sensing and adhesion of Escherichia coli controlled by the Cpx-signalling pathway. Proc. Natl. Acad. Sci. USA 2002, 99, 2287–2292. [Google Scholar] [CrossRef] [Green Version]
- Vidovic, S.; An, R.; Rendahl, A. Molecular and physiological characterization of fluoroquinolone-highly resistant Salmonella enteritidis strains. Front. Microbiol. 2019, 10, 729. [Google Scholar] [CrossRef]
- Choi, U.; Lee, C.R. Distinct roles of outer membrane porins in antibiotic resistance and membrane integrity in Escherichia coli. Front. Microbiol. 2019, 10, 953. [Google Scholar] [CrossRef]
- Vergalli, J.; Bodrenko, I.V.; Masi, M.; Moynié, L.; Acosta-Gutiérrez, S.; Naismith, J.H.; Davin-Regli, A.; Ceccarelli, M.; van den Berg, B.; Winterhalter, M.; et al. Porins and small-molecule translocation across the outer membrane of Gram-negative bacteria. Nat. Rev. Microbiol. 2020, 18, 164–176. [Google Scholar] [CrossRef]
- Futoma-Koloch, B.; Bugla-Płoskońska, G.; Dudek, B.; Dorotkiewicz-Jach, A.; Drulis-Kawa, Z.; Gamian, A. Outer membrane proteins of Salmonella as potential markers of resistance to serum, antibiotics and biocides. Curr. Med. Chem. 2019, 26, 1960–1978. [Google Scholar] [CrossRef]
- Liu, X.; Omar, M.; Abrahante, J.E.; Nagaraja, K.V.; Vidovic, S. Insights into the oxidative stress response of Salmonella enterica serovar Enteritidis revealed by the next generation sequencing approach. Antioxidants 2020, 9, 849. [Google Scholar] [CrossRef] [PubMed]
- Imlay, J.A. The molecular mechanisms and physiological consequences of oxidative stress: Lessons from a model bacterium. Nat. Rev. Microbiol. 2013, 11, 443–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holmgren, A. Antioxidant function of thioredoxin and glutaredoxin systems. Antioxid. Redox Signals. 2000, 2, 811–820. [Google Scholar] [CrossRef] [PubMed]
- Serata, M.; Lino, T.; Yasuda, E.; Sako, T. Roles of thioredoxin and thioredoxin reductase in the resistance to oxidative stress in Lactobacillus casei. Microbiology 2012, 158, 953–962. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tattawasart, U.; Hann, A.C.; Maillard, J.Y.; Furr, J.R.; Russell, A.D. Cytological changes in chlorhexidine-resistant isolates of Pseudomonas stutzeri. J. Antimicrob. Chemother. 2000, 45, 145–152. [Google Scholar] [CrossRef] [Green Version]
- Fraikin, N.; Goormaghtigh, F.; van Melderen, L. Type II toxin-antitoxin systems: Evolution and revolutions. J. Bacteriol. 2020, 202, e00763-19. [Google Scholar] [CrossRef] [Green Version]
- van Melderen, L.; Bernard, P.; Couturier, M. Lon-dependent proteolysis of CcdA is the key control for activation of CcdB in plasmid-free segregate bacteria. Mol. Microbiol. 1994, 11, 1151–1157. [Google Scholar] [CrossRef]
- Soo, V.W.; Wood, T.K. Antitoxin MqsA represses curli formation through the master biofilm regulator CsgD. Sci. Rep. 2013, 3, 3186. [Google Scholar] [CrossRef] [Green Version]
- Christensen, S.K.; Mikkelsen, M.; Pedersen, K.; Gerdes, K. RelE, a global inhibitor of translation, is activated during nutritional stress. Proc. Natl. Acad. Sci. USA 2001, 98, 14328–14333. [Google Scholar] [CrossRef] [Green Version]
- Helaine, S.; Cheverton, A.M.; Watson, K.G.; Faure, L.M.; Matthews, S.A.; Holden, D.W. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 2014, 343, 204–208. [Google Scholar] [CrossRef]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef] [Green Version]
- Shitty, D.; Abrahante, J.E.; Chekabab, S.M.; Wu, X.; Korber, D.R.; Vidovic, S. Role of CpxR in biofilm development: Expression of key fimbrial, O-antigen and virulence operons of Salmonella enteritidis. Int. J. Mol. Sci. 2019, 20, 5146. [Google Scholar] [CrossRef] [Green Version]
- Sato, T. A modified method for lead staining of thin sections. J. Electron Microsc. 1986, 17, 158–159. [Google Scholar]
- Anderson, K.J.; Vermillion, K.L.; Jagtap, P.; Johnson, J.E.; Griffin, T.J.; Andrews, M.T. Proteogenomic analysis of a hibernating mammal indicates contribution of skeletal muscle physiology to the hibernation phenotype. J. Proteome Res. 2016, 15, 1253–1261. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, S.; Elder, J.; Medihala, P.; Lawrence, J.R.; Predicala, B.; Zhang, H.; Korber, D.R. ZnO nanoparticles impose a panmetabolic toxic effect along with strong necrosis, inducing activation of the envelope stress response in Salmonella enterica serovar Enteritidis. Antimicrob. Agents Chemother. 2015, 59, 3317–3328. [Google Scholar] [CrossRef] [Green Version]
- Lin-Moshier, Y.; Sebastian, P.J.; Higgins, L.; Sampson, N.D.; Hewitt, J.E.; Marchant, J.S. Re-evaluation of the role of calcium homeostasis endoplasmic reticulum protein (CHERP) in cellular calcium signalling. J. Biol. Chem. 2013, 288, 355–367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oberg, A.L.; Mahoney, D.W.; Eckel-Passow, J.E.; Malone, C.J.; Wolfinger, R.D.; Hill, E.G.; Cooper, L.T.; Onuma, O.K.; Spiro, C.; Therneau, T.M.; et al. Statistical analysis of relative labeled mass spectrometry data from complex samples using ANOVA. J. Proteome Res. 2008, 7, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, S.; Liu, X.; An, R.; Mendoza, K.M.; Abrahante, J.E.; Johny, A.K.; Reed, K.M. Transcriptional profiling and molecular characterization of the yccT mutant link: A novel STY1099 protein with the peroxide stress response and cell division of Salmonella enterica serovar Enteritidis. Biology 2019, 8, 86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocol. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]





| KO Identifiers | Gene Name | Accession Number | Mol. wt. kDa | Benjamini Hochberg p Value | Fold Change | Description |
|---|---|---|---|---|---|---|
| Peptidoglycan Biosynthesis and Degradation | ||||||
| K00075 | murB | WP_000149793.1 | 38 | 0.00012 | 1.27 | UDP-N-acetylmuramate dehydrogenase |
| K01928 | murE | WP_000775071.1 | 53 | 0.001 | 1.15 | UDP-N-acetylmuramoyl-l-alanyl-d-glutamate--2,6-diaminopimelate ligase |
| K01776 | murL | WP_031619622.1 | 31 | 0.013 | 1.12 | Glutamate racemase |
| K01297 | ldcA | WP_000051603.1 | 33 | 0.00078 | 1.23 | Muramoyltetrapeptide carboxypeptidase |
| K01921 | ddl | WP_000763905.1 | 33 | 0.001 | 1.26 | d-alanine-d-alanine ligase |
| K03806 | ampD | WP_000936324.1 | 21 | 0.003 | 1.20 | N-acetyl-anhydromuramoyl-l-alanine amidase |
| K06078 | lpp | WP_001082307.1 | 8 | 0.021 | 1.18 | Murein lipoprotein |
| K08306 | mltC | WP_000976287.1 | 40 | 0.006 | 1.32 | Membrane-bound lytic murein transglycosylase C |
| K19236 | ycfS | WP_001708681.1 | 33 | 0.002 | 2.22 | l,d-transpeptidase YcfS |
| K01448 | amiA | WP_069040808.1 | 32 | 0.002 | 0.81 | N-acetylmuramoyl-l-alanine amidase |
| K01775 | alr | WP_001147296.1 | 39 | 0.0016 | 0.90 | Alanine racemase |
| K07258 | dacD | WP_000925044.1 | 43 | 0.002 | 0.40 | Serine-type d-Ala-d-Ala carboxypeptidase |
| K18988 | ampH | WP_000830784.1 | 42 | 0.004 | 0.46 | Serine-type d-Ala-d-Ala endopeptidase |
| K21470 | ltdT | WP_000925899.1 | 68 | 0.002 | 0.72 | l,d-transpeptidase |
| Lipopolysaccharide Biosynthesis | ||||||
| K00677 | lpxA | WP_000565950.1 | 28 | 0.003 | 1.32 | UDP-N-acetylglucosamine acyltransferase |
| K02517 | lpxL | WP_000163977.1 | 35 | 0.01 | 1.21 | Kdo2-lipid IVA lauroyltransferase/acyltransferase |
| K02535 | lpxC | WP_000595474.1 | 34 | 0.012 | 1.23 | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase |
| K12979 | lpxO | WP_000457031.1 | 35 | 0.012 | 1.50 | Lipid A hydroxylase LpxO |
| K02841 | rfaC | WP_076915342.1 | 35 | 0.00031 | 1.57 | Lipopolysaccharide heptosyltransferase |
| K03760 | eptA | WP_038427769.1 | 62 | 0.013 | 1.25 | Lipid A ethanolaminephosphotransferase |
| K10011 | arnA | WP_038425381.1 | 73 | 0.02 | 1.10 | UDP-4-amino-4-deoxy-l-arabinose formyltransferase/UDP-glucuronic acid dehydrogenase |
| K09953 | lpxR | WP_001046434.1 | 35 | 0.002 | 0.44 | Lipid A 3-O-deacylase |
| Glycosyltransferases | ||||||
| K02841 | cobK | WP_076915342.1 | 35 | 0.00032 | 1.27 | Lipopolysaccharide heptosyltransferase |
| K02852 | rffM | WP_000183613.1 | 28 | 0.012 | 1.32 | Lipopolysaccharide N-acetylmannosaminouronosyltransferase |
| K12992 | rfbN | WP_000705151.1 | 36 | 0.00012 | 1.16 | O antigen biosynthesis rhamnosyltransferase |
| Transporters | ||||||
| K02000 | proV | WP_069057539.1 | 44 | 0.017 | 1.23 | Glycine betaine/proline transporter |
| K02068 | fetA | WP_000166987.1 | 25 | 0.004 | 1.21 | Iron ABC transporter ATP-binding protein |
| K02071 | metN | WP_079837681.1 | 32 | 0.00018 | 1.30 | Methionine ABC transporter |
| K02598 | nirC | WP_000493575.1 | 29 | 0.004 | 1.69 | Nitrite transporter |
| K02759 | chbA | WP_001732541.1 | 11 | 0.001 | 1.23 | Cellobiose PTS system EIIA component |
| K03475 | ulaA | WP_001721663.1 | 47 | 0.004 | 1.50 | Ascorbate PTS system EIIC component |
| K03561 | exbB | WP_000527859.1 | 26 | 0.004 | 1.28 | Biopolymer transporter ExbB |
| K03562 | tolQ | WP_000131318.1 | 26 | 0.001 | 1.33 | Biopolymer transport protein TolQ |
| K03832 | tonB | WP_001517937.1 | 26 | 0.006 | 1.19 | TonB system transport protein TonB |
| K04759 | feoB | WP_000736978.1 | 84 | 0.015 | 1.15 | Ferrous iron transport protein B |
| K05685 | macB | WP_000125899.1 | 71 | 0.003 | 1.22 | Macrolide ABC transporter permease MacB |
| K05776 | modF | WP_079983686.1 | 55 | 0.001 | 1.27 | Molybdate transporter ATP-binding protein |
| K06155 | gntT | WP_001131737.1 | 46 | 0.012 | 1.77 | Gluconate transporter |
| K07091 | lptF | WP_000584130.1 | 40 | 0.005 | 1.29 | LPS export ABC transporter permease LptF |
| K07127 | hiuH | WP_080199438.1 | 15 | 0.004 | 1.32 | 5-hydroxyisourate hydrolase |
| K07278 | tamA | WP_001120233.1 | 65 | 0.00016 | 1.26 | Translocation and assembly module TamA |
| K07308 | dmsB | WP_132631290.1 | 55 | 0.00045 | 1.34 | Dimethylsulfoxide reductase subunit B |
| K08351 | bisC | WP_023227718.1 | 86 | 0.006 | 1.17 | Biotin/methionine sulfoxide reductase |
| K09013 | sufC | WP_001580259.1 | 28 | 0.0004 | 1.20 | Fe-S cluster assembly ATP-binding protein |
| K09810 | lolD | WP_001033714.1 | 25 | 0.004 | 1.35 | Lipoprotein-releasing ABC transporter |
| K09812 | ftsE | WP_000617729.1 | 24 | 0.002 | 1.26 | Cell division transporter ATP-binding protein |
| K09817 | znuC | WP_000203014.1 | 28 | 0.001 | 1.22 | Zinc ABC transporter ATP-binding protein |
| K09997 | artJ | WP_000756583.1 | 27 | 0.002 | 1.11 | Arginine ABC transporter |
| K10001 | gltI | WP_000588819.1 | 34 | 0.003 | 1.30 | Glutamate/aspartate transporter |
| K10017 | hisP | WP_000986780.1 | 29 | 0.012 | 1.49 | Histidine transporter ATP-binding protein |
| K10439 | rbsB | WP_023254734.1 | 31 | 0.001 | 1.13 | Ribose transporter substrate-binding protein |
| K10540 | mglB | WP_069057568.1 | 36 | 0.00056 | 1.27 | Methyl-galactoside transporter |
| K10542 | mglA | WP_000535907.1 | 56 | 0.004 | 1.64 | Methyl-galactoside transporter |
| K11073 | potF | WP_000125769.1 | 41 | 0.004 | 1.43 | Putrescine transporter |
| K11720 | lptG | WP_001182241.1 | 40 | 0.009 | 1.26 | LPS export ABC transporter permease |
| K13893 | yejA | WP_135416368.1 | 69 | 0.0049 | 1.39 | Microcin C substrate-binding protein |
| K16012 | cydC | WP_001202251.1 | 63 | 0.0055 | 1.39 | Cysteine/glutathione ABC transporter |
| K18138 | adeB | WP_001132506.1 | 114 | 0.001 | 1.27 | Multidrug efflux RND transporter |
| K19226 | sapA | WP_001241619.1 | 62 | 0.001 | 1.27 | Peptide ABC transporter substrate-binding |
| K23991 | ptsP | WP_079829802.1 | 85 | 0.003 | 3.46 | Multiphosphoryl transfer protein |
| K24163 | nhaK | WP_001696843.1 | 60 | 0.016 | 1.10 | Na+/H+ antiporter |
| K01999 | livK | WP_000676957.1 | 39 | 0.003 | 0.48 | Branched-chain amino acid ABC transporter |
| K02017 | modC | WP_000891715.1 | 39 | 0.00037 | 0.61 | Molybdate transporter ATP-binding protein |
| K02036 | pstB | WP_000063118.1 | 29 | 0.00017 | 0.82 | Phosphate ABC transporter |
| K02064 | tbpA | WP_000915326.1 | 36 | 0.00083 | 0.88 | Thiamine ABC transporter |
| K02445 | glpT | WP_080195040.1 | 44 | 0.004 | 0.25 | Glycerol-3-phosphate transporter |
| K02575 | narK | WP_000019850.1 | 50 | 0.001 | 0.55 | Nitrate transporter NarK |
| K02774 | gatB | WP_000723161.1 | 10 | 0.004 | 0.19 | PTS sugar transporter subunit IIB |
| K02777 | crr | WP_000522253.1 | 18 | 0.014 | 0.87 | PTS glucose transporter subunit IIA |
| K02779 | ptsG | WP_000475705.1 | 50 | 0.001 | 0.78 | PTS glucose transporter subunit IIBC |
| K02782 | srlE | WP_000199033.1 | 34 | 0.007 | 0.57 | PTS glucitol/sorbitol transporter subunit IIB |
| K02784 | ptsH | WP_000487600.1 | 9 | 0.002 | 0.76 | Phosphocarrier protein Hpr |
| K03284 | corA | WP_000947139.1 | 37 | 0.014 | 0.86 | Magnesium/cobalt transporter CorA |
| K04758 | feoA | WP_061451033.1 | 8 | 0.01 | 0.73 | Ferrous iron transporter A |
| K05517 | tsx | WP_000752021.1 | 33 | 0.004 | 0.74 | Nucleoside-specific channel-forming protein |
| K05816 | ugpC | WP_000907837.1 | 39 | 0.004 | 0.80 | Sn-glycerol-3-phosphate import protein UgpC |
| K05845 | opuC | WP_000155871.1 | 33 | 0.002 | 0.40 | ABC transporter substrate-binding protein |
| K07122 | mlaB | WP_000188843.1 | 11 | 0.018 | 0.79 | Lipid asymmetry maintenance protein MlaB |
| K07306 | dmsA | WP_077917388.1 | 89 | 0.004 | 0.10 | Anaerobic dimethyl sulfoxide reductase A |
| K07795 | tctC | WP_000744418.1 | 30 | 0.002 | 0.40 | Tricarboxylic transport membrane protein |
| K08154 | emrD | WP_000828735.1 | 42 | 0.007 | 0.51 | Multidrug efflux MFS transporter EmrD |
| K08353 | phsB | WP_001015351.1 | 21 | 0.003 | 0.47 | Thiosulfate reductase electron transport protein |
| K09475 | ompC | WP_000758335.1 | 41 | 0.002 | 0.50 | Outer membrane pore protein C |
| K10555 | lsrB | WP_079920817.1 | 37 | 0.002 | 0.73 | AI-2 transport substrate-binding protein |
| K11732 | pheP | WP_000786283.1 | 51 | 0.02 | 0.75 | Phenylalanine-specific permease |
| K11738 | ansP | WP_000857110.1 | 54 | 0.0015 | 0.58 | l-asparagine permease |
| K14062 | ompN | WP_000824321.1 | 42 | 0.019 | 0.35 | Outer membrane protein N |
| K18141 | acrE | WP_000160380.1 | 41 | 0.02 | 0.74 | Efflux RND transporter periplasmic adaptor |
| K23188 | fepC | WP_023227378.1 | 29 | 0.004 | 0.51 | Iron-enterobactin ABC transporter |
| Gene Name | Accession Number | Mol. wt. kDa | Benjamini Hochberg p Value | Log 2 Fold Change | Fold Change | Description |
|---|---|---|---|---|---|---|
| emtA | WP_000776974.1 | 22 | 0.006 | −0.85 | 0.55 | Membrane-bound lytic murein transglycosylase |
| asmA | WP_023227411.1 | 75 | 0.006 | −1.43 | 0.37 | Assembly of outer membrane proteins |
| frlB | WP_023206877.1 | 37 | 0.006 | −3.08 | 0.12 | Fructoselysine-6-P-deglycase |
| arnC | WP_000458893.1 | 37 | 0.001 | −0.84 | 1.55 | Undecaprenyl-phosphate 4-deoxy-4-formamido-l-arabinose transferase |
| pepE | WP_000421776.1 | 25 | 0.018 | −1.55 | 0.34 | Dipeptidase |
| fdxH | WP_000061599.1 | 32 | 0.001 | −1.13 | 0.46 | Formate dehydrogenase subunit beta |
| hemX | WP_000138954.1 | 42 | 0.001 | −1.83 | 0.28 | Uroporphyrinogen-III C-methyltransferase |
| nfsB | WP_000355870.1 | 24 | 0.006 | −4.38 | 0.05 | Oxygen-insensitive NAD(P)H nitroreductase |
| WP_065618791.1 | 27 | 0.006 | −1.01 | 0.50 | Phosphatase PAP2 family protein | |
| WP_000750393.1 | 8 | 0.006 | −1.13 | 0.46 | YgdI/YgdR family lipoprotein | |
| WP_000748128.1 | 11 | 0.006 | −3.64 | 0.08 | EexN family lipoprotein | |
| WP_001095011.1 | 58 | 0.006 | −2.36 | 0.19 | Membrane protein | |
| WP_001240360.1 | 39 | 0.00017 | −1.6 | 0.33 | Membrane protein | |
| rfbB | WP_000697848.1 | 41 | 0.006 | 3.87 | 14.62 | dTDP-glucose 4,6-dehydratase |
| malS | WP_000761323.1 | 76 | 0.006 | 1.17 | 2.25 | Periplasmic alpha-amylase |
| Gene Name. | Protein Function | Primer Pair | Sequence of Primers |
|---|---|---|---|
| acrB | Multidrug efflux RND transporter AcrB | acrB–F | CACGAAACCAATCTGCGTAAAG |
| acrB–R | CTTCGCCGTCCTGCTTATT | ||
| cutC | Copper homeostasis protein CutC | cutC–F | GAATACTCGTCCGCCTGTATATC |
| cutC–R | GAGTACGGGAAGTACACAGTTC | ||
| phsA | Thiosulfate reductase PhsA | phsA–F | ATCTCATCGCCGGTCTTAATG |
| phsA–R | GACGCAGTACGTACCTTTACTC | ||
| tdcA | Transcriptional regulator TdcA | tdcA–F | GATTCAGACCAGGAAAGCAGTA |
| tdcA–R | AGCGATGTTGAGGCCTATTT | ||
| hlyD | Secretion protein HlyD | hlyD–F | GATAATGCGCAGGCGATAGA |
| hlyD–R | GGTCGTCCTGACAAACCTTAC | ||
| rfaC | Lipopolysaccharide heptosyltransferase RfaC | rfaC–F | GCAACACCGGATTACGGATAAA |
| rfaC–R | GCGGTAACACATCCACATAGTC | ||
| lbpB | Heat shock chaperone IbpB | ibpB–F | CCAGATCTTCCTGACGGAAAC |
| ibpB–R | AGAGCTTCCCGCCCTATAA | ||
| dedD | Cell division protein DedD | dedD–F | CCGACGAGAATACGGGTTATTT |
| dedD–R | CGCGGATAAAGTCAACGAGATA | ||
| fumD | Fumarate hydratase FumD | fumD–F | CTTAACACGCCCGCAATAAC |
| fumD–R | AACTGTACCGCGAGATGTG | ||
| pspB | Envelope stress response protein PspB | pspB–F | TCGCTTTGCGACAGTTCT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Omar, M.; Nagaraja, K.V.; Goyal, S.M.; Vidovic, S. Novel Insight into the Effects of CpxR on Salmonella enteritidis Cells during the Chlorhexidine Treatment and Non-Stressful Growing Conditions. Int. J. Mol. Sci. 2021, 22, 8938. https://doi.org/10.3390/ijms22168938
Liu X, Omar M, Nagaraja KV, Goyal SM, Vidovic S. Novel Insight into the Effects of CpxR on Salmonella enteritidis Cells during the Chlorhexidine Treatment and Non-Stressful Growing Conditions. International Journal of Molecular Sciences. 2021; 22(16):8938. https://doi.org/10.3390/ijms22168938
Chicago/Turabian StyleLiu, Xiaoying, Misara Omar, Kakambi V. Nagaraja, Sagar M. Goyal, and Sinisa Vidovic. 2021. "Novel Insight into the Effects of CpxR on Salmonella enteritidis Cells during the Chlorhexidine Treatment and Non-Stressful Growing Conditions" International Journal of Molecular Sciences 22, no. 16: 8938. https://doi.org/10.3390/ijms22168938
APA StyleLiu, X., Omar, M., Nagaraja, K. V., Goyal, S. M., & Vidovic, S. (2021). Novel Insight into the Effects of CpxR on Salmonella enteritidis Cells during the Chlorhexidine Treatment and Non-Stressful Growing Conditions. International Journal of Molecular Sciences, 22(16), 8938. https://doi.org/10.3390/ijms22168938






